
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Cell. Infect. Microbiol. , 03 April 2025
Sec. Clinical Infectious Diseases
Volume 15 - 2025 | https://doi.org/10.3389/fcimb.2025.1499154
This article is part of the Research Topic Recent Advancements in the Research Models of Infectious Diseases View all 6 articles
 Yongjie Xu1,2,3,4,5,6†
Yongjie Xu1,2,3,4,5,6† Minjie Tang1,2,3,4,5,6
Minjie Tang1,2,3,4,5,6 Zhaopei Guo1,2,3,4,5†
Zhaopei Guo1,2,3,4,5† Yanping Lin7†
Yanping Lin7† Hongyan Guo1,2,3,4,5
Hongyan Guo1,2,3,4,5 Fengling Fang1,2,3,4,5
Fengling Fang1,2,3,4,5 Lin Lin1,2,3,4,5
Lin Lin1,2,3,4,5 Yue Shi1,2,3,4,5
Yue Shi1,2,3,4,5 Lu Lai1,2,3,4,5
Lu Lai1,2,3,4,5 Yan Pan1,2,3,4,5
Yan Pan1,2,3,4,5 Xiangjun Tang1,2,3,4,5
Xiangjun Tang1,2,3,4,5 Weiquan You1,2,3,4,5†
Weiquan You1,2,3,4,5† Zishun Li7
Zishun Li7 Jialin Song8*
Jialin Song8* Liang Wang9*
Liang Wang9* Weidong Cai1,2,3,4,5,6
Weidong Cai1,2,3,4,5,6 Ya Fu1,2,3,4,5*
Ya Fu1,2,3,4,5*COVID-19 caused by the coronavirus SARS-CoV-2 has resulted in a global pandemic. Considering some patients with COVID-19 rapidly develop respiratory distress and hypoxemia, early assessment of the prognosis for COVID-19 patients is important, yet there is currently a lack of research on a comprehensive multi-marker approach for disease prognosis assessment. Here, we utilized a large sample of hospitalized individuals with COVID-19 to systematically compare the clinical characteristics at admission and developed a nomogram model that was used to predict prognosis. In all cases, those with pneumonia, older age, and higher PT-INR had a poor prognosis. Besides, pneumonia patients with older age and higher PT-INR also had a poor prognosis. A nomogram model incorporating presence of pneumonia, age and PT-INR could evaluate the prognosis in all patients with SARS-CoV-2 infections well, while a nomogram model incorporating age and PT-INR could evaluate the prognosis in those with pneumonia well. Together, our study establishes a prognostic prediction model that aids in the timely identification of patients with poor prognosis and helps facilitate the improvement of treatment strategies in clinical practice in the future.
COVID-19, short for a novel coronavirus disease, is a severe acute respiratory syndrome caused by the coronavirus SARS-CoV-2, resulting in a global pandemic defined by the Director-General of the World Health Organization (WHO) on March 11, 2020 (Rothan and Byrareddy, 2020; Rubin et al., 2020). The pathogen of this disease belongs to the beta genus of coronaviruses and is mainly transmitted through respiratory droplets and close contact, and the general population is susceptible to it (Wang et al., 2020).
Although most patients have a good prognosis, some patients rapidly develop respiratory distress and hypoxemia after the onset of the disease, leading to the development of acute respiratory distress syndrome (ARDS), and even multiple organ failure, of which the mortality rate is relatively high (Gilbert et al., 2020). Based on the severity of the condition, COVID-19 can be classified into four categories: mild, moderate, severe, and critical. Early identification of potential severe cases, preventing the progression from mild or moderate to severe or critical, as well as assessing the prognosis and predicting the outcome of COVID-19, pose significant challenges in clinical practice.
Several indicators have been reported to be associated with the prognosis of COVID-19. Throughout the course of infection, the persistently low count of eosinophils could have fatal consequences (Chen et al., 2020; Zhao et al., 2021). Significantly elevated levels of IL-6 were associated with adverse clinical outcomes (Coomes and Haghbayan, 2020). Additionally, age, gender, and hypertension were also associated with the severity of the disease (Shi et al., 2020). Nevertheless, there is currently a lack of research on a comprehensive multi-marker approach for disease prognosis assessment. Although there are already various early warning scoring systems that can comprehensively assess the severity of patients’ conditions and predict clinical outcomes, such as the National Early Warning Score version 2 (NEWS2) and the Acute Physiology and Chronic Health Evaluation II (APACHE II) scoring systems, prognostic prediction models based on specific clinical indicators are equally important and effective. These models can serve as supplements to early warning scoring systems, thereby enhancing the accuracy of predictions.
Therefore, this study aims to utilize data from a large cohort of hospitalized COVID-19 patients to systematically compare the clinical characteristics of different prognostic groups at the time of admission. Our objective is to develop a standardized prognostic prediction model that incorporates patients’ age and PT-INR data upon admission, in order to provide accurate predictive capabilities for different prognostic groups. Through this research, we hope to establish an effective prognostic prediction tool that will assist clinicians in timely identifying patients with poor prognoses, ultimately improving treatment strategies in the future.
