
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Cell. Infect. Microbiol. , 01 April 2025
Sec. Veterinary and Zoonotic Infection
Volume 15 - 2025 | https://doi.org/10.3389/fcimb.2025.1575365
This article is part of the Research Topic Unveiling Host-Pathogen Interactions: Insights into Animal Cellular Immunity and Novel Diagnostics - Volume II View all articles
Introduction: Streptococcus suis (S. suis) and Glaesserella parasuis (G. parasuis) are prevalent pathogens in pig populations and are often associated with co-infections, leading to substantial economic losses in the swine industry. However, there is currently a shortage of rapid detection methods. In this study, a dual loop-mediated isothermal amplification combined with lateral flow dipstick (LAMP-LFD) assay was developed for the simultaneous and convenient detection of S. suis and G. parasuis.
Methods: The assay utilized primers targeting the conserved regions of the gdh gene of S. suis and the infB gene of G. parasuis. Optimal primer sets were identified, and reaction conditions, including temperature, time, and primer concentration ratios, were optimized using single-variable control method. The LAMP-LFD assay was established with biotin and digoxin or biotin and 6-FAM-labeled FIP/BIP primers, combined with LFD.
Results: The assay was most effective at a reaction temperature of 62°C, a primer concentration ratio of 1:4, and a reaction time of 40 minutes. The minimum detection limits were 22 and 18 copies/μL for recombinant plasmids and 19 and 20 CFU for bacterial samples of S. suis and G. parasuis, respectively. The assay showed no cross-reactivity with other pathogens and exhibited high adaptability across various thermal platforms, including PCR instruments, metal baths, and water baths. Clinical testing of 106 samples revealed positive rates of 11.32% (12/106) for S. suis, 25.47% (27/106) for G. parasuis, and 2.83% (3/106) for mixed infections.
Discussion: This simple, rapid, specific, and sensitive dual LAMP-LFD assay provides robust technical support for the prevention and control of swine streptococcosis and Glässer's disease.
Streptococcus suis (S. suis, SS) and Glaesserella parasuis (G. parasuis, GPS) are significant and prevalent pathogens in swine, causing similar clinical symptoms and pathological lesion (Guo et al., 2024; Santoya et al., 2024). Both pathogens can lead to septicemia, polyserositis, meningitis, arthritis, and pneumonia (Jiang et al., 2024; Yue et al., 2024). S. suis is classified into 35 serotypes based on capsular polysaccharides, with serotypes 1, 2, 1/2, 7, 9, and 14 being particularly pathogenic in swine, and serotype 2 being the most widespread and severe (Fan et al., 2024; Petrocchi et al., 2024; Xia et al., 2024). In contrast, G. parasuis has 15 serotypes, with serotypes 4, 5, and 12 most frequently isolated from clinical cases, though serotypes 1, 2, 6, 7, 9, and 14 have also been reported (Schuwerk et al., 2020; Wu et al., 2023; Yan et al., 2023). Cross-protection between serotypes is limited (Bujold et al., 2023). Co-infections of S. suis and G. parasuis are common and pose a significant clinical challenge due to overlapping symptoms. In recent years, S. suis and G. parasuis infections have been widely prevalent in the global swine industry, emerging as major pathogens that severely impact pig health. In high-density pig farming regions of Italy, S. suis and G. parasuis were responsible for 18.0% and 20.3% of bacterial arthritis cases in weaned piglets, respectively (Salogni et al., 2022). Epidemiological studies conducted in Austria (2016–2021) and the United States (2017–2022) reported S. suis and G. parasuis co-infection rates of 3% and 5%, respectively (Renzhammer et al., 2023; Silva et al., 2023). In contrast, the prevalence in China appears to be higher. A study in the eastern region of China revealed S. suis and G. parasuis detection rates of 52.3% and 33.2%, with a co-infection rate of 33.2% (Zhu et al., 2021). Additionally, a study in Heilongjiang province using multiplex real-time quantitative PCR found that S. suis serotype 2 and G. parasuis detection rates exceeded 60%, with co-infection rates ranging from 78% to 96%. However, the limited sample size may not accurately reflect the broader epidemiological situation (Li et al., 2024). These findings indicate that S. suis and G. parasuis co-infections are common in the swine industry, highlighting the critical need to develop rapid, sensitive, and specific diagnostic methods for effective disease prevention and control.
Current diagnostic methods for S. suis and G. parasuis include bacterial isolation, molecular biology techniques, and serological assays (Nedbalcova et al., 2022; de Jong et al., 2023). However, bacterial isolation is labor-intensive, time-consuming, and less practical in clinical settings (Scherrer et al., 2024). Molecular techniques such as conventional PCR and quantitative PCR provide high specificity and sensitivity but require expensive equipment and skilled personnel, limiting their application in field conditions (Goto et al., 2023; Wang et al., 2024). Serological assays, including agglutination tests and ELISA, are hindered by high material costs and their inability to simultaneously detect both pathogens (Guo et al., 2010; Chidkoksung et al., 2024).
Compared to conventional PCR and qPCR methods, LAMP technology does not require expensive thermal cyclers, offering lower costs and shorter detection times, while maintaining high sensitivity and strong specificity. Particularly, when combined with LFD, LAMP enables rapid on-site detection, facilitating timely intervention and disease control. This makes the LAMP-LFD approach highly promising for applications in disease diagnostics (Pilchova et al., 2020; Bai et al., 2024). To date, no dual LAMP-LFD detection method has been developed for S. suis and G. parasuis. This study aims to establish a dual LAMP-LFD assay for the simultaneous detection of these pathogens in pigs, improving diagnostic accuracy, facilitating epidemiological investigations, and enhancing the prevention and control of swine streptococcosis and Glässer’s disease.
S. suis serotype 1 CVCC2937 strain, serotype 2 CVCC9740 strain, serotype 7 CVCC563 strain, serotype 9 CVCC989 strain, serotype 14 CVCC212 strain, serotype 16 CVCC223 strain, and Mycoplasma hyopneumoniae CVCC679, and G. parasuis serotype 4 CVCC156 strain, serotype 5 CVCC167 strain, serotype 12 CVCC134 strain, and Enterococcus faecalis CVCC1927 strain, Streptococcus agalactiae CVCC586 strain, Pasteurella multocida CVCC390 strain, Actinobacillus pleuropneumoniae CVCC259 strain, Streptococcus pyogenes CVCC593 strain, and Streptococcus pneumoniae CVCC1929 strain were stored and supplied by China Institute of Veterinary Drug Control. From July 2023 to December 2024, 106 clinical samples (pleural effusion, lung tissue, nasal swabs) with respiratory disease and arthritis symptoms were collected from pig farms in Henan province.
Primers targeting the conserved regions of the S. suis gdh gene (AM946016.1) and the G. parasuis infB gene (CP071489.1) were designed following the principles of LAMP primer design by using Primer Explorer V5 software (http://primerexplorer.jp/lampv5e/index.html) (Kirkoyun et al., 2024). The fluorescent dye method was employed to identify the most specific and effective primer sets.
