
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Genet. , 19 March 2025
Sec. Genomics of Plants and the Phytoecosystem
Volume 16 - 2025 | https://doi.org/10.3389/fgene.2025.1532956
This article is part of the Research Topic Precision Trait Mapping and Molecular Breeding in High-Impact Crop Plants View all 3 articles
 Upendra Kumar1,2*‡
Upendra Kumar1,2*‡ Sourav Panigrahi1‡
Sourav Panigrahi1‡ Rita Goswami1‡
Rita Goswami1‡ Yogita Singh1†
Yogita Singh1† Priyanka Balyan3
Priyanka Balyan3 Prexha Kapoor1
Prexha Kapoor1 Sundip Kumar4
Sundip Kumar4 Krishna Pal Singh5,6
Krishna Pal Singh5,6 Farkhandah Jan7
Farkhandah Jan7 Reyazul Rouf Mir7
Reyazul Rouf Mir7Introduction: To address the urgent demand for biofortified wheat enriched with health-beneficial dietary fibres such as β-glucan, this study employed meticulous crossbreeding between established wheat cultivars and the β-glucan-rich wild relative Aegilops kotschyi accession “AK-3790”.
Methods: Within this context, a derivative line encompassing a pair of 7U chromosomes from Ae. Kotschyi, denoted as 63-2-13, was identified. The presence of the 7U chromosome in this line was confirmed through comprehensive molecular marker and genomic in situ hybridization (GISH) analyses. With the aim of increasing the β-glucan content in hexaploid wheat, two distinct backcross populations were developed utilizing the 63-2-13 line as the donor parent and two separate recurrent parents (WH1105 and HD3086). These populations underwent an exact selection regimen, encompassing parent-like phenotypes, heightened yield, and robust resistance to yellow rust, meticulously tracked across successive generations until the BC2F2:3 stage.
Results and Discussion: Notably, among the outcomes, selected BC2F2:3 progenies presented remarkable increases in β-glucan levels, with a notable increase (BC2F2:3 23-5) resulting in an impressive increase in the 1.76% grain β-glucan content. Despite a discernible reduction in yield compared with their high-yielding counterparts, BC2F2:3 23-5 demonstrated a harmonious trait profile, encompassing heightened β-glucan content and moderate yellow rust resistance, thus positioning it as a compelling candidate for subsequent refinement endeavors. This research notably underscores the substantial potential of precise introgression strategies for increasing the β-glucan content in wheat, thereby underscoring the imperative of adept trait optimization to ensure both yield stability and nutritional enhancement.
The contemporary emphasis on fostering healthier dietary patterns has underscored the imperative of devising nutritionally fortified food products. Among the pivotal constituents of a health-promoting diet, dietary fibre, a fundamental component abundant in plant cell walls, has garnered particular attention. As dietary fibre consumption increases, evidence has emerged linking it to a reduced susceptibility to chronic maladies, including heart disease, type 2 diabetes, breast cancer, and colon cancer (Reynolds et al., 2019). Notably, cereal-derived fibres, constituting a significant share of dietary fibre intake, have gained prominence. Within this dietary fibre landscape, 1–3,1–4 mixed-linkage β-D-glucan, a prominent soluble fibre predominantly found in cereal grains such as barley and oats, has garnered special interest. This fiber exerts a multifaceted influence by mitigating risk factors associated with type 2 diabetes and cardiovascular diseases, primarily by modulating glycemic responses, reducing cholesterol levels, and attenuating insulin-related effects (Wood, 2007; Sibakov et al., 2013; Cui and Liu, 2013).
Among the diverse array of cereal crops, wheat (Triticum aestivum L.) plays a pivotal role as a global staple, substantially contributing to daily caloric and protein intake (FAO, 2020). However, the content of soluble β-glucans in wheat grains is notably modest, particularly in comparison to the more abundant concentrations present in oat and barley grains (Beresford and Stone, 1983; Havrlentová and Kraic, 2006; Collins et al., 2010). Despite dedicated endeavors, the primary and secondary gene pools of bread wheat have exhibited constrained genetic diversity conducive to the expression of high β-glucan contents (Pritchard et al., 2011; Marcotuli et al., 2017).
Moreover, neither bread wheat (T. aestivum L. 2n = 6x = 42) nor its closest relatives exhibit genetic diversity for high grain β-glucan levels (Friebe et al., 1996). This scarcity of genetic diversity extends to both the primary and secondary gene pools, limiting the feasibility of establishing a breeding program aimed at increasing the β-glucan content in cultivated wheat varieties. This conclusion stems from comprehensive studies involving over 500 wheat germplasms of diverse origins, including primitive, elite, and synthetic lines of hexaploid, tetraploid, and diploid wheat, as well as related species (Pritchard et al., 2011; Marcotuli et al., 2019).
To alter the quality of wheat grains, a strategic avenue involves incorporating genes from less closely related relatives, which may belong to the tertiary wheat gene pool and encompass both wild and cultivated species. This is achieved through chromosome engineering and interspecific hybridization (Cseh et al., 2011; 2013; Rakszegi et al., 2017; Danilova et al., 2018; Türkösi et al., 2018). Among these tertiary gene pool members, barley has the highest β-glucan concentration (up to 11%), making it an effective candidate for interspecific hybridization. Notably, certain species of Aegilops, which are also part of the wheat tertiary gene pool, present significant β-glucan contents (up to 7.1%) in their grains. These species include diploid types such as Ae. Umbellulata (Zhuk.), as well as various tetraploid species containing U and C genomes (Rakszegi et al., 2017; Marcotuli et al., 2019). In contrast, most bread wheat (T. aestivum L.) cultivars and durum wheat (Triticum turgidum L. ssp. durum) exhibit lower levels of β-glucan in their grains (Marcotuli et al., 2016; Mohebbi et al., 2018).
