
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Cell. Infect. Microbiol. , 03 April 2025
Sec. Bacteria and Host
Volume 15 - 2025 | https://doi.org/10.3389/fcimb.2025.1511141
This article is part of the Research Topic Advancing Understanding of Neonatal Bacterial Infections View all 6 articles
Objective: To investigate the specific characteristics and differences in the placental microbiome of preterm infants with and without pneumonia.
Methods: Fifty-nine infants born at 32–36 weeks’ gestation were enrolled in this study. Among them, 33 developed pneumonia within 48 hours of birth, while the remaining 26 did not. Placental swabs were collected at birth for DNA extraction, from which the placental microbial composition was analyzed using a bioinformatics pipeline following PCR amplification of genetic material and subsequent sequencing of bacterial 16S rRNA.
Results: Significant differences in both the alpha and beta diversities were found between the two groups (P<0.05). Proteobacteria, Firmicutes, and Actinobacteriota were identified as the predominant phyla in the placenta, while predominant identified genera included Brevundimonas, Caulobacter, Lactobacillus, and Citrobacter. There were no significant inter-group differences in the relative abundances of the predominant phyla and genera except Lactobacillus(P>0.05). Compared to infants without pneumonia, those with pneumonia demonstrated a decreased abundance of Lactobacillus, and an increased abundance of Ureaplasma and Staphylococcus (P<0.05). The relative abundance of Ureaplasma was positively correlated with that of Staphylococcus, and negatively correlation with that of Lactobacillus (P<0.05). Notably, we observed significant disparities in the metabolic pathways and phenotypes between the two groups (P<0.05).
Conclusion: Overall, this study suggests that alterations in the placental microbiome may be linked to the onset of pneumonia in preterm infants. Further investigations are required to elucidate the relationship between microbiota and disease pathogenesis.
Preterm birth, a common and serious complication of pathological pregnancies, is a significant contributor to neonatal mortality rates worldwide, exerting direct implications for population health outcomes (Bhandekar et al., 2024). Over one million premature births occur annually in China alone (Zhu et al., 2021). Advances in perinatal and neonatal care have led to noticeable improvements in survival rates among preterm infants (Wu et al., 2019); however, affected infants remain susceptible to various degrees of pulmonary injury, leading to heightened respiratory complications compared to their full-term counterparts (Huang et al., 2024). Pneumonia is an enduring consequence of premature birth, causing sustained lung damage, and jeopardizing the survival prospects of vulnerable newborns, consequently affecting their long-term quality of life. Management of preterm infants with pneumonia should include extended pulmonary rehabilitation (Wang et al., 2024). According to the life course theory, factors in various stages of human development can have long-term cumulative and interactive effects on both human development and disease. Changes in environmental factors during early life and pregnancy may also influence the occurrence of diseases in later life. The human microecosystem is defined as a complex network of the microbiotic communities and their habitats located within and on the body (Valentine et al., 2018). Evidence from prior studies has confirmed the presence of microorganisms in the placental tissue; however, it is important to note that these colonized organisms are not associated with perinatal infections (Piersigilli et al., 2020). Increasingly, studies have begun to characterize the dynamic microecosystem of pregnant women, which has been shown to undergo complex physiological changes closely linked to maternal and fetal health (Zheng et al., 2017a; La et al., 2022; Powell et al., 2024).
In recent years, a growing emphasis has been placed on the interplay between maternal, placental, and fetal microecologies. Vertical transmission of bacteria from the mother to fetus during pregnancy has also been scientifically validated, while the presence of symbiotic bacteria and lactic acid bacterial products in the meconium has been firmly established (Martin et al., 2016). The microbial colonization pattern in each neonate is unique, and subject to variability among individuals and anatomical locations (Wu et al., 2023). The human respiratory tract hosts a microbial community even before birth, while the airway microbiota of neonates delivered vaginally or via cesarean section are strikingly similar. It has thus been hypothesized that the fetal airway microbiota may have dual origins in both the placenta and amniotic fluid (Lal et al., 2016). However, research on the effects of placental microbiome on pneumonia in preterm infants remains limited.
To address this knowledge gap, the present study aimed to characterize and compare the placental microbiome of preterm infants with and without pneumonia.
Preterm infants born between 32 and 36 weeks’ gestational age at the Chengdu Women and Children’s Central Hospital between June and December 2021 were enrolled in this study. The exclusion criteria included the offspring of mothers aged ≥ 35 years, received antibiotics during pregnancy, with a history of vaginitis or immune diseases during pregnancy, or pregnancies complicated by internal and surgical diseases. Additionally, infants with congenital developmental malformations (including complex congenital heart disease and pulmonary dysplasia), inherited metabolic diseases, chromosomal diseases, and those whose parents did not provide consent were also excluded. The indications for Cesarean section were based on an expert consensus on Cesarean section surgery published in 2014 (Obstetrics Group of the Chinese Association of Obstetrics and Gynecology, the Chinese Association of Obstetrics and Gynecology, and the Chinese Medical Association, 2014).
Apgar scores were determined by a qualified healthcare professional, in accordance with established protocols. An Apgar score of less than 7 was defined as low (American Academy of Pediatrics Committee on Fetus and Newborn and American College of Obstetricians and Gynecologists Committee on Obstetric Practice, 2015).Neonatal pneumonia was defined in accordance with the relevant clinical guidelines (Nissen, 2007).
The study protocol was approved by Chengdu Women and Children’s Central Hospital (2021 [123]). Written informed consent was obtained from the parents or guardians of the children who participated in this study. The study was conducted in accordance with The Code of Ethics of the World Medical Association (Declaration of Helsinki).
Immediately following preterm delivery, aseptic swabs were obtained by gently scraping the fetal surface of the placenta three times in sterile manner. Collected samples were promptly stored in a -80°C refrigerator within 30 minutes.
DNA extraction kits from Omega Bio-Tek (Madison, WI United) were used to ensure effective and high-quality DNA extraction. Following extraction, a target fragment library was constructed, sequencing PCR primers targeting conserved regions within the 16S rRNA gene were designed. Amplification of V3-V4 regions within the 16S rRNA gene was achieved through PCR utilizing the following primer pair: forward primer -341F (5’-CCTACGGGNGGCWGCAG-3’) and reverse primer -806R(5’-GACTACHVGGGTATCTAATCC-3’). Following thirty-five cycles of PCR amplification, sequencing adapters and barcodes were incorporated into the samples before detection by agarose gel electrophoresis at a concentration of 1.5%. Target fragments underwent recovery utilizing AxyPrep™PCR Cleanup Kits from Beckman Coulter Genomics based in Danvers Massachusetts, while purification involved employing the Quant-iT PicoGreen dsDNA Assay Kit from Invitrogen in America.
Additionally, Amplicon pools were prepared prior to assessment for quality assessment using both an Agilent 2100 Bioanalyzer and a Library Quantification Kit specifically designed for Illumina by Kapa Biosciences located in Woburn Massachusetts. The final step involved subjection of the library to sequencing on an Illumina NovaSeq PE250 platform, which resulted in the generation of paired-end reads measuring at 250 base pairs.
