
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Cell. Infect. Microbiol. , 14 March 2025
Sec. Intestinal Microbiome
Volume 15 - 2025 | https://doi.org/10.3389/fcimb.2025.1535204
This article is part of the Research Topic Harnessing the Medicinal Potential of Gut Microbiota for Human Health View all 4 articles
 Wenhao Liao1,2
Wenhao Liao1,2 Lei Cao1,2
Lei Cao1,2 Xuemei Jiang1,2
Xuemei Jiang1,2 Lianqiang Che1,2
Lianqiang Che1,2 Zhengfeng Fang1,2
Zhengfeng Fang1,2 Shengyu Xu1,2
Shengyu Xu1,2 Yan Lin1,2
Yan Lin1,2 Yong Zhuo1,2
Yong Zhuo1,2 Lun Hua1,2
Lun Hua1,2 Jian Li1,2
Jian Li1,2 Guangmang Liu1
Guangmang Liu1 Mengmeng Sun3
Mengmeng Sun3 De Wu1,2
De Wu1,2 Hairui Wang4*
Hairui Wang4* Bin Feng1,2*
Bin Feng1,2*The intestinal microbiota is important for the health of the host and recent studies have shown that some genes of the host regulated the composition of the intestinal microbiota. Group 10 phospholipase A2 (PLA2G10) is a member of the lipolytic enzyme family PLA2, which hydrolyze the ester bond at the sn-2 position of phospholipids to produce free fatty acids and lysophospholipids. PLA2G10 is secreted into the intestinal lumen, but its impact on the gut microbiota remains unclear. In this study, we generated intestine-specific Pla2g10 knock-in mice, and used 16S RNA sequencing to compare their gut microbiota with that of their wild-type (WT) littermates. Results showed that gut-specific Pla2g10 knock-in induced both PLA2G10 mRNA and protein levels in the colon. Moreover, intestinal Pla2g10 overexpression reduced the α-diversity of the gut microbiota relative to that of WT mice. The abundance of Bacteroidetes was lower in the Pla2g10 knock-in mice than that in the control mice, while the ratio of Firmicutes/Bacteroidetes was higher. Furthermore, the abundance of the genus Allobaculum was reduced, whereas the abundance of beneficial bacteria genera, including Enterorhabdus, Dubosiella, and Lactobacillus, was increased by host intestinal Pla2g10 overexpression. In summary, intestinal Pla2g10 overexpression increased the proportions of beneficial bacterial in the colonic chyme of mice, providing a potential therapeutic target for future improvement of the gut microbiota.
In recent years, our understanding of the gut microbiota has greatly improved. Our gastrointestinal tract, particularly the colon, contains a great variety of microbes, harbors a wide variety of bacteria, viruses and other microorganisms. These gut microbes interact with the host’s epithelial and immune cells, forming a biological barrier in the gut and coordinating the primary and secondary metabolism on the surface of the intestinal mucosa (Durack and Lynch, 2019). In addition, the gut microbiota coexists symbiotically with its host, participating in numerous physiological activities and playing crucial roles in the host’s health and the development of diseases. The numbers, species, abundance, and evenness of the microorganisms that make up this complex ecosystem affect the host’s health, and the ecosystem is in a state of dynamic homeostasis, and any disruption of its balance increases the risk of disease in the host (Chen et al., 2021; Clemente et al., 2018; Durack and Lynch, 2019).
The phospholipase A2 (PLA2) family belongs to the lipolytic enzymes and its members hydrolyze the sn-2 ester bond of phospholipids to produce free fatty acids and lysophospholipids. Secretory PLA2 (sPLA2) constitutes the largest subfamily of PLA2, with 10 catalytically active sPLA2s identified in mammals: PLA2G1B, -2A, -2C, -2D, -2E, -2F, -3, -5, -10, and -12 (Dennis et al., 2011). Among the secretory phospholipases, PLA2G10 contains a histidine/asparagine catalytic dyad and a highly conserved Ca2+-binding loop, and conserved disulfide bonds contribute to its structural stability (Murakami et al., 2023). PLA2G10 is synthesized as a zymogen and converted into an active mature enzyme by the proteolytic cleavage of the N-terminal propeptide. This processing can occur within the cell before its secretion, by protease-like conversion enzymes, or outside the cell after its secretion, as is the case for various digestive enzymes in the gastrointestinal tract (Jemel et al., 2011; Layne et al., 2015; Masuda et al., 2005).
PLA2G10 is highly expressed in the Paneth cells of the colon (Schewe et al., 2016) and it has a greater capacity than other PLAs to hydrolyze phosphatidylcholine (PC), the major phospholipid component of the plasma membrane, producing lysophosphatidylcholine (LPC) and free fatty acids (such as polyunsaturated fatty acids of arachidonic acid, eicosapentaenoic acid, docosapentaenoic acid and docosahexaenoic acid) (Bezzine et al., 2000; Hanasaki et al., 1999; Kudo et al., 2022; Murase et al., 2016). Reports have suggested that dietary supplementation of polyunsaturated fatty acids could change the diversity and abundance of gut microbiota (Cao et al., 2019; Feng et al., 2020; Mujico et al., 2013; Tang et al., 2021; Zhuang et al., 2017). Additional, several studies have demonstrated that various secretory phospholipases alter the composition and abundance of the gut microbiota in mice. For instance, the relative abundance of Bacteroides correlates positively with the expression of Pla2g5, Pla2g2a, and Pla2g2f, whereas the abundances of norank_f_ Erysipelotrichaceae and Roseburia correlate negatively with these three enzymes. Pla2g2a−/− mice have reduced Coriobacteriaceae FCS020 and Ruminococcaceae UCG-013, whereas Coriobacteriaceae UCG-008 is significantly elevated compared with its abundance in Pla2g2a+/+ mice (Taketomi et al., 2022; Zhang et al., 2023). However, the researches on the effect of Pla2g10 on gut microbiota are still few. To further investigate the effect of intestinal Pla2g10 overexpression on gut microbiota, we generated intestine-specific Pla2g10 knock-in mice, which specifically overexpressed Pla2g10 in intestine. Results showed that overexpression of Pla2g10 in intestine altered the composition, diversity and function of the gut microbiota in mice.
All animal protocols were reviewed and approved by the Animal Care and Use Committee of Sichuan Agricultural University (SICAU-2020-280, 12 May 2020). All animal studies were performed in accordance with the Use and Care of Laboratory Animals. Pla2g10loxp+/− mice were constructed with C57BL/6 background Embryonic stem cells (Figure 1A). C57BL/6J-background tomoxifen-dependent Lgr5cre+ mice (T003768) were purchased from GemPharmatech LLC., China. Pla2g10 knock-in (KI, Pla2g10loxp+/− Lgr5cre+/−, n = 6) mice were generated by mating Pla2g10loxp+/− mice with Lgr5cre+ mice, and Pla2g10loxp+/− mice (n = 6) were used as the control. All the mice were housed under the individual ventilated caging system, maintained in a 12-h light/12-h dark environment, with a room temperature maintained of 22°C and humidity of 60%, with free access to food and water.