The study population included hospitalized patients diagnosed with COVID-19 at our hospital. The training cohort consisted of patients who were discharged or deceased between December 1, 2022, and January 31, 2023, while the validation cohort included patients who were discharged or deceased between April 1, 2023, and April 30, 2023. Data collection was conducted through the hospital’s electronic medical record system, and the relevant data were compiled into spreadsheets and reviewed by senior physicians. The collected information included the patients’ age, gender, diagnosis at the time of admission, presence of pneumonia, laboratory test results, and prognosis. For the sample size calculation, we set the significance level at 0.05, the statistical power at 0.80, the expected effect size at 0.8, and the loss percentage at 20%, resulting in a final sample size of 554 participants. According to the guidelines and standards set by the World Health Organization (WHO) regarding COVID-19 (O. World Health, 2021), the diagnostic criteria for confirmed cases include the presence of one of the following microbiological evidence based on suspected cases: 1) The patient exhibits acute respiratory infection symptoms, such as fever, cough, fatigue, shortness of breath, sore throat, muscle or joint pain, headache, and loss of smell or taste; 2) Positive results for the novel coronavirus nucleic acid via real-time fluorescent RT-PCR testing; 3) Viral gene sequencing showing high homology with known novel coronaviruses; 4) A history of contact with confirmed COVID-19 cases within the past 14 days, or travel or residence in areas experiencing severe outbreaks; 5) Chest X-ray or CT scans revealing pneumonia or other lung lesions associated with COVID-19. The criteria for excluding COVID-19 typically include the following aspects: 1) Clinical Symptoms: Patients who do not exhibit COVID-19-related symptoms (such as fever, cough, shortness of breath, fatigue, muscle or joint pain, sore throat, headache, loss of taste or smell, etc.) usually do not meet the criteria for a COVID-19 diagnosis. 2) Test Results: If PCR or rapid antigen test results are negative, and the patient’s symptoms or clinical history do not align with COVID-19, then a COVID-19 diagnosis should also be excluded. 3) Other Causes: If the patient’s symptoms can be explained by other known causes (such as influenza, other respiratory viral infections, or bacterial pneumonia), and relevant tests confirm this, then a diagnosis of COVID-19 can be ruled out. 4) Epidemiological Background: An absence of epidemiological risk factors for COVID-19 (such as no contact with confirmed cases or recent travel to areas with high incidence of the disease) can also serve as a basis for exclusion from the diagnosis. 5) Imaging Studies: If X-ray or CT imaging does not show typical lesions associated with COVID-19, the patient may also be considered for exclusion from a COVID-19 diagnosis. Based on the clinical treatment outcomes, we categorized the patients into four groups: Cured Group: Patients who had a complete resolution of their condition after clinical treatment. Improved Group: Patients who showed significant improvement in their symptoms following treatment. Unimproved Group: This group includes a portion of patients whose condition did not improve and actually worsened after treatment. In these cases, family members chose to discontinue treatment, leading to the patient’s discharge against medical advice. Deceased Group: Patients who passed away during the course of clinical treatment. In our study, patients were categorized into two groups based on their clinical outcomes following treatment: “Effective Treatment” and “Ineffective Treatment.” Patients classified as having received “Effective Treatment” were those who demonstrated significant improvement in symptoms and met the discharge criteria during our observation/data collection period (this includes patients from both the “Cured” and “Improved” groups). In contrast, “Ineffective Treatment” refers to patients who did not meet these criteria within the specified observation period (this includes patients from both the “Unimproved” and “Deceased” groups). The written consent was obtained from each patient and the study was approved by the Ethics Committee of the First Affiliated Hospital of Fujian Medical University. Ethics Committee Approval Number: MTCA, ECFAH of FMU〔2015〕No. 084-2.
In the training cohort, the one-way analysis of variance (ANOVA) was used to compare the differences in various clinical indicators among the cured, improved, unimproved, and dead groups. Indicators with significantly statistical differences were further filtered by lasso regression and logistic regression analysis. We further simplified the complex logistic regression model into a visualized nomogram by using the rms package of R. Subsequently, the efficiency of the visualized nomogram was evaluated by calibration curve and receiver operating characteristic (ROC) curve. The calibration curve was used to compare the association between actual outcomes and predicted probabilities. The ROC curve was used to assess the discriminative ability of the nomogram and then the area under the curve (AUC). Nomogram scores are utilized to predict the patient’s 30-day clinical outcomes during hospitalization. Cumulative events of effective treatment within 30 days of hospitalization based on cox regression analysis were visualized by using the survminer and ggplot2 packages of R.
Statistical differences were evaluated by one-way analysis of variance (ANOVA), chi-squared test (Fisher’s exact test was used when needed), lasso regression, logist regression, and cox regression with IBM SPSS Statistics software (Version 22.0.0; IBM, Armonk, New York, USA) and R (version 4.1.0 http://www.r-project.org). All P values were two-tailed. P < 0.05 was considered to be statistically significant.
The study enrolled a total of 1134 hospitalized patients who were infected with COVID-19, including 823 cases in the training set and 311 cases in the validation set (Figure 1). Among these 823 patients in the training set, 89 succumbed to the disease, 293 successfully recovered, while the remaining 441 individuals were still undergoing treatment at the time of their enrollment in the study. The clinical data of patients with different prognoses at admission in the training set are shown in Table 1. By anlysing the differences in the clinical characteristics among the cured, improved, unimproved and dead groups, significant differences in 38 indexes by ANOVA were observed across above 4 groups. The results shown that gender was associated with prognosis (P = 0.0297), and male patients have a poorer prognosis compared with female ones (Dead: male vs female, 73.03% vs 26.97%. Unimproved: male vs female, 70.59% vs 29.41%). Moreover, patients with pneumonia in this study also had the poorer prognosis compared with those without pneumonia (P < 0.0001. Dead: pneumonia vs non-pneumonia, 91.01% vs 8.99%). In addition, the elevated levels of 18 factors were found correlated to the poorer prognosis, while the declining levels of the other 18 factors were correlated to the poorer prognosis, In addition, the elevated levels of 18 biochemical and clinical factors were found to be correlated to the poorer prognosis as detailed in table.