Paired-end reads were assigned to samples based on their unique barcodes, and were truncated by removing the barcode and primer sequences. Subsequently, the paired-end reads were merged using FLASH (V1.2.8). Quality filtering of the raw reads was conducted under specific conditions to obtain high-quality clean tags, following the QIIME (V1.9.1) quality control process. Chimeric sequences were filtered using Vsearch (V2.3.4). Effective tags were further obtained after dereplication using a divisive amplicon-denoising algorithm. Alpha and beta diversities were further calculated using QIIME2 (V.2019.7). An equal number of sequences was also randomly extracted by reducing the sequence count to match that of the sample with the fewest sequences, followed by calculation of the relative abundance (X bacterial count/total count) for bacterial taxonomy analysis. Images were generated using R software (V3.5.2). Species annotation sequence alignment was performed using BLAST with the SILVA and NT-16S alignment databases. Fisher’s exact test was applied to assess the differences in species at each taxonomic level.
Measurement data conforming to normal distribution are presented as x ± s, while non-normally distributed measurement data are presented as P50 (P25–P75). Comparisons between the two groups were conducted using the t-test and Mann-Whitney U test. The chi-square test was applied to compare count data. Spearman’s rank correlation analysis was used to examine the associations between different bacterial genera. All statistical tests were two-sided, and the results were considered statistically significant at P < 0.05. SAS 9.3 (SAS Institute Inc., Cary, NC, USA) was used for the data analysis.
A total of 59 preterm infants with gestational ages ranging from 32 to 35 weeks were enrolled in this study. Among them, 33 developed pneumonia within 48 hours of birth while the remaining 26 did not. The cohort comprised 40 male and 19 female infants (Table 1). There were 11 instances of spontaneous labor and 48 cesarean sections. Participants who developed pneumonia within 48 h after birth were assigned to Group A, whereas those who did not develop pneumonia were assigned to Group B. There were no significant differences in gestational age, 1-minute Apgar score, body weight, body length, chest circumference, or abdominal circumference between the two groups (P>0.05). However, the head circumference of infants in Group A was smaller than that of infants in Group B (P=0.026). Compared with Group B, Group A also tended to have longer hospital stay (30.24 + 17.98 days VS 9.04 + 6.0 days, P<0.001). There was no significant difference in the cesarean section rate between the two groups (84.45% VS 76.92%, P>0.05). However, in Group A, the most prevalent indications for cesarean section were fetal distress and severe preeclampsia, whereas in Group B, severe intrahepatic cholestasis of pregnancy and IVF-ET were the most prevalent indications. White blood cell counts within 24 hours after birth, neutrophils and eosinophils remained within the normal range. The difference in C-reactive protein between the two groups was statistically significant (1.0(1.0∼7.1)ng/ml VS 1.0(1.0∼1.0)ng/ml P < 0.05).
A higher richness of microbial populations was found in the APS (A placental swab) group (P<0.05) (Figures 1A, B). The sequencing depth exhibited statistically significant differences between the two groups, as measured by Good’s coverage (P<0.05, Figure 1C). The evenness of the species distribution determined by Pielou-e showed no significant differences between the two samples (P>0.05, Figure 1D).
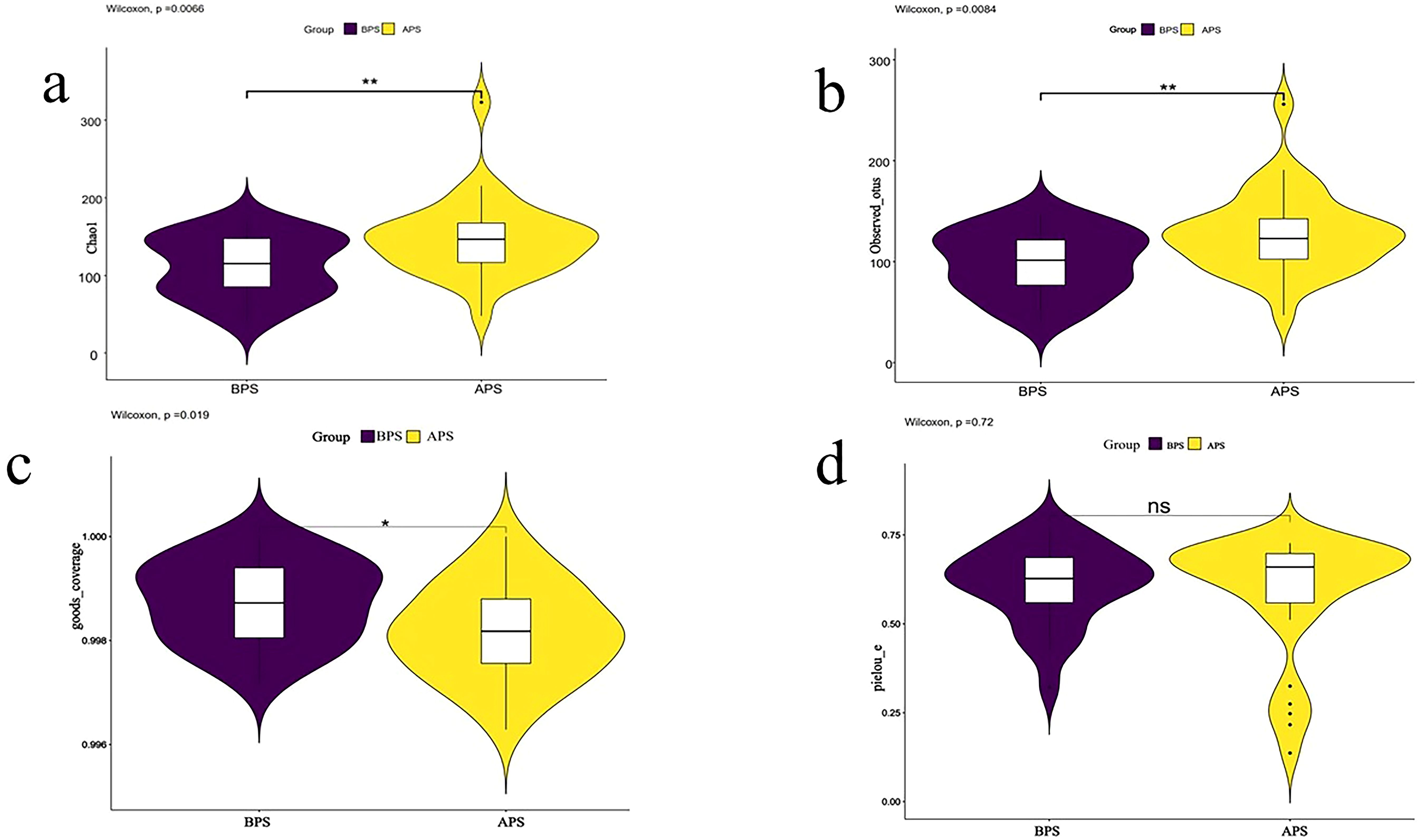
Figure 1. Results of alpha diversity analysis. Two groups of placenta swab 16S rRNA detection were divided into two groups: A placenta swab (APS) and B placenta swab (BPS). Alpha diversity is applied in analyzing complexity of species diversity for a sample through 4 indices, including (A) Chao1, (B)observed-species, (C) Goods_coverage, (D) pielou-e. Community richness can be assessed through the Chao1 and observed_species. Compared to BPS. *p<0.05; **p<0.01.