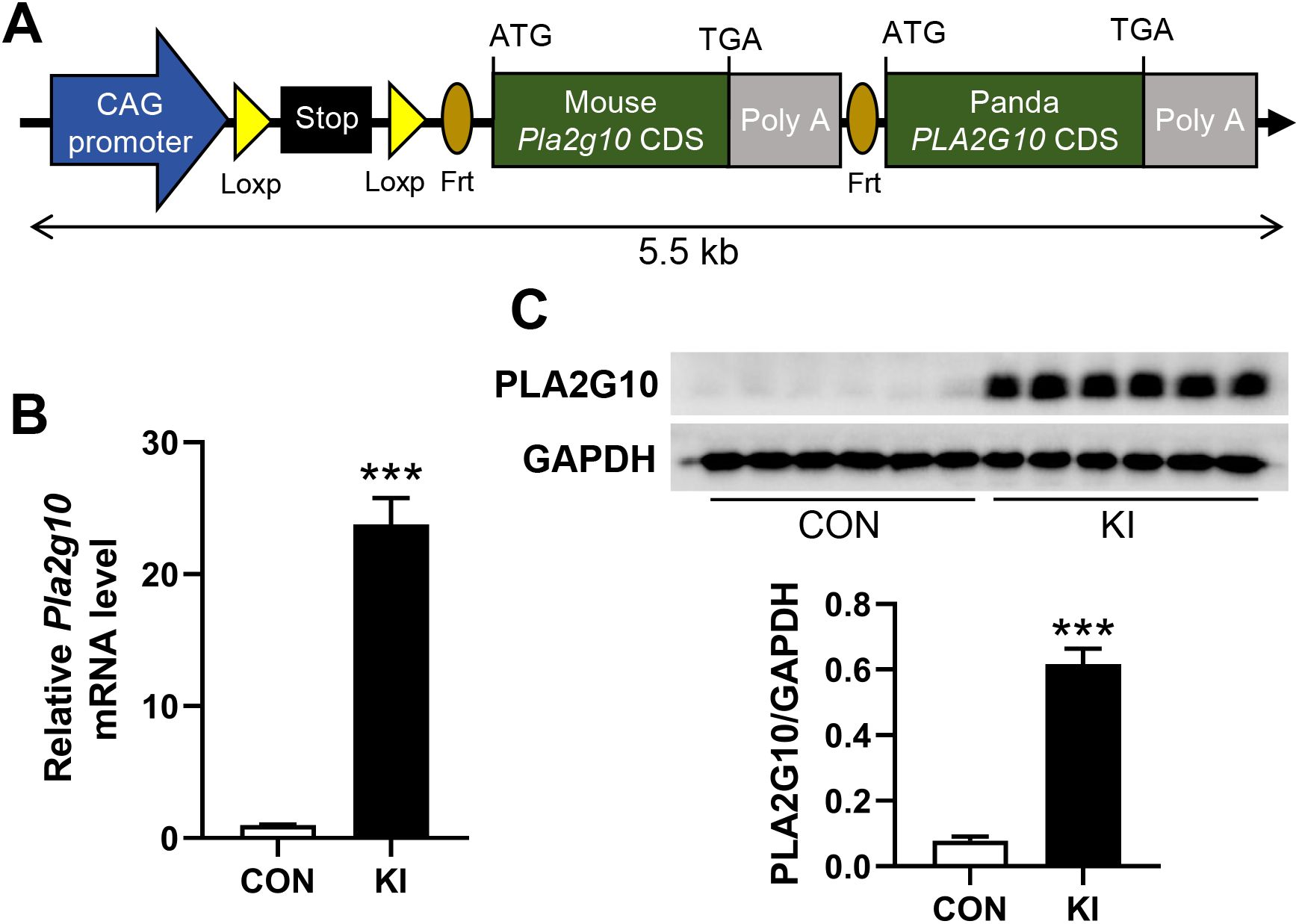
Figure 1. Generation of intestine-specific Pla2g10 knock-in mice. (A) Schematic for construction of mouse Pla2g10 loxp mice. Coding sequence of panda PLA2G10 was inserted for future study. (B) qPCR analysis for Pla2g10 gene expression in mice colon (n = 6). (C) Western blot analysis for PLA2G10 protein expression in mice colon. Data is presented as mean ± SEM. CON, control group; KI, Pla2g10 konck-in group. ***P < 0.001.
Mice with the same genotype were housed together, with 2 mice per cage. At an age of 4 months, the mice were weighted and injected with 100 mg/kg tamoxifen (#S1238, Selleck, Shanghai, China) once daily for a week to induce Cre expression. After injection, their body weight were recorded weekly for 4 weeks, and fed blood glucose levels were measured twice weekly, and food intake were recorded daily for 4 days. 4 weeks after tamoxifen injection, mice were euthanized with CO2. The colons were collected, and the chyme in the colons was collected for microbial DNA extraction.
The initial weight of food in each cage on the first day was recorded at 5 pm, while the weight of the remaining food was measured at 5 pm on the following days. The food intake was calculated as (initial food weight - remaining food weight)/2.
Total RNA was extracted from tissues using RNAiso Plus (#9109, TaKaRa Bio, Dalian, China) following the manufacturer’s instructions. 1 μg of total RNA was used to synthesize cDNA using the PrimeScript FAST RT reagent Kit with gDNA Eraser (#RR092A, TaKaRa Bio). The cDNA was then subjected to qPCR using the iTaq Universal SYBR Green Supermix (#1725124, Bio-Rad, Shanghai, China) with a 384-well real-time PCR machine (QuantStudio 5, Thermo Fisher, Shanghai, China). The following primers were used: Gapdh F: 5’-AGGTCGGTGTGAACGGATTTG-3′, R: 5′-TGTAGACCATGTAGTTGAGGTCA-3′; Pla2g10 F: 5’-GTGCAGGTGTGACGAGGAG-3′, R:5′-CACTTGGGAGAGTCCTTCTCA-3′.
After homogenized in Western ReProbe (TM) PLUS (#C500035, Sangon Biotech, Shanghai, China) supplemented with PMSF (#abs812852, Absin, Shanghai, China), the protein samples were centrifuged at 12,000 rpm for 30 min. The protein lysates were quantified using a BCA protein assay kit (#23227, Thermo Fisher, Shanghai, China). Protein samples with equivalent concentrations were separated by SDS-PAGE. The following antibodies were used: rabbit anti-GAPDH Ab (#60004-1-lg, Proteintech, Wuhan, China) and mouse anti-PLA2G10 Ab (#sc-514324, Santa cruz, Shanghai, China).
Genomic DNA (gDNA) was extracted from the colonic chyme using a soil genomic DNA extraction kit (#DP336, Tiangen biotech, Beijing, China). The gDNA was used as a template to amplify the highly-variable V3–V4 region of the 16S RNA gene by PCR, using the forward primer 341F (5’-ACTCCTACGGGAGGCAGCAG-3’) and the reverse primer 806R (5’-GGACTACHVGGGTWTCT AAT-3’). All PCRs were performed in 15μL of Phusion High-Fidelity PCR Master Mix (New England Biolabs, Manchester, UK), with 0.2 μM of both the forward and reverse primers, and approximately 10 ng of template DNA. Thermal cycling consisted of an initial denaturation at 98°C for 1 minute, followed by 30 cycles of denaturation at 98°C for 10s, annealing at 50°C for 30s, and elongation at 72°C for 30s, and finally an extension at 72°C for 5 min.
The PCR products were purified using magnetic beads. The samples were mixed in equal-density ratios based on the concentrations of the PCR products. After the PCR products were thoroughly mixed, they were detected and purified using a universal DNA purification and recovery kit (#DP214-03, Tiangen Biotech, Beijing, China) to isolate the nucleic acids. The nucleic acids were subjected to construct libraries using a library construction kit (#E7645S, New England BioLabs, Manchester, UK). The libraries were sequenced using the Illumina NovaSeq 6000 (PE250) system.
Paired-end reads were merged using the FLASH (v1.2.11) to obtain raw tags (Magoč and Salzberg, 2011). The Cutadapt software (v3.3) was then used to match the reverse primer sequence and trim the remaining sequences to prevent their interference in subsequent analyses. Quality filtering of the raw tags was performed using the fastp software (v0.23.1) to obtain high-quality clean tags (Bokulich et al., 2013). To detect chimeric sequences, the tags were compared with the reference database (Silva database (16S/18S), and the effective tags were obtained by removing the chimeric sequences using the vsearch package (v2.16.0) (Edgar et al., 2011). To obtain the initial amplicon sequence variants (ASVs), the DATA2 program in the QIIME2 software (v202202) was performed to denoise the previous effective tags (Y. Wang et al., 2021). The species annotation was performed with QIIME2 software.
