
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
REVIEW article
Front. Cell Dev. Biol. , 27 March 2025
Sec. Cancer Cell Biology
Volume 13 - 2025 | https://doi.org/10.3389/fcell.2025.1559059
CTNND2 gene is located on the short arm of human chromosome 5 and encodes δ-catenin protein, which interacts with different proteins and plays different cell functions. Studies have demonstrated that δ-catenin plays an important role in regulating synaptic maturation and neuronal integrity. The CTNND2 gene is closely associated with a variety of neurological diseases, including Cri-du-Chat syndrome, Autism spectrum disorders, Alzheimer’s disease, and Epilepsy. Furthermore, an increasing number of studies have demonstrated that CTNND2 is involved in various cancers and may serve as a novel biomarker for the diagnosis and treatment for these diseases. In this review, we will focus on the signaling regulatory functions of CTNND2 and its encoded protein δ-catenin in neuro-related diseases and cancers, and discuss the limitations of previous investigative studies and the challenges of the future researches on CTNND2 and δ-catenin signaling.
The CTNND2 gene is situated on the short arm of human chromosome 5 and encodes a protein called delta-catenin/δ-catenin (referred as δ-catenin in this article) belonging to the armadillo/β-catenin superfamily (Paffenholz and Franke, 1997; Medina et al., 2000) and neural plakophilin-related armadillo repeat protein (NPRAP). The δ-catenin protein plays a variety of roles within cell, including the regulation of cell adhesion (Koutras and Lévesque, 2011; Koutras et al., 2011; Hatzfeld, 2005), neurodevelopment (Pauly et al., 2024; Tuncay et al., 2022; Wang L. Y. et al., 2024; Xu et al., 2023a; He et al., 2012; Vaz et al., 2023), and particularly formation and maintenance of dendritic spines and synapses (Yuan et al., 2015; Halder et al., 2015; Hassani et al., 2020; Adegbola et al., 2020; Wang S. et al., 2021; Li et al., 2021; Assendorp et al., 2024a; Ide et al., 1999; Izawa et al., 2002). Alterations in δ-catenin expression have been linked to the occurrence and progression of numerous pathological conditions, including autism (Zhang et al., 2016; Коваленко et al., 2020; Wang L. et al., 2024; Turner et al., 2015; Asadollahi et al., 2014; Lu et al., 2016; Zhou et al., 2019), schizophrenia (Chen et al., 2023; Vrijenhoek et al., 2008; Nivard et al., 2014), intellectual disability (Asadollahi et al., 2014; Einarsdottir et al., 2017; Hofmeister et al., 2015), and cancers (Oh et al., 2023; Wang et al., 2009; Burger et al., 2002; Huang et al., 2006; Zheng et al., 2004; Frattini et al., 2013).
In recent years, the study of δ-catenin has made remarkable progress due to the rapid development of molecular biology technology. It has been demonstrated that δ-catenin plays a crucial role in the regulation of synaptic maturation and neuronal integrity. The loss of function of δ-catenin may result in synaptic loss and neurodevelopmental disorders. Furthermore, the interaction between δ-catenin and the Wnt signaling pathway may play a significant role in tumor proliferation and drug resistance, offering a novel perspective for the treatment of cancers.
The aim of this article is to provide a timely review on the structure and function of CTNND2 gene and its encoded protein δ-catenin, its relationship with human diseases, and the potential regulatory mechanisms in the physiological and pathological processes. Revealing the relationship between δ-catenin and diseases could provide a referable therapeutic direction for the treatment of related diseases.
The CTNND2 gene is situated on the short arm of human chromosome 5, within the 5p15.2 region, and spans approximately 1.2 Mb (10,971,836–11,904,446 bp). The nucleic acid structure of the CTNND2 gene is shown schematically in Figure 1A, where one of the most widely studied transcripts contains 22 exons. Like all members of the subfamily of armadillo proteins (Keil et al., 2013; Peifer et al., 1992), δ-catenin protein possessed a central domain comprising a series of repeating motifs with approximately 40–45 amino acids in length, which is referred to as an arm repeat sequence (Paffenholz and Franke, 1997; Huang et al., 2021). The structure diagram is presented in Figure 1B. The arm repeat domain provides a multifunctional scaffold for protein-protein interactions, thereby enabling the participation in protein folding and stability, as well as interactions with other proteins (Huang et al., 2021). Consequently, some members of the armadillo protein-related protein subfamily, such as δ-catenin, can function in a variety of biological processes, including intracellular signaling, intercellular adhesion, and cytoskeletal organization. The C-terminal of δ-catenin protein contains a PDZ-binding motif, which contribute to its interaction with proteins containing PDZ domain, such as PSD95 and ABP/GRIP proteins. These interactions are critical for synaptic function and neurodevelopment (Yuan et al., 2015).
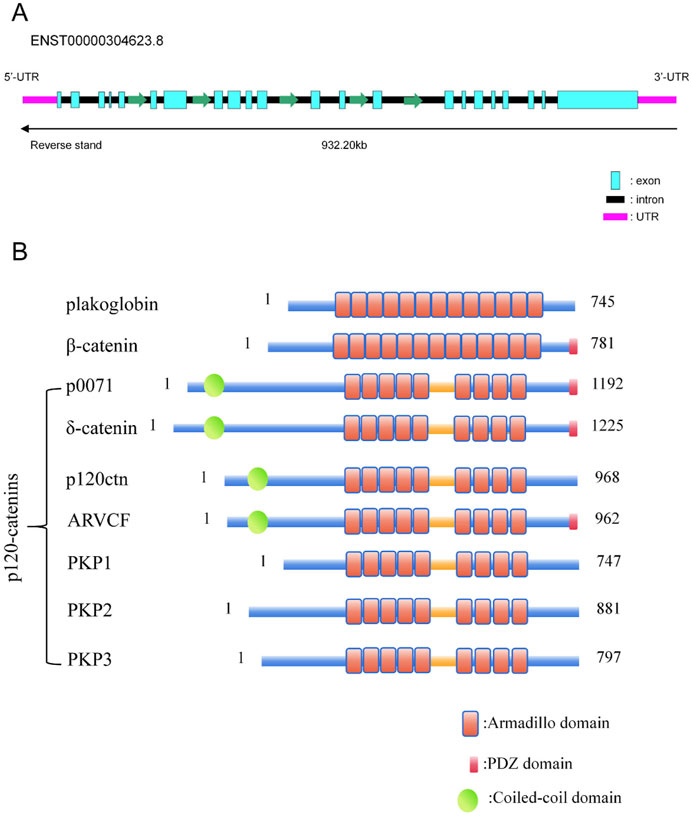
Figure 1. CTNND2 Gene and the p120 protein family proteins. (A) Gene structure diagram of the transcript ENST00000304623.8 of the CTNND2 gene. (B) Diagram of the structures of proteins with Armadillo and PDZ structural domains. The structure diagram of plakoglobin, β-catenin, and p120 catenins were summarized. All of the p120 protein family proteins, including p0071, δ-catenin, p120-catenin, ARVCF, PKP1, PKP2, and PKP3, contain a central armadillo structural domain, which consists of nine tandem incomplete 42-amino acid repeat sequences. p0071, Plakophilin-4 (PKP4); ARVCF, Armadillo Repeat Protein Deleted in Velo-Cardio-Facial Syndrome; PKP1, Plakophilin 1; PKP2, Plakophilin 2; PKP3, Plakophilin 3.