The LAMP reaction was performed in a 25 μL system comprising 5 μL of 5 × LAMP Reaction Mix, 2 μL of Bst II DNA polymerase (Harbin Tianyuehao Biotechnology Co., Ltd), 0.45 μL of TS LAMP Green (20×), 1.6 μM of FIP/BIP, 0.4 μM of B3/F3, 2 μL of DNA template, and ddH2O to a final volume of 25 μL. The reaction was carried out at 62°C for 40 cycles, with fluorescence signals recorded every minute. Two optimal primer pairs were selected and labeled at the 5’ ends with Biotin and 6-FAM, and Biotin and Digoxigenin, respectively (Table 1). Additionally, primers for standard plasmids were designed using Primer Premier 5 software (Table 1). All primers, including the labeled ones, were synthesized by Beijing Liuhehuada Gene Technology Co., Ltd.
Genomic DNA from S. suis and G. parasuis was used as a template to design two pairs of specific primers for single PCR amplification. The PCR products were analyzed by 1% agarose gel electrophoresis, purified using the EasyPure® Quick Gel Extraction Kit (TransGen Biotech), and subsequently cloned into the pMD18-T vector to construct recombinant plasmids, designated as pMD-SS and pMD-GPS. The recombinant plasmids were verified by PCR and sequencing. Plasmid extraction was performed using a plasmid extraction kit, and the plasmid concentrations were accurately determined. The number of plasmid copies was calculated using the following equation: (Plasmid copies/µL = (6.02×1023) × [X* ng/µL×10−9)/constructed plasmid length (bp) × 660] (Wang et al., 2023).
X* means recombinant plasmid concentration.
The single-variable control method was employed to optimize reaction conditions, including temperature, time, and primer concentration ratio. Reaction temperatures were tested at 60.0, 60.3, 61.0, 62.0, 63.2, 64.2, 64.7, and 65.0°C. Reaction times were evaluated at 20, 25, 30, 35, 40, and 45 minutes. Primer concentration ratios for internal to external primers (F3:FIP and B3:BIP) were assessed at 1:1, 1:2, 1:4, 1:6, 1:8, and 1:10. After each reaction, the products were analyzed by 1.5% agarose gel electrophoresis to identify and confirm the optimal conditions.
The nucleic acids of S. suis serotype 1, 2, 7, 9, 14, 16, Mycoplasma hyopneumoniae, G. parasuis serotype 4, serotype 5, serotype 12, Enterococcus faecalis, Streptococcus agalactiae, Pasteurella multocida, Actinobacillus pleuropneumoniae, Streptococcus pyogenes, and Streptococcus pneumoniae were used as templates for dual LAMP-LFD detection. Taking the mixture of recombinant plasmids pMD-SS and pMD-GPS as positive control, the specificity of the dual LAMP-LFD method was verified.
The two positive plasmids (pMD-SS: 2.2×1010 copies/μL; pMD-GPS: 1.8×1010 copies/μL) from 1010 copies/μL to 100 copy/μL, and the minimum copy number of the plasmids was determined by selecting 106-100 copies/μL for the double LAMP-LFD assay.
S. suis was cultured in 10 mL of tryptic soy broth (TSB) supplemented with 5% neonatal bovine serum, while G. parasuis was cultured in 10 mL of TSB supplemented with 5% neonatal bovine serum and 0.1% NAD. Both bacterial cultures were incubated at 37°C with shaking at 180 rpm for 12 hours and then serially diluted up to 10-7. From each dilution (10-5, 10-6, 10-7), 100 μL of the bacterial suspension was plated onto tryptic soy agar (TSA) containing 5% neonatal bovine serum and 0.1% NAD. The bacterial suspensions were evenly spread using a sterile spreader, and colony counts were recorded after 12 hours of incubation at 37°C. This procedure was performed in triplicate. Additionally, 0.25 g of healthy porcine lung tissue was homogenized, and 100 μL of the diluted bacterial suspension was added to the homogenate. Genomic DNA was extracted using a commercial DNA extraction kit and subsequently analyzed using the dual LAMP-LFD assay.
Three batches of diagnostic reagents were prepared to evaluate inter- and intra-batch sensitivity, specificity, and reproducibility. Thermostatic reactions were performed using these reagents across three platforms: a PCR instrument, a metal bath, and a water bath, to assess the compatibility and suitability of each instrument for the assays.
106 clinical samples, including pleural effusion, lung tissue, nasal swabs, and others collected from pigs, were tested using the LAMP-LFD method. These samples were also tested using an association standard (GB/T 19915.5—2005, Protocol of multiplex PCR identification of Streptococcus suis type 2; GB/T 34750—2017, Detection methods for haemophilus parasuis) to validate the feasibility of the LAMP-LFD method (Xin et al., 2023).
S. suis and G. parasuis genomic DNA were used as templates for PCR amplification with two pairs of specific primers designed for each pathogen, respectively. The PCR products were then ligated into the pMD18-T vector to generate the recombinant plasmids pMD-SS or pMD-GPS. These plasmids were verified by PCR amplification and DNA sequencing, confirming that the amplified target fragments matched the expected sequences (Supplementary Figure S1). The concentrations of the recombinant plasmids were determined to be 97.42 ng/μL and 79.96 ng/μL, corresponding to 2.2 × 1010 copies/μL and 1.8 × 1010 copies/μL, respectively. The plasmids were aliquoted and stored at -20°C for future use.
Each of the two sets of LAMP primers designed in this study was tested separately using a fluorescence quantitative PCR instrument. The results demonstrated that all four primer sets produced positive amplification curves, while negative controls showed no amplification (Figure 1). Under the same conditions, primer set 1 for S. suis (Figure 1A) and primer set 1 for G. parasuis (Figure 1B) exhibited smoother curves with lower Ct values and higher amplification efficiency. Consequently, S. suis primer set 1 and G. parasuis primer set 1 were selected for use in the dual LAMP-LFD assay (Table 1).