The absolute abundance of ASVs was normalized using a standard of sequence number corresponding to the sample with the fewest sequences. The subsequent analyses of α-diversity and β-diversity were all performed based on the output normalized data. The top 10 taxa at the phylum level and top 22 taxa at the genus level of each sample were selected to plot the distribution histogram of relative abundance using Perl software (v5.26.2). The Venn diagram was produced in R software (v4.0.3) using the Venn Diagram function, and the α-diversity was calculated using Chao1 and Shannon indices in QIIME2 software to analyze the diversity, richness and uniformity. The β-diversity was calculated by the unweighted Unifrac distance using QIIME2 software, and was visualized by principal coordinate analysis (PCoA) and non-metric multidimensional scaling (NMDS) analysis using R software (v4.0.3). The Linear Discriminant Analysis Effect Size (LEfSe) was applied to analyze the differences at each level using R software (v4.0.3). The functional genes and Kyoto Encyclopedia of Genes and Genomes (KEGG) orthologs were predicted by the Phylogenetic Investigation of Communities by Reconstruction of Unobserved States (PICRUSt2) (v2.3.0). The network plot was produced using Chiplot (https://www.chiplot.online/).
Data are presented as mean ± standard error of the mean (SEM). Growth data were preliminarily analyzed using Microsoft Excel 2019 and then analyzed statistically using the SPSS 21.0 software. Normality of the data was evaluated by Shapiro-Wilk test using the SPSS 21.0 software. Microbial data were organized and analyzed statistically using the R software (v4.0.3). Data were visualized using the GraphPad Prism (v8.0) and R software (v4.0.3) The P - values used to compare two groups were calculated with Student’s t-test. A trend was deemed present when 0.05 < P < 0.1; the significance of differences was defined as *P < 0.05.
To explore the effects of intestinal Pla2g10 overexpression on the homeostasis of gut microbiota, Pla2g10loxp+/− mice were mated with tamoxifen-dependent Lgr5cre+ mice to generate intestine-specific Pla2g10 knock-in (KI) mice. Both the gene expression and protein levels of PLA2G10 were significantly higher in the colons of the Pla2g10 KI mice than in those of the control mice (Figures 1B, C). In the subsequent 4 weeks after CRE induced, the body weights and glucose levels did not differ significantly between the two groups (Figures 2A, B). Although there was no significant difference in the food intake of the two groups, the Pla2g10 KI mice consistently consumed 1 g more than the control mice (Figure 2C). A PCoA with an unweighted UniFrac distance algorithm revealed differences in the microbial compositions of the two groups (Figure 3A). The first principal axis (PCoA1) could explain 30.44% of the sample difference, and the second principal axis (PCoA2) could explained 18.45% of the sample difference. The NMDS stress value was 0.039, indicating that the subsequent analytical results accurately reflected the differences between the samples with high confidence (Figure 3B). We assessed the α-diversity of the gut microbiota by calculating the Chao1 and Shannon indices, both of which were significantly lower in the Pla2g10 KI mice than in the control group (Figures 3C, D), indicating that the total number of gut bacteria and their richness and evenness were reduced after the overexpression of Pla2g10. A Venn diagram confirmed this, showing that the number of operational taxonomic units (OTUs) was lower in the Pla2g10 KI mice than in the control group (Supplementary Figure 1A).
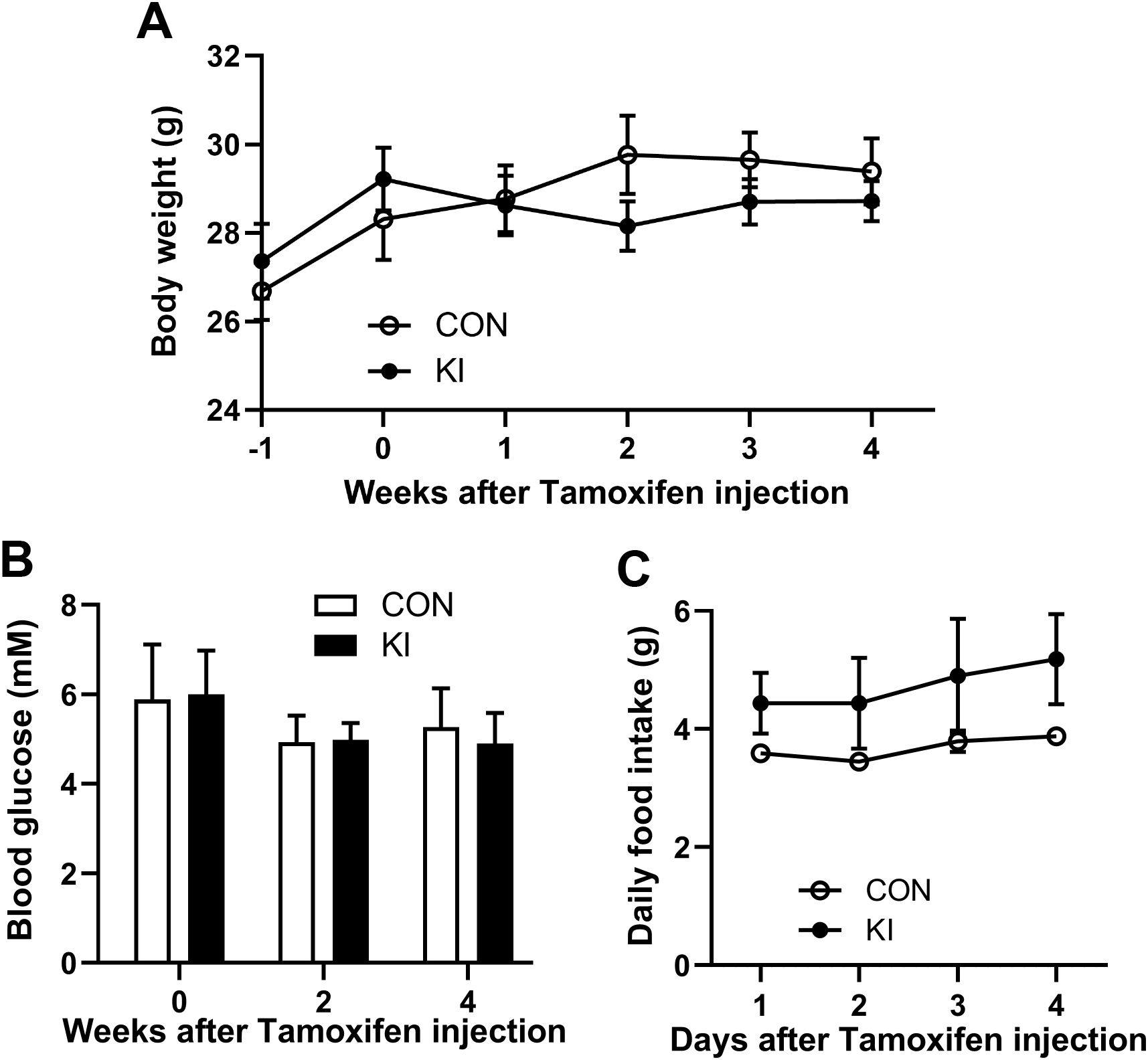
Figure 2. Pla2g10 overexpression did not alter the body weight, food intake, and blood glucose levels of mice. (A) The body weight (n = 6). (B) Blood glucose levels (n = 6). (C) Food intake (n = 3). Mice with the same genotype were housed together, with 2 mice per cage. The initial weight of food in each cage on the first day was recorded at 5 pm, while the weight of the remaining food was measured at 5 pm on the following days. The food intake was calculated as (initial food weight - remaining food weight)/2. Data is presented as mean ± SEM. CON, control group; KI, Pla2g10 konck-in group.