The ecological distances differed significantly between APS and BPS (B placental swab) by principal coordinate analysis (PCoA), as well as by nonmetric multidimensional scaling (NMDS) (P<0.05) (Figures 2A, B). Analysis of similarity (ANOSIM) is a non-parametric test widely applied to compare differences between groups with differences within groups. The ANOSIM results showed that the similarities within the groups were greater than those between the groups (P<0.05).
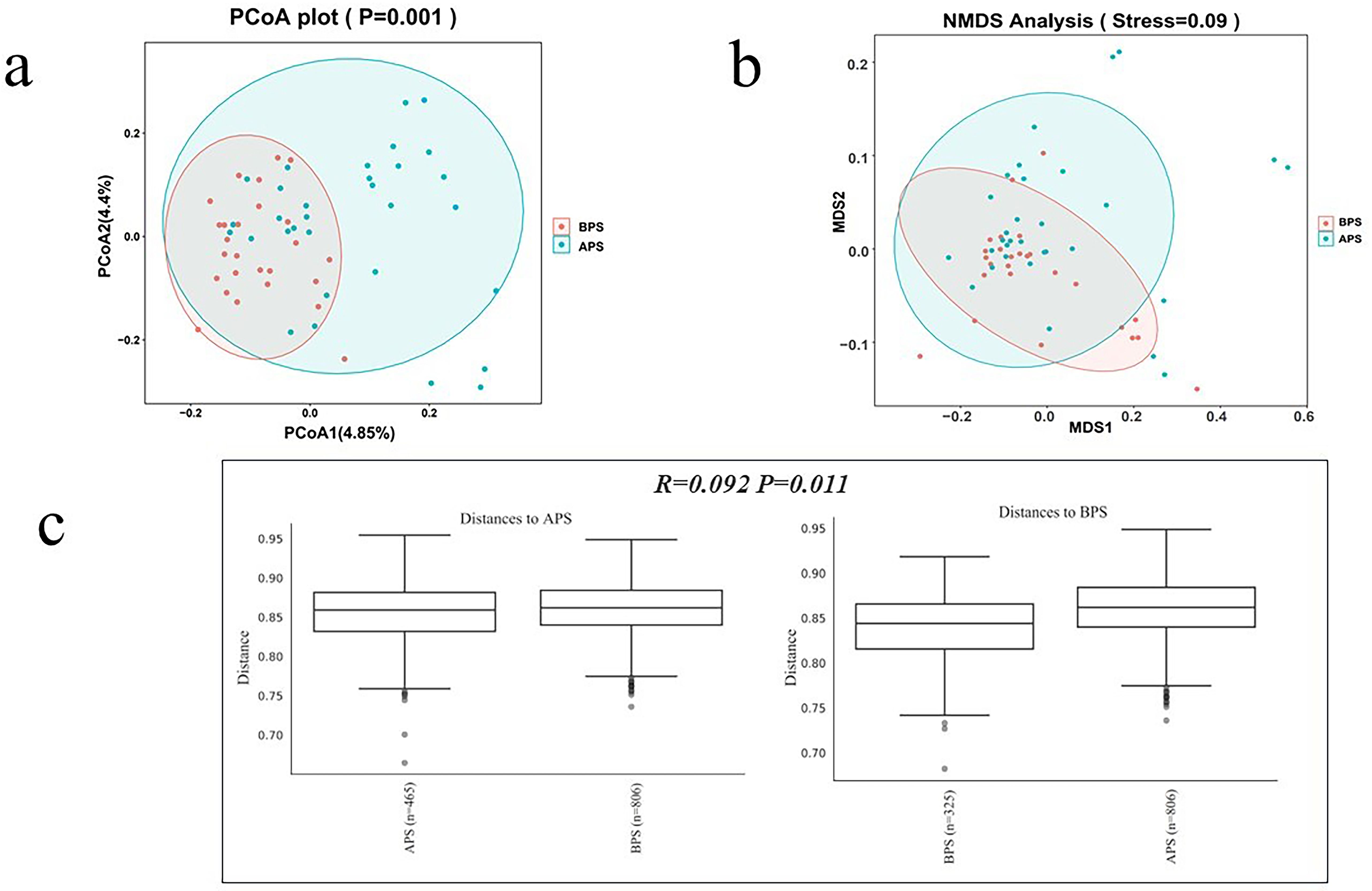
Figure 2. Bacterial beta diversity analysis. (a) PCoA analysis, (b) NMDS analysis, (c) ANOSIM analysis.
Beta diversity analysis was applied to evaluate the differences in species complexity. PCoA was performed to obtain the principal coordinates and to visualize complex multidimensional data. Non-metric multidimensional scaling (NMDS) is an indirect gradient analysis approach that produces an ordination based on a distance or dissimilarity matrix. The results of these analyses are presented in Figure 2. In Figure 2A, points of the same color denote they are from a common group, presented as a circle plot (≥4 biologic replicates per group), with 95% confidence intervals. Axis 1 represents the principal coordinate that explains the largest data change, while axis 2 represents the coordinate that accounts for the largest proportion of the remaining data changes. In Figure 2B, the spatial distance between sample points represents the difference between the samples. The axes were essentially arbitrary but displayed data in a manner that best represented their dissimilarity. Following the initial ordination, it was imperative to analyze the stress values generated by the algorithm. Stress values below 0.1 were considered to indicate good ordination with little risk of misinterpretation (Figure 2B). In Figure 2C, the test statistic R was bounded within the range of -1 to 1. Positive values indicate greater similarities within groups, whereas values equal to zero suggest no between-group differences or within-group similarities. Negative values suggest greater similarities between groups than within groups, potentially leading to incorrect sample assignments. In this study, the R-value was 0.092 (P<0.05).
The microbiota of the APS and BPS were analyzed, encompassing a wide taxonomic range from phyla to genera. Proteobacteria, Firmicutes, and Actinobacteriota were the dominant phyla in both groups (Figure 3A), with discernible differences observed in dominant phyla between the two groups. At the genus level, the top five dominant genera in APS were Brevundimonas, Caulobacter, Lactobacillus, Citrobacter, and Ureaplasma, while those in the BPS were Caulobacter, Lactobacillus, Citrobacter, Phenylobacterium, and Brevundimonas (Figure 3B).