The practice of harnessing genetic traits from distantly related species to augment wheat’s genetic robustness and resilience has been previously demonstrated, notably in the context of conferring resistance against various biotic stresses (Friebe et al., 1996; 2000; Marais et al., 2005; Qi et al., 2007; Kuraparthy et al., 2007a; b; Schneider et al., 2008). In this study, we present a comprehensive exploration of the process and molecular insights underlying the successful introgression of Ae. Kotschyi chromosomes, thereby paving the way for an enriched β-glucan content in wheat grains.
The Ae. Kotschyi accession AK-3790 was chosen for our breeding program because of its known high grain β-glucan concentration (up to 2.4%) (Marcotuli et al., 2019). The seeds of a wheat-Ae. kotschyi derivative line (63-2–13) (Tiwari et al. 2010) were sourced from the Wheat Germplasm Collection at Eternal University, Baru Sahib, Himachal Pradesh, India. This derivative line resulted from a cross between Ae. Kotschyi (accession AK-3790) as the male ‘donor’ and the recipient Chinese spring PhI. To develop two backcross populations, the derivative line 63-2-13 was employed as the donor parent in a backcrossing program, utilizing the wheat cultivars WH1105 and HD3086 as recurrent parents to generate BC2F1 plants during the winter of 2020–21. BC2F2 plants were subsequently generated, followed by self-pollination and the production of BC2F2:3 lines via the single-seed descent method in 2022–23. These lines were grown via an augmented design alongside three reference checks: Chinese spring (CS), WH1105, and HD3086, following standard wheat agronomic conditions. The breeding program was carried out at the farm of the Department of Molecular Biology and Biotechnology, while subsequent generation advancement was performed in the field of Wheat and Barley Section, Chaudhary Charan Singh Haryana Agricultural University, Hisar. On the basis of the performance of the genotypes in both generations, the BC2F2:3 population underwent screening for traits such as yield, yellow rust resistance, and parental backgrounds. The seeds of BC2F2:3 lines of selected agronomical superior plants were screened for grain β-glucan content, aiming to identify progeny lines with significantly elevated β-glucan levels comparable to those of checks WH1105 and HD3086.
At maturity, a range of morphological and phenotypic traits were observed and recorded in the field. These traits included plant height, the number of productive tillers, thousand-grain weight, and grain yield per plant. Additionally, the yellow rust score was documented during the anthesis stage according to the modified Cobb scale of 1-5 provided by Peterson et al. (1948), along with the severity of infection as a percentage.
Since the BC2F2:3 population is representative of the BC2F2 population, the mean data obtained from the populations were subjected to analysis via R (version 4.2.2). The descriptive statistics were computed, and frequency distribution graphs were plotted. The ANOVA of the traits taken in the study was carried out via the package ‘augmentedRCBD’. The correlations between the traits were also evaluated via SPSS 2.0, along with the significance of the correlations.
Ten-gram seed samples from each of the BC2F2:3 plants selected on the basis of phenotyping were analysed to determine the grain moisture content, 1,000 kernel weight (TKW), and β-glucan levels. For β-glucan analysis, the sampled grains were finely milled into 0.5 mm particles via a tissuelyser (Retsch MM400). The dry weight basis β-glucan content of the harvested seeds was determined via a β-glucan assay kit (Megazyme, Ireland) specifically designed for mixed linkage (1–3,1–4) β-D-glucan in cereal grains, following the guidelines of the AACC Method 32-23.01. This analysis was performed in triplicate using whole-grain flour. The measurement of the grain moisture content was also conducted in triplicate via a DA 7250 NIR Analyser (Perten, Sweden).
Leaves from young plants of BC2F2:3 progenies were used for DNA extraction, followed by the CTAB method of DNA extraction (Murray and Thompson, 1980). The SSR markers covering the entire length of each of the chromosomes in wheat were selected from the wheat genetic maps (Roder et al., 1998; Pestsova et al., 2000; Somers et al., 2004). The selected markers were screened for transferability to Ae. Kotschyi. A total of 135 SSR markers were selected on the basis of the screening for transferability. These markers, which exhibit polymorphisms between Aegilops kotschyi accession AK-3790 and the recipient wheat cultivars, were used for molecular marker analysis of the BC2F2:3 lines. In addition to these markers, new primers have also been designed for chromosome 7. The details of the designed primers are given in Online Resource 1. These SSR markers were utilized to validate the presence of Ae. Kotschyi chromosomal introgression in situations where plausible candidate chromosome(s) were detected.
The genomic DNAs extracted from Ae. Longissima (SlSl) and Ae. Umbellulata (UU) were employed to generate genomic probes for use in genomic in situ hybridization (GISH) experiments. Actively growing root tips from germinating seeds were subjected to a 24-h treatment with ice water to accumulate metaphases. The root tips were subsequently fixed in a mixture of 3 parts ethanol and 1 part glacial acetic acid. Following fixation, the root tips were stained with 1% acetocarmine and then gently squashed in a solution containing 45% acetic acid. For the preparation of the genomic probes, sheared genomic DNA (ranging from 0.2 kb to 0.6 kb) from Ae. Longissima (SlSl) and Ae. Umbellulata (UU) was utilized. For the preparation of the genomic probe of the Sl-genome, the genomic DNA of Ae. Longissima was labelled with fluorescein-11-dUTP (green) (Roche Applied Science, Indianapolis, IN), whereas for the preparation of the genomic probe of the U-genome, the genomic DNA of Ae. Umbellulata was labelled with tetramethylrhodamine-5-dUTP (red) (Roche Applied Science, Indianapolis, IN) through nick translation. The labelled probes were purified via the QIA quick Nucleotide Removal Kit (Qiagen, Valencia, CA). Blocking DNA, consisting of unlabelled sheared genomic DNA from Chinese spring wheat (ranging from 100 bp to 1 kb), was employed at a ratio of 1 ng of labelled probe (Sl- and U-genomes) to 100 ng of blocking DNA. The hybridization conditions and posthybridization washes followed the protocols outlined by Zhang et al. (2001). To enhance visualization, chromosomes were counterstained with 4,6-diamidino-2-phenylindole (DAPI). The slides were analysed via an Epifluorescence Zeiss Axioimager M1 microscope.