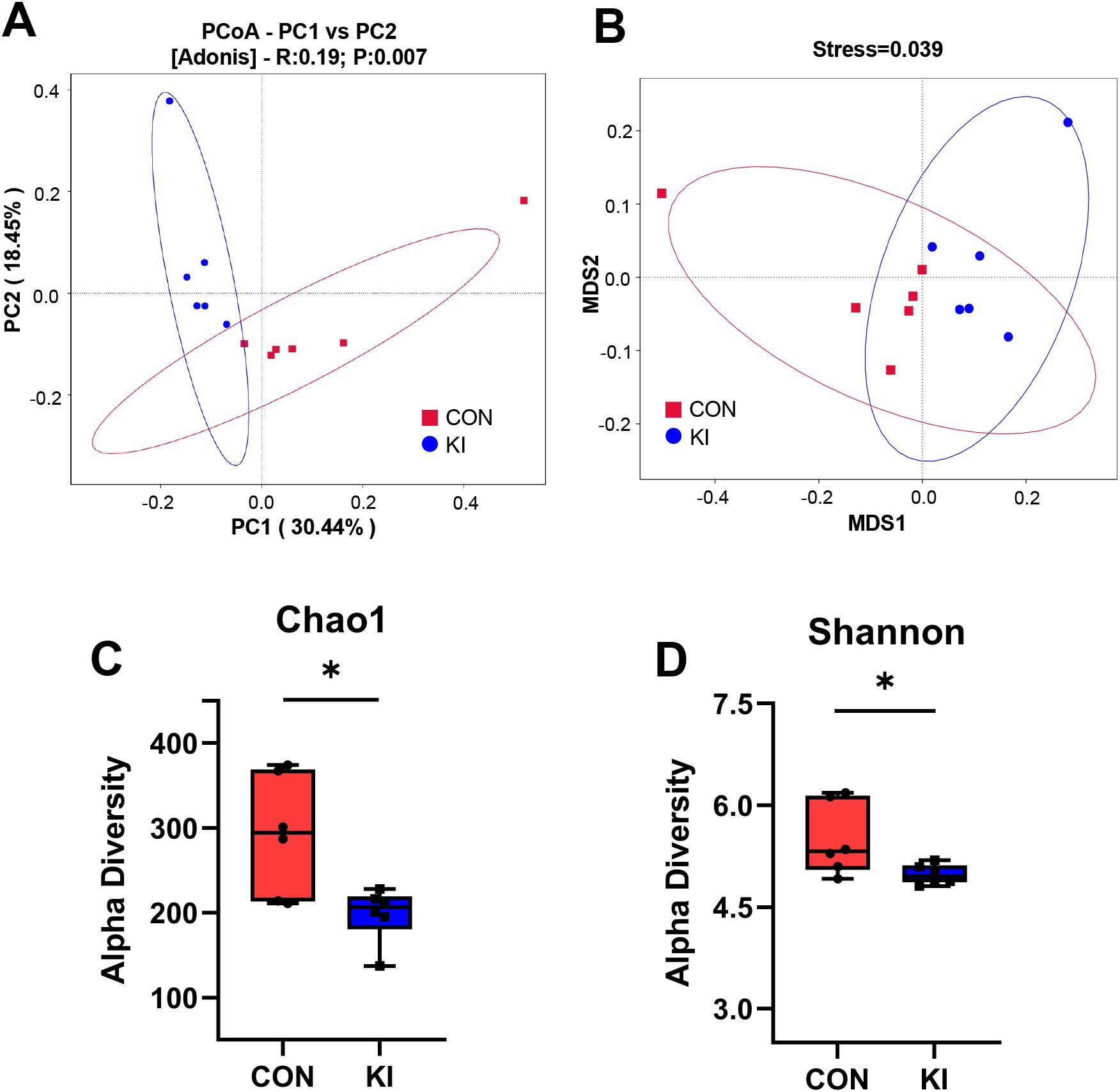
Figure 3. Pla2g10 overexpression altered the composition of the gut microbiota in mice. (A, B) PCoA and NMDS plot, distance algorithm: unweighted_unifrac. (C, D) Shannon and Chao1 index of α-diversity. Data is presented as mean ± SEM. *P < 0.05. CON, control group; KI, Pla2g10 konck-in group.
Analysis of the community composition and species differences between the Pla2g10 KI and control mice showed that the overexpression of Pla2g10 altered the composition of the gut microbiota at the phylum (Figure 4A) and genus levels (Figure 4B). The detailed distributions of the gut microbiotal compositions of the two groups are shown in Supplementary Table 1. The abundance of Actinobacteria in the colonic chyme of the Pla2g10 KI group had an upward trend compared with the control group, whereas the abundance of Bacteroidetes was significantly lower (Figure 5A), and the Firmicutes/Bacteroidetes (F/B) ratio also showed an increasing trend in the Pla2g10 KI mice (Supplementary Figure 1B). At the genus level, the relative abundances of Dubosiella (P = 0.020), Enterorhabdus (P = 0.030), Lactococcus (P = 0.033) and Lactobacillus (P = 0.045) were significantly elevated in the Pla2g10 KI mice compared to control group (Figure 5B), while Faecalibaculum only showed an increasing trend (P = 0.066). One genus, Allobaculum, showed a significant decline in abundance in the gut microbiota of the Pla2g10 KI mice compared with control mice. Differences in bacteria at other levels between the Pla2g10 KI and control groups are shown in Supplementary Figures 1C–E.
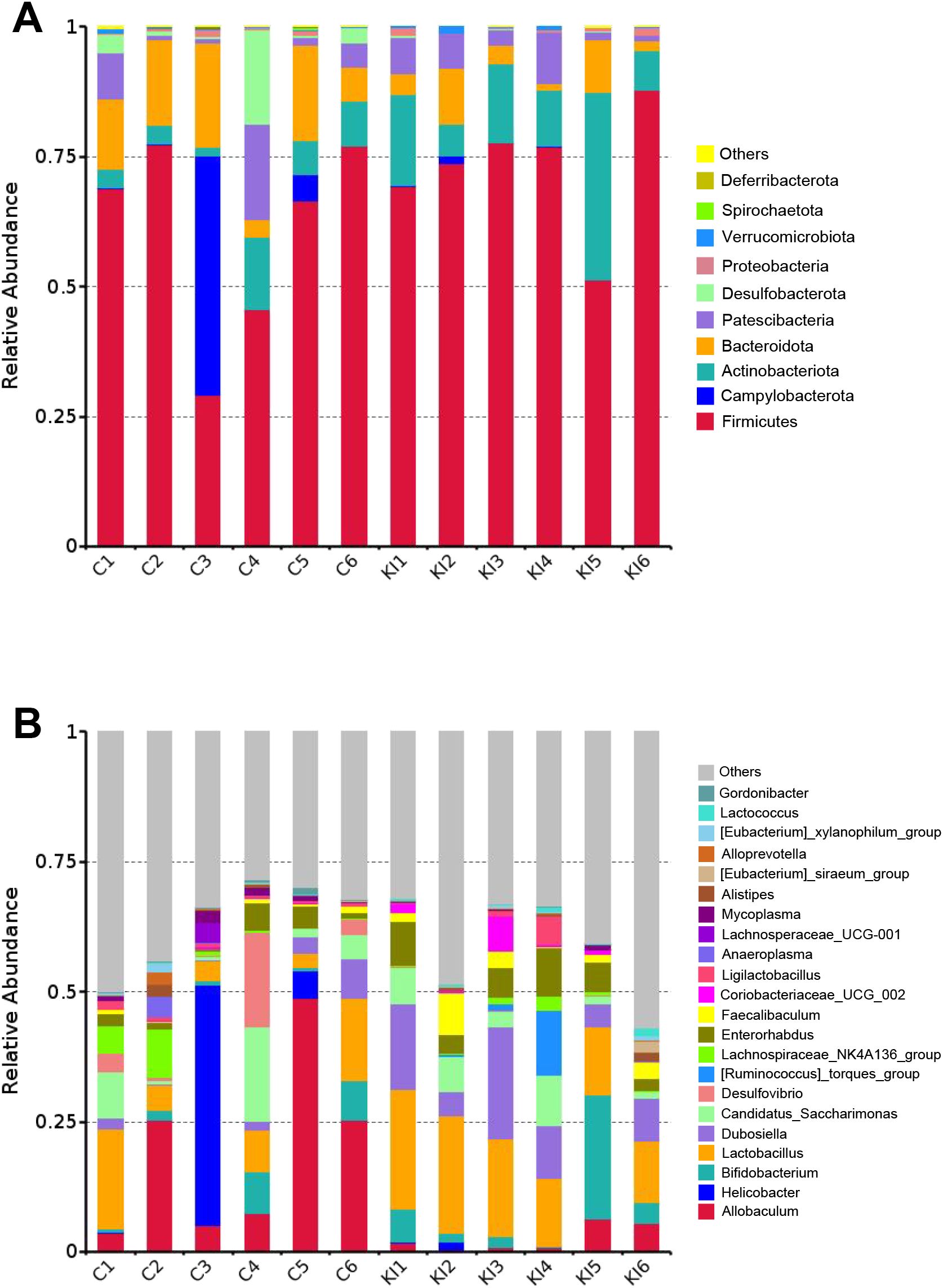
Figure 4. Taxonomic comparison of gut microbiota in Pla2g10 KI and control mice. Taxonomic composition of the gut microbiota at the phylum (A) and genus (B) levels. C, control group; KI, Pla2g10 konck-in group.