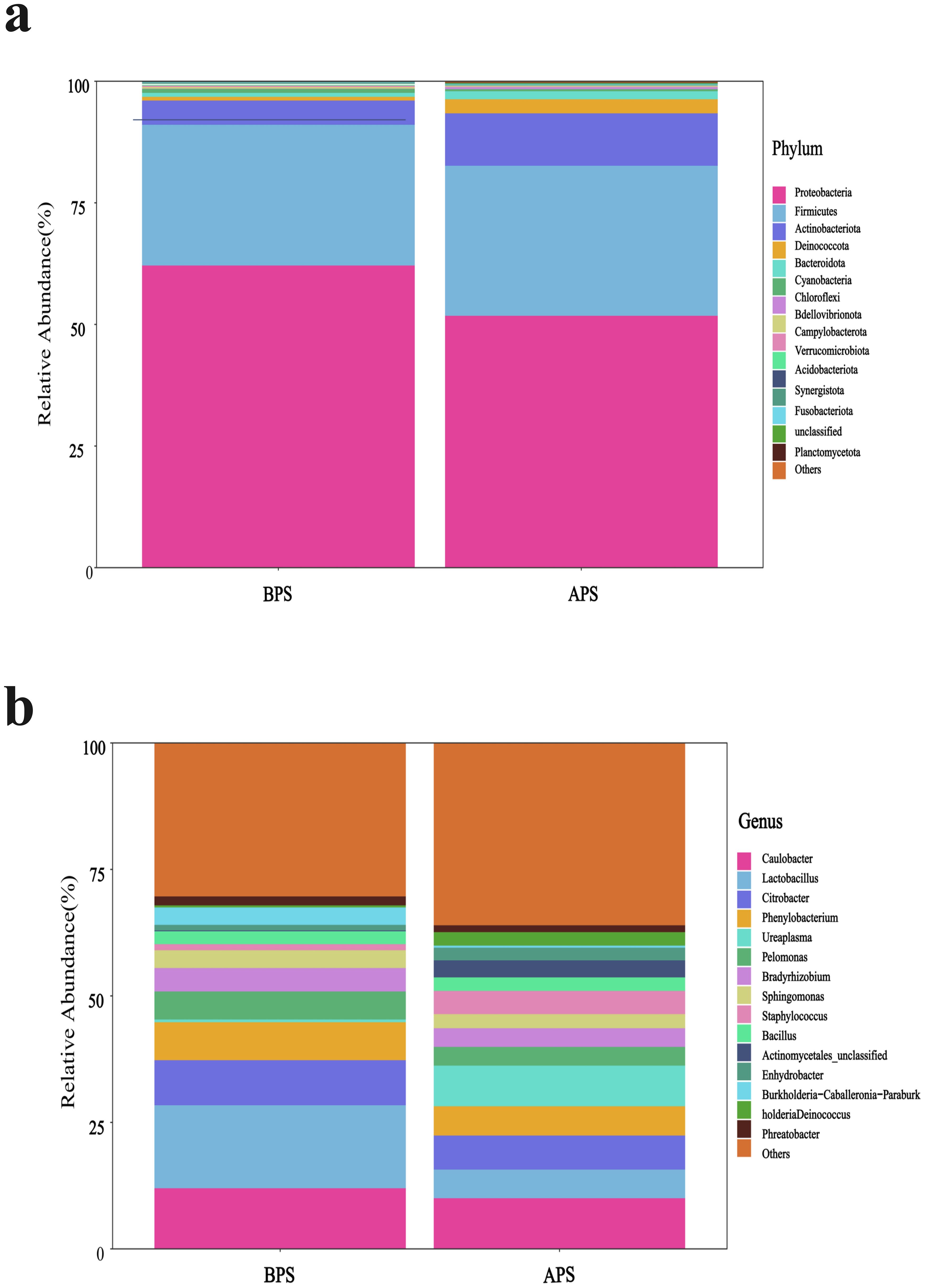
Figure 3. Microbiota composition, shown as the relative abundance plot of the top 15 species from placental swabs at the phylum (A) and genus levels (B).
There were no significant inter-group differences in the relative abundances of the predominant phyla and genera except Lactobacillus(P>0.05).Compared to BPS, we observed a decrease in Lactobacillus, and an increase in Ureaplasma and Staphylococcus in APS, all of which belonged to Firmicutes (P<0.05) (Figure 4A). Fisher’s exact test was further applied to examine the disparity in species at the genus level, which revealed a significant reduction in Lactobacillus and a notable increase in Staphylococcus in APS(P < 0.05) (Figure 4B).
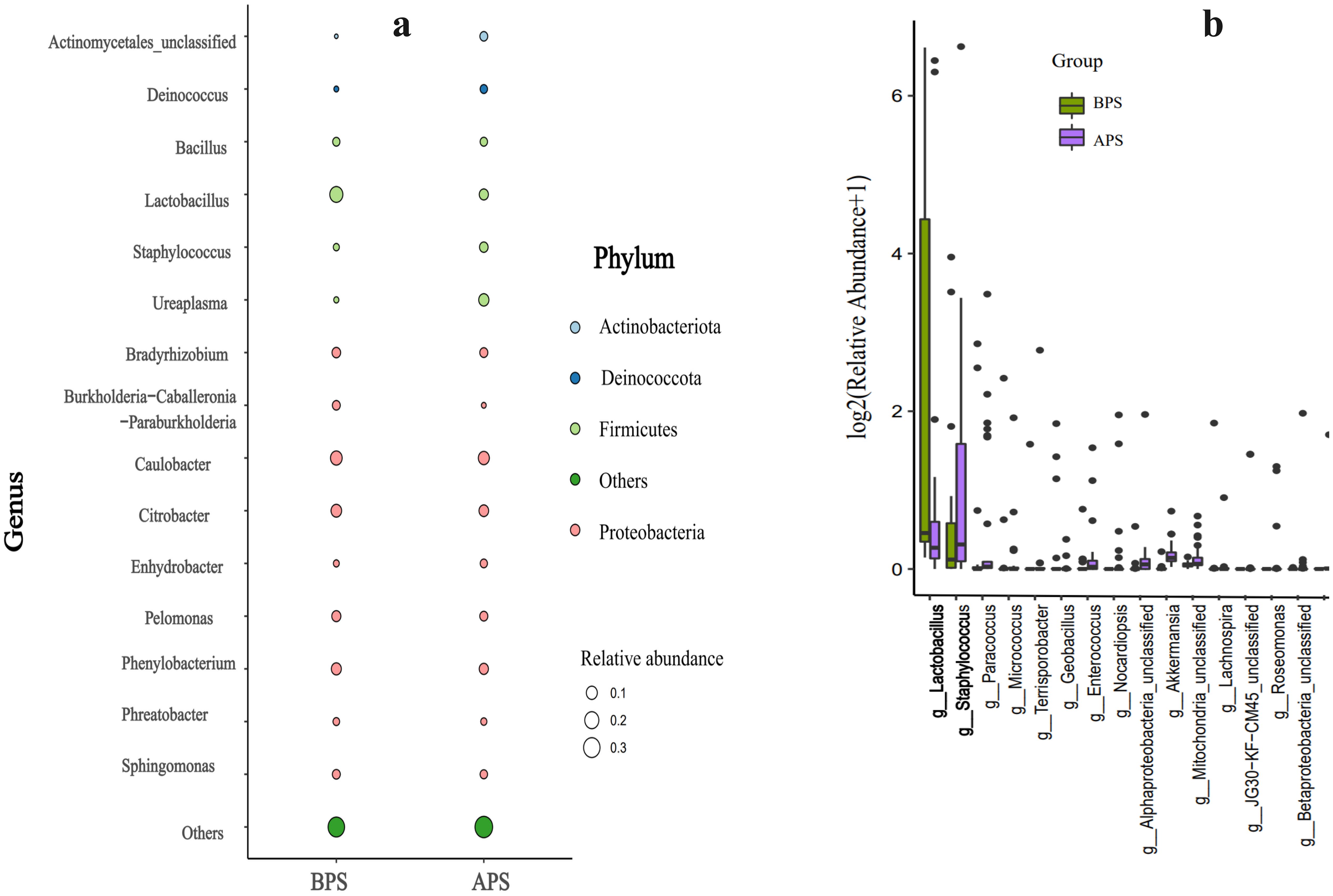
Figure 4. Results of microbiota significant difference analysis .Taxa significant difference analysis at the phylum (a) and genus (b) levels.