In normal human tissues, δ-catenin is predominantly expressed in the brain, with minimal expression observed in other normal human tissues. δ-catenin is predominantly localized to the cytoplasm (Lu, 2010; Wang et al., 2011) with potential nuclear localization capabilities (Koutras and Lévesque, 2011; Koutras et al., 2011), which may be associated with its involvement in transcriptional regulation. A comprehensive review of the existing literature is timely important to reveal a consistent association between the specific location of δ-catenin and a number of neurological disorders. The analysis of the protein structural domains specific to δ-catenin informs their participation and function in a variety of life activities.
δ-catenin is a protein that plays a crucial role in cell adhesion (Koutras et al., 2011; Izawa et al., 2002), signal conduction (Bareiss et al., 2010; He et al., 2023; Dunbar et al., 2021), and neurodevelopment (Pauly et al., 2024; Hassani et al., 2020). δ-catenin can form a complex with other components of adhesion junctions and regulate adhesion molecules, including cadherin and β-catenin, during the process of cell adhesion, which further regulate the histomorphogenetic processes (Izawa et al., 2002; Bareiss et al., 2010; Lu et al., 1999; Ligon et al., 2020; Kim et al., 2002). In addition, δ-catenin has been identified as a nervous system adhesion protein that undergoes dynamic reorientation during neurodevelopment (Ho et al., 2000). δ-catenin proteins can be modified by different post-translational modifications, including phosphorylation (Li et al., 2021; Zhou et al., 2019; Bareiss et al., 2010; Chen et al., 2021; Qu et al., 2024), methylation (Arpón et al., 2019; Singh et al., 2015), and palmitoylation (Zhang X. L. et al., 2018), during the process of cell signaling. Therefore, the localization, structure, interaction, and function of δ-catenin can be changed after these modifications.
δ-catenin plays a significant role in neurodevelopment by regulating the maturation of dendritic spines, synaptic maintenance, and neuronal excitability. Assendorp et al. have shown that δ-catenin can impede the maturation of synaptic connections and simultaneously promote the integrity of neuronal cells. In addition, δ-catenin can regulate neuronal excitability during postnatal development; and maintain synaptic integrity in adulthood. δ-catenin plays a crucial role in regulating neuronal excitability and synaptic plasticity (Assendorp et al., 2024b). Deficiency of CTNND2 may lead to impairment of synaptic function. For example, the knockout of CTNND2 gene induces sleep wake disorders in mice (Xu et al., 2023b). CTNND2-KO mice exhibit typical autism-like behaviors, suggesting that CTNND2 gene is important in the processes of spatial learning and memory, as well as in the mechanisms of Rictor-mediated actin polymerization and synaptic plasticity (Wang X. et al., 2021). Another study demonstrated that CTNND2-KO mouse models exhibit behaviors consistent with autism spectrum disorder (ASD) and exhibit decreased dendritic spine density in the hippocampus. Melatonin can improve the synaptic function of gamma-aminobutyric neurons by activating the PI3K/Akt signaling pathway, which may be related to the improvement of social behavior deficits and dendritic spine damage in CTNND2-KO mice (Wang L. Y. et al., 2024). These studies provide possible therapeutic options for disorders related to neurodevelopmental defects, such as autism caused by CTNND2 deletion.
Additionally, δ-catenin may play a role in the evolution of the human brain. δ-catenin may promote synaptic regeneration through the regulation of SRGAP2C in human evolution, suggesting that δ-catenin may promote the development and function of the human brain by regulating the maturation rate of synapses (Assendorp et al., 2024b). δ-catenin also plays a role in spatial learning and memory through rictor-mediated actin aggregation and synaptic plasticity (Wang S. et al., 2021). Moreover, abnormal δ-catenin expression or function has been linked to a range of neurological disorders, including intellectual disabilities (Hofmeister et al., 2015), autism spectrum disorder (Tuncay et al., 2022; Wang L. et al., 2024; Turner et al., 2015; Xu et al., 2023b), and schizophrenia (Chen et al., 2023; Vrijenhoek et al., 2008). The following section will address the relationship between δ-catenin and neurosystem-related diseases.
The CTNND2 gene is located in close proximity to the lesions of the neurodevelopmentally related disease Cri-du-Chat syndrome. Alterations in CTNND2 gene are strongly associated with severe mental retardation in Cri-du-Chat syndrome. Furthermore, recent studies have substantiated the correlation between CTNND2 and other neurological disorders, including Epilepsy, Autism, Alzheimer’s disease, and Myopia. An increasing number of studies have indicated that δ-catenin may play an important role in the progression of multiple types of cancer. Therefore, we shall present an overview on the existing studies related to CTNND2, outlining the relationship between CTNND2 gene and certain types of diseases and providing suggestions on the treatment of related diseases according to the different changes of CTNND2 in different diseases.
Cri-du-Chat syndrome is a chromosomal deletion syndrome, in which a portion of chromosome 5 is absent (Chhaya and Chan, 2013; Almeida et al., 2023). The deletion typically encompasses the 5p15.2–5p15.3 region, where the CTNND2 gene is situated (Sardina et al., 2014). In fact, the deletion of the CTNND2 gene is linked to a range of abnormalities, including facial deformities, microcephaly, cerebellar hypoplasia, lateral ventricle enlargement and severe psychomotor development delays, which are hallmarks of Cri-du-Chat syndrome (Sheth et al., 2012). In Cri-du-Chat syndrome, deletion or abnormal function of the CTNND2 gene can lead to severe intellectual impairment and developmental delays in the nervous system (Medina et al., 2000). In Cri-du-Chat syndrome, the deletion of the CTNND2 gene not only affects the development of the nervous system, but may also impact the development of other organs, resulting in a complex set of clinical symptoms. For example, in one clinical case, a full-term female infant presented with ischemic retina and retinal hemorrhage (Chhaya and Chan, 2013). Therefore, the function of the δ-catenin in Cri-du-Chat syndrome is multifaceted, including specific neuronal proteins that affect brain development, which in turn affect normal brain development, and its deletion ultimately results in developmental delays in the nervous system.
The relationship between the CTNND2 gene and autism spectrum disorder is gradually appreciated. δ-catenin plays an important role in the development of the nervous system, particularly in the formation and functionality of synapses (Wang L. Y. et al., 2024). The CTNND2 gene is regarded as a novel autism gene and its encoded δ-catenin protein is a neuron-specific protein and plays a role in cell adhesion and dendritic branching. Abnormal dendritic spines have been observed in the cerebral cortex of both patients and mouse models of autism spectrum disorder, which is related to the dysfunction of the CTNND2 gene (Wang L. et al., 2024; Ligon et al., 2020). The CTNND2 gene knockout mice exhibited such symptoms as social behavior disorder and decreased dendrite spine density, which may be related to an imbalance of excitatory and inhibitory neurotransmitters (Wang L. Y. et al., 2024; Xu et al., 2023a; Wang X. et al., 2021). Moreover, some studies have corroborated the existence of tandem duplication (Miller et al., 2020), missense, and dose sequence variation (Tuncay et al., 2022; Turner et al., 2015) in the genes of patients with autism, behavioral problems and malformation characteristics, indicating that anomalous CTNND2 level may be associated with the severity of autism and the cognitive phenotype. Therefore, normal CTNND2 gene and its δ-catenin expression are critical for neurodevelopment, and aberrant δ-catenin expression is evident in patients with Autism Spectrum Disorder.