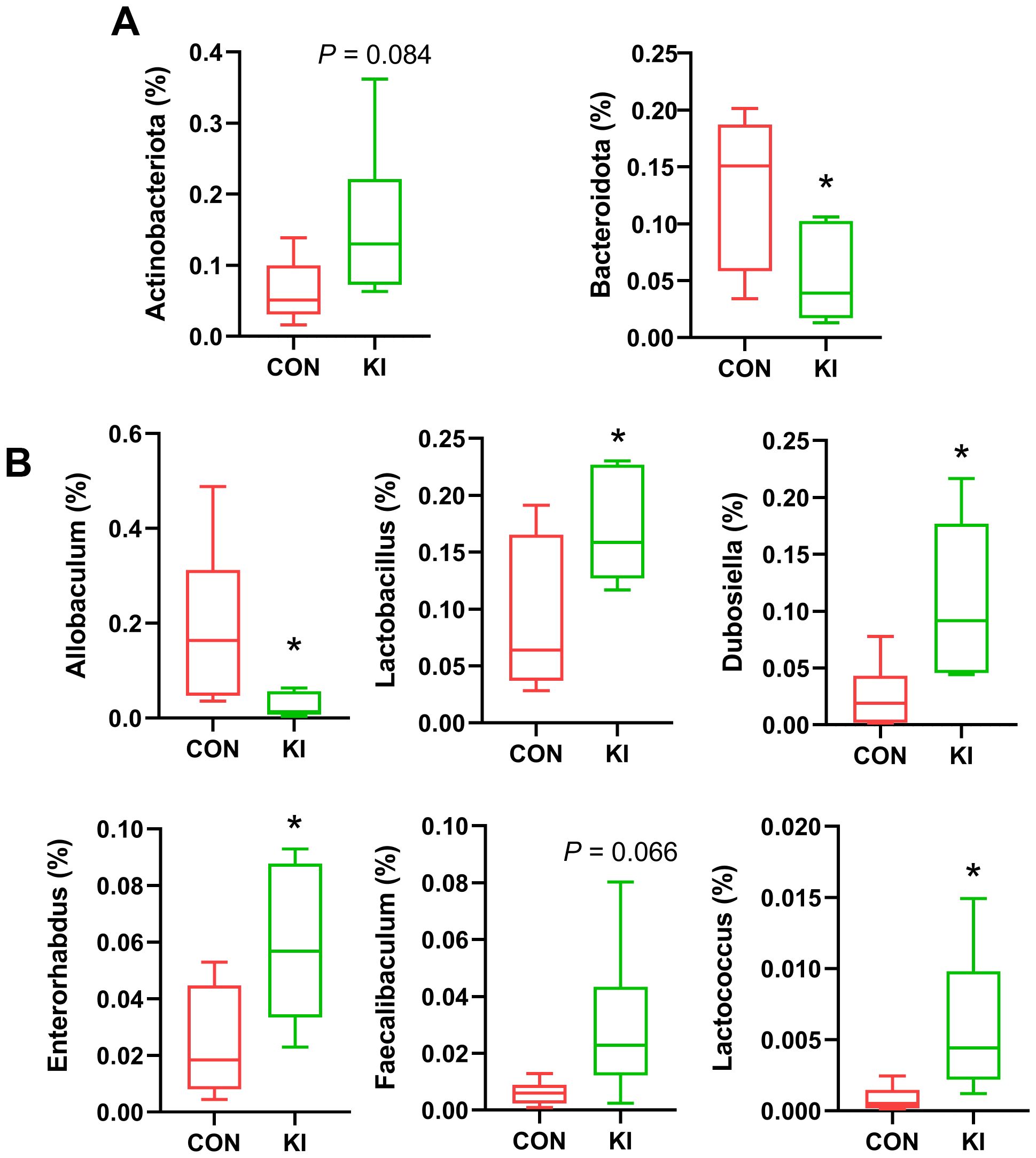
Figure 5. Bacterial taxa in the chyme of Pla2g10 KI and control mice. Significantly changed bacterial taxa at the phylum (A, B) genus levels. Data is presented as mean ± SEM. CON, control group; KI, Pla2g10 konck-in group. *P < 0.05.
To determine the effect of PLA2G10 on the gut microbial community, a linear discriminant analysis (LDA) and its effect size (LEfSe) analysis were performed to identify the biomarkers in the microbiomes of the two groups (Figure 6A and Supplementary Figure 2A). A logarithmic LDA cutoff score of 3.0 was set to identify important taxonomic differences between the two groups. We then calculated the scores that reflected the effect size of each differentially enriched taxon from these results. The LDA value histogram for the samples at the genus level (Figure 6B) showed that the gut microbiota of the control group contained more Allobaculum and Desulfovibrio than that of the Pla2g10 KI group, whereas Dubosiella and Enterorhabdus were dominant in the Pla2g10 KI group. Taken together, these results suggest that Pla2g10 overexpression altered the abundances of the specific gut bacteria.
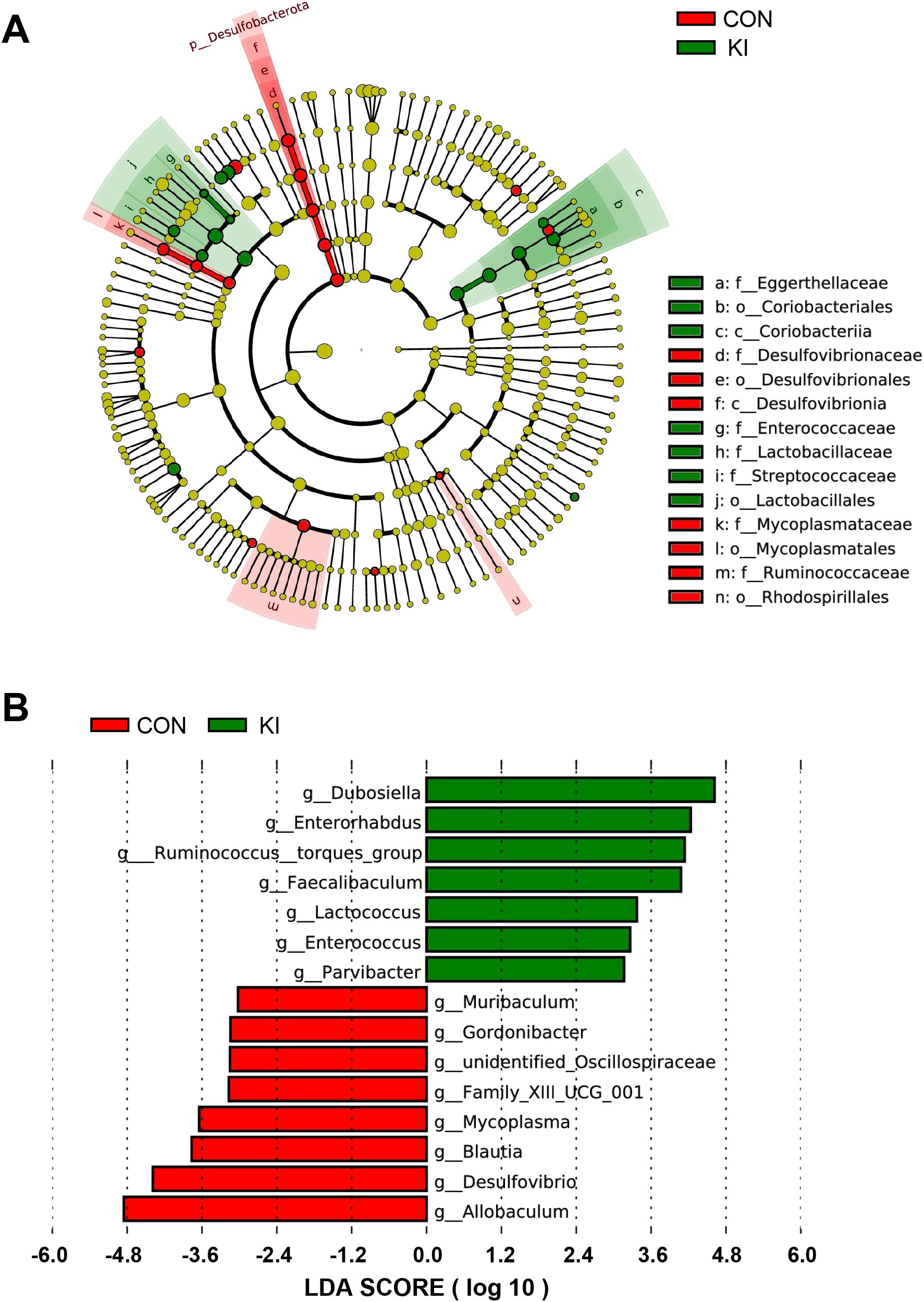
Figure 6. Analysis of the difference of gut microbiota between Pla2g10 KI and control groups. (A) Cladogram generated by linear discriminant analysis (LDA) effect size (LEfSe) method. (B) LDA score reflecting the effect size of each differentially enriched taxon at genus level. CON, control group; KI, Pla2g10 konck-in group.