The distribution of taxa at the phylum and genus levels in APS and BPS is displayed in Figure 4A. Differences between APS and BPS at the genus level, and the top 15 species (P < 0.05) were selected for display in the bar chart in Figure 4B.
Significant rank correlations were observed among the dominant genera. Notably, relative abundance of Ureaplasma positively correlated with that of Staphylococcus, and negatively correlated with that of Lactobacillus(P < 0.05) (Figure 5).
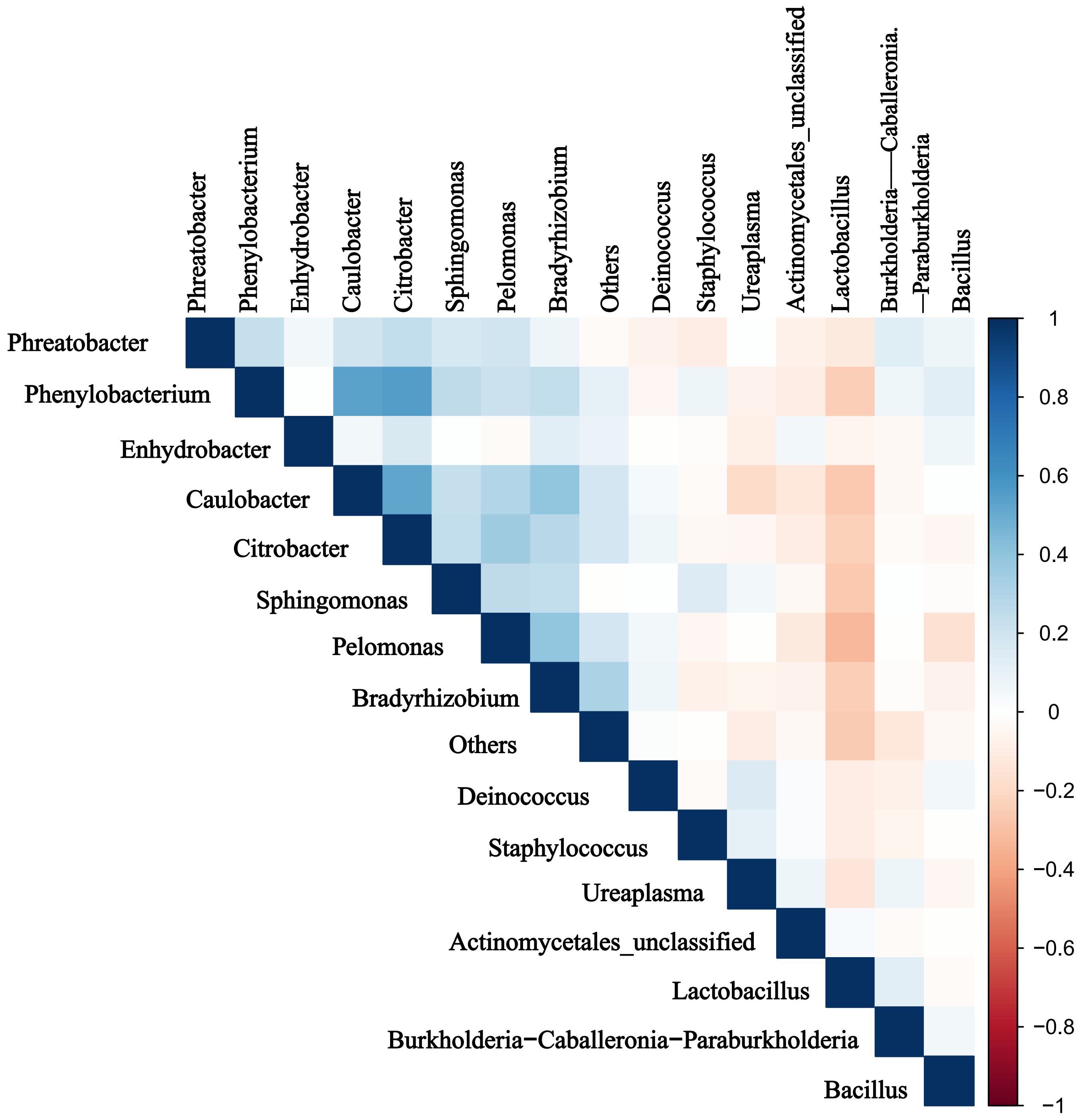
Figure 5. Results of the microbiota correlation analysis. Correlation heatmap of the top 15 dominant genera. Blue and red represent positive and negative correlations, respectively.
Compared to BPS, the metabolic pathways related to glycogen degradation I (bacterial), the superpathway of menaquinol biosynthesis, the superpathway of hexitol degradation (bacteria), adenine and adenosine salvage, enterobactin biosynthesis, and glycogen degradation I (bacterial) were all found to be enriched in APS using PICRUS, based on the Clusters of Orthologous Genes (COG) database (Figure 6A). BugBase is a microbiome analysis tool that determines high-level phenotypes present in microbiome samples. Overall, there were nine identified phenotypes associated with aerobic, anaerobic, facultatively anaerobic, potentially pathogenic, stress tolerance, containing mobile element, biofilms formation, gram-negative bacteria and gram-positive bacteria. The dominant phenotypes in APS were aerobic, biofilms formation and gram negative at phylum level, so were in BPS. The relative abundances of the anaerobic and mobile element phenotypes were significantly different between the two groups (P<0.05) (Figure 6B).