To evaluate factors associated with prognosis post COVID-19 infection, patients in this study were classified into effective-treatment group versus ineffective-treatment group. The effective group included cured and improved patients, while the ineffective group included unimproved and dead patients. One case was excluded from the analysis due to missing the data about duration of hospitalization. The lasso regression was used to initially screen the prognosis-related factors from above 38 indexes with significant difference among the cured, improved, unimproved and dead patients (Table 1, Figures 2A, B). Consequently, 19 of 38 indexes were selected for the subsequent analysis, including gender, presence/absence of pneumonia, age, PT-INR, APTT, Fg, #NEUT, %MONO, %EOS, #LUC, PLT, ALB, AST, LDH, CK, GLU, TCHO, CA, CO2CP. The logist regression was employed to analyze the probability of using these 19 indexes for predicting the prognosis. It was shown that except #LUC, 18 of 19 indexed were all significantly different between effective-treatment and ineffective-treatment groups by univariate analysis of cox regression, while only 3 indexes —— presence/absence of pneumonia(mean OR = 3.783, P < 0.001), age (mean OR = 1.029, P = 0.001) and PT-INR (mean OR = 3.286, P = 0.007) —— were significantly different between effective-treatment and ineffective-treatment groups by multivariate analysis of cox regression (Figure 2C). Further, these aforementioned 3 predictors were integrated to develop a nomogram model that could be used to evaluate the prognosis (Figure 2D). The observed and predicted values of this model exhibited a high level of agreement, indicating a reliable performance of the model (Figure 2E). Using the scores marked by the nomogram model, the receiver operating characteristic (ROC) curves were generated between the effective-treatment and ineffective-treatment groups (Figure 2F). The areas under the curve (AUC) of model scores, presence/absence of pneumonia, age and PT-INR were respectively 0.802, 0.727, 0.697 and 0.710, suggesting that the model could effectively predict the prognosis post COVID-19 infection. The cox regression analysis was employed to further enhance the reliability of the model in predicting patient prognosis by evaluating the cumulative events of treatment effectiveness within 30 days of hospitalization (Figures 2G-J). The findings indicated that younger age (low vs high, mean HR = 0.70, P < 0.001), lower PT-INR levels (low vs high, mean HR = 0.64, P < 0.001), and the absence of pneumonia (pneumonia vs absence of pneumonia, mean HR = 2.13 or absence of pneumonia vs pneumonia, mean HR = 1/2.13 = 0.47, P < 0.001) were correlated with a more favorable prognosis within 30 days of hospitalization. Furthermore, patients with lower model scores also exhibited improved outcomes within 30 days of hospitalization (low vs high, mean HR = 0.45, P < 0.001).
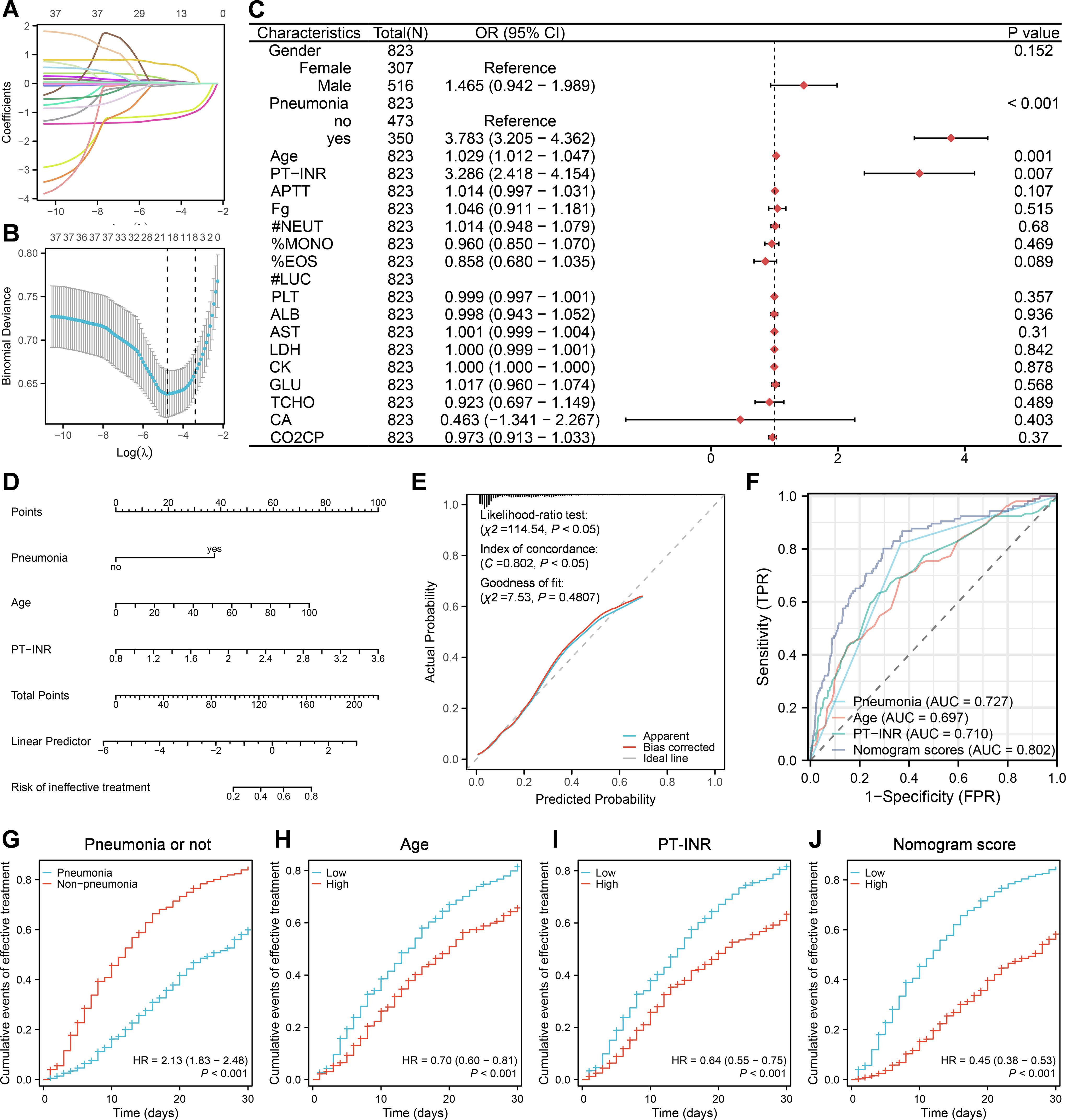