Alterations in δ-catenin expression, predominantly in the nervous system of the brain, are associated with a range of neurological disorders. For instance, CTNND2 has been implicated in the development of myopia. Adegbola et al. (2020) found that the destruction of the CTNND2 gene is correlated with the development of attention deficit hyperactivity disorder and myopia, through genetic analysis in several members belongs to a family. Yu et al. observed significant differences in the distribution of variants of SNP rs1479617, located within the CTNND2 gene, between the pathological and control groups (Yu et al., 2012). Polymorphisms in the specific CTNND2 gene and 11q24.1 genomic region were significantly associated with pathological myopia in the Chinese population (Yu et al., 2012). Lu et al. demonstrated a robust correlation between CTNND2 polymorphism and myopia using Sanger sequencing (Lu et al., 2011). Li et al. found a robust association between CTNND2 and high myopia in an Asian cohort (Li et al., 2011b). Wen et al. demonstrated that extrachromosomal circular DNA (eccDNA) levels of CTNND2 were significantly increased in the anterior lens capsule of highly myopic patients (Wen et al., 2023). Liu et al. conducted a comprehensive meta-analysis of myopic patients and controls, and revealed a significant association between myopia and two specific genetic variants, rs6885224 and rs12716080, of the CTNND2 gene (Liu and Zhang, 2014). In addition, some researchers have also shown a strong correlation between CTNND2 and high myopia through animal studies (Yin et al., 2019; Srinivasalu et al., 2018) and data from clinical studies in different regions (Liu J. et al., 2021; Li et al., 2011a). These findings suggested that the CTNND2 gene may play an important role in the development of high myopia.
CTNND2 is one of the genes that regulate the development of brain neurons. Four single-nucleotide polymorphisms (SNPs) of the CTNND2 gene have been identified as being associated with schizophrenia (SZ), indicating that CTNND2 gene be involved in the susceptibility to SZ (Chen et al., 2023), and CTNND2 and some other genes important in neuronal function were disrupted in schizophrenia patients (Vrijenhoek et al., 2008). CTNND2 has been identified as the causative gene of Dutch family corticoclonic tremor and epilepsy, since CTNND2 missense mutations have been found in samples from patients with familial cortical myoclonic tremor and epilepsy (van Rootselaar et al., 2017).
Additionally, genetic variations in the CTNND2 gene may contribute to the development of cortical cataracts of midlife patients and the subsequent structural and functional changes in the brain of Alzheimer’s disease (AD) patients (Jun et al., 2012). A rare missense mutation (G810R) located within the CTNND2 gene alters the intracellular distribution of δ-catenin and increase secretion of amyloid beta protein (Aβ1-42) in nerve cell cultures, while amyloid-β (Aβ) is a neuropathological biomarker of AD (Jun et al., 2012). Moreover, Moncaster et al. have also proposed a potential association between CTNND2 and AD (Moncaster et al., 2022). A review of the extant literature reveals a correlation between δ-catenin expression malfunction and certain psychiatric disorders. This correlation may be attributable to the location of and the unique structural domain of the δ-catenin protein. Further research on δ-catenin could offer novel insights into the treatment of neurological diseases.
With the development of sequencing technology, there is increasing evidence that variants in the CTNND2 gene are strongly associated with a number of diseases. Kang et al. applied whole exon sequencing to identify variants in the CTNND2 gene, and discovered that these variants were linked to an increased risk of early-onset depression (Kang et al., 2021). Belcaro et al. reported two cases of internal deletion of the CTNND2 gene detected by molecular karyotyping in patients who presented with isolated intellectual disabilities clinically, and pointed out that the deletion of the CTNND2 gene was associated with intellectual disability (Belcaro et al., 2015). Hofmeister et al. combining human genetic and in vivo data with zebrafish embryos, found that impaired neuronal migration resulting from insufficient doses of the CTNND2 gene may underlie cognitive dysfunction in patients with borderline intelligence and learning problems within the dyslexia spectrum (Hofmeister et al., 2015). Medina et al. found a strong correlation between haploid deletion of the CTNND2 gene and the severity of intellectual disability, which suggests that when only one copy of the CTNND2 gene is present, it plays an important role in the intellectual impairment of Cri-du-Chat syndrome (Medina et al., 2000). Therefore, the aberrant expression of δ-catenin is frequently observed in numerous neurologic diseases, indicating a potential correlation between δ-catenin and disease pathophysiology.
Cancer is a leading cause of death worldwide due to the abnormal proliferation and structural heterogeneity of tumors, as well as the high tendency to metastasize. Earlier studies have shown that δ-catenin is involved in neurodevelopment. However, an increasing number of studies have demonstrated that δ-catenin plays a crucial role in the progression of neuro-related cancers. For example, Wang et al. identified the presence of δ-catenin protein in the cytoplasm of astrocytoma cells, and established a correlation between δ-catenin expression and the malignant progression of astrocytoma (Wang et al., 2011). Furthermore, they demonstrated that δ-catenin promotes the invasion of astrocytoma cells by upregulating Rac1 activity. Frattini et al. demonstrates that loss-of-function mutations in CTNND2 target neurospecific genes and are associated with the transformation of glioma cells along a highly aggressive mesenchymal phenotype (Frattini et al., 2013). Shimizu et al. demonstrated that overexpression of δ-catenin increased the ex vivo invasiveness of glioma cells, whereas knockdown of δ-catenin decreased the invasiveness of glioma cells, suggesting δ-catenin as a potential therapeutic target for the treatment of aggressive glioma when used in combination with bevacizumab (Shimizu et al., 2019). Hu et al. demonstrated that the expression level of δ-catenin in tumor tissues of medulloblastoma patients was significantly elevated in comparison to normal tissues (Hu et al., 2022). This may potentially impede the invasion of medulloblastoma cells by inhibiting the epithelial-mesenchymal transformation (EMT) pathway, and may also predict a favorable prognosis of medulloblastoma patients. Collectively, the aberrant expression of δ-catenin is frequently observed in neural-related cancers. However, the current research on δ-catenin in nerve-related cancers is limited and underlying mechanism remains to be elucidated. The more studies are needed to explore the clinical significance of δ-catenin as a biomarker for diagnosis and treatment of those cancers.
In addition to nerve-related cancers, aberrant expression of δ-catenin is also seen in other cancers. Huang et al. (2018) demonstrated that the removal of the CTNND2 gene resulted in a notable reduction in the tumorigenesis and metastatic ability of Lewis lung cancer cells. Conversely, the overexpression of δ-catenin enhanced the subcutaneous tumorigenesis and distant. The expression levels of a group of linking genes, including CTNND2, were significantly correlated with the overall survival rate of lung adenocarcinoma patients (Xie et al., 2024). Wang et al. demonstrated that δ-catenin (CTNND2) may serve as a prognostic biomarker for lung adenocarcinoma (Wang et al., 2020). Therefore, δ-catenin may play an important role in the pathogenesis of lung adenocarcinoma.