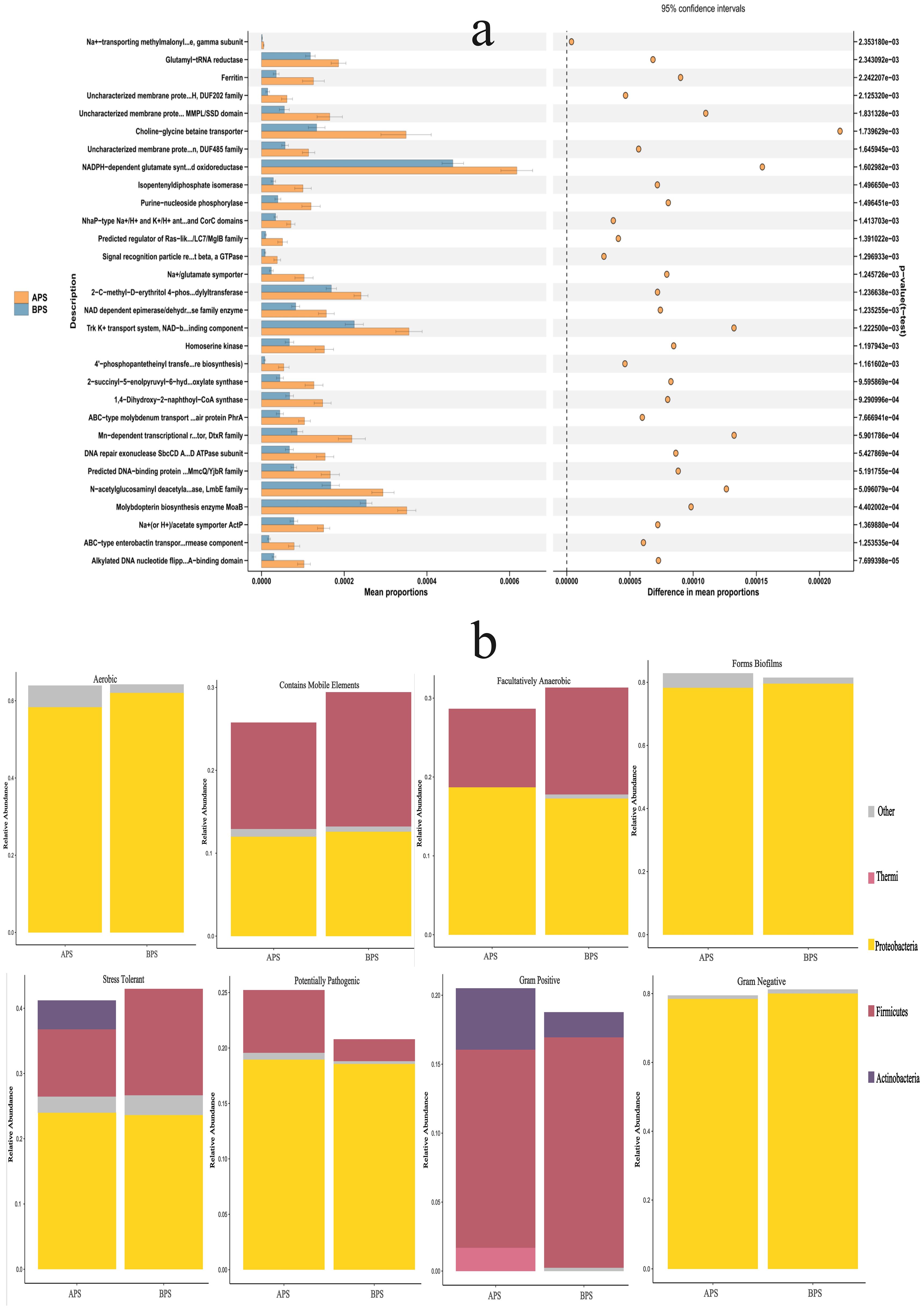
Figure 6. Results of functional prediction and phenotypes analysis. (a) Functional prediction of PICRUS, (b) Phenotypes analysis of BugBase.
The top 30 metabolic functions with a p-value <0.05 are shown in Figure 6A, while the relative abundances of phyla showing different phenotypes are presented in Figure 6B.
In this study, the duration of hospital stay in group A was found to be prolonged, while infants diagnosed with pneumonia showed an elevated risk of requiring ventilator support. No significant differences were observed between the two groups in terms of gestational age, sex, birth weight, body length, or 1-minute Apgar score. The distribution of reasons for Cesarean section differed significantly between the two groups. The incidence of premature rupture of membranes and severe eclampsia increased in Group A, with the former identified as a risk factor for preterm birth. Neonatal infections and pneumonia have been shown to occur in infants born to mothers with chorioamnionitis (Jiang et al., 2018). One study investigating the bacterial communities in the placental tissues of 1391 women demonstrated that pregnant women with severe chorioamnionitis have an increased bacterial load and reduced species richness in the placenta (Prince et al., 2016). Sneathia sanguinengens and Peptostreptococcus anaerobius were also found to be associated with a smaller infant size. Further, another study observed dysbiosis of the gut microbiota in preeclamptic patients, which was characterized by significant reductions in the bacteria that produce short-chain fatty acids (Jin et al., 2022). Overall, these results suggest that gut microbiota dysbiosis plays an important role in preeclampsia development.
In the present study, the top three phyla identified in the placental microbiota of preterm infants were Proteobacteria, Firmicutes, and Actinobacteriota. This finding aligns closely with that of a previous study, which also identified Proteobacteria, Actinobacteriota, and Bacteroidetes as the predominant phyla (Aagaard et al., 2014). We hypothesized that the minor discrepancies between the two studies could be attributed to variations in ethnicity and regional factors. The relative abundances of the dominant phyla showed no significant inter-group differences. Further, Brevundimonas, Caulobacter, Lactobacillus, Citrobacter were all found to be prevalent at the genus level in both groups. These results differ from those of other studies (Seferovic et al., 2019; Zheng et al., 2017b; Benny et al., 2021), which may be explained by previous postulations that variations in the diseases under investigation contribute to disparities in placental microbiome composition. In the present study, we found that Lactobacillus were decreased, while Ureaplasma and Staphylococcus were increased significantly in APS, and that the abundances of Ureaplasma and Staphylococcus were negatively correlated with that of Lactobacillus. Lactobacillus confers numerous advantages to the human body, exerting antioxidative effects and antimicrobial activity, while promoting cholesterol metabolism, immunomodulation, anti-allergenic effects, and tumor suppression (Slattery et al., 2019). In one study, the researchers. compared vaginal samples and placental tissues from preterm and term mothers (Aslam et al., 2022), with results indicating that a decrease in Lactobacillus and an increase in Ureaplasma were correlated with preterm delivery. Dysbiosis of Lactobacillus and Ureaplasma in the placenta contributes to early onset pulmonary infections in preterm neonates. Staphylococcus is a prevalent bacterium in intrauterine infections and is a frequent causative agent of neonatal pneumonia (Mabena et al., 2024). In the present study, Staphylococcus was found to be more abundant in patients with APS. Farooqi et al. further explored the diversity of microbiota in full-term and preterm placental samples (Farooqi et al., 2022), finding that 16/68 (23.5%) full-term and 4/16 (25%) preterm placental samples were positive for Staphylococcus aureus. Nevertheless, the pathophysiological mechanisms underlying Staphylococcus-induced preterm birth remain unknown. As such, further research is required to prevent the transition of Staphylococcus from symbiotic microbes to pathogenic microorganisms and minimize its impact on neonates.