Figure 2. A nomogram model built based on pneumonia, age and PT-INR at admission for predicting prognosis post COVID-19 infection. (A, B) Lasso regression analysis. (C) Multivariate logistic regression analysis. (D) Construction of nomogram model based on multivariate logistic regression analysis. (E) Construction of calibration curves for the nomogram model. (F) Construction of ROC curves for nomogram scores, presence/absence of pneumonia, age, and PT-INR, respectively. (G-J) Cumulative events of effective treatment within 30 days of hospitalization based on cox regression analysis.
To further obtain factors associated with recovery post COVID-19 infection in 350 patients with pneumonia, the differences in the clinical characteristics were analyzed among the cured, improved, unimproved and dead groups, and a total of 23 indicators were found to have statistically significant differences (Table 2). The lasso regression was also used to screen the recovery-related factors from aforementioned 23 indexes with significant difference by ANOVA among the cured group, uncured group and dead group (Figures 3A, B). 11 of 23 indexes were selected for the subsequent analysis, including age, PT-INR, APTT, WBC, %MONO, %EOS, %LUC, LDH, UREA/CREA, CA and CO2CP (Figure 3C). The logistic regression was used to analyze the feasibility of using these 11 indexes for assessing the probability of recovery post COVID-19 infection, and the results demonstrated that except #LUC, 10 of 11 indexed were all significantly different between effective-treatment and ineffective-treatment groups by univariate analysis of cox regression, while only 2 indexes; age (mean OR = 1.042, P < 0.001) and PT-INR (mean OR = 2.742, P = 0.04); were significantly different between effective-treatment and ineffective-treatment groups by multivariate analysis of cox regression (Figure 3C). Subsequently, a nomogram model incorporating age and PT-INR was constructed to assess the prognosis (Figure 3D). The calibration curve showed that the observed and predicted values of this model exhibited a high level of agreement, indicating a reliable performance of the model (Figure 3E). Using the scores marked by the nomogram model, ROC curves were generated between the effective-treatment and ineffective-treatment groups (Figure 3F). The areas under the curve (AUC) of model scores, age and PT-INR were respectively 0.701, 0.639 and 0.665. The cox regression analysis was also employed to evaluate the cumulative events of treatment effectiveness within 30 days of hospitalization in patients with pneumonia (Figures 3G-I). The findings indicated that younger age (low vs high, mean HR = 0.73, P = 0.030) and lower PT-INR levels (low vs high, mean HR = 0.70, P = 0.013) were correlated with a more favorable prognosis within 30 days of hospitalization in patients with pneumonia. Furthermore, pneumonia patients with lower model scores also exhibited improved outcomes within 30 days of hospitalization (low vs high, mean HR = 0.66, P < 0.001).