In human prostate cancer, δ-catenin displays exon mutations and promotes cancer cell survival, adaptation and metabolic reprogramming (Nopparat et al., 2015). Nucleotide polymorphisms in the 5′untranslated region (UTR) of the CTNND2 gene were linked to a high Gleason score and poorly differentiated prostate adenocarcinoma (Wang et al., 2009). Furthermore, functional nonsense mutations in CTNND2 have been identified as a promotor in the development of prostate cancer. The rearrangement of CTNND2 loci, including gene replication, is a prevalent phenomenon in clinically significant prostate cancer (Zhang P. et al., 2018). This may serve as a potential underlying mechanism for δ-catenin overexpression. Lu et al. identified a coordinated regulation of δ-catenin expression by E2F1 and Hes1 in prostate cancer progression (Lu et al., 2010). Both the activating transcription factor E2F1 and repressive transcription factor Hes1 regulated the expression of δ-catenin (Lu et al., 2010). Other studies have found that the expression levels of PSMA and δ-catenin in prostate cancer tissue are significantly higher than those in normal prostate tissue, pointing that these two proteins could serve as potential diagnostic biomarker for prostate cancer (Burger et al., 2002).
In liver cancer, a number of studies have proven that δ-catenin may play a role in the progression of liver cancer. Hypoxia can induce the expression of δ-catenin, thereby promoting hepatocellular carcinoma (HCC) progression through the Wnt signaling pathway (Huang et al., 2019). Hypoxia can also induce the expression of EIF3J-AS1 and δ-catenin, and result in the downregulation of miR-122-5p in HCC cells and enhanced proliferation, migration, and invasion of HCC cells (Yang et al., 2019). Zhu et al. identified 10 genes associated with iron -dependent cell death, including δ-catenin (Zhu et al., 2022). Yue et al. proposed that δ-catenin may represent a promising therapeutic target for the treatment of intrahepatic cholangiocarcinom (Yue et al., 2023). Additionally, recent studies have indicated that δ-catenin may play a role in the progression of esophageal cancer and may serve as a potential biomarker for esophageal cancer through single-cell RNA sequencing analysis from esophageal adenocarcinoma patients and matched normal-adjacent tissue (Maity et al., 2022). Singh et al. employed genome-wide methylation analysis to ascertain that CTNND2 promoter was hypomethylated and overexpressed in esophageal cancer (Singh et al., 2015). A combination of bioinformatics analysis and experimental verification has demonstrated that δ-catenin is significantly correlated with the prognosis of patients with pancreatic ductal adenocarcinoma (Zhang Y. L. et al., 2018). In breast cancer, δ-catenin is a potential contributor to the progression of breast cancer. δ-catenin may be a direct target of HOXB7, which is overexpressed in breast cancer (Heinonen et al., 2015).
Malignant transformation is frequently accompanied by loss of cell polarity and changes in cell morphology and the alterations in the connections between cancer cells and neighboring cells. δ-catenin is a cell junction protein and its change in expression is likely involved in many processes of cancer cell changes. δ-catenin as a biomarker for early diagnosis of cancers deserves further and in-depth investigation.
Most of the existing literature on δ-catenin has been focused on neurological related diseases and cancer. However, δ-catenin is not only closely associated with neuro-related diseases and cancers, but also associated with inflammation, pain, kidney disease, diarrhea, diabetes, dental health, stroke, and so on. First, δ-catenin is upregulated in patients with asthma, and δ-catenin is the target of miR-218-5p. The mum-mir-218-5p expression was also decreased with δ-catenin expression increased in the airway of mouse model of allergic airway inflammation (Liang et al., 2020). Furthermore, δ-catenin is markedly abundant in cadherin-mediated pathways that regulate inflammation of cerebrovascular endothelial (Mackinnon et al., 2016).
Second, pancreatitis patients, especially chronic pancreatitis and recurrent acute pancreatitis, suffer different degrees of pain and persistent inflammation. δ-catenin was associated with all forms of pain and all causes of pancreatitis, and three SNPs of CTNND2 gene was linked to the pain of pancreatitis by using candidate genetic association investigation (Dunbar et al., 2021). The gene region of CTNND2 is associated with postoperative analgesic orthognathic surgery (Nishizawa et al., 2022).
Third, the δ-catenin expression level was higher in kidney tissue from scleroderma renal crisis patients by immunostaining analysis (Stern et al., 2020). δ-catenin is a potential biomarker that contribute to the formation of small-vessel stroke (Cárcel-Márquez et al., 2024). The δ-catenin level from peripheral blood may be potential diagnostic biomarker for ischemic strokes (Simats et al., 2020).
Forth, Nrf2 can regulate Wnt signaling by targeting δ-catenin, which explains the negative effects of gestational diabetes on lung development in late embryonic development and may be used clinically to prevent and treat lung developmental abnormalities caused by gestational diabetes (He et al., 2023). Besides, δ-catenin may be a biomarker for insulin-related disease risk (Arpón et al., 2019).
Finally, δ-catenin is associated with malaria (Maity et al., 2022), dental health (Lauritano et al., 2019; Ma et al., 2015), placental abruption (Workalemahu et al., 2018), buffalo milk production (Erdoğan et al., 2024; Du et al., 2019), and pig farming (Liu H. et al., 2021; Zhao et al., 2022). Thus, it is clear that δ-catenin, which regulates cellular adhesion and nerves, is involved in a number of biological processes, and is not only associated with disease, but also has the ability to influence many aspects of biological traits. A comprehensive summary of δ-catenin functional studies is presented in Table 1.
δ-catenin also participates in the Wnt/β-catenin signaling pathway and further affect cell function. Wnt/β-catenin signaling pathway is frequently associated with the onset and progression of tumors. Notably, δ-catenin has a variety of post-translational modifications, including phosphorylation (Luckert et al., 2011), methylation (Arpón et al., 2019), and palmitoylation (Zhang X. L. et al., 2018), which can alter the localization, structure, interaction, and function of the protein. Moreover, the prior research on δ-catenin focuses on neurodevelopment primarily and synaptic maturation and differentiation. The following section will discuss the epigenetic modification processes associated with CTNND2 and the relevant signaling mechanisms.
In the prostate cancer cells, the tyrosine residues of δ-catenin can be phosphorylated after basic fibroblast growth factor (bFGF) interacting with fibroblast growth factor receptor 1 (FGFR1) (Chen et al., 2021). After phosphorylation, the binding between δ-catenin and glycogen synthase kinase 3β (GSK3β) was weakened, while the stability of δ-catenin was enhanced, facilitating the nuclear transport of β-catenin. δ-catenin phosphorylation promoted E-cadherin process and increased the total protein expression level, and result in the nuclear redistribution of β-catenin, thus enhancing the proliferation and migration of prostate cancer cells. Furthermore, the function of δ-catenin in the Wnt/β-catenin signaling pathway can also be regulated by phosphorylation. Phosphorylated δ-catenin enhanced the intracellular transport of β-catenin and the transcriptional activity of the downstream genes, by interacting with β-catenin. δ-catenin phosphorylation can also be mediated by other kinases, including Src family kinases, focal adhesion kinase (FAK), Janus kinase (JAK) (Chen et al., 2021), and MAPK JNK (Edbauer et al., 2009). The Ser-447 in δ-catenin is phosphorylated by MAPK JNK in a synaptic activity-dependent manner in neurons (Edbauer et al., 2009). A δ-catenin mutant defective in Ser-447 phosphorylation showed enhanced ability to promote dendrite branching in cultured neurons (Edbauer et al., 2009).