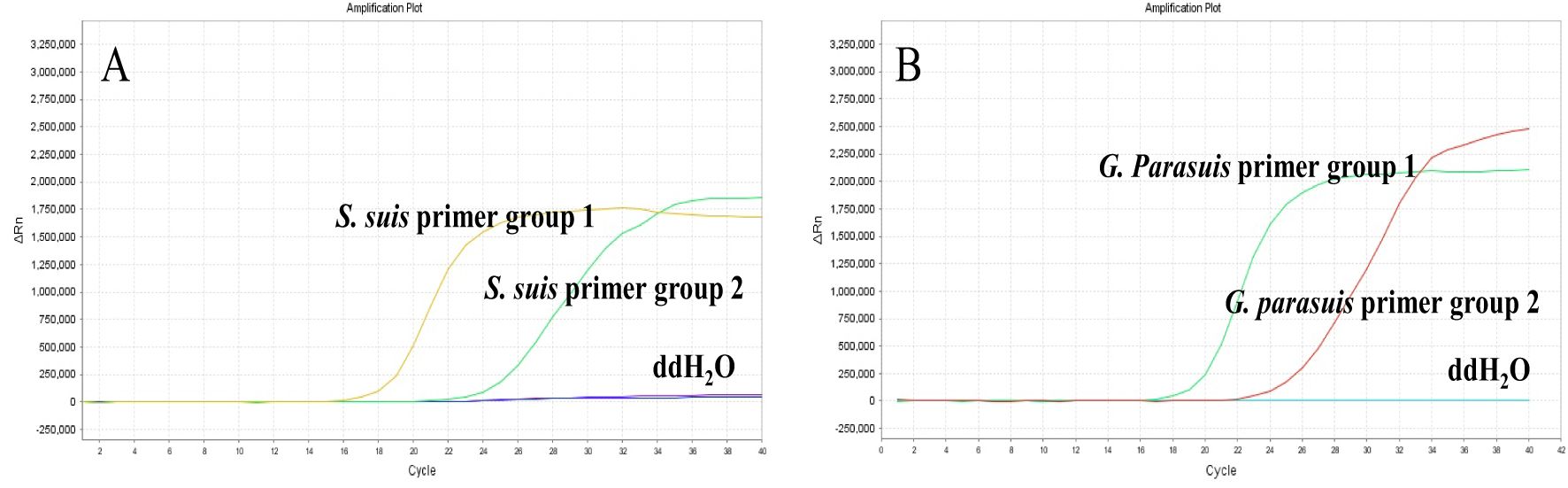
Figure 1. S. suis and G. parasuis LAMP primer screening. (A) the amplification curves of S. suis based on primers set 1 and set 2; (B) the amplification curves of G. parasuis based on primers set 1 and set 2.
The reaction temperature, reaction time, and primer concentration ratio were optimized using the single control variable method. The results indicated that at a reaction temperature of 62°C, the LAMP amplification products consistently exhibited clear, typical trapezoidal bands (Figures 2A, B). When the reaction time was 40 minutes or longer, the LAMP amplification products became stable, with clear bands observed (Figures 2C, D). Additionally, a primer concentration ratio of 1:4 yielded the most effective and clearly defined LAMP amplification bands (Figures 2E, F). The LAMP reaction system (25 μL) includes: 5× LAMP Reaction Mix 5 μL, Bst II DNA Polymerase 2 μL, SS-F3 (10 μM) 1 μL, SS-B3 (10 μM) 1 μL, SS-FIP (100 μM) 0.4 μL, SS-BIP (10 μM) 0.4 μL, GPS-F3 (10 μM) 1 μL, GPS-B3 (10 μM) 1 μL, GPS-FIP (100 μM) 0.4 μL, GPS-BIP (10 μM) 0.4 μL, Template DNA 2 μL, ddH2O 10.4 μL. The reaction was carried out at a constant temperature of 62°C for 40 minutes. After the reaction, 5-10 μL of the LAMP amplification product was diluted 20 times with ddH2O, mixed well, and 80 μL of the diluted reaction product was dropped onto the sample hole. The results in the detection zone were recorded within 15 minutes. The color changes in the T line and C line of the test strip were observed (Figure 3; Table 2).

Figure 2. Optimization of S. suis (A, C, E) and G. parasuis (B, D, F) LAMP reaction conditions. M: DL 2000 DNA Marker; (A) and (B) 1-8: 60, 60.3, 61, 62, 63.2, 64.2, 64.7, 65°C; 9: ddH2O; C and D: 1-6: 20, 25, 30, 35, 40, 45min; 7: ddH2O; (C) and (D) 1-6: 1:1, 1:2, 1:4, 1:6, 1:8, 1:10.
Genomic DNA of S. suis seotypes 1, 2, 7, 9, 14, and 16; Mycoplasma hyopneumoniae; G. parasuis types 4, 5, and 12; Enterococcus faecalis; Streptococcus agalactiae; Pasteurella multocida; Actinobacillus pleuropneumoniae; Streptococcus pyogenes and Streptococcus pneumoniae were tested using the established dual LAMP-LFD method. The results indicated only the S. suis and G. parasuis were detectable, with no cross-reactivity observed with any of the other porcine pathogens (Figure 4), suggesting the method is high specificity.
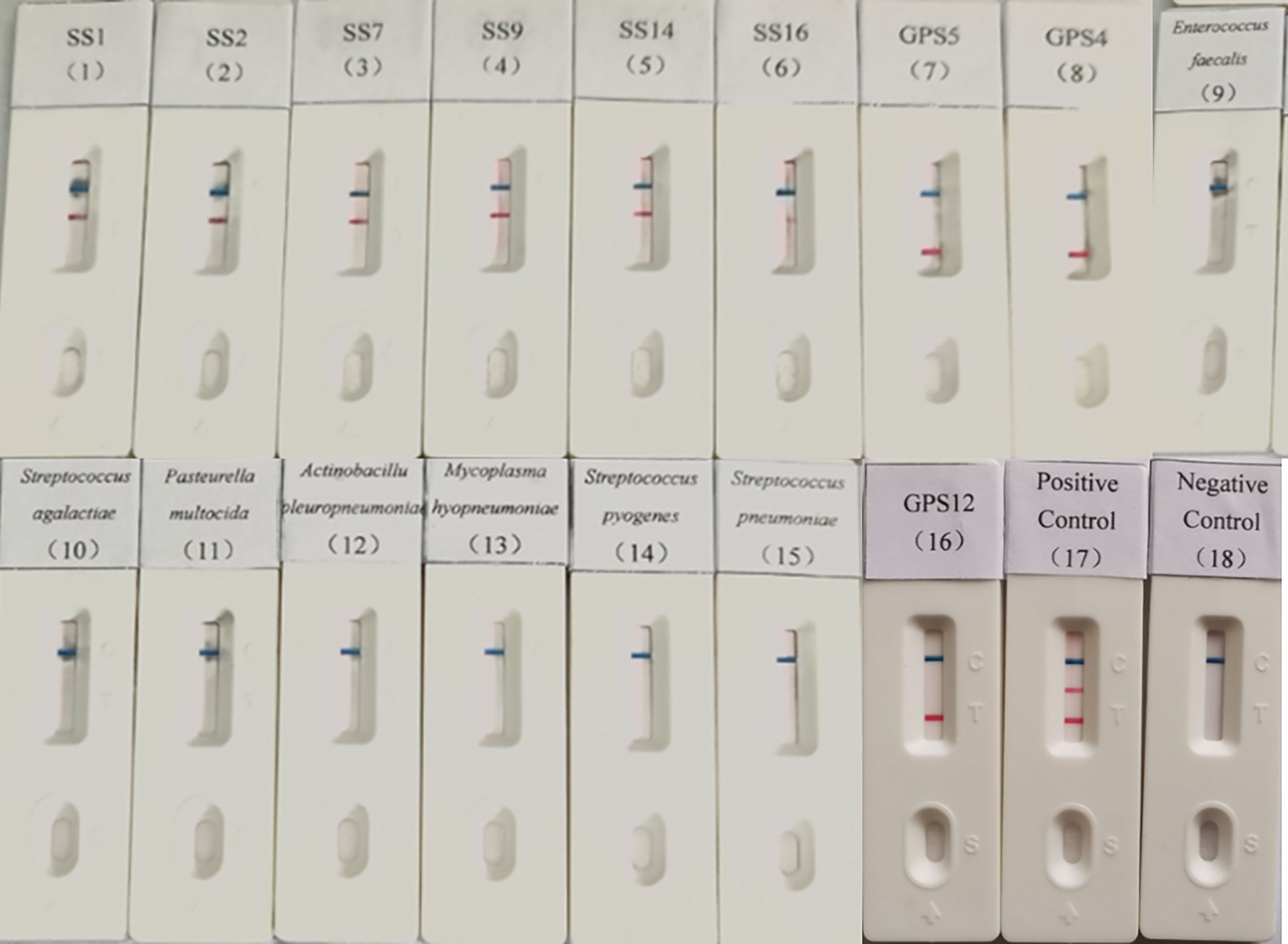
Figure 4. Specificity validation of dual LAMP-LFDs. 1: SS1; 2: SS2; 3: SS7; 4: SS9; 5: SS14; 6: SS16; 7: GPS5; 8: GPS4; 9: Enterococcus faecalis; 10: Streptococcus agalactiae; 11: Pasteurella multocida; 12: Actinobacillus pleuropneumoniae; 13: Mycoplasma hyopneumoniae; 14: Streptococcus pyogenes; 15: Streptococcus pneumoniae; 16: GPS12; 17: Positive control; 18: Negative control.