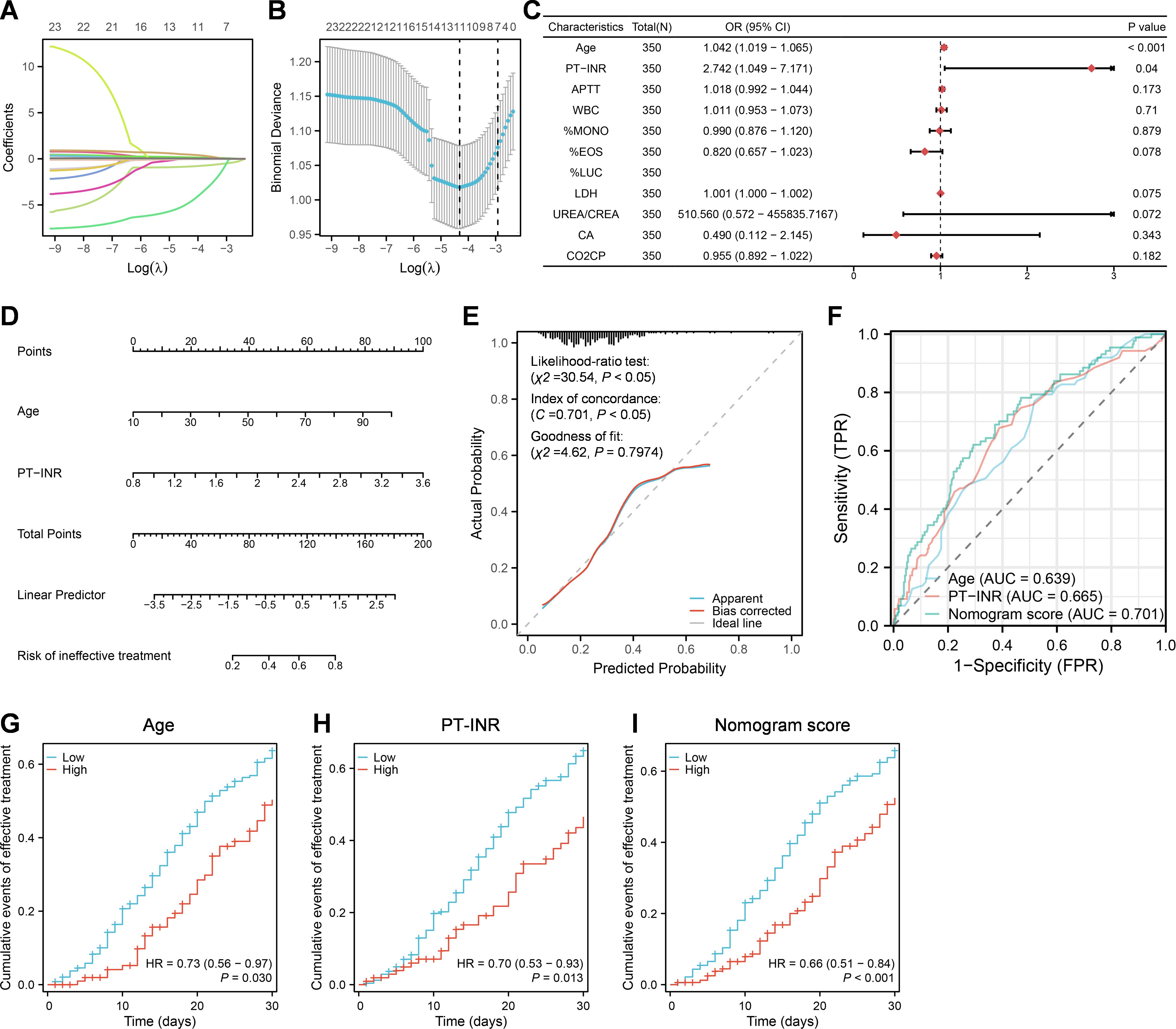
Figure 3. A nomogram model built based on age and PT-INR at admission for predicting prognosis post COVID-19 infection in patients with pneumonia. (A, B) Lasso regression analysis. (C) Multivariate logistic regression analysis. (D) Construction of nomogram model based on multivariate logistic regression analysis. (E) Construction of calibration curves for the nomogram model. (F) Construction of ROC curves for nomogram scores, age, and PT-INR, respectively. (G-I) Cumulative events of effective treatment within 30 days of hospitalization based on cox regression analysis.
Furthermore, we utilized the constructed nomogram models to evaluate the prognosis of 311 patients in the validation set within 30 days of hospitalization. The cox regression analysis was employed to analyze prognosis by evaluating the cumulative events of treatment effectiveness within 30 days of hospitalization (Figure 4). The findings indicated that in all 311 patients, younger age (low vs high, mean HR = 0.57, P < 0.001), lower PT-INR levels (low vs high, mean HR = 0.69, P = 0.003), and the absence of pneumonia (pneumonia vs absence of pneumonia, mean HR = 1.74 or absence of pneumonia vs pneumonia, mean HR = 1/1.74 = 0.57, P < 0.001) were correlated with a more favorable prognosis within 30 days of hospitalization (Figure 4A). Furthermore, patients with lower model scores exhibited improved outcomes within 30 days of hospitalization (low vs high, mean HR = 0.54, P < 0.001) (Figure 4A). Moreover, in patients with pneumonia, except PT-INR, the younger age (low vs high, mean HR = 0.59, P < 0.001) and lower model scores (low vs high, mean HR = 0.55, P < 0.001) were also correlated with a more favorable prognosis within 30 days of hospitalization (Figure 4B).