Self-pollination of BC2F1 plants resulted in the generation of 128 distinct BC2F2 lines, each exhibiting a wide range of phenotypic characteristics. Various yield-related traits, such as per-plant grain yield, thousand-grain weight, spike length, and productive tiller count, were evaluated, along with resistance to yellow rust. The BC2F2 lines were selfed to obtain the BC2F2:3 generation lines. Multiple yield-related traits, such as per-plant grain yield, thousand-grain weight, spike length, and productive tiller count, were evaluated, along with resistance to yellow rust (Online Resource 2.1).
ANOVA of the 128 BC2F2:3 genotypes revealed that the variation among the genotypes for each of the traits evaluated in the study was highly significant (Online Resource 2.2). Although the frequency distribution revealed that most of the genotypes presented a relatively low yield per plant, with relatively few productive tillers per plant, a relatively high thousand-grain weight and relatively small spikes, few genotypes presented yield and other yield attributes that were similar to those of the high-yield parents HD3086 and WH1105 (Figure 1). The yield per plant was highly correlated with the number of productive tillers per plant and was significantly negatively correlated with the severity of yellow rust, as expected. Similarly, a significant correlation was observed between thousand grain weight and yield but was negatively correlated with yellow rust infection. Surprisingly, plant height was significantly positively correlated with thousand-grain weight (Table 1).
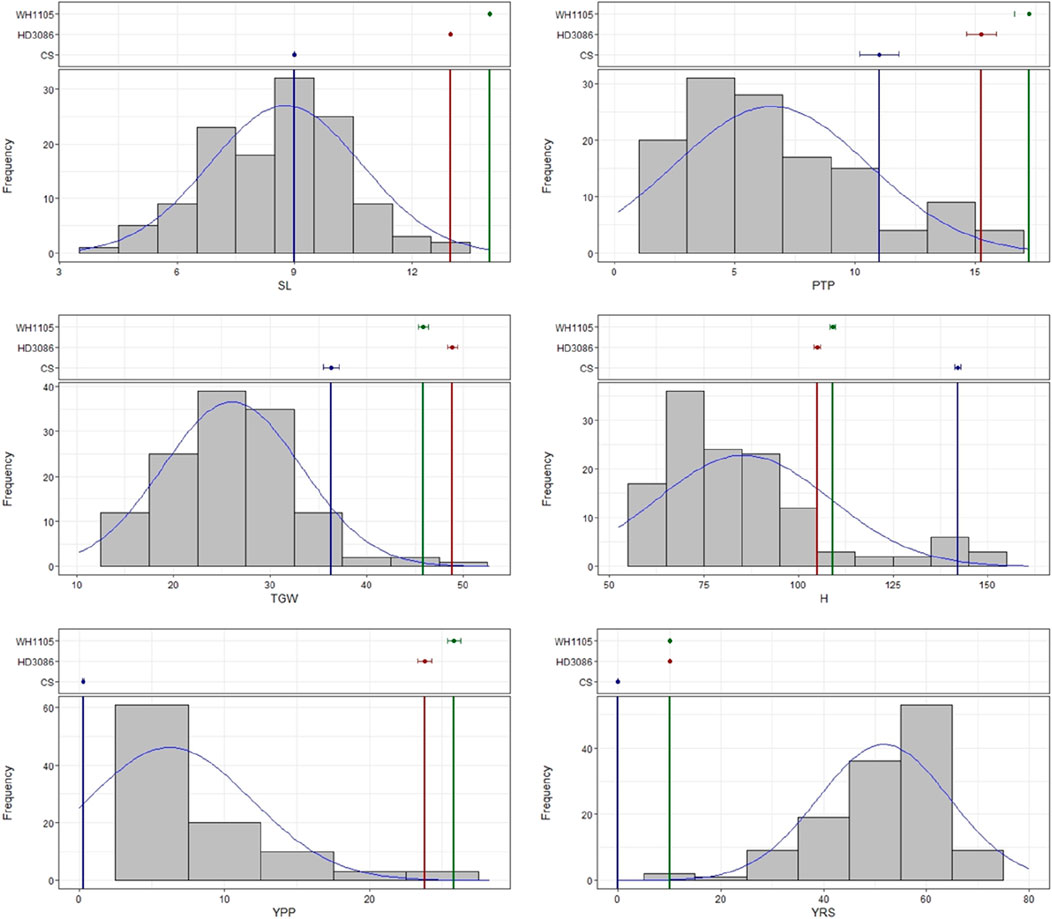
Figure 1. Frequency distributions of the BC2F2:3 population and parents for the traits spike length (SL), productive tillers/plant (PTP), thousand grain weight (TGW), plant height (H), yield per plant (YPP) and yellow rust severity (YRS).