The dual LAMP-LFD assay was performed using plasmid concentrations ranging from 106 copies/μL to 100 copy/μL as templates to assess the minimum detectable amount of plasmid standards. The results showed that the lowest detectable copy numbers of S. suis and G. parasuis by the double LAMP-LFD method were 22 and 18 copies/μL, respectively (Figure 5). Bacterial counts from cultured samples were determined, revealing concentrations of 1.9 × 10⁸ CFU/mL for S. suis and 6 × 10⁸ CFU/mL for G. parasuis (Supplementary Table S1). The results of the dual LAMP-LFD assay demonstrated that the lowest detectable numbers of bacteria were 19 CFU/mL for S. suis and 20 CFU/mL for G. parasuis (Figure 6).
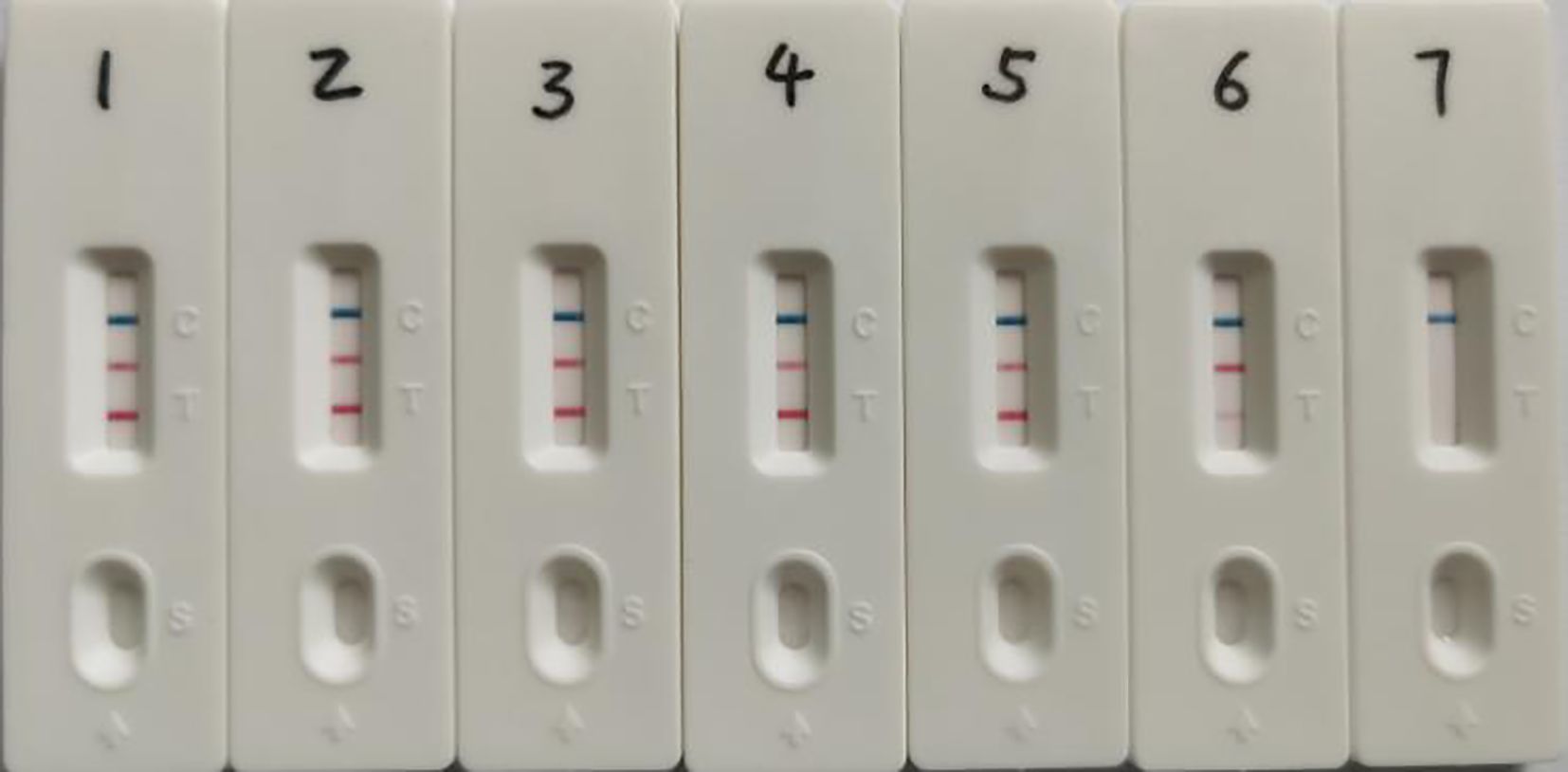
Figure 5. Recombinant plasmid standards as templates for dual LAMP-LFD sensitivity assay. 1-7: pMD-SS concentrations from 2.2×106-2.2×100 copies/μL, pMD-GPS concentrations from 1.8×106-1.8×100 copies/μL.