In the field of ecology, alpha diversity is a metric which represents the average species diversity within a specific site at a local scale, whereas beta diversity represents the ratio of regional to local species diversity. In the present study, we observed a significant difference in the alpha and beta diversities between the two groups. Some researchers have further noted that alterations in placental microbiota, which impact the outcome of delivery, are concomitant with changes in microbiota-mediated metabolic pathways (Prince et al., 2016; Aagaard et al., 2014; Antony et al., 2015). Antony et al. previously observed that the placentas of term-delivering women without chorioamnionitis exhibited a significant positive correlation with functional pathways related to the metabolism of various cofactors and vitamins, including riboflavin, nicotinate, pantothenate, and CoA (Antony et al., 2015). These pathways were similar to those of Lactobacillus in preterm women without chorioamnionitis. Virgiliou et al. previously investigated the metabolites in amniotic fluid samples taken from women who delivered preterm and at term, revealing significant differences in energy metabolism-related metabolites (pyruvic acid, glutamic acid, and glutamine) between these groups (Virgiliou et al., 2017). Our study similarly revealed variations in metabolic pathways and phenotypes across different diseases, suggesting that the placental microbiota may contribute to the development of the fetal innate immune system, and exert an impact on lung diseases in preterm infants. The transplacental mechanisms by which maternal taxonomy influences the offspring currently remains poorly understood (Piersigilli et al., 2020).
This study had several limitations. First, the sample size was relatively small. Therefore, large-scale studies are required. Secondly, the study sample did not include extremely preterm infants. As such, these findings are not representative of the entire population of preterm infants. Third, our study primarily focused on the quantification of bacterial abundance through 16S rRNA analysis, without delving into investigations pertaining to viruses and fungi. A comprehensive analysis of the bacterial, viral, and fungal microbiota, such as metagenomics could further enhance our understanding of host-microbiota interactions. Nevertheless, our results have the potential to significant contribute to the field of research.
Overall, this study represents the first attempt to investigate the impact of placental microecology on pulmonary diseases in preterm infants within this region. Throughout our investigation, we investigated the composition of the placental microbiota in preterm infants. Consequently, our research team aimed to conduct a comprehensive analysis of placental microecology and metabolomics across multiple regions with a large sample size to derive more precise conclusions regarding the influence of placental microecology on pulmonary infections in preterm infants. These results are important as they suggest that the placental microbiome may be linked to the onset of pneumonia in preterm infants, which could provide a basis for further research into novel methods to prevent this condition.
The datasets presented in this study can be found in online repositories. The names of the repository/repositories and accession number(s) can be found in the article/supplementary material.
The studies involving humans were approved by The Chengdu Women and Children’s Central Hospital ethics committee. The studies were conducted in accordance with the local legislation and institutional requirements. Written informed consent for participation in this study was provided by the participants’ legal guardians/next of kin. Written informed consent was obtained from the minor(s)’ legal guardian/next of kin for the publication of any potentially identifiable images or data included in this article.
LeZ: Conceptualization, Data curation, Formal Analysis, Funding acquisition, Investigation, Methodology, Project administration, Resources, Software, Supervision, Validation, Visualization, Writing – original draft, Writing – review & editing. ZC: Conceptualization, Data curation, Formal Analysis, Investigation, Resources, Writing – original draft. SH: Conceptualization, Data curation, Project administration, Writing – review & editing. HL: Data curation, Investigation, Resources, Writing – review & editing. FL: Data curation, Formal Analysis, Resources, Writing – review & editing. YF: Project administration, Visualization, Writing – review & editing. ML: Data curation, Formal Analysis, Resources, Software, Writing – original draft. LiZ: Data curation, Investigation, Methodology, Software, Writing – review & editing.
The author(s) declare that financial support was received for the research and/or publication of this article. The Chengdu Municipal Health Commission of Sichuan Province, China (No. 2021107), and Sichuan Medical Association of China (S21101).
We would like to thank all the subjects and their guardians who agreed to participate in this study.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Aagaard, K., Ma, J., Antony, K. M., Ganu, R., Petrosino, J., Versalovic, J. (2014). The placenta harbors a unique microbiome. Sci. Transl. Med. 6, 237ra65. doi: 10.1126/scitranslmed.3008599
American Academy of Pediatrics Committee on Fetus and Newborn, American College of Obstetricians and Gynecologists Committee on Obstetric Practice (2015). The apgar score. Pediatrics. 136, 819–822. doi: 10.1542/peds
Antony, K. M., Ma, J., Mitchell, K. B., Racusin, D. A., Versalovic, J., Aagaard, K. (2015). The preterm placental microbiome varies in association with excess maternal gestational weight gain. Am. J. Obstet Gynecol 212, 653.e1–653.16. doi: 10.1016/j.ajog.2014.12.041
Aslam, S., Sayeed Saraf, V., Saleem, S., Saeed, S., Javed, S., Junjua, M., et al. (2022). Lactobacillus species signature in association with term and preterm births from low-income group of Pakistan. J. Matern Fetal Neonatal Med. 35, 2843–2852. doi: 10.1080/14767058.2020.1810660
Benny, P. A., Al-Akwaa, F. M., Dirkx, C., Schlueter, R. J., Wolfgruber, T. K., Chern, I. Y., et al. (2021). Placentas delivered by pre-pregnant obese women have reduced abundance and diversity in the microbiome. FASEB J. 35, e21524. doi: 10.1096/fj.202002184RR