These BC2F2:3 lines were also subjected to meticulous screening, with a focus on various agronomic attributes, including resistance to yellow rust and overall yield potential. During the selection process, desirable parental traits, particularly those associated with high yield, which aim to retain robust yield performance while concurrently inheriting traits related to the β-glucan content in seeds, were also considered. Following a comprehensive disease severity evaluation, the BC2F2:3 lines were given a disease score of 3 according to the modified Cobb scale of 1–5 provided by Peterson et al. (1948), exhibiting a severity percentage of <60%, displaying a moderate level of tolerance to yellow rust, and a moderate to high per plant yield (≥10.0 g) was prioritized for further analysis. Specifically, this subset of plants (Online Resource 3) was chosen for the estimation of grain β-glucan content, as these criteria ensured a balanced combination of disease resistance and favorable yield potential.
The estimation of β-glucan content in seeds was conducted on a selection of 18 BC2F2:3 plants chosen on the basis of their distinct phenotypic traits. The data on the β-glucan concentration in the grains were collected in triplicate, and their means were computed to ensure accuracy. Notably, the majority of the plants subjected to β-glucan content analysis presented higher levels of β-glucan content than did the three parental reference strains, namely, Chinese spring (0.77%), WH1105 (0.75%), and HD3086 (0.71%). Among these analysed lines, 9 lines whose β-glucan content exceeded 0.9% are listed in Table 2. Interestingly, one specific line, designated BC2F2:3 23-5, presented a notably elevated β-glucan content (1.77%), which surpassed the levels observed in both the other lines and the wheat parental references. However, it is important to note that while this content was significantly high, it did not reach the same level as that found in Ae. Kotschyi. Given this remarkable finding, further molecular analysis was initiated for the 9 BC2F2:3 lines whose β-glucan content exceeded 0.9% to investigate potential introgression from Ae. Kotschyi, which might provide insights into the observed high β-glucan content.
The BC2F2:3 lines were investigated because of their suspected involvement in carrying vital segments related to high β-glucan content in the Ae. Kotschyi genome. To explore this further, a comprehensive molecular marker analysis was performed at the whole-genome level. These molecular markers, which spanned all chromosomes, were amplified via DNA isolated from various sources: Chinese spring (CS), Ae. Kotschyi, WH1105, HD3086 and the 9 BC2F2:3 progenies. The amplified products were then meticulously examined via 10% urea-PAGE to ensure the precise resolution of the amplicons. Verification of Ae. Kotschyi genetic introgression into one of the BC2F2:3 lines, namely, 23-5, was achieved through specific molecular markers, namely, gwm111 and barc349. These markers exhibited variations in both Ae. Kotschyi and BC2F2:3 23-5 in comparison with those in CS, WH1105, and HD3086 (see Figures 2a, b). Notably, the marker locations were precisely identified on the 2B and 7D chromosomes. These findings confirmed that the introduction of the group 2 and 7 chromosomes originated from Ae. Kotschyi into the wheat-Ae. Kotschyi derivative line BC2F2:3 23-5. To delve deeper into the analysis, selected markers from both arms of these chromosome centromeric and telomeric regions were utilized. This analysis covered the parent lines as well as the derivative BC2F2:3 23-5. An important observation was made: a specific primer, bglu-7DSt, originating from the telomeric region of the 7D chromosome’s short arm, displayed variations in both Ae. Kotschyi and BC2F2:3 23-5 (Figure 2c). This significant insight further confirmed the introgression of the group 7 chromosome from Ae. Kotschyi (Online resource 4).
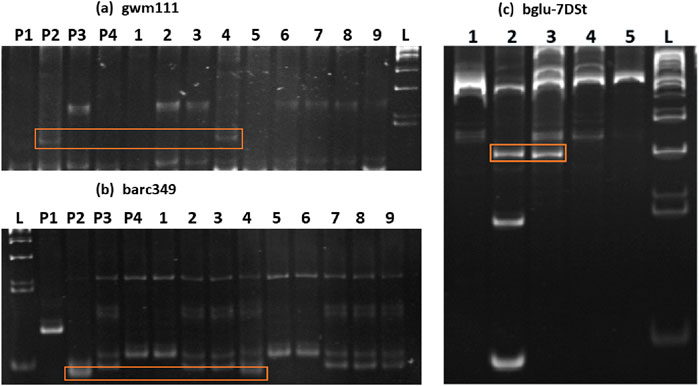
Figure 2. 10% urea-PAGE of markers showing introgression of genomic regions from Ae. Kotschyi to the progeny BC2F2:3 line 23-5. The bands that are specific to Ae. Kotschyi and BC2F2:323-5 are marked. The wells for the SSR markers gwm111 (a) and barc349 (b) were numbered P1- Chinese spring PhI, P2- Ae. Kotschyi, P3- WH1105, P4- HD3086, and 1- (21-4), 2- (22-5), 3- (22-6), 4- (23-5), 5- (25-2), 6- (27-7), 7- (32-11), 8- (32-15), 9- (67-7) and L- 50 bp ladders. The wells for the gene specific marker bglu-7DSt (c) were numbered as follows: 1- Chinese spring PhI, 2- Ae. Kotschyi, 3- BC2F2:323-5, 4- WH1105, 5- HD3086, and L- 100 bp ladder.