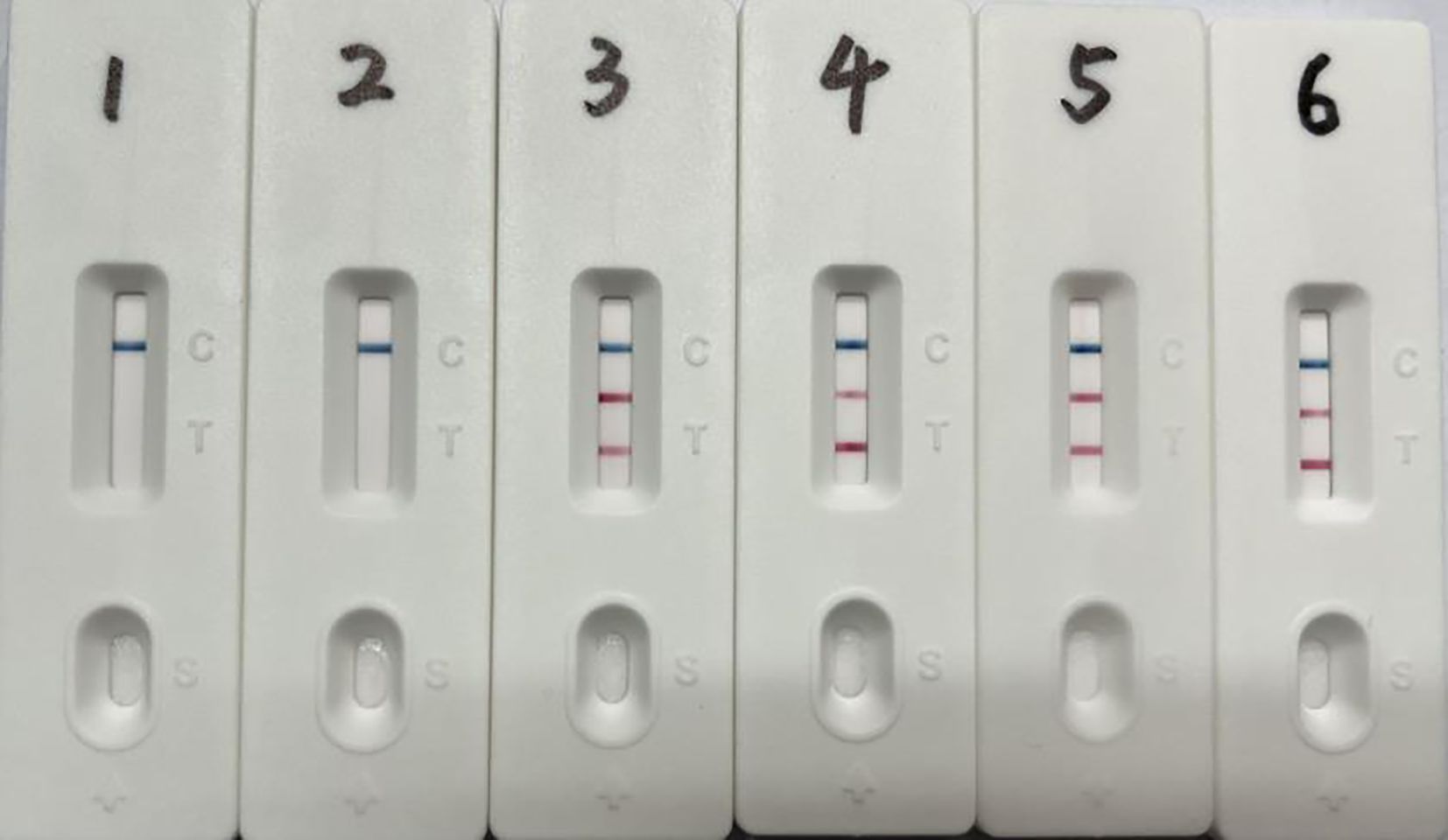
Figure 6. Dual LAMP-LFD detection of simulated tissue samples. 1: Pig lung tissue free of S. suis and G. parasuis; 2-6: S. suis 19 to 1.9 × 105 CFU and G. parasuis 20 to 2.0 × 105 CFU.
Three batches of diagnostic reagents were prepared and tested to assess both inter-batch and intra-batch reproducibility. Reactions using the same batch of reagents were performed on a PCR instrument, metal bath, and water bath at a temperature of 62°C to evaluate the performance across these three platforms. The results showed consistent performance in both inter- and intra-batch tests, confirming the method’s reproducibility. Additionally, the diagnostic reagents produced consistent reaction outcomes across the PCR instrument, metal bath, and water bath (Figure 7).
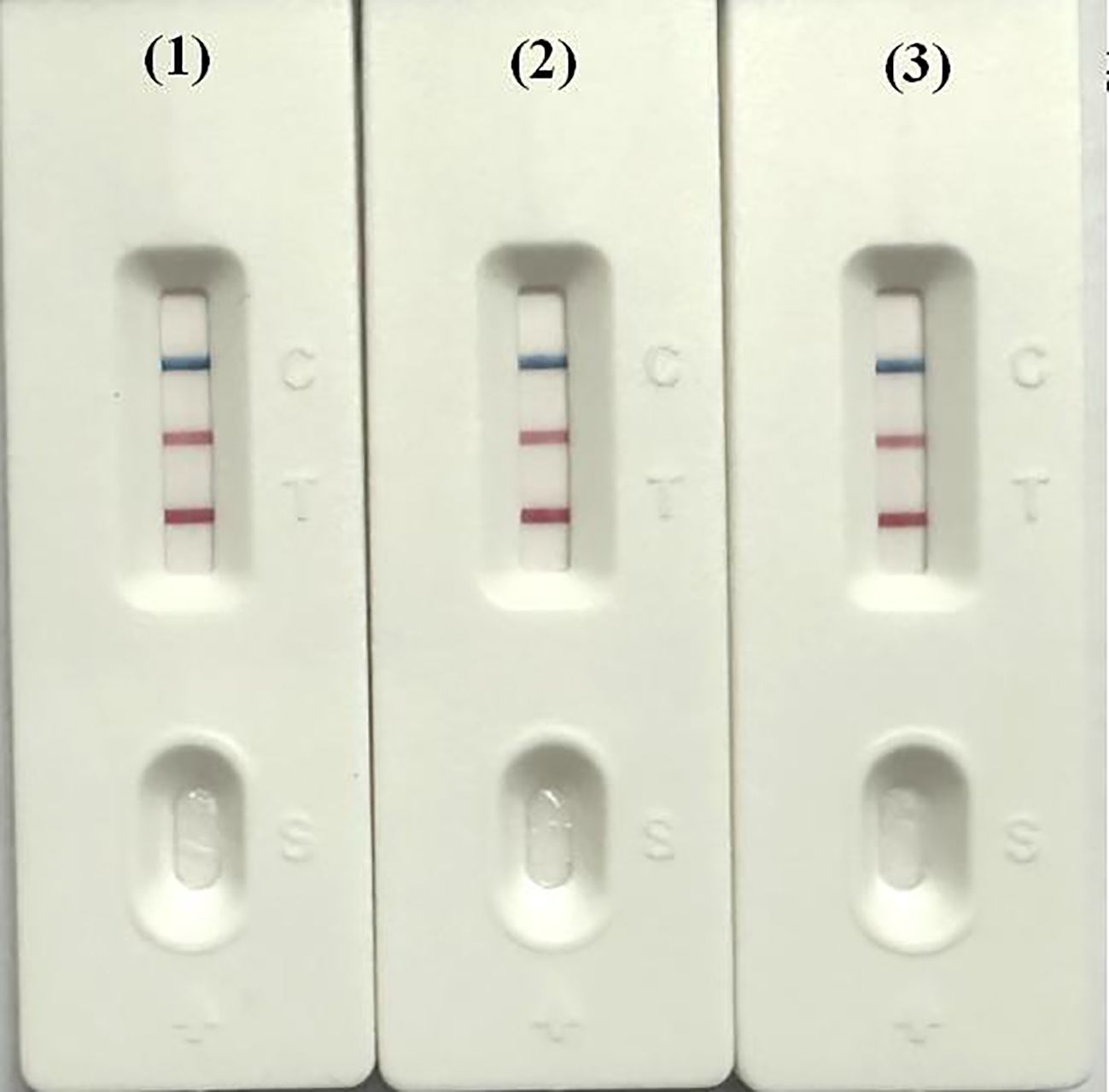
Figure 7. Detection of dual LAMP-LFD in different instruments. 1: PCR instrument (62°C); 2: metal bath (62°C); 3: water bath (62°C).