To further investigate the changes in the metabolic functions of the gut microbiota associated with Pla2g10 overexpression, we performed a functional prediction analysis based on the KEGG orthology (KO) database, using the PICRUSt2 software. Combination of the PICRUSt2 functional clustering heat map (Figure 7A) with a t test analysis (Supplementary Figure 2B) clearly showed that Pla2g10 overexpression significantly altered various functions of the gut microbiota in mice, and most of the differentially expressed functions were more abundant in the Pla2g10 KI mice than in the control mice, as shown in Supplementary Table 2. We then constructed a network plot of the 30 most differentially abundant functions identified in the KEGG analysis and the 10 most abundant genera in both groups of mice. The results revealed that the main genera whose functions were affected by Pla2g10 overexpression in mice were Dubosiella, Lactobacillus, X Ruminococcus_torques_group, and Allobaculum (Figure 7B).

Figure 7. Functional prediction analysis of the gut microbiota based on the KEGG orthology database. (A) PICRUSt2 functional clustering heatmap of top 30 KO between two groups. (B) Network plot of the 30 most differentially abundant functions identified in the KEGG analysis and the 10 most abundant genera in both groups of mice. C, control group; KI, Pla2g10 konck-in group.
In the current study, the value of body weight decreased, while value of food intake increased in the Pla2g10 KI group compared with control group, however neither of them reached significance. It is generally believed that a high Firmicutes/Bacteroidetes (F/B) ratio improves energy absorption efficiency, leading to obesity related functional disorders (Murphy et al., 2010). The current study showed that the F/B ratio in the Pla2g10 KI mice showed an increasing trend compared the control mice. Thus, the change of gut microbiota might play a role in the regulation of host nutrient utilization. However, further study will be performed to confirmed the effect of intestinal Pla2g10 overexpression on nutrition metabolism.
Many recent studies have demonstrated that host’s genes significantly affect the diversity and functions of the gut microbiota, the abundance of specific taxa, and the genetic diversity of gut microorganisms (Lopera-Maya et al., 2022; Wang et al., 2016; Yang et al., 2022; Zhernakova et al., 2024). And in the present study, Pla2g10 overexpression reduced the abundance of Allobaculum, but significantly increased the abundance of other beneficial bacteria, such as Dubosiella, Enterorhabdus, Lactococcus and Lactobacillus (Figure 5). A recent study showed that deficiency of Pla2g10 decreased the abundance of several Clostridium species, and reduced levels of SCFAs, while these phenotypes could be reversed by dietary supplementation with ω-3 PUFAs (Sato et al., 2024). Interestingly, in this study, the abundance of SCFAs-producing bacteria decreased by Pla2g10 knockout, while in our study, the abundance of SCFAs-producing bacteria like Dubosiella increased by Pla2g10 knock-in. This indicated that host PLA2G10 might increase the Dubosiella abundance through producing ω-3 PUFAs.
The analysis of α-diversity indicated that Pla2g10 overexpression significantly reduced the quantity and richness of the microbial community. This supports our speculation that other gut microbes compensated for the gut imbalance in the Pla2g10-overexpressing mice. The abundance of Actinobacteria in the gut of Pla2g10 KI mice had an upward trend compared to the control group. Actinobacteria, like Firmicutes, are Gram-positive bacteria but have a higher GC content (Belizário and Napolitano, 2015). As shown in Figure 5B, the genus Enterorhabdus, which belongs to the phylum Actinobacteria, was significantly increased in the Pla2g10 KI mice, which may explain the high abundance of Actinobacteria. It has been reported that Enterorhabdus plays a positive role in intestinal homeostasis and is negatively correlated with intestinal diseases such as ulcerative colitis and inflammatory bowel disease (IBD) (Cheng et al., 2023; Liu et al., 2022). Enterorhabdus is also closely involved in the regulation of glucose metabolism, as well as bile acid, fatty acid, and amino acid production by the gut microbiota in mice (Li et al., 2023; Wang et al., 2022). Therefore, PLA2G10 might also regulate host nutrient utilization through gut Enterorhabdus. However this will be further studied.
Dubosiella is a bacterium that produces short-chain fatty acids (SCFAs). It correlates positively with the production of butyrate and mediates the protective effect of movement on the development of nonalcoholic fatty liver disease (Wan et al., 2021; Ye et al., 2023). There has been relatively little research on Dubosiella before, but in recent years, a new species, Dubosiella newyorkensis, has been discovered, which balances the Treg/Th17 response through the production of propionic acid and L-lysine, thus exerting an immune regulatory effect on the gut (Zhang et al., 2024). It also has antioxidant and anti-aging capabilities superior to those of resveratrol, and improves the gut microbiota by increasing beneficial bacteria, such as Lactobacillus, Bifidobacterium, and Akkermansia (Liu T. H. et al., 2023). In our study, Dubosiella and Lactobacillus are positively correlated with the expression of various functional genes present in microbes, and intestinal Pla2g10 overexpression increased the abundance of these two genera, indicating that the differences in functional gene expression of the gut microbiota in Pla2g10 KI mice are mainly attributable to Dubosiella and Lactobacillus. Considering Dubosiella’s potential ability to regulate beneficial bacteria, we speculate that PLA2G10 regulates the abundance of microbes such as Lactobacillus by stimulating the proliferation of Dubosiella.
Similar to Dubosiella, Allobaculum can physiologically utilize carbohydrates to produce butyrate (Greetham et al., 2004). It has also been reported that Allobaculum constitutes the greatest proportion of bacteria in the gut microbiota, accounting for 40.42% of the total number of bacteria at the genus level (Wang et al., 2018), which is consistent with the results obtained in this study. Two contradictory conclusions have been drawn from research on Allobaculum. Some researchers believe that it can stimulate the proliferation of epithelial cells and improve the integrity of the intestinal barrier by producing volatile fatty acids (Fan et al., 2015), whereas another report showed that Allobaculum isolated from patients with IBD exacerbated colitis in mice and caused antigen-specific mucosal and systemic antibody responses (Rice et al., 2022). The role of Allobaculum the current study needs further study to be illustrated.
As mentioned earlier, there may be receptors for PLA2G10 on gut microbiota, and the PC has also been found to be exposed on the surfaces of some microorganisms, such as Streptococcus pneumoneae (de Faire and Frostegård, 2009), and these substances can be enzymatic targets of PLA2G10 secreted into the intestinal lumen. Consequently, we suspect that the changes in the gut microbiota caused by the overexpression of Pla2g10 occur in multiple ways, but how PLA2G10 exerts its regulatory effect on the gut microbiota requires further research.