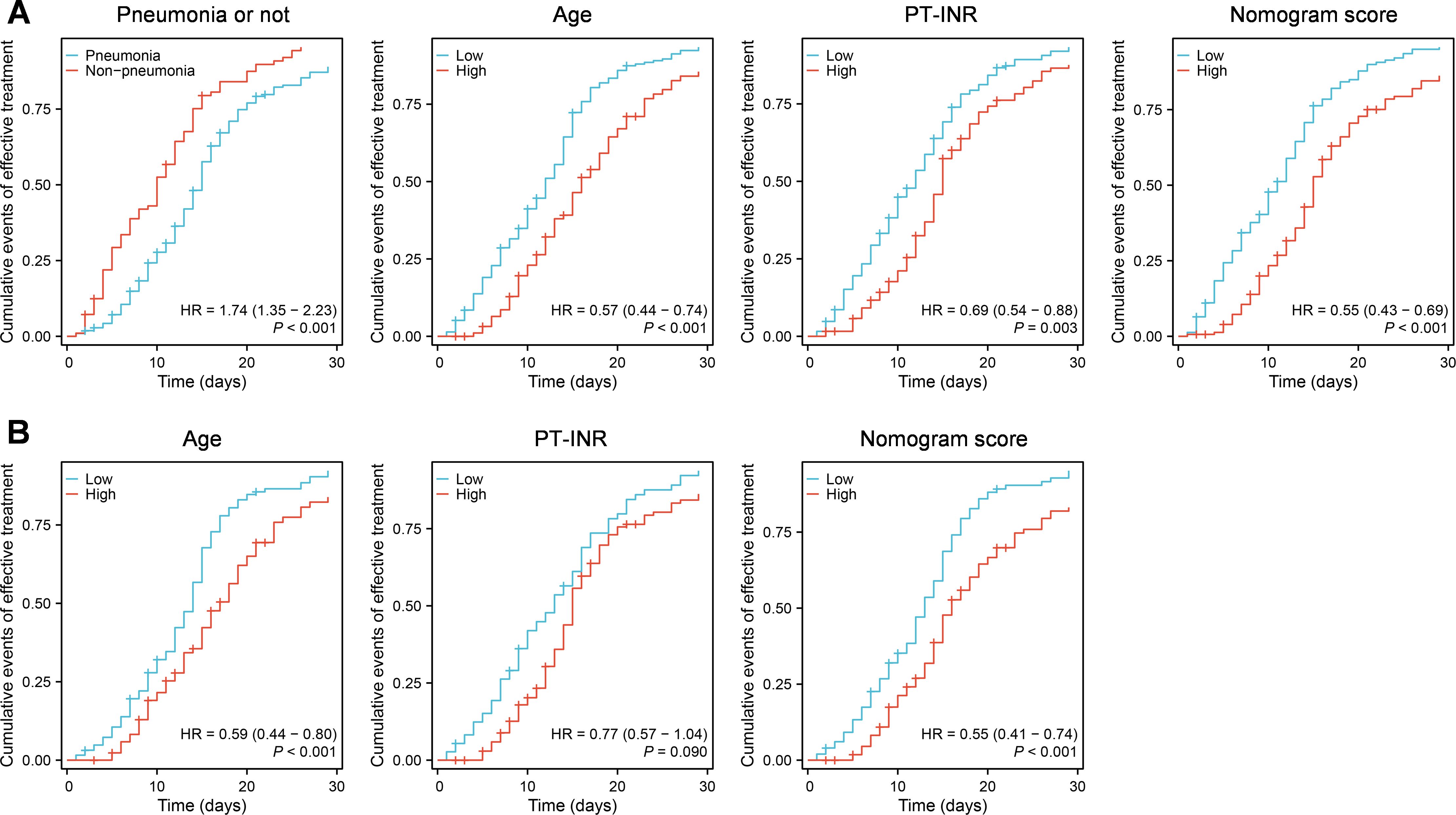
Figure 4. The validation of the constructed nomogram models for evaluating prognosis. (A) The validation of the constructed nomogram model incorporating the presence of presence/absence of pneumonia, age and PT-INR. (B) The validation of the constructed nomogram model incorporating age and PT-INR.
The outbreak of coronavirus disease 2019 (COVID-19) caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has resulted in high morbidity and mortality rates worldwide. As of December 19, 2021, COVID-19 has impacted 273 million people and resulted in over 5.3 million deaths (Zhang et al., 2022). SARS-CoV-2 infection may initially be asymptomatic, until severe pneumonia, respiratory distress, organ dysfunction, and even death occur (Li et al., 2020), raising questions regarding the risks and protective factors of COVID-19. In this study, we utilized large-scale clinical data to compare the differences in clinical characteristics among different prognostic groups at admission and attempted to develop a model for predicting patient outcomes.
This study compared the clinical characteristics of different prognostic groups at admission and found a correlation between the presence of presence/absence of pneumonia, gender, age, and 35 laboratory biomarkers with clinical outcomes (Table 1). In addition, to further explore death - related factors, we divided the subjects in Tables 1 and 2 into a death group and a non-death group (including the cured, improved, and unimproved groups), and created Tables 3 and 4 as Supplementary Materials. Our current study demonstrated that older age, male gender, and the presence of pneumonia were closely associated with poor prognosis, consistent with previous research findings (Fauci et al., 2020). COVID-19 has been reported to cause coagulation dysfunction, characterized by significant elevation of D-dimer and fibrinogen, mild thrombocytopenia, and mild prolongation of PT/APTT (Lim and McRae, 2021). In this study, we found elevation of fibrinogen and PT-INR, reduction of PLT, and prolongation of PT, APTT and TT in patients with a poor prognosis. Some researchers have reported that COVID-19 could also cause alterations in hemogram of patients (Wang et al., 2020; Zhao et al., 2021). The current study displayed that significant reduction of %EOS, #EOS, %LYMPH, and %MONO but significant elevation of %NEUT and #NEUT at admission in dead cases. Besides, we also identified several biochemical indicators at admission, including TP, ALB, ALT, AST, LDH, CK, CKMB, UREA, CREA, GLU, TC, LDL-C, APOA1, CA, and CO2CP, that exhibited significant alterations in patients with adverse prognosis, particularly in fatal cases. These biochemical changes suggested that multi-organ involvement might be a major contributing factor to poor prognosis in patients. Together, these observations provide a detailed analysis of the clinical characteristic differences at admission among different prognostic groups, offering a data-driven support for clinicians in assessing the severity of patients’ conditions.