A total of 106 clinical samples, including pleural effusion, lung tissue, nasal swabs, and others collected from pigs, were tested using the established dual LAMP-LFD method, as well as standard reference methods. The results showed a positive rate of 11.32% (12/106) for S. suis, 25.47% (27/106) for G. parasuis, and a mixed infection rate of 2.83% (3/106). Different levels of positivity were detected for S. suis and G. parasuis in various tissues, including serum, heart, liver, spleen, lungs, kidneys, tonsils, thymus, and abdominal and femoral lymph nodes. The results were consistent with those obtained using national standard detection methods, confirming the accuracy of the dual LAMP-LFD method (Table 3).
Streptococcal and Glässer’s diseases pose significant challenges to the livestock industry, with annual increases in the incidence of bronchopneumonia caused by S. suis and G. parasuis of approximately 6% and 4.3%, respectively, along with a 23% annual rise in S. suis-related endocarditis (Silva et al., 2023). Surveys conducted in China between 2017 and 2021 reported detection rates of 63.50% for S. suis and 28.54% for G. parasuis (Sun et al., 2022). S. suis and G. parasuis was found to be major pathogens of porcine respiratory disease in Guangxi province, with prevalence rates of 65.21% and 48.19%, respectively, and a mixed infection rate of 13.10% (Rao et al., 2023). These statistics highlight the urgent need for rapid and effective diagnostic methods to manage and control these infections. The dual LAMP-LFD method developed in this study addresses this need by offering advantages such as simplicity, rapid results, ease of use, and no requirement for specialized equipment or personnel. These features make it particularly suitable for on-site diagnosis of S. suis and G. parasuis infections, enabling timely decisions regarding treatment and vaccination.
The selection of target genes is a key factor in effective LAMP detection. In this study, the S. suis gdh gene and the G. parasuis infB gene were chosen for the dual LAMP-LFD method. The gdh gene encodes glutamate dehydrogenase, a key virulence factor, and exhibits high nucleotide sequence conservation among different S. suis serotypes (96% to 100%) (Okwumabua et al., 2001; Okwumabua et al., 2003; Xu et al., 2021). While 16S rRNA is commonly used for G. parasuis detection, it lacks specificity in distinguishing G. parasuis from closely related species such as Actinobacillus (Turni et al., 2010; Yang et al., 2010; Zhang et al., 2012). In contrast, the infB gene, as reported by Hedegaard et al., serves as a reliable genetic marker for species identification and can effectively distinguish G. parasuis from related species (Hedegaard et al., 2000). Turni et al. and Pilchová et al. confirmed the suitability of the infB gene for real-time fluorescent quantitative PCR and LAMP detection methods, enhancing the specificity of the dual LAMP-LFD method (Turni et al., 2010; Pilchova et al., 2020).
The design and optimization of LAMP primers were critical to the success of this method. Initial primer selection was performed using Primer Explorer V5 software, followed by screening with a quantitative PCR instrument to identify primers that exhibited optimal performance, as indicated by lower Ct values, smoother curves, earlier peaks, and the absence of non-specific amplification. Optimization revealed that a reaction temperature of 62°C and a primer concentration ratio of 1:4 provided the best amplification results. Additionally, a 40 minute reaction time effectively stabilized amplification products.
Previous studies have reported varying detection limits for LAMP methods. Li et al. achieved a detection limit of 1 fg for the S. suis ermB and mefA genes (Li et al., 2022). Zhang et al. found that gel electrophoresis and SYBR Green I methods for S. suis type 2 cps2J-LAMP products demonstrated the highest sensitivity, with a detection limit of 7.16 copies/μL, although dye methods are prone to non-specific amplification (Zhang et al., 2013; Guang et al., 2023). Pilchová et al. established a LAMP method for G. parasuis infB with a detection limit of 10 fg/μL, while Guang et al. developed a LAMP-LFD method for the same gene with a detection limit of 1.285 × 10-¹² ng/μL (Silva et al., 2023). Unlike these methods, which are limited to single-pathogen detection, the dual LAMP-LFD method developed here can simultaneously detect both S. suis and G. parasuis, with minimum detection limits of 22 and 18 copies/μL for recombinant plasmids, respectively. The consistency of results obtained from testing 106 clinical samples compared to national standard methods confirms the high sensitivity and effectiveness of the method for clinical application. S. suis and G. parasuis were detected in tissues such as the lungs, tonsils, and blood of pigs, underscoring the importance of enhanced health monitoring in pig herds. Particular attention should be given to S. suis due to its potential zoonotic risk, especially in high-exposure environments such as slaughterhouses.
A rapid and convenient dual LAMP-LFD detection method for the simultaneous identification of S. suis and G. parasuis has been successfully developed and validated. This method exhibits high sensitivity, strong specificity, and excellent reproducibility. Preliminary testing of clinical samples yielded promising results, confirming the method’s effectiveness. This study presents a novel diagnostic tool that enhances the rapid identification and epidemiological surveillance of S. suis and G. parasuis, offering significant advantages for the prevention, control, and timely treatment of streptococcal and Glässer’s diseases.
The original contributions presented in the study are included in the article/Supplementary Material. Further inquiries can be directed to the corresponding authors.
HW: Writing – original draft, Validation. CD: Writing – original draft, Validation. XT: Validation, Writing – original draft. YP: Validation, Writing – original draft. LW: Writing – original draft, Validation. TA: Writing – review & editing, Data curation, Formal Analysis, Resources. LZ: Writing – review & editing, Data curation, Formal Analysis, Resources.
The author(s) declare that financial support was received for the research and/or publication of this article. The research was supported by grants from the National Key Research Development Program of China (2022YFD1800300, 2022YFD1800903 and 2022YFD1800703), and the Innovation Program of the Chinese Academy of Agricultural Sciences (CAAS-CSLPDCP-202301).
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fcimb.2025.1575365/full#supplementary-material
Bai, W., Chen, J., Chen, D., Zhu, Y., Hu, K., Lin, X., et al. (2024). Sensitive and rapid detection of three foodborne pathogens in meat by recombinase polymerase amplification with lateral flow dipstick (RPA-LFD). Int. J. Food Microbiol. 422, 110822. doi: 10.1016/j.ijfoodmicro.2024.110822
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Bujold, A. R., Barre, A., Kunkel, E., MacInnes, J. I. (2023). Strain-dependent interactions of Streptococcus suis and Glaesserella parasuis in co-culture. Can. J. Vet. Res. 87, 245–253.