To increase our understanding of the wheat-Ae. Kotschyi derivative line BC2F2:3 23-5, in which the integration of group 2 and 7 chromosomes (either 2S, 2U, 7U or 7S) from Ae. Kotschyi was previously validated through molecular marker analysis, we employed genomic in situ hybridization (GISH). The primary goal of GISH is to provide conclusive evidence regarding the genome of the introgressed chromosome. For the GISH experiments, genomic probes specific to the S and U genomes were employed. Notably, the genomic probe derived from Ae. Umbellulata genomic DNA (U-genome) generated a robust hybridization signal on a specific pair of chromosomes of the BC2F2:3 23-5 line (Figure 3). This significant hybridization outcome confirmed the presence of the 7U chromosome from Ae. Kotschyi in the BC2F2:3 23-5 line.
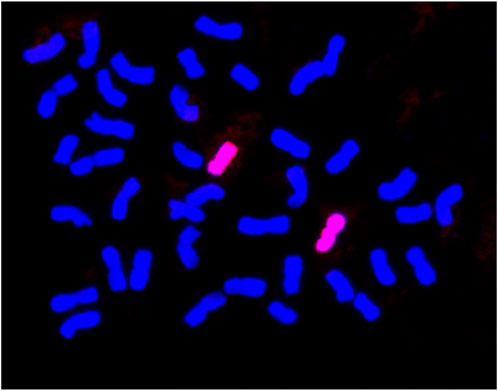
Figure 3. Genomic in situ hybridization of BC2F2:3 23-5 showing introgression from Ae. Kotschyii chromosome 7U.
The majority of cultivated bread wheat (T. aestivum L.), durum wheat (T. turgidum L. ssp. durum), are known to possess relatively low levels of β-glucan in their grain composition. The β-glucan content in bread wheat was less than 1% on a dry weight basis. In contrast, tetraploid wheat genotypes presented a broader range of β-glucan contents, ranging from 0.39% to 0.7% (McCartney et al., 2005). Therefore, in this study, we conducted a thorough investigation into the intricate processes and molecular mechanisms involved in the successful integration of Ae. Kotschyi chromosomes. This integration represents a parallel endeavor aimed at cultivating lines inherently equipped to accumulate elevated levels of β-glucan within wheat grains.
Numerous studies have consistently highlighted that the distant relatives of wheat contain notably greater amounts of β-glucan in their grains than cultivated wheat does. Among these wild relatives of wheat, certain members of the Aegilops species, specifically Ae. Biuncialis, Ae. Umbellulata, and Ae. Neglecta, presented remarkably elevated β-glucan contents, reaching values as high as 7.1% (Marcotuli et al., 2019). Furthermore, research conducted by (Rakszegi et al., 2017) supported this trend, confirming considerably higher β-glucan contents across all accessions of Aegilops sp. In comparison with cultivated wheat genotypes. Specifically, grain β-glucan levels varied within the range of 3%–5% for Ae. Biuncialis and Ae. Geniculate. Intriguingly, introducing chromosome pairs of 5Ug, 5 Mg and 7 Mg from Ae. Geniculate and 5Ub, 5Mb and 7Mb from Ae. Biuncialis into the bread wheat genotype Chinese spring resulted in a noteworthy increase in the β-glucan content (Rakszegi et al., 2017). In this study, we similarly tried to introduce chromosomes from another wild genotype of wheat, Ae. Kotschyi, to improve the β-glucan content in wheat while sustaining the increased productivity of the wheat cultivars.
The findings of this study aligned with earlier reports, where they confirmed the introgression of chromosome 7U from Ae. Kotschyi in one of the parental derivative lines (63-2–13) of BC2F2:3 23-5, which also presented elevated grain Fe and Zn contents (Tiwari et al., 2010). Furthermore, these results are in agreement with those of previous studies (Marcotuli et al., 2017) that identified three significant QTLs for β-glucan content located on chromosomes 2A and 2B. These QTLs were identified through the analysis of recombinant inbred lines (RILs) derived from the Duilio × Avonlea durum wheat cross. Collectively, this body of research emphasizes the likelihood that several genomic regions associated with β-glucan accumulation in wheat grains are predominantly situated within chromosomes 2 and 7. Similarly, a noteworthy QTL on chromosome 7H in barley, which is responsible for 39% of the β-glucan content variation in barley grain (Li et al., 2008), further supports the influence of chromosome seven on the grain β-glucan content. This was evident when the 7H chromosome from barley was introduced into the wheat genome, resulting in a substantial increase in the β-glucan content in wheat grains (Colasuonno et al., 2020). Additionally, Manickavelu et al. (2011) identified β-glucan content QTLs on chromosomes 3A, 1B, 5B, and 6D in wheat via RILs derived from a hybrid between Chinese spring and spelt wheat. Similarly, Marcotuli et al. (2016) employed an array of 230 tetraploid wheat genotypes to predict β-glucan content QTLs. These findings revealed QTLs on chromosomes 1A, 2A, 2B, 5B, and 7A while also identifying a subset of putative genes exerting either direct or indirect influences on the grain β-glucan content. This emphasis on chromosomes 2 and 7 reinforces their pivotal role in β-glucan accumulation, which is consistent with earlier research.
Various yield-related attributes, such as yield per plant, thousand-grain weight, and the number of productive tillers, were considered alongside the parental characteristics of the high-yield parents to ensure that yield levels were maintained while passing through the trait related to the β-glucan content in the seeds. During the BC1F1 generation, most of the plants were tall. However, within the BC2F2 and BC2F2:3 populations, notable variation in plant height was evident. Some plants grew exceptionally tall, even surpassing the height of the Chinese spring parent, whereas a smaller portion of plants presented shorter stature.