δ-catenin can regulate the expression of MTA2, D1 and MMP7 in a methylation-dependent manner via Kaiso, a δ-catenin-bound transcription factor. Methylation-specific PCR and ChIP analysis revealed that δ-catenin regulated gene expression via Kaiso and DNA methylation, e.g., regulation of MTA2 expression by δ-catenin through Kaiso depends on the methylation status of the MTA2 promoter (Dai et al., 2011). In addition to DNA and protein methylation, the methylation of CTNND2 mRNA may play a significant role in regulating its expression and function, given that m6A methylation is a prevalent RNA modification that can influence mRNA stability, translation efficiency, and localization. CTNND2 methylation and δ-catenin interactions with other proteins involved in methylation are also important in signaling and gene regulation, including Wnt/β-catenin signaling (Vershinin et al., 2016) and Wnt signaling (Lu et al., 2015).
The palmitoylation of δ-catenin is mainly regulated by DHHC5, a palmitoyl transferase. DHHC5 enhances the hydrophobicity of δ-catenin by conjugating palmitic acid to the cysteine residue of δ-catenin, thereby promoting its localization and function on the cell membrane. Following enhanced synaptic activity, the cadherin-binding protein δ-catenin is subjected to a brief palmitoylation process by DHHC5, which enhances the interaction between δ-catenin and cadherin at the synapse (Brigidi et al., 2014). The palmitoylation of δ-catenin plays a crucial role in synaptic plasticity and memory formation. Increased synaptic activity induces palmitoylation of δ-catenin, a process that increases the interaction of δ-catenin with N-cadherin, thereby stabilizing synaptic connections and promoting extension of the postsynaptic spine. Specifically, palmitoylated δ-catenin contributes to the stabilization of N-cadherin in the synapse and promotes the insertion of GluA1 and GluA2 subunits into the synaptic membrane, thereby increasing the amplitude of the small excitatory postsynaptic currents (Brigidi et al., 2014). Furthermore, palmitoylation of δ-catenin has been demonstrated to be a critical factor in the pathogenesis of neuropathic pain. In a rat model of neuropathic pain, there was an increase in the levels of palmitoylated delta-catenin and palmitoyltransferase DHHC3 in sensory neurons of the dorsal root ganglion (DRG). Inhibiting palmitoyl transferase or reducing the abundance of delta-catenin in DRG may alleviate neuropathic pain caused by oxaliplatin or nerve damage. The palmitoylation of δ-catenin induced by the inflammatory cytokine TNF-α promotes its interaction with the voltage-gated sodium channel Nav1.6 and the driver protein KIF3A. This interaction facilitates the transport of Nav1.6 across the plasma membrane of DRG neurons, which in turn leads to mechanical hypersensitivity and abnormal pain (Zhang X. L. et al., 2018)
The Wnt/β-catenin signaling pathway plays a crucial role in a multitude of biological processes, including cell proliferation, migration, invasion, EMT, angiogenesis, immune escape, phenotypic transition, clonal evolution and drug resistance (Ramakrishna et al., 2023). As a regulator of the Wnt/β-catenin signaling pathway, the abnormal expression and function of δ-catenin in cancers may be closely related to cancer development and progression. δ-catenin may enhance survival and adaptation of cancer cells by promoting activation of the Wnt/β-catenin signaling pathway and metabolic reprogramming, thereby supporting the growth and progression of tumor.
δ-catenin may play a significant role in the malignant progression through the activation of canonical Wnt signaling and the maintenance of cancer stem cells (Huang et al., 2018). In prostate cancer cells, δ-catenin is present in complexes with E-cadherin, p120 and α-catenin and β-catenin. Increased expression level in δ-catenin results in its further stabilization, as well as the stabilization and upregulation of its binding partners, including E-cadherin, p120, α-catenin and β-catenin. Anti-degradation and overexpression of δ-catenin isomers activate the Wnt signaling pathway by increasing nuclear β-catenin levels and subsequently stimulating Tcf/Lef transcriptional targets (Zhang P. et al., 2018). Hypoxia induced δ-catenin to promote the progression of liver cancer in mice through the Wnt signaling pathway. The hypoxia-induced transcription factor HIF1α can directly upregulate the expression of δ-catenin. The increased expression of δ-catenin then leads to an increase in its binding partner, which further leads to an increase in the expression of β-catenin, and leads to activation of the Wnt signaling pathway (Huang et al., 2019). Furthermore, nuclear factor erythroid 2-related factor 2 regulate the Wnt signaling pathway by targeting δ-catenin by RNA sequencing and luciferase reporter gene analysis (He et al., 2023).
In addition, studies have found that CTNND2 gene mutation can also participate in the Wnt/β-catenin signaling pathway. Functional nonsense mutations in CTNND2 promote the development of prostate cancer, and these mutations affect the activity of the Wnt/β-catenin signaling pathway. This suggests that δ-catenin may contribute to the progression of prostate cancer by affecting the Wnt/β-catenin signaling pathway. The mechanisms underlying δ-catenin-mediated regulation of WNT/β-catenin signaling pathway are summarized in Figure 2.
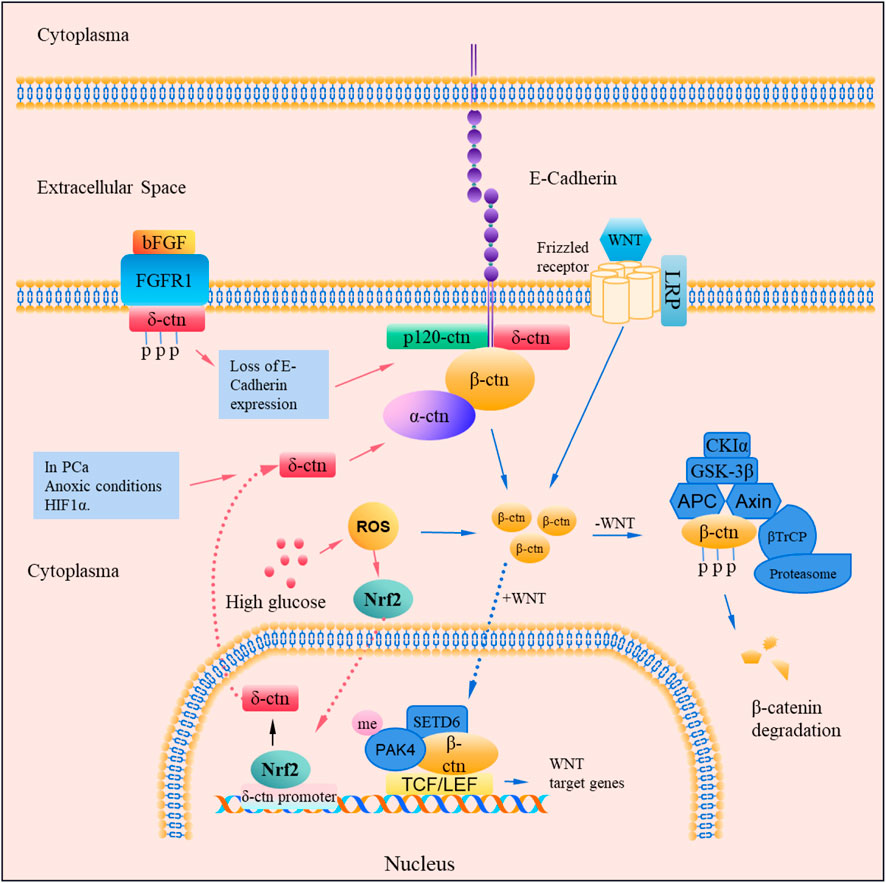
Figure 2. Mechanisms of CTNND2 involved in the WNT/β-catenin pathway. δ-catenin exists in protein complexes with E-cadherin, p120, α-catenin and β-catenin. Increased expression of δ-catenin leads to upregulation and stabilization of its binding partners and can activate the Wnt signaling pathway by increasing the nuclear β-catenin level which in turn can stimulate the activation of TCF/LEF transcriptional targets. In prostate cancer cells, bFGF can phosphorylate the tyrosine residues of δ-catenin mediated by FGFR1, and enhance fragmentation of E-Cadherin, and promote intranuclear redistribution of β-catenin. Nrf2 increases the expression of CTNND2, which regulates the WNT/β-catenin signaling pathway. In the WNT signaling pathway, β-catenin stabilizes and binds to TCF/LEF in the nucleus to regulate target genes; and β-catenin can be phosphorylated and degraded by the GSK-3β protein complex.