Chidkoksung, K., Parakasikron, N., Nuanualsuwan, S., Khantasup, K. (2024). Development of a latex agglutination test based on VH antibody fragment for detection of Streptococcus suis serotype 2. PloS One 19, e0299691. doi: 10.1371/journal.pone.0299691
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
de Jong, A., Morrissey, I., Rose, M., Temmerman, R., Klein, U., Simjee, S., et al. (2023). Antimicrobial susceptibility among respiratory tract pathogens isolated from diseased cattle and pigs from different parts of Europe. J. Appl. Microbiol. 134, 1–14. doi: 10.1093/jambio/lxad132
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Fan, Q., Wang, H., Wang, Y., Yi, L., Wang, Y. (2024). Evaluation of the protective efficacy of three novel identified membrane associated proteins of Streptococcus suis serotype 2. Microb. Pathog. 193, 106759. doi: 10.1016/j.micpath.2024.106759
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Goto, Y., Fukunari, K., Tada, S., Ichimura, S., Chiba, Y., Suzuki, T. (2023). A multiplex real-time RT-PCR system to simultaneously diagnose 16 pathogens associated with swine respiratory disease. J. Appl. Microbiol. 134 (11), lxad263. doi: 10.1093/jambio/lxad263
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Guang, M., Jin, H. Z., Peng, K. Y., Li, H. M., Xiang, W., Chen, R. G., et al. (2023). Combined loop-mediated isothermal amplification and transverse flow test paper for visual detection of Haemophilus parasuis. J. China Agric. Univ. 28, 180–189. doi: 10.11841/j.issn.1007-4333
Guo, D., Tang, C., Hai, Q., Shao, G., Yue, H. (2010). Development of a universal plate-agglutination test for detecting Haemophilus parasuis. J. Vet. Sci. 11, 355–357. doi: 10.4142/jvs.2010.11.4.355
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Guo, M., Zhang, J., Wang, Q., Tang, J., Li, Y., Zhou, H., et al. (2024). Porcine circovirus type 2 and Glaesserella parasuis serotype 4 co-infection activates Snail1 to disrupt the intercellular junctions and facilitate bacteria translocation across the tracheal epithelium. Vet. Microbiol. 288, 109954. doi: 10.1016/j.vetmic.2023.109954
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Hedegaard, J., Hauge, M., Fage-Larsen, J., Mortensen, K. K., Kilian, M., Sperling-Petersen, H. U., et al. (2000). Investigation of the translation-initiation factor IF2 gene, infB, as a tool to study the population structure of Streptococcus agalactiae. Microbiol. (Reading) 146, 1661–1670. doi: 10.1099/00221287-146-7-1661
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Jiang, Z. J., Hong, J. C., Tang, Q. X., Lin, B. W., Zhang, W. Q., Xia, H., et al. (2024). Streptococcus suis meningoencephalitis diagnosed with metagenomic next-generation sequencing: A case report with literature review. J. Infect. Chemother. 30, 544–547. doi: 10.1016/j.jiac.2023.11.017
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Kirkoyun, U. H., Eryildiz, M., Demirci, M. (2024). Fabrication of a microfluidic test device with a 3D printer and its combination with the loop mediated isothermal amplification method to detect streptococcus pyogenes. Micromachines (Basel) 15 (3), 365. doi: 10.3390/mi15030365
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Li, L., Ren, J., Zhang, Q., Luo, Y., Zhang, Y., Qi, J., et al. (2022). Development of Two Loop-Mediated Isothermal Amplification Assays for Rapid Detection of ermB and mefA Genes in Streptococcus suis. Foodborne Pathog. Dis. 19, 817–822. doi: 10.1089/fpd.2022.0034
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Li, K., Zhang, Y., Luo, T., Li, C., Yu, H., Wang, W., et al. (2024). Development of a Triplex qPCR Assay Based on the TaqMan Probe for the Detection of Haemophilus parasuis, Streptococcus suis Serotype 2 and Pasteurella multocida. Microorganisms 12 (10), 2017. doi: 10.3390/microorganisms12102017
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Nedbalcova, K., Kucharovicova, I., Zouharova, M., Matiaskova, K., Kralova, N., Brychta, M., et al. (2022). Resistance of Streptococcus suis Isolates from the Czech Republic during 2018-2022. Antibiotics (Basel) 11 (9), 1214. doi: 10.3390/antibiotics11091214
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Okwumabua, O., O'Connor, M., Shull, E. (2003). A polymerase chain reaction (PCR) assay specific for Streptococcus suis based on the gene encoding the glutamate dehydrogenase. FEMS Microbiol. Lett. 218, 79–84. doi: 10.1111/j.1574-6968.2003.tb11501.x
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Okwumabua, O., Persaud, J. S., Reddy, P. G. (2001). Cloning and characterization of the gene encoding the glutamate dehydrogenase of Streptococcus suis serotype 2. Clin. Diagn. Lab. Immunol. 8, 251–257. doi: 10.1128/CDLI.8.2.251-257.2001
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Petrocchi, R. M., Gutierrez, M. C., Acebes, F. V., Aguaron, T. A., Gonzalez, F. A., Miguelez, P. R., et al. (2024). Streptococcus suis research update: serotype prevalence and antimicrobial resistance distribution in swine isolates recovered in Spain from 2020 to 2022. Vet. Sci. 11 (1), 40. doi: 10.3390/vetsci11010040
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Pilchova, V., Seinige, D., Hennig-Pauka, I., Buttner, K., Abdulmawjood, A., Kehrenberg, C. (2020). Development and validation of a loop-mediated isothermal amplification (LAMP) assay for rapid detection of glaesserella (Haemophilus) parasuis. Microorganisms 9 (1), 41. doi: 10.3390/microorganisms9010041
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Rao, J., Wei, X., Li, H., Zhang, Z., Liu, J., Lian, M., et al. (2023). Novel multiplex PCR assay and its application in detecting prevalence and antibiotic susceptibility of porcine respiratory bacterial pathogens in Guangxi, China. Microbiol. Spectr. 11, e0397122. doi: 10.1128/spectrum.03971-22
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Renzhammer, R., Auer, A., Loncaric, I., Entenfellner, A., Dimmel, K., Walk, K., et al. (2023). Retrospective analysis of the detection of pathogens associated with the porcine respiratory disease complex in routine diagnostic samples from Austrian swine stocks. Vet. Sci. 10 (10), 601. doi: 10.3390/vetsci10100601
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Salogni, C., Capucchio, M. T., Colombino, E., Pozzi, P., Pasquali, P., Alborali, G. L. (2022). Bacterial polyarthritis in post-weaning pigs in a high-density swine breeding area in Italy. J. Vet. Diagn. Invest. 34, 709–711. doi: 10.1177/10406387221090903
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Santoya, E. L., Moran, D. D., Diaz, A. P., Rodriguez, A. E. (2024). Streptococcus suis meningitis. Medicina (B Aires) 84, 329–332.