Although our experiments have demonstrated the effects of PLA2G10 on the composition and function of the gut microbiota, we did not determine the PUFAs levels in the intestinal lumen. So, it is uncertain whether PLA2G10 affects the composition of microbiota through the production of PUFA. Further sequencing of the gut microbiota is required after single-bacterial culture to detect the presence of PLA2G10 receptors on microorganisms.
In summary, overexpression of Pla2g10 altered the composition of gut microbiota in mice. Although the α-diversity of the microbiome reduced in the Pla2g10 knock-in mice, the abundance of beneficial microbes such as Enterorhabdus, Dubosiella and Lactobacillus increased, so our study provides a therapeutic target for the improvement of the gut microbiota in the future.
The original contributions presented in the study are publicly available. This data can be found here: NCBI PRJNA1222569.
The animal study was approved by the Institutional Animal Care and Use Committee at Sichuan Agricultural University. The study was conducted in accordance with the local legislation and institutional requirements.
WL: Writing – original draft, Data curation, Formal analysis, Investigation, Methodology. LCa: Data curation, Methodology, Writing – review & editing. XJ: Writing – review & editing, Data curation, Formal analysis. LCh: Writing – review & editing, Data curation, Investigation. ZF: Writing – review & editing, Investigation. SX: Writing – review & editing, Investigation. YL: Writing – review & editing, Investigation, Methodology. YZ: Writing – review & editing, Formal analysis, Methodology. JL: Writing – review & editing, Methodology. GL: Writing – review & editing, Formal analysis, Investigation. MS: Writing – review & editing, Formal analysis, Methodology. DW: Writing – review & editing, Conceptualization, Supervision. HW: Writing – review & editing, Funding acquisition, Validation, Visualization. BF: Writing – review & editing, Conceptualization, Funding acquisition, Project administration, Resources, Supervision, Validation, Visualization. LC: Data curation, Methodology, Writing – review & editing. LH: Formal analysis, Methodology, Writing – review & editing.
The author(s) declare that financial support was received for the research and/or publication of this article. This study was supported by Fok Ying Tung Education Foundation (171019) and Research Foundation from the Giant Panda Breeding (2020CPB-C10).
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fcimb.2025.1535204/full#supplementary-material
Belizário, J. E., Napolitano, M. (2015). Human microbiomes and their roles in dysbiosis, common diseases, and novel therapeutic approaches. Front. Microbiol. 6. doi: 10.3389/fmicb.2015.01050
Bezzine, S., Koduri, R. S., Valentin, E., Murakami, M., Kudo, I., Ghomashchi, F., et al. (2000). Exogenously added human group X secreted phospholipase A(2) but not the group IB, IIA, and V enzymes efficiently release arachidonic acid from adherent mammalian cells. J. Biol. Chem. 275, 3179–3191. doi: 10.1074/jbc.275.5.3179
Bokulich, N. A., Subramanian, S., Faith, J. J., Gevers, D., Gordon, J. I., Knight, R., et al. (2013). Quality-filtering vastly improves diversity estimates from Illumina amplicon sequencing. Nat. Methods 10, 57–59. doi: 10.1038/nmeth.2276
Cao, W., Wang, C., Chin, Y., Chen, X., Gao, Y., Yuan, S., et al. (2019). DHA-phospholipids (DHA-PL) and EPA-phospholipids (EPA-PL) prevent intestinal dysfunction induced by chronic stress. Food Funct. 10, 277–288. doi: 10.1039/c8fo01404c
Chen, Y., Zhou, J., Wang, L. (2021). Role and mechanism of gut microbiota in human disease. Front. Cell Infect. Microbiol. 11. doi: 10.3389/fcimb.2021.625913
Cheng, H., Zhang, D., Wu, J., Liu, J., Tan, Y., Feng, W., et al. (2023). Atractylodes macrocephala Koidz. volatile oil relieves acute ulcerative colitis via regulating gut microbiota and gut microbiota metabolism. Front. Immunol. 14. doi: 10.3389/fimmu.2023.1127785
Clemente, J. C., Manasson, J., Scher, J. U. (2018). The role of the gut microbiome in systemic inflammatory disease. Bmj 360, j5145. doi: 10.1136/bmj.j5145
de Faire, U., Frostegård, J. (2009). Natural antibodies against phosphorylcholine in cardiovascular disease. Ann. N. Y. Acad. Sci. 1173, 292–300. doi: 10.1111/j.1749-6632.2009.04748.x
Dennis, E. A., Cao, J., Hsu, Y. H., Magrioti, V., Kokotos, G. (2011). Phospholipase A2 enzymes: physical structure, biological function, disease implication, chemical inhibition, and therapeutic intervention. Chem. Rev. 111, 6130–6185. doi: 10.1021/cr200085w
Durack, J., Lynch, S. V. (2019). The gut microbiome: Relationships with disease and opportunities for therapy. J. Exp. Med. 216, 20–40. doi: 10.1084/jem.20180448
Edgar, R. C., Haas, B. J., Clemente, J. C., Quince, C., Knight, R. (2011). UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27, 2194–2200. doi: 10.1093/bioinformatics/btr381
Fan, P., Li, L., Rezaei, A., Eslamfam, S., Che, D., Ma, X. (2015). Metabolites of dietary protein and peptides by intestinal microbes and their impacts on gut. Curr. Protein Pept. Sci. 16, 646–654. doi: 10.2174/1389203716666150630133657
Feng, R., Ma, L. J., Wang, M., Liu, C., Yang, R., Su, H., et al. (2020). Oxidation of fish oil exacerbates alcoholic liver disease by enhancing intestinal dysbiosis in mice. Commun. Biol. 3, 481. doi: 10.1038/s42003-020-01213-8
Greetham, H. L., Gibson, G. R., Giffard, C., Hippe, H., Merkhoffer, B., Steiner, U., et al. (2004). Allobaculum stercoricanis gen. nov., sp. nov., isolated from canine feces. Anaerobe 10, 301–307. doi: 10.1016/j.anaerobe.2004.06.004
Hanasaki, K., Ono, T., Saiga, A., Morioka, Y., Ikeda, M., Kawamoto, K., et al. (1999). Purified group X secretory phospholipase A(2) induced prominent release of arachidonic acid from human myeloid leukemia cells. J. Biol. Chem. 274, 34203–34211. doi: 10.1074/jbc.274.48.34203
Jemel, I., Ii, H., Oslund, R. C., Payré, C., Dabert-Gay, A. S., Douguet, D., et al. (2011). Group X secreted phospholipase A2 proenzyme is matured by a furin-like proprotein convertase and releases arachidonic acid inside of human HEK293 cells. J. Biol. Chem. 286, 36509–36521. doi: 10.1074/jbc.M111.268540
Kudo, K., Miki, Y., Carreras, J., Nakayama, S., Nakamoto, Y., Ito, M., et al. (2022). Secreted phospholipase A(2) modifies extracellular vesicles and accelerates B cell lymphoma. Cell Metab. 34, 615–633.e618. doi: 10.1016/j.cmet.2022.02.011
Layne, J. D., Shridas, P., Webb, N. R. (2015). Ectopically expressed pro-group X secretory phospholipase A2 is proteolytically activated in mouse adrenal cells by furin-like proprotein convertases: implications for the regulation of adrenal steroidogenesis. J. Biol. Chem. 290, 7851–7860. doi: 10.1074/jbc.M114.634667
Li, Y., Zhu, Y., Li, D., Liu, W., Zhang, Y., Liu, W., et al. (2023). Depletion of gut microbiota influents glucose metabolism and hyperandrogenism traits of mice with PCOS induced by letrozole. Front. Endocrinol. (Lausanne). 14. doi: 10.3389/fendo.2023.1265152