In addition to its involvement in the Wnt/β-catenin pathway, δ-catenin regulates neurodevelopment and signaling in vivo through other pathways. For example, δ-catenin can slow synaptic maturation and promote neuronal integrity; human specific protein SRGAP2C can enhance synaptic accumulation of δ-catenin in human neurons. Besides, CTNND2 deficiency can lead to loss of SYNGAP1 synapse. δ-catenin sets the pace of synaptic maturation and contributes to synaptic regeneration of human neurons through regulation of the human-specific protein SRGAP2C. δ-catenin is essential for the synaptic accumulation of SYNGAP1 and the formation of ID/Autism Spectrum Disorder-related protein complexes, which are uniquely regulated in humans and shape the synaptic developmental trajectories (Assendorp et al., 2024b).
During the differentiation of primary neurons, δ-catenin levels are increased while REST and TRIM28 protein levels are decreased, and the δ-catenin expression may be co-regulated by REST and TRIM28 protein (Lee et al., 2016). Moreover, δ-catenin binds to the last PDZ domain of S-SCAM through its carboxyl terminal, suggesting that δ-catenin regulate the molecular organization associated with synaptic connection through its interaction with S-SCAM (Ide et al., 1999). δ-catenin is also a potential iron death-related gene and subject to post-transcriptional regulation by circRNA/miRNA and m1A/m5C/m6A modifications in HCC (Zhu et al., 2022).
There are many lines of evidence suggest the association between CTNND2 and autism spectrum disorder. CTNND2 gene knockout mouse models exhibit behaviors characteristic of autism spectrum disorder and display a reduction in the density of dendritic spines in the hippocampus. The deletion of exon 2 of the CTNND2 gene is linked to a number of neurological deficits, including social impairment in the prefrontal cortex of the brain, loss of dendritic spines, damage to inhibitory neurons, and the inhibition of the phosphatidylinositol-3-kinase (PI3K)/protein kinase B (Akt) signaling pathway. Melatonin enhances the synaptic function of gamma-aminobutyric neurons by triggering the PI3K/Akt signaling cascade, which provides a potential therapeutic target for neurodevelopmental disorders caused by CTNND2 deficiency (Wang L. Y. et al., 2024). Figure 3 summarizes the currently available studies with a focus on CTNND2-related neuronal disorders and potential underlying mechanisms.
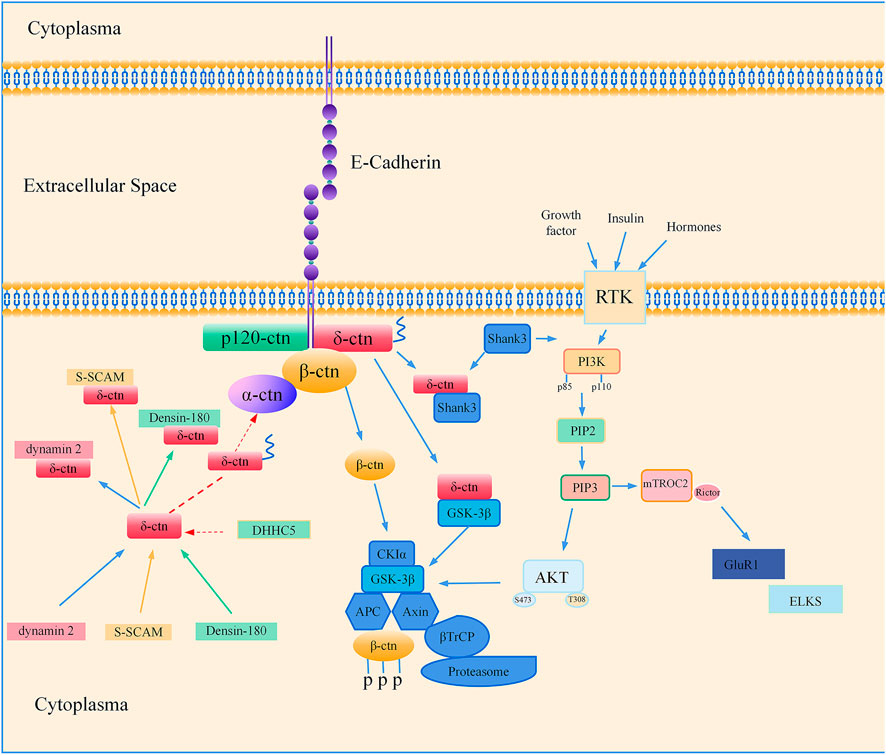
Figure 3. Regulatory mechanisms of δ-catenin associated with neural development and function. Co-localization and direct interaction of dynamin 2/NPRAP in neuroblastoma in vivo; NPRAP/δ-catenin is involved in synaptic junctions through its carboxy-terminal binding to the last PDZ structural domain of S-SCAM; Densin-180 co-localizes with δ-catenin/NPRAP; δ-catenin is transiently palmitoylated by DHHC5, and palmitoylation increases δ-catenin-adhesion protein interactions at the synapse; δ-catenin interacts directly with Shank3, which allows targeting of postsynaptic sites. Shank3 is an upstream node of the PI3K/AKT signaling pathway, and Rictor is a key regulatory and structural subunit of mTORC2 signaling. Downregulation of Rictor leads to altered expression of postsynaptic proteins such as GluR1 and ELKS; GSK-3β forms a complex with delta-catenin.
There are also many studies on the role of δ-catenin in cancer development and progression through a range of molecular actions, including Wnt/β-catenin signaling pathway. The expression of δ-catenin can be regulated by the repressive transcription factor Hes1 and activating transcription factor E2F1 during the progression of prostate cancer (Lu et al., 2010). δ-catenin may impede the invasion of medulloblastoma cells by inhibiting the EMT pathway, and has predicts a positive prognosis of medulloblastoma patients (Hu et al., 2022). δ-catenin signaling in neural diseases and cancer is summarized in Table 2.
The aberrant expression or mutation of δ-catenin has been linked to a range of diseases, including cat-call syndrome (CdCS), autism spectrum disorder (ASD), and certain types of cancer. Most recent research advancements have shed light on therapeutic developments in the context of δ-catenin signaling.