Scherrer, S., Biggel, M., Schneeberger, M., Cernela, N., Rademacher, F., Schmitt, S., et al. (2024). Genetic diversity and antimicrobial susceptibility of Streptococcus suis from diseased Swiss pigs collected between 2019 - 2022. Vet. Microbiol. 293, 110084. doi: 10.1016/j.vetmic.2024.110084
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Schuwerk, L., Hoeltig, D., Waldmann, K. H., Strutzberg-Minder, K., Valentin-Weigand, P., Rohde, J. (2020). Serotyping and pathotyping of Glaesserella parasuis isolated 2012-2019 in Germany comparing different PCR-based methods. Vet. Res. 51, 137. doi: 10.1186/s13567-020-00862-1
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Silva, A., Almeida, M., Michael, A., Rahe, M. C., Siepker, C., Magstadt, D. R., et al. (2023). Detection and disease diagnosis trends (2017-2022) for Streptococcus suis, Glaesserella parasuis, Mycoplasma hyorhinis, Actinobacillus suis and Mycoplasma hyosynoviae at Iowa State University Veterinary Diagnostic Laboratory. BMC Vet. Res. 19, 268. doi: 10.1186/s12917-023-03807-w
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Sun, Q., Yu, X., He, D., Ku, X., Hong, B., Zeng, W., et al. (2022). Investigation and analysis of etiology associated with porcine respiratory disease complex in China from 2017 to 2021. Front. Vet. Sci. 9. doi: 10.3389/fvets.2022.960033
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Turni, C., Pyke, M., Blackall, P. J. (2010). Validation of a real-time PCR for Haemophilus parasuis. J. Appl. Microbiol. 108, 1323–1331. doi: 10.1111/j.1365-2672.2009.04526.x
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Wang, H., Xin, L., Wu, Y., Liu, Y., Yao, W., Zhang, H., et al. (2023). Construction of a one-step multiplex real-time PCR assay for the detection of serogroups A, B, and E of Pasteurella multocida associated with bovine pasteurellosis. Front. Vet. Sci. 10, 1193162. doi: 10.3389/fvets.2023.1193162
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Wang, G., Zhu, H., Zhan, C., Chen, P., Wu, B., Peng, Z., et al. (2024). Establishment and application of a quadruplex real-time reverse-transcription polymerase chain reaction assay for differentiation of porcine reproductive and respiratory syndrome virus, porcine circovirus type 2, porcine circovirus type 3, and streptococcus suis. Microorganisms 12. doi: 10.3390/microorganisms12030427
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Wu, C. F., Hsu, C. Y., Chou, C. C., Wang, C. M., Huang, S. W., Kuo, H. C. (2023). Serotypes, virulence factors and multilocus sequence typing of Glaesserella parasuis from diseased pigs in Taiwan. PeerJ 11, e15823. doi: 10.7717/peerj.15823
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Xia, Y., Wang, Z., Hu, Y., Zhao, P., Li, J., Zhang, L., et al. (2024). Isolation, identification, genomic diversity, and antimicrobial resistance analysis of streptococcus suis in Hubei Province of China from 2021 to 2023. Microorganisms 12 (5), 917. doi: 10.3390/microorganisms12050917
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Xin, L., Wang, H., Hu, Y., Liu, Y., Yao, W., Wang, X., et al. (2023). The establishment and application of a one-step multiplex real-time polymerase chain reaction assay for the detection of Streptococcus suis, Streptococcus suis serotype 2, and Glaesserella parasuis. Anim. Res. One Health, 1–12. doi: 10.14202/vetworld.2024.946-955
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Xu, Q., Chen, H., Sun, W., Zhu, D., Zhang, Y., Chen, J. L., et al. (2021). Genome-wide analysis of the synonymous codon usage pattern of Streptococcus suis. Microb. Pathog. 150, 104732. doi: 10.1016/j.micpath.2021.104732
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Yan, P., Jia, Y. C., Zhang, X. L., Zhou, Y. Y., Guo, Y., Yin, R. L., et al. (2023). Virulence assessment of four Glaesserella parasuis strains isolated in Liaoning province of China. Res. Vet. Sci. 158, 226–234. doi: 10.1016/j.rvsc.2023.03.021
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Yang, W., Ying, F., Yingyu, L., Pin, C., Wentao, L., Shuqing, L., et al. (2010). Development and evaluation of loop-mediated isothermal amplification for rapid detection of Haemophilus parasuis. FEMS Microbiol. Lett. 313, 54–60. doi: 10.1111/j.1574-6968.2010.02126.x
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Yue, C., Li, J., Zhang, S., Ma, R., Suo, M., Chen, Y., et al. (2024). Activation of the NLRP3-CASP-1 inflammasome is restrained by controlling autophagy during Glaesserella parasuis infection. Vet. Microbiol. 295, 110160. doi: 10.1016/j.vetmic.2024.110160
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Zhang, J. M., Shen, H. Y., Liao, M., Ren, T., Guo, L. L., Xu, C. G., et al. (2012). Detection of Haemophilus parasuis isolates from South China by loop-mediated isothermal amplification and isolate characterisation. Onderstepoort J. Vet. Res. 79, E1–06. doi: 10.4102/ojvr.v79i1.383
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Zhang, J., Zhu, J., Ren, H., Zhu, S., Zhao, P., Zhang, F., et al. (2013). Rapid visual detection of highly pathogenic Streptococcus suis serotype 2 isolates by use of loop-mediated isothermal amplification. J. Clin. Microbiol. 51, 3250–3256. doi: 10.1128/JCM.01183-13
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Zhu, H., Chang, X., Zhou, J., Wang, D., Zhou, J., Fan, B., et al. (2021). Co-infection analysis of bacterial and viral respiratory pathogens from clinically healthy swine in Eastern China. Vet. Med. Sci. 7, 1815–1819. doi: 10.1002/vms3.533
PubMed Abstract | PubMed Abstract | Crossref Full Text | Google Scholar
Keywords: Streptococcus suis, Glaesserella parasuis, LAMP, LFD, rapid detection
Citation: Wang H, Dong C, Tian X, Pan Y, Wang L, An T and Zhu L (2025) Development and application of a dual LAMP-LFD assay for the simultaneous detection of Streptococcus suis and Glaesserella parasuis. Front. Cell. Infect. Microbiol. 15:1575365. doi: 10.3389/fcimb.2025.1575365
Received: 12 February 2025; Accepted: 17 March 2025;
Published: 01 April 2025.
Edited by:
Xiaoyuan Wei, University of South Florida, United StatesReviewed by:
Yang Wang, Henan University of Science and Technology, ChinaCopyright © 2025 Wang, Dong, Tian, Pan, Wang, An and Zhu. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Liangquan Zhu, MTM2NzM5MTg5NEBxcS5jb20=; Tonqging An, YW50b25ncWluZ0BjYWFzLmNu
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.