Liu, T. H., Wang, J., Zhang, C. Y., Zhao, L., Sheng, Y. Y., Tao, G. S., et al. (2023). Gut microbial characteristical comparison reveals potential anti-aging function of Dubosiella newyorkensis in mice. Front. Endocrinol. (Lausanne). 14. doi: 10.3389/fendo.2023.1133167
Liu, B., Ye, D., Yang, H., Song, J., Sun, X., Mao, Y., et al. (2022). Two-sample mendelian randomization analysis investigates causal associations between gut microbial genera and inflammatory bowel disease, and specificity causal associations in ulcerative colitis or crohn’s disease. Front. Immunol. 13. doi: 10.3389/fimmu.2022.921546
Lopera-Maya, E. A., Kurilshikov, A., van der Graaf, A., Hu, S., Andreu-Sánchez, S., Chen, L., et al. (2022). Effect of host genetics on the gut microbiome in 7,738 participants of the Dutch Microbiome Project. Nat. Genet. 54, 143–151. doi: 10.1038/s41588-021-00992-y
Magoč, T., Salzberg, S. L. (2011). FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963. doi: 10.1093/bioinformatics/btr507
Masuda, S., Murakami, M., Takanezawa, Y., Aoki, J., Arai, H., Ishikawa, Y., et al. (2005). Neuronal expression and neuritogenic action of group X secreted phospholipase A2. J. Biol. Chem. 280, 23203–23214. doi: 10.1074/jbc.M500985200
Mujico, J. R., Baccan, G. C., Gheorghe, A., Díaz, L. E., Marcos, A. (2013). Changes in gut microbiota due to supplemented fatty acids in diet-induced obese mice. Br. J. Nutr. 110, 711–720. doi: 10.1017/s0007114512005612
Murakami, M., Sato, H., Taketomi, Y. (2023). Modulation of immunity by the secreted phospholipase A(2) family. Immunol. Rev. 317, 42–70. doi: 10.1111/imr.13205
Murase, R., Sato, H., Yamamoto, K., Ushida, A., Nishito, Y., Ikeda, K., et al. (2016). Group X secreted phospholipase A2 releases ω3 polyunsaturated fatty acids, suppresses colitis, and promotes sperm fertility. J. Biol. Chem. 291, 6895–6911. doi: 10.1074/jbc.M116.715672
Murphy, E. F., Cotter, P. D., Healy, S., Marques, T. M., O’Sullivan, O., Fouhy, F., et al. (2010). Composition and energy harvesting capacity of the gut microbiota: relationship to diet, obesity and time in mouse models. Gut 59, 1635–1642. doi: 10.1136/gut.2010.215665
Rice, T. A., Bielecka, A. A., Nguyen, M. T., Rosen, C. E., Song, D., Sonnert, N. D., et al. (2022). Interspecies commensal interactions have nonlinear impacts on host immunity. Cell Host Microbe 30, 988–1002.e1006. doi: 10.1016/j.chom.2022.05.004
Sato, H., Taketomi, Y., Murase, R., Park, J., Hosomi, K., Sanada, T. J., et al. (2024). Group X phospholipase A(2) links colonic lipid homeostasis to systemic metabolism via host-microbiota interaction. Cell Rep. 43, 114752. doi: 10.1016/j.celrep.2024.114752
Schewe, M., Franken, P. F., Sacchetti, A., Schmitt, M., Joosten, R., Böttcher, R., et al. (2016). Secreted phospholipases A2 are intestinal stem cell niche factors with distinct roles in homeostasis, inflammation, and cancer. Cell Stem Cell 19, 38–51. doi: 10.1016/j.stem.2016.05.023
Taketomi, Y., Miki, Y., Murakami, M. (2022). Old but new: group IIA phospholipase A(2) as a modulator of gut microbiota. Metabolites 12, 352. doi: 10.3390/metabo12040352
Tang, X., Wang, W., Hong, G., Duan, C., Zhu, S., Tian, Y., et al. (2021). Gut microbiota-mediated lysophosphatidylcholine generation promotes colitis in intestinal epithelium-specific Fut2 deficiency. J. BioMed. Sci. 28, 20. doi: 10.1186/s12929-021-00711-z
Wan, F., Han, H., Zhong, R., Wang, M., Tang, S., Zhang, S., et al. (2021). Dihydroquercetin supplement alleviates colonic inflammation potentially through improved gut microbiota community in mice. Food Funct. 12, 11420–11434. doi: 10.1039/d1fo01422f
Wang, Y., Guo, H., Gao, X., Wang, J. (2021). The intratumor microbiota signatures associate with subtype, tumor stage, and survival status of esophageal carcinoma. Front. Oncol. 11. doi: 10.3389/fonc.2021.754788
Wang, J., Thingholm, L. B., Skiecevičienė, J., Rausch, P., Kummen, M., Hov, J. R., et al. (2016). Genome-wide association analysis identifies variation in vitamin D receptor and other host factors influencing the gut microbiota. Nat. Genet. 48, 1396–1406. doi: 10.1038/ng.3695
Wang, S., Wang, S., Wang, X., Xu, Y., Zhang, X., Han, Y., et al. (2022). Effects of icariin on modulating gut microbiota and regulating metabolite alterations to prevent bone loss in ovariectomized rat model. Front. Endocrinol. (Lausanne). 13. doi: 10.3389/fendo.2022.874849
Wang, Z., Zhang, W., Wang, B., Zhang, F., Shao, Y. (2018). Influence of Bactrian camel milk on the gut microbiota. J. Dairy. Sci. 101, 5758–5769. doi: 10.3168/jds.2017-13860
Yang, H., Wu, J., Huang, X., Zhou, Y., Zhang, Y., Liu, M., et al. (2022). ABO genotype alters the gut microbiota by regulating GalNAc levels in pigs. Nature 606, 358–367. doi: 10.1038/s41586-022-04769-z
Ye, X., Sun, P., Lao, S., Wen, M., Zheng, R., Lin, Y., et al. (2023). Fgf21-Dubosiella axis mediates the protective effects of exercise against NAFLD development. Life Sci. 334, 122231. doi: 10.1016/j.lfs.2023.122231
Zhang, Y., Tu, S., Ji, X., Wu, J., Meng, J., Gao, J., et al. (2024). Dubosiella newyorkensis modulates immune tolerance in colitis via the L-lysine-activated AhR-IDO1-Kyn pathway. Nat. Commun. 15, 1333. doi: 10.1038/s41467-024-45636-x
Zhang, H., Zhang, Y., Mu, T., Cao, J., Liu, X., Yang, X., et al. (2023). Response of gut microbiota and ileal transcriptome to inulin intervention in HFD induced obese mice. Int. J. Biol. Macromol. 225, 861–872. doi: 10.1016/j.ijbiomac.2022.11.151
Zhernakova, D. V., Wang, D., Liu, L., Andreu-Sánchez, S., Zhang, Y., Ruiz-Moreno, A. J., et al. (2024). Host genetic regulation of human gut microbial structural variation. Nature 625, 813–821. doi: 10.1038/s41586-023-06893-w
Zhuang, P., Shou, Q., Lu, Y., Wang, G., Qiu, J., Wang, J., et al. (2017). Arachidonic acid sex-dependently affects obesity through linking gut microbiota-driven inflammation to hypothalamus-adipose-liver axis. Biochim. Biophys. Acta Mol. Basis. Dis. 1863, 2715–2726. doi: 10.1016/j.bbadis.2017.07.003
Keywords: PLA2G10, gut microbiota, intestine, Allobaculum, Dubosiella
Citation: Liao W, Cao L, Jiang X, Che L, Fang Z, Xu S, Lin Y, Zhuo Y, Hua L, Li J, Liu G, Sun M, Wu D, Wang H and Feng B (2025) Intestinal overexpression of Pla2g10 alters the composition, diversity and function of gut microbiota in mice. Front. Cell. Infect. Microbiol. 15:1535204. doi: 10.3389/fcimb.2025.1535204
Received: 27 November 2024; Accepted: 26 February 2025;
Published: 14 March 2025.
Edited by:
Sidharth Prasad Mishra, University of South Florida, United StatesReviewed by:
Amanda Carroll-Portillo, University of New Mexico, United StatesCopyright © 2025 Liao, Cao, Jiang, Che, Fang, Xu, Lin, Zhuo, Hua, Li, Liu, Sun, Wu, Wang and Feng. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Bin Feng, ZmVuZ2JpbkBzaWNhdS5lZHUuY24=; Hairui Wang, cGFuZGFoYXJyeUBwYW5kYS5vcmcuY24=
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.