Based on the analysis of variance, lasso regression, and logistic regression, we found that the presence of pneumonia, older age, and higher PT-INR at admission were the most important prognostic indicators for COVID-19 patients. Moreover, older age and higher PT-INR were also important prognostic indicators for COVID-19 patients with pneumonia. Although the prognostic indicators for patients with COVID-19 and pneumonia show limited predictive capability, with individual AUC values for age and PT-INR falling below 0.7, our study demonstrates that by constructing a multivariable model that incorporates both age and PT-INR, the overall predictive ability of the model exceeds 0.7. This indicates that while the predictive power of certain individual factors may be limited, their combination in a multivariable model can significantly enhance predictive performance. This underscores the importance of utilizing multivariable models in clinical predictions. Clearly, existing studies have shown that the presence of pneumonia is one of the main causes of mortality in COVID-19 patients (Wiersinga et al., 2020; Zhang et al., 2022). The findings of O’Driscoll et al. demonstrated that in the population studied, the fatality rate of SARS-CoV-2 infection increased with age, beginning as early as 5 years old (O’Driscoll et al., 2021), which highlighted the significant influence of age on the prognosis of SARS-CoV-2 infection and was consistent with the experimental outcomes in our study. Similar to our findings, several studies reported a significant increase in PT-INR in COVID-19 patients with poor prognosis (Aminasnafi et al., 2022; Ceci et al., 2023; Wu et al., 2023), suggesting that liver involvement might be a major factor contributing to COVID-19 mortality. Collectively, these findings highlight the close relationship between age, PT-INR, and poor prognosis in COVID-19 patients, and it is the first to establish a predictive model based on age and PT-INR. Several studies (Martin-Rodriguez et al., 2022; Wibisono et al., 2022) have demonstrated that the National Early Warning Score version 2 (NEWS2) can effectively predict clinical deterioration and hospitalization outcomes in COVID-19 patients, particularly in emergency and inpatient settings. Its simplicity and ease of use make it an ideal tool for initial screening in clinical practice. However, based on the analyses presented in Supplementary Figures S1 and S2, we found that the ROC score of the NEWS2 assessment from our observed case records was lower than that of our model, indicating that our study’s model has relatively good predictive performance. Our study has several limitations. It is currently a single-center retrospective study, and further validation of our model’s efficacy will require a multicenter approach. Additionally, due to incomplete data collection, we were unable to conduct a correlation analysis between the symptoms of COVID-19 patients and the severity of their condition.
In conclusion, this study emphasizes the strong correlation between age, PT-INR at hospital admission, and patient prognosis. The prognostic model developed based on age and PT-INR can effectively identify patients at higher risk of poor outcomes. Our findings enhance the understanding of COVID-19 disease characteristics and provide valuable guidance for clinical management and treatment.
The raw data supporting the conclusions of this article will be made available by the authors, without undue reservation.
The studies involving humans were approved by the Ethics Committee of the First Affiliated Hospital of Fujian Medical University. The studies were conducted in accordance with the local legislation and institutional requirements. Written informed consent for participation was not required from the participants or the participants’ legal guardians/next of kin in accordance with the national legislation and institutional requirements.
YX: Formal Analysis, Investigation, Writing – original draft. MT: Data curation, Validation, Resources, Writing – review & editing. ZG: Formal Analysis, Writing – original draft. YL: Funding acquisition, Investigation, Writing – original draft. HG: Writing – review & editing. FF: Writing – review & editing. LLi: Data curation, Writing – review & editing. YS: Data curation, Writing – review & editing. LLa: Software, Writing – review & editing. YP: Validation, Writing – review & editing. XT: Validation, Writing – review & editing. WY: Software, Writing – review & editing. ZL: Visualization, Writing – review & editing. JS: Writing – review & editing. LW: Writing – review & editing. WC: Data curation, Validation, Resources, Writing – review & editing. YF: Funding acquisition, Methodology, Writing – review & editing.
The author(s) declare that financial support was received for the research and/or publication of this article. This work was supported by grants from the National Natural Science Foundation (82202596), Joint Funds for the Innovation of Science and Technology, Fujian Province (2023Y9022), and Fujian Provincial Natural Science Foundation of China (2020J05281, 2022D009).