Bhandekar, H., Bansode Bangartale, S., Arora, I. (2024). Evaluating the clinical risk index for babies (CRIB) II score for mortality prediction in preterm newborns: A prospective observational study at a tertiary care hospital. Cureus. 16, e58672. doi: 10.7759/cureus.58672
Farooqi, H. M. U., Kim, K. H., Kausar, F., Muhammad, J., Bukhari, H., Choi, K. H. (2022). Frequency and molecular characterization of staphylococcus aureus from placenta of mothers with term and preterm deliveries. Life (Basel). 12, 257. doi: 10.3390/life12020257
Huang, J., Shen, W., Wu, F., Mao, J., Liu, L., Chang, Y., et al. (2024). Risk factors for severe bronchopulmonary dysplasia in a Chinese cohort of very preterm infants. Saudi Med. J. 45, 369–378. doi: 10.15537/smj.2024.45.4.20230741
Jiang, M., Mishu, M. M., Lu, D., Yin, X. (2018). A case control study of risk factors and neonatal outcomes of preterm birth. Taiwan J. Obstet Gynecol. 57, 814–818. doi: 10.1016/j.tjog.2018.10.008
Jin, J., Gao, L., Zou, X., Zhang, Y., Zheng, Z., Zhang, X., et al. (2022). Gut dysbiosis promotes preeclampsia by regulating macrophages and trophoblasts. Circ. Res. 131, 492–506. doi: 10.1161/CIRCRESAHA.122.320771
La, X., Wang, Y., Xiong, X., Shen, L., Chen, W., Zhang, L., et al. (2022). The composition of placental microbiota and its association with adverse pregnancy outcomes. Front. Microbiol. 13. doi: 10.3389/fmicb.2022.911852
Lal, C. V., Travers, C., Aghai, Z. H., Eipers, P., Jilling, T., Halloran, B., et al. (2016). The airway microbiome at birth. Sci. Rep. 6, 31023. doi: 10.1038/srep31023
Mabena, F. C., Olwagen, C. P., Phosa, M., Ngwenya, I. K., van der Merwe, L., Khan, A., et al. (2024). Bacterial and candida colonization of neonates in a regional hospital in South Africa. Pediatr. Infect. Dis. J. 43, 263–270. doi: 10.1097/INF.0000000000004177
Martin, R., Makino, H., Cetinyurek Yavuz, A., Ben-Amor, K., Roelofs, M., Ishikawa, E., et al. (2016). Early-life events, including mode of delivery and type of feeding, siblings and gender, shape the developing gut microbiota. PLoS One 11, e0158498. doi: 10.1371/journal.pone.0158498
Nissen, M. D. (2007). Congenital and neonatal pneumonia. Paediatr. Respir. Rev. 8, 195–203. doi: 10.1016/j.prrv.2007.07.001
Obstetrics Group of the Chinese Association of Obstetrics and Gynecology, the Chinese Association of Obstetrics and Gynecology, and the Chinese Medical Association. (2014). The expert consensus on cesarean delivery operation, (2014). Zhonghua Fu Chan Ke Za Zhi. 49, 721–724. doi: 10.3760/cma.j.issn.0529-567X.2014.10.001
Piersigilli, F., Van Grambezen, B., Hocq, C., Danhaive, O. (2020). Nutrients and microbiota in lung diseases of prematurity: the placenta-gut-lung triangle. Nutrients. 12, 469. doi: 10.3390/nu12020469
Powell, A. M., Ali Khan, F. Z., Ravel, J., Elovitz, M. A. (2024). Untangling associations of microbiomes of pregnancy and preterm birth. Clin. Perinatol. 51, 425–439. doi: 10.1016/j.clp.2024.02.009
Prince, A. L., Ma, J., Kannan, P. S., Alvarez, M., Gisslen, T., Harris, R. A., et al. (2016). The placental membrane microbiome is altered among subjects with spontaneous preterm birth with and without chorioamnionitis. Am. J. Obstet Gynecol. 214, 627.e1–627.e16. doi: 10.1016/j.ajog.2016.01.193
Seferovic, M. D., Pace, R. M., Carroll, M., Belfort, B., Major, A. M., Chu, D. M., et al. (2019). Visualization of microbes by 16S in situ hybridization in term and preterm placentas without intraamniotic infection. Am. J. Obstet Gynecol. 221, 146.e1–146.e23. doi: 10.1016/j.ajog.2019.04.036
Slattery, C., Cotter, P. D., O’Toole, P. W. (2019). Analysis of health benefits conferred by lactobacillus species from kefir. Nutrients. 11, 1252. doi: 10.3390/nu11061252
Valentine, G., Chu, D. M., Stewart, C. J., Aagaard, K. M. (2018). Relationships between perinatal interventions, maternal-infant microbiomes, and neonatal outcomes. Clin. Perinatol. 45, 339–355. doi: 10.1016/j.clp.2018.01.008
Virgiliou, C., Gika, H. G., Witting, M., Bletsou, A. A., Athanasiadis, A., Zafrakas, M., et al. (2017). Amniotic fluid and maternal serum metabolic signatures in the second trimester associated with preterm delivery. J. Proteome Res. 16, 898–910. doi: 10.1021/acs.jproteome.6b00845
Wang, X., Li, Y., Shi, T., Bont, L. J., Chu, H. Y., Zar, H. J., et al. (2024). Global disease burden of and risk factors for acute lower respiratory infections caused by respiratory syncytial virus in preterm infants and young children in 2019: a systematic review and meta-analysis of aggregated and individual participant data. Lancet. 403, 1241–1253. doi: 10.1016/S0140-6736(24)00138-7
Wu, F., Liu, G., Feng, Z., Tan, X., Yang, C., Ye, X., et al. (2019). Short-term outcomes of extremely preterm infants at discharge: a multicenter study from Guangdong province during 2008-2017. BMC Pediatr. 19, 405. doi: 10.1186/s12887-019-1736-8
Wu, S., Ren, L., Li, J., Shen, X., Zhou, Q., Miao, Z., et al. (2023). Breastfeeding might partially contribute to gut microbiota construction and stabilization of propionate metabolism in cesarean-section infants. Eur. J. Nutr. 62, 615–631. doi: 10.1007/s00394-022-03020-9
Zheng, J., Xiao, X., Zhang, Q., Mao, L., Yu, M., Xu, J., et al. (2017a). Correlation of placental microbiota with fetal macrosomia and clinical characteristics in mothers and newborns. Oncotarget. 8, 82314–82325. doi: 10.18632/oncotarget.19319
Zheng, J., Xiao, X., Zhang, Q., Mao, L., Yu, M., Xu, J., et al. (2017b). The placental microbiota is altered among subjects with gestational diabetes mellitus: A pilot study. Front. Physiol. 8. doi: 10.3389/fphys.2017.00675
Keywords: placental, microbiota, preterm infants, pneumonia, 16S rRNA
Citation: Zhang L, Chen Z, Hu S, Liu H, Lai F, Fan Y, Li M and Zhou L (2025) Assessment of the placental microbiota of preterm infants with pneumonia: a case control study. Front. Cell. Infect. Microbiol. 15:1511141. doi: 10.3389/fcimb.2025.1511141
Received: 14 October 2024; Accepted: 28 January 2025;
Published: 03 April 2025.
Edited by:
Shelley M. Lawrence, The University of Utah, United StatesReviewed by:
Hennie Lombaard, Baylor College of Medicine, United StatesCopyright © 2025 Zhang, Chen, Hu, Liu, Lai, Fan, Li and Zhou. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Lei Zhang, NTM0MTY3MzEzQHFxLmNvbQ==; Hong Liu, MjQ0NzAzOTI0Mzk5QHFxLmNvbQ==
†These authors have contributed equally to this work
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.