Similar diversity was observed for other traits, including resistance to yellow rust, yield per plant, number of productive tillers, and grain β-glucan content. Some plants presented exceptionally high yields and tiller numbers, comparable to or even exceeding those of the high-yield parents HD3086 and WH1105. Conversely, some plants were highly susceptible to yellow rust, in contrast to the parents, whereas others exhibited nearly complete resistance. This wide range of traits among the progeny suggests that these characteristics may be controlled by loci present in the chromosomes introgressed from Aegilops kotschyi (Marais et al., 2005). The study of yellow rust resistance revealed a diverse spectrum of infections, ranging from highly susceptible to highly resistant plants within the population, which primarily consisted of landraces. This diversity underscores the quantitative nature of yellow rust resistance, as elucidated by Vikram et al., 2021. In the case of our BC2F2:3 population, differences among the progenies can be attributed to the segregation of parental genotypes resulting from genetic recombination. Morphological traits, which are inherently quantitative in nature, exhibit variability in their expression due to the combination of different genes and alleles distributed throughout the genome, including the introgressed chromosomes, as observed in our population.
Said et al. (2021) reported that Ae. Umbellulata is composed of two significant translocations originating from the short arm of the group 7 chromosome of wheat to chromosome 7UL, followed by a paracentric inversion. Additionally, the second translocation originated from the wheat group 3 short arm and relocated to the 7UL chromosome. The presence of a QTL on chromosome 3 A in wheat has also been reported (Manickavelu et al., 2011). It is feasible that the translocation of wheat group 3 chromosomes could lead to the duplication of genes involved in β-glucan synthesis. This duplication could contribute to increased β-glucan content in wild wheat relatives containing the 7UL chromosome, such as Ae. Umbellulata, Ae. Kotschyi, and others, including the addition/substitution line generated in this study, namely, 23-5. Importantly, while the U genome of Ae. Kotschyi has experienced significant diversification, it still retains similarities to the D genome of wheat (Kilian et al., 2011). This similarity poses challenges in finding SSR markers that exhibit polymorphism, making it difficult to distinguish between these genomes without relying on highly dense markers.
The newly developed Ae. Kotschyi introgressed line generated in this study carries a significant amount of unwanted linkage drag. This unintended genetic baggage has led to reduced yield and heightened vulnerability to yellow rust disease, possibly stemming from the disruption of genes responsible for yellow rust resistance. To address these issues, this genotype could be employed in future breeding efforts by initiating a backcrossing program with high-yielding and yellow rust-resistant varieties. This process should involve careful consideration of the transfer of specific chromosomal segments, which can be facilitated through techniques such as irradiation and recombination between homeologous chromosome pairs.
In summary, this study successfully demonstrated the introgression of Ae. Kotschyi chromosomes into wheat cultivars, resulting in significantly elevated grain β-glucan contents. Confirmation of this introgressed region was achieved through the use of two SSR markers, gwm111 and barc349, as well as one gene specific marker, bglu-7DSt. Notably, these markers are located on chromosomes two and seven of the wheat genome and are known to harbor QTLs related to the grain β-glucan content. The findings of this research offer valuable insights into the mechanism governing β-glucan synthesis and deposition in wheat grains. This knowledge can serve as a foundation for further analyses and predictions in this field. Moreover, the information presented in this study can be harnessed for future breeding programs aimed at developing wheat varieties with high β-glucan contents. These programs could involve backcrossing the lines established in this study with high-yielding and rust-resistant cultivars, paving the way for the creation of improved wheat varieties with enhanced nutritional profiles.
The original contributions presented in the study are included in the article/Supplementary Material, further inquiries can be directed to the corresponding author.
UK: Conceptualization, Data curation, Formal Analysis, Funding acquisition, Investigation, Methodology, Project administration, Resources, Software, Supervision, Validation, Visualization, Writing–original draft, Writing–review and editing. SP: Formal Analysis, Methodology, Validation, Writing–original draft, Writing–review and editing. RG: Data curation, Formal Analysis, Writing–original draft, Writing–review and editing. YS: Data curation, Formal Analysis, Methodology, Resources, Software, Writing–review and editing. PB: Formal Analysis, Methodology, Resources, Software, Writing–review and editing. PK: Formal Analysis, Methodology, Software, Writing–review and editing. SK: Data curation, Formal Analysis, Methodology, Resources, Software, Writing–review and editing. KS: Formal Analysis, Software, Visualization, Writing–review and editing. FJ: Formal Analysis, Methodology, Writing–review and editing. RM: Formal Analysis, Software, Supervision, Validation, Writing–review and editing.
The author(s) declare that financial support was received for the research and/or publication of this article. This work was supported by the Department of Biotechnology, Govt. Of India as SRF to SP[Fellow ID: DBT/2020/CCSHAU/1,479].