Firstly, gene replacement therapy for cat-call syndrome (CdCS). CdCS is caused by heterozygous deletion of the short arm of chromosome 5 (5p15.2), and CTNND2 deficiency and leads to mental retardation and synaptic developmental abnormalities. The gene replacement therapy restored CTNND2 expression and improves cognitive function in a rat model of CdCS constructed by CRISPR/Cas9 and successfully elevated δ-catenin levels in the brain, alleviated neuroinflammation and synaptic pathology, and significantly ameliorated cognitive deficits using an adeno-associated virus (AAVPHP.eB) vector (Shen et al., 2025).
Second, the development of small-molecule drugs targets neurodevelopmental disorders associated with CTNND2 deletion. CTNND2 knockout mouse models have demonstrated that its deletion affects synaptic protein synthesis through the downregulation of the PI3K/Akt/mTOR pathway, resulting in cerebellar developmental deficits and abnormalities in motor function. These findings provide a foundation for the potential development of PI3K/Akt/mTOR pathway activators or compounds that enhance synaptic plasticity (e.g., NMDA receptor modulators) (Wang L. et al., 2024). Conversely, the inflammatory response in the prefrontal cortex and hippocampus in the CdCS model suggests that combined anti-inflammatory agents (e.g., JAK/STAT inhibitors) may enhance efficacy (Shen et al., 2025).
Third, targeted therapy against CTNND2 fusion gene-driven cancer. In non-small cell lung cancer, the CTNND2-ROS1 fusion gene has been identified as a critical driver of tumor growth, operating through the activation of the ROS1 tyrosine kinase. This pathway is susceptible to ALK inhibitors, such as crizotinib and loratinib, highlighting a targeted therapeutic approach with potential efficacy enhancement. Therefore, the development of more efficient second-generation inhibitors (e.g., Repotinib, Taletrectinib) against CTNND2-ROS1 fusion is crucial to overcome the resistance mutation.
Fourth, the potential of CTNND2-based therapy for neurodegenerative diseases. δ-catenin plays a crucial role in synaptic stability and neuronal survival, and its aberrant expression has been linked to diseases such as Alzheimer’s disease (Koutras and Lévesque, 2011). A promising therapeutic strategy involves increasing δ-catenin expression through the use of small molecules that either activate CTNND2 transcription or inhibit its degradation, such as ubiquitinase inhibitors. Alternatively, the use of histone deacetylase inhibitors in the context of epigenetic modifications has been explored as a means to upregulate δ-catenin expression.
The field of CTNND2/δ-catenin research is rapidly developing, bringing many exciting discoveries, but also many challenges. Here are some key directions and potential difficulties for future research.
δ-catenin plays an important role in the development and function of the nervous system. However, the precise mechanisms of action remain unclear, and further researches are required to elucidate the exact mechanisms that δ-catenin influences neurodevelopment, especially in terms of synaptic formation and maturation. For instance, further investigation is required to ascertain how δ-catenin regulates synaptic maturation and neuronal excitability, whether δ-catenin is associated with other aspects of the nervous system, and how these processes are linked to the development of disorders, such as intellectual disability and autism spectrum disorder. In addition, cancer is often accompanied by dysregulation of neuro-regulation, and whether the role of δ-catenin in cancer pathogenesis is through neural regulation, such as regulating neurons or axonal progression.
Second, CTNND2 gene variations. Alterations of CTNND2 are associated with a range of conditions, including intellectual disability, autism, attention deficit hyperactivity disorder, and cancer. However, whether CTNND2 can be exploited to serve as potential therapeutic target remains unclear. Further researches are required to determine the role of CTNND2 variations in larger populations and their exact association with these diseases and to explore potential therapeutic targets. A better understanding of the function and mechanism on δ-catenin would help explore therapeutic strategies targeting CTNND2 expression or function to treat the related diseases, e.g., the development of small molecule drugs, gene therapies, or cell therapies.
Third, technical challenges on CTNND2 researches. It is very difficult to precisely manipulate of δ-catenin expression and to simulate δ-catenin function in complex biological systems. Novel tools and models need to be developed to elucidate the function of δ-catenin in vivo, particularly focus on its role in the brain. CTNND2 is one of the genes unique to humans that may have played a pivotal role in the evolution of the human brain. Further studies are required to elucidate the role of CTNND2 in human evolution, particularly focus on its effects on the characteristics of cortical synapses, including prolonged maturation and neoteny of synapses. And all of those investigations require more advanced technology.
In summary, the field of CTNND2 gene research is full of opportunities and challenges. With the advancement of science and technology and the innovation of research methods, we are expected to have a deeper understanding of CTNND2 in the next few years and provide new strategies for the diagnosis and treatment of related diseases. However, in-depth studies are clearly warranted in the future to understand the definitive roles of CTNND2/δ-catenin signaling in physiology and pathology, e.g., neural disorders and cancers, and to translate findings into clinical management of those diseases.
YZ: Conceptualization, Writing – original draft. KX: Conceptualization, Funding acquisition, Supervision, Writing – review and editing. TJ: Conceptualization, Funding acquisition, Writing – original draft, Writing – review and editing.
The author(s) declare that financial support was received for the research and/or publication of this article. The work is partly supported by National Natural Science Foundation of China (#82072632) and Guangzhou Municipality Bureau of Science and Technology, Guangzhou, China (#202102010033) and Natural Science Foundation of Guangdong Province, China (#2022A1515012585).
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Adegbola, A., Lutz, R., Nikkola, E., Strom, S. P., Picker, J., and Wynshaw-Boris, A. (2020). Disruption of CTNND2, encoding delta-catenin, causes a penetrant attention deficit disorder and myopia. Hum. Genet. Genomics Adv. 1, 100007. doi:10.1016/j.xhgg.2020.100007
Almeida, V. T., Chehimi, S. N., Gasparini, Y., Nascimento, A. M., Carvalho, G. F. S., Montenegro, M. M., et al. (2023). Cri-du-Chat syndrome: revealing a familial atypical deletion in 5p. Mol. Syndromol. 13, 527–536. doi:10.1159/000524371
Arpón, A., Santos, J. L., Milagro, F. I., Cataldo, L. R., Bravo, C., Riezu-Boj, J. I., et al. (2019). Insulin sensitivity is associated with lipoprotein lipase (LPL) and catenin delta 2 (CTNND2) DNA methylation in peripheral white blood cells in non-diabetic young women. Int. J. Mol. Sci. 20, 2928. doi:10.3390/ijms20122928
Asadollahi, R., Oneda, B., Joset, P., Azzarello-Burri, S., Bartholdi, D., Steindl, K., et al. (2014). The clinical significance of small copy number variants in neurodevelopmental disorders. J. Med. Genet. 51, 677–688. doi:10.1136/jmedgenet-2014-102588
Assendorp, N., Fossati, M., Libé-Philippot, B., Christopoulou, E., Depp, M., Rapone, R., et al. (2024a). CTNND2 moderates the pace of synaptic maturation and links human evolution to synaptic neoteny. Cell Rep. 43, 114797. doi:10.1016/j.celrep.2024.114797
Assendorp, N., Fossati, M., Libe-Philippot, B., Christopoulou, E., Depp, M., Rapone, R., et al. (2024b). CTNND2 moderates the pace of synaptic maturation and links human evolution to synaptic neoteny. Cell Rep. 43, 114797. doi:10.1016/j.celrep.2024.114797
Bareiss, S., Kim, K., and Lu, Q. (2010). Delta-catenin/NPRAP: a new member of the glycogen synthase kinase-3beta signaling complex that promotes beta-catenin turnover in neurons. J. Neurosci. Res. 88, 2350–2363. doi:10.1002/jnr.22414
Belcaro, C., Dipresa, S., Morini, G., Pecile, V., Skabar, A., and Fabretto, A. (2015). CTNND2 deletion and intellectual disability. Gene 565, 146–149. doi:10.1016/j.gene.2015.03.054
Brigidi, G. S., Sun, Y., Beccano-Kelly, D., Pitman, K., Mobasser, M., Borgland, S. L., et al. (2014). Palmitoylation of δ-catenin by DHHC5 mediates activity-induced synapse plasticity. Nat. Neurosci. 17, 522–532. doi:10.1038/nn.3657
Brown, J., Bothma, H., Veale, R., and Willem, P. (2011). Genomic imbalances in esophageal carcinoma cell lines involve Wnt pathway genes. World J. Gastroenterol. 17, 2909–2923. doi:10.3748/wjg.v17.i24.2909
Burger, M. J., Tebay, M. A., Keith, P. A., Samaratunga, H. M., Clements, J., Lavin, M. F., et al. (2002). Expression analysis of delta-catenin and prostate-specific membrane antigen: their potential as diagnostic markers for prostate cancer. Int. J. Cancer 100, 228–237. doi:10.1002/ijc.10468
Cárcel-Márquez, J., Muiño, E., Gallego-Fabrega, C., Cullell, N., Lledós, M., Llucià-Carol, L., et al. (2024). Sex-stratified genome-wide association study in the Spanish population identifies a novel locus for lacunar stroke. Stroke 55, 2462–2471. doi:10.1161/STROKEAHA.124.047833
Cerruti Mainardi, P. (2006). Cri du Chat syndrome. Orphanet J. Rare Dis. 1, 33. doi:10.1186/1750-1172-1-33
Chen, G., An, N., Zhu, Y., Zhou, R., Noh, M. G., Kim, H., et al. (2021). bFGF-mediated phosphorylation of δ-catenin increases its protein stability and the ability to induce the nuclear redistribution of β-catenin. Am. J. Cancer Res. 11, 3877–3892.