We thank all the participants who agreed to take part in this study.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fcimb.2025.1499154/full#supplementary-material
Aminasnafi, A., Heidari, S., Alisamir, M., Mirkarimi, M., Namehgoshayfard, N., Pezeshki, S. M. S. (2022). Hematologic evaluation of children with COVID-19 infection: mortality biomarkers. Clin. Lab. 68 (4). doi: 10.7754/Clin.Lab.2021.210746
Ceci, F. M., Ferraguti, G., Lucarelli, M., Angeloni, A., Bonci, E., Petrella, C., et al. (2023). Investigating biomarkers for COVID-19 morbidity and mortality. Curr. topics medicinal Chem. 23, 1196–1210. doi: 10.2174/1568026623666230222094517
Chen, R., Sang, L., Jiang, M., Yang, Z., Jia, N., Fu, W., et al. (2020). Longitudinal hematologic and immunologic variations associated with the progression of COVID-19 patients in China. J. Allergy Clin. Immunol. 146, 89–100. doi: 10.1016/j.jaci.2020.05.003
Coomes, E. A., Haghbayan, H. (2020). Interleukin-6 in Covid-19: A systematic review and meta-analysis. Rev. Med. Virol. 30, 1–9. doi: 10.1002/rmv.v30.6
Fauci, A. S., Lane, H. C., Redfield, R. R. (2020). Covid-19 — Navigating the uncharted. New Engl. J. Med. 382, 1268–1269. doi: 10.1056/NEJMe2002387
Gilbert, M., Pullano, G., Pinotti, F., Valdano, E., Poletto, C., Boelle, P. Y., et al. (2020). Preparedness and vulnerability of African countries against importations of COVID-19: a modelling study. Lancet (London England) 395, 871–877. doi: 10.1016/S0140-6736(20)30411-6
Li, G., Fan, Y., Lai, Y., Han, T., Li, Z., Zhou, P., et al. (2020). Coronavirus infections and immune responses. J. Med. Virol. 92, 424–432. doi: 10.1002/jmv.25685
Lim, M. S., McRae, S. (2021). COVID-19 and immunothrombosis: Pathophysiology and therapeutic implications. Crit. Rev. Oncology/Hematol. 168, 103529. doi: 10.1016/j.critrevonc.2021.103529
Martin-Rodriguez, F., Sanz-Garcia, A., Ortega, G. J., Delgado-Benito, J. F., Garcia Villena, E., Mazas-Perez-Oleaga, C., et al. (2022). One-on-one comparison between qCSI and NEWS scores for mortality risk assessment in patients with COVID-19. Ann. Med. 54, 646–654. doi: 10.1080/07853890.2022.2042590
O’Driscoll, M., Ribeiro Dos Santos, G., Wang, L., Cummings, D. A. T., Azman, A. S., Paireau, J., et al. (2021). Age-specific mortality and immunity patterns of SARS-CoV-2. Nature 590, 140–145. doi: 10.1038/s41586-020-2918-0
O. World Health (2021). COVID-19 clinical management: living guidance, 25 January 2021 (Geneva: World Health Organization).
Rothan, H. A., Byrareddy, S. N. (2020). The epidemiology and pathogenesis of coronavirus disease (COVID-19) outbreak. J. Autoimmun. 109, 102433. doi: 10.1016/j.jaut.2020.102433
Rubin, E. J., Baden, L. R., Morrissey, S. (2020). Audio interview: A look at covid-19 prevention and care in 2020. New Engl. J. Med. 383, e147. doi: 10.1056/NEJMe2036225
Shi, Y., Yu, X., Zhao, H., Wang, H., Zhao, R., Sheng, J. (2020). Host susceptibility to severe COVID-19 and establishment of a host risk score: findings of 487 cases outside Wuhan. Crit. Care 24, 108. doi: 10.1186/s13054-020-2833-7
Wang, D., Hu, B., Hu, C., Zhu, F., Liu, X., Zhang, J., et al. (2020). Clinical characteristics of 138 hospitalized patients with 2019 novel coronavirus-infected pneumonia in Wuhan, China. JAMA 323, 1061–1069. doi: 10.1001/jama.2020.1585
Wibisono, E., Hadi, U., Bramantono, Arfijanto, M. V., Rusli, M., Rahman, B. E., et al. (2022). National early warning score (NEWS) 2 predicts hospital mortality from COVID-19 patients. Ann. Med. Surg. (Lond) 76, 103462. doi: 10.1016/j.amsu.2022.103462
Wiersinga, W. J., Rhodes, A., Cheng, A. C., Peacock, S. J., Prescott, H. C. (2020). Pathophysiology, transmission, diagnosis, and treatment of coronavirus disease 2019 (COVID-19). Jama 324, 782. doi: 10.1001/jama.2020.12839
Wu, P. J., Feng, I. C., Lai, C. C., Ho, C. H., Kan, W. C., Sheu, M. J., et al. (2023). The mortality of hospitalized patients with COVID-19 and non-cirrhotic chronic liver disease: a retrospective multi-center study. PeerJ 11, e16582. doi: 10.7717/peerj.16582
Zhang, J.-j., Dong, X., Liu, G.-h., Gao, Y.-d. (2022). Risk and protective factors for COVID-19 morbidity, severity, and mortality. Clin. Rev. Allergy Immunol. 64, 90–107. doi: 10.1007/s12016-022-08921-5
Keywords: PT-INR, age, predictor, mortality, SARS-CoV-2
Citation: Xu Y, Tang M, Guo Z, Lin Y, Guo H, Fang F, Lin L, Shi Y, Lai L, Pan Y, Tang X, You W, Li Z, Song J, Wang L, Cai W and Fu Y (2025) A model based on PT-INR and age serves as a promising predictor for evaluating mortality risk in patients with SARS-CoV-2 infection. Front. Cell. Infect. Microbiol. 15:1499154. doi: 10.3389/fcimb.2025.1499154
Received: 20 September 2024; Accepted: 03 March 2025;
Published: 03 April 2025.
Edited by:
Krupakar Parthasarathy, Sathyabama Institute of Science and Technology, IndiaReviewed by:
Pei Lu, University of Texas Southwestern Medical Center, United StatesCopyright © 2025 Xu, Tang, Guo, Lin, Guo, Fang, Lin, Shi, Lai, Pan, Tang, You, Li, Song, Wang, Cai and Fu. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Ya Fu, aHp4ZnlAZmptdS5lZHUuY24=; Liang Wang, NTIwNjk4NzhAcXEuY29t; Jialin Song, amlhbGluc29uQGZqbXUuZWR1LmNu
†These authors have contributed equally to this work
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.