The authors are thankful to the Head of the Department of Molecular Biology and Biotechnology and Dean, College of Biotechnology, and the Director of Research, CCS Haryana Agricultural University, Hisar, India, for providing the necessary facilities.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declared that they were an editorial board member of Frontiers, at the time of submission. This had no impact on the peer review process and the final decision.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fgene.2025.1532956/full#supplementary-material
Beresford, G., and Stone, B. A. (1983). (1→3), (1→4)-β-D-glucan content of Triticum grains. J. Cereal Sci. 1, 111–114. doi:10.1016/S0733-5210(83)80028-9
Colasuonno, P., Marcotuli, I., Cutillo, S., Simeone, R., Blanco, A., and Gadaleta, A. (2020). Effect of barley chromosomes on the β-glucan content of wheat. Genet. Resour. Crop Evol. 67, 561–567. doi:10.1007/s10722-019-00829-y
Collins, H. M., Burton, R. A., Topping, D. L., Liao, M., Bacic, A., and Fincher, G. B. (2010). REVIEW: variability in fine structures of noncellulosic cell wall polysaccharides from cereal grains: potential importance in human health and nutrition. Cereal Chem. 87, 272–282. doi:10.1094/CCHEM-87-4-0272
Cseh, A., Kruppa, K., Molnár, I., Rakszegi, M., Doležel, J., and Molnár-Láng, M. (2011). Characterization of a new 4BS.7HL wheat-barley translocation line using GISH, FISH, and SSR markers and its effect on the β-glucan content of wheat. Genome 54, 795–804. doi:10.1139/g11-044
Cseh, A., Soõs, V., Rakszegi, M., Tūrkösi, E., Balázs, E., and Molnár-Láng, M. (2013). Expression of HvCslF9 and HvCslF6 barley genes in the genetic background of wheat and their influence on the wheat β-glucan content. Ann. Appl. Biol. 163, 142–150. doi:10.1111/AAB.12043
Cui, S., and Liu, R. H. (2013). “Health benefits of oat phytochemicals,” in Oats nutrition and technology (John Wiley and Sons, Ltd), 171–194.
Danilova, T. V., Friebe, B., Gill, B. S., Poland, J., and Jackson, E. (2018). Development of a complete set of wheat–barley group-7 Robertsonian translocation chromosomes conferring an increased content of β-glucan. Theor. Appl. Genet. 131, 377–388. doi:10.1007/s00122-017-3008-z
FAO, (2020). Food and Agriculture Organisation. Available at: https://www.fao.org/home/en)
Friebe, B., Jiang, J., Raupp, W. J., McIntosh, R. A., and Gill, B. S. (1996). Characterization of wheat-alien translocations conferring resistance to diseases and pests: current status. Euphytica 91, 59–87. doi:10.1007/bf00035277
Friebe, B., Qi, L. L., Nasuda, S., Zhang, P., Tuleen, N. A., and Gill, B. S. (2000). Development of a complete set of Triticum aestivum-Aegilops speltoides chromosome addition lines. Theor. Appl. Genet. 101, 51–58. doi:10.1007/s001220051448
Havrlentová, M., and Kraic, J. Á. N. (2006). Content of β-D-glucan in cereal grains. J. Food Nutr. Res. 45, 97–103.
Kilian, B., Mammen, K., Millet, E., Sharma, R., Graner, A., Salamini, F., et al. (2011). “Aegilops,” in Wild crop relatives: genomic and breeding resources. Editor Kole, C. (Berlin, Heidelberg: Springer), 1–76. doi:10.1007/978-3-642-14228-4_1
Kuraparthy, V., Chhuneja, P., Dhaliwal, H. S., Kaur, S., Bowden, R. L., and Gill, B. S. (2007a). Characterization and mapping of cryptic alien introgression from Aegilops geniculata with new leaf rust and stripe rust resistance genes Lr57 and Yr40 in wheat. Theor. Appl. Genet. 114, 1379–1389. doi:10.1007/s00122-007-0524-2
Kuraparthy, V., Sood, S., Chhuneja, P., Dhaliwal, H. S., Kaur, S., Bowden, R. L., et al. (2007b). A cryptic wheat–Aegilops triuncialis translocation with leaf rust resistance gene Lr58. Crop Sci. 47, 1995–2003. doi:10.2135/CROPSCI2007.01.0038
Li, J., Båga, M., Rossnagel, B. G., Legge, W. G., and Chibbar, R. N. (2008). Identification of quantitative trait loci for β-glucan concentration in barley grain. J. Cereal Sci. 48:647–655. doi:10.1016/j.jcs.2008.02.004
Manickavelu, A., Kawaura, K., Imamura, H., Mori, M., and Ogihara, Y. (2011). Molecular mapping of quantitative trait loci for domestication traits and β-glucan content in a wheat recombinant inbred line population. Euphytica 177, 179–190. doi:10.1007/s10681-010-0217-9
Marais, G. F., McCallum, B., Snyman, J. E., Pretorius, Z. A., and Marais, A. S. (2005). Leaf rust and stripe rust resistance genes Lr54 and Yr37 transferred to wheat from Aegilops kotschyi. Plant Breed. 124, 538–541. doi:10.1111/J.1439-0523.2005.01116.X
Marcotuli, I., Colasuonno, P., Cutillo, S., Simeone, R., Blanco, A., and Gadaleta, A. (2019). β-glucan content in a panel of Triticum and Aegilops genotypes. Genet. Resour. Crop Evol. 66, 897–907. doi:10.1007/s10722-019-00753-1
Marcotuli, I., Gadaleta, A., Mangini, G., Signorile, A. M., Zacheo, S. A., Blanco, A., et al. (2017). Development of a high-density SNP-based linkage map and detection of QTL for β-glucans, protein content, grain yield per spike and heading time in durum wheat. Int. J. Mol. Sci. 18, 1329. doi:10.3390/IJMS18061329