Chen, Z. N., Li, X. J., Cui, X. Z., Zhang, L. W., Liu, Q., Lu, Y. L., et al. (2023). Association of CTNND2 gene polymorphism with schizophrenia: two-sample case-control study in Chinese Han population. Int. J. Psychiat Med. 58, 433–448. doi:10.1177/00912174231164669
Chhaya, N., and Chan, T. (2013)2021). Retinopathy in a full-term infant with cri-du-chat syndrome. R. I. Med. J. 104, 37–39.
Dai, S. D., Wang, Y., Zhang, J. Y., Zhang, D., Zhang, P. X., Jiang, G. Y., et al. (2011). Upregulation of δ-catenin is associated with poor prognosis and enhances transcriptional activity through Kaiso in non-small-cell lung cancer. Cancer Sci. 102, 95–103. doi:10.1111/j.1349-7006.2010.01766.x
Du, C., Deng, T., Zhou, Y., Ye, T., Zhou, Z., Zhang, S., et al. (2019). Systematic analyses for candidate genes of milk production traits in water buffalo (Bubalus Bubalis). Anim. Genet. 50, 207–216. doi:10.1111/age.12739
Dunbar, E. K., Greer, P. J., Amann, S. T., Alkaade, S., Banks, P., Brand, R., et al. (2021). Pain experience in pancreatitis: strong association of genetic risk loci for anxiety and PTSD in patients with severe, constant, and constant-severe pain. Am. J. Gastroenterol. 116, 2128–2136. doi:10.14309/ajg.0000000000001366
Edbauer, D., Cheng, D. M., Batterton, M. N., Wang, C. F., Duong, D. M., Yaffe, M. B., et al. (2009). Identification and characterization of neuronal mitogen-activated protein kinase substrates using a specific phosphomotif antibody. Mol. Cell Proteomics 8, 681–695. doi:10.1074/mcp.M800233-MCP200
Einarsdottir, E., Peyrard-Janvid, M., Darki, F., Tuulari, J. J., Merisaari, H., Karlsson, L., et al. (2017). Identification of NCAN as a candidate gene for developmental dyslexia. Sci. Rep. 7, 9294. doi:10.1038/s41598-017-10175-7
Erdoğan, M., Çinkaya, S., Brenig, B., Çelikeloğlu, K., Demirtaş, M., Sarıibrahimoğlu, S., et al. (2024). Genome-wide association studies for milk production traits and persistency of first calving Holstein cattle in Türkiye. Front. Vet. Sci. 11, 1461075. doi:10.3389/fvets.2024.1461075
Frattini, V., Trifonov, V., Chan, J. M., Castano, A., Lia, M., Abate, F., et al. (2013). The integrated landscape of driver genomic alterations in glioblastoma. Nat. Genet. 45, 1141–1149. doi:10.1038/ng.2734
Halder, D., Mandal, C., Lee, B. H., Lee, J. S., Choi, M. R., Chai, J. C., et al. (2015). PCDHB14- and GABRB1-like nervous system developmental genes are altered during early neuronal differentiation of NCCIT cells treated with ethanol. Hum. Exp. Toxicol. 34, 1017–1027. doi:10.1177/0960327114566827
Hassani, N. F., Woike, D., Martens, V., Klüssendorf, M., Hönck, H. H., Harder, S., et al. (2020). Targeting of δ-catenin to postsynaptic sites through interaction with the Shank3 N-terminus. Mol. Autism 11, 85. doi:10.1186/s13229-020-00385-8
Hatzfeld, M. (2005). The p120 family of cell adhesion molecules. Eur. J. Cell Biol. 84, 205–214. doi:10.1016/j.ejcb.2004.12.016
He, M. Y., Zhang, P., Shi, N. N., Li, T. T., Wang, J. Y., He, L. K., et al. (2023). Nrf2 is involved in hyperglycemia-induced abnormal lung development through both antioxidation-dependent and antioxidation-independent activation. Am. J. Resp. Cell Mol. 69, 197–209. doi:10.1165/rcmb.2022-0345OC
He, W. Z., Liu, W. Q., Zhong, X. Q., Chen, X. L., Li, S. Y., Zhang, H. M., et al. (2012). Analysis of de novo copy number variations in a family affected with autism spectrum disorders using high-resolution array-based comparative genomic hybridization. Zhonghua Yi Xue Yi Chuan Xue Za Zhi 29, 266–269. doi:10.3760/cma.j.issn.1003-9406.2012.03.004
Heinonen, H., Lepikhova, T., Sahu, B., Pehkonen, H., Pihlajamaa, P., Louhimo, R., et al. (2015). Identification of several potential chromatin binding sites of HOXB7 and its downstream target genes in breast cancer. Int. J. Cancer 137, 2374–2383. doi:10.1002/ijc.29616
Ho, C., Zhou, J. H., Medina, M., Goto, T., Jacobson, M., Bhide, P. G., et al. (2000). delta-catenin is a nervous system-specific adherens junction protein which undergoes dynamic relocalization during development. J. Comp. Neurol. 420, 261–276. doi:10.1002/(sici)1096-9861(20000501)420:2<261::aid-cne8>3.0.co;2-q
Hofmeister, W., Nilsson, D., Topa, A., Anderlid, B. M., Darki, F., Matsson, H., et al. (2015). CTNND2-a candidate gene for reading problems and mild intellectual disability. J. Med. Genet. 52