Marcotuli, I., Houston, K., Schwerdt, J. G., Waugh, R., Fincher, G. B., Burton, R. A., et al. (2016). Genetic diversity and genome wide association study of β-glucan content in tetraploid wheat grains. PLoS One 11, e0152590. doi:10.1371/journal.pone.0152590
McCartney, C. A., Somers, D. J., Humphreys, D. G., Lukow, O., Ames, N., Noll, J., et al. (2005). Mapping quantitative trait loci controlling agronomic traits in the spring wheat cross RL4452 x “AC Domain.”. Genome 48, 870–883. doi:10.1139/G05-055
Mohebbi, Z., Homayouni, A., Azizi, M. H., and Hosseini, S. J. (2018). Effects of beta-glucan and resistant starch on wheat dough and prebiotic bread properties. J. Food Sci. Technol. 55, 101–110. doi:10.1007/s13197-017-2836-9
Murray, M. G., and Thompson, W. F. (1980). Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res. 8, 4321–4325. doi:10.1093/NAR/8.19.4321
Pestsova, E., Ganal, M. W., and Roder, M. S. (2000). Isolation and mapping of microsatellite markers specific for the D genome of bread wheat. Genome 43, 689–697. doi:10.1139/g00-042
Peterson, R. F., Campbell, A. B., and Hannah, A. E. (1948). A diagrammatic scale for estimating rust intensity on leaves and stems of cereals. Can. J. Res. 26c, 496–500. doi:10.1139/cjr48c-033
Pritchard, J. R., Lawrence, G. J., Larroque, O., Li, Z., Laidlaw, H. K., Morell, M. K., et al. (2011). A survey of β-glucan and arabinoxylan content in wheat. J. Sci. Food Agric. 91, 1298–1303. doi:10.1002/JSFA.4316
Qi, L., Friebe, B., Zhang, P., and Gill, B. S. (2007). Homoeologous recombination, chromosome engineering and crop improvement. Chromosome Res. 15, 3–19. doi:10.1007/s10577-006-1108-8
Rakszegi, M., Molnár, I., Lovegrove, A., Darkó, É., Farkas, A., Láng, L., et al. (2017). Addition of aegilops U and M chromosomes affects protein and dietary fiber content of wholemeal wheat flour. Front. Plant Sci. 8, 1529. doi:10.3389/fpls.2017.01529
Reynolds, A., Mann, J., Cummings, J., Winter, N., Mete, E., and Te Morenga, L. (2019). Carbohydrate quality and human health: a series of systematic reviews and meta-analyses. Lancet 393, 434–445. doi:10.1016/S0140-6736(18)31809-9
Röder, M. S., Korzun, V., Wendehake, K., Plaschke, J., Tixier, M. H., Leroy, P., et al. (1998). A microsatellite map of wheat. Genetics 149, 2007–2023. doi:10.1093/GENETICS/149.4.2007
Said, M., Holušová, K., Farkas, A., Ivanizs, L., Gaál, E., Cápal, P., et al. (2021). Development of DNA markers from physically mapped loci in Aegilops comosa and Aegilops umbellulata using single-gene FISH and chromosome sequences. Front. Plant Sci. 12, 689031–689117. doi:10.3389/fpls.2021.689031
Schneider, A., Molnár, I., and Molnár-Láng, M. (2008). Utilisation of Aegilops (goatgrass) species to widen the genetic diversity of cultivated wheat. Euphytica 163, 1–19. doi:10.1007/s10681-007-9624-y
Sibakov, J., Lehtinen, P., and Poutanen, K. (2013). “Cereal brans as dietary fibre ingredients,” in Fibre-rich and wholegrain foods: improving quality (Cambridge:Woodhead Publishing Ltd), 170–192.
Somers, D. J., Isaac, P., and Edwards, K. (2004). A high-density microsatellite consensus map for bread wheat (Triticum aestivum L.). Theor. Appl. Genet. 109, 1105–1114. doi:10.1007/s00122-004-1740-7
Tiwari, V. K., Rawat, N., Neelam, K., Kumar, S., Randhawa, G. S., and Dhaliwal, H. S. (2010). Substitutions of 2S and 7U chromosomes of Aegilops kotschyi in wheat enhance grain iron and zinc concentration. Theor. Appl. Genet. 121, 259–269. doi:10.1007/s00122-010-1307-8
Türkösi, E., Darko, E., Rakszegi, M., Molnár, I., Molnár-Láng, M., and Cseh, A. (2018). Development of a new 7BS.7HL winter wheat-winter barley Robertsonian translocation line conferring increased salt tolerance and (1,3;1,4)-β-D-glucan content. PLoS One 13, e0206248. doi:10.1371/JOURNAL.PONE.0206248
Vikram, P., Sehgal, D., Sharma, A., Bhavani, S., Gupta, P., Randhawa, M., et al. (2021). Genome-wide association analysis of Mexican bread wheat landraces for resistance to yellow and stem rust. PLoS One 16, e0246015. doi:10.1371/journal.pone.0246015
Wood, P. J. (2007). Cereal β-glucans in diet and health. J. Cereal Sci. 46, 230–238. doi:10.1016/j.jcs.2007.06.012
Keywords: β-glucan, introgression, Aegilops kotschyi, in situ hybridization, SSR, backcross
Citation: Kumar U, Panigrahi S, Goswami R, Singh Y, Balyan P, Kapoor P, Kumar S, Singh KP, Jan F and Mir RR (2025) Enhancing wheat β-glucan content through precision crossbreeding: development and evaluation of biofortified lines with improved nutritional and agronomic traits. Front. Genet. 16:1532956. doi: 10.3389/fgene.2025.1532956
Received: 22 November 2024; Accepted: 28 February 2025;
Published: 19 March 2025.
Edited by:
Luis Herrera-Estrella, Texas Tech University, United StatesReviewed by:
Vikas Sharma, Sant Baba Bhag Singh University, IndiaCopyright © 2025 Kumar, Panigrahi, Goswami, Singh, Balyan, Kapoor, Kumar, Singh, Jan and Mir. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Upendra Kumar, YmFsaXlhbi51cGVuZHJhQGdtYWlsLmNvbQ==
†Present address: Yogita Singh, Stockbridge School of Agriculture, University of Massachusetts, Amherst, MA, United States
‡These authors have contributed equally to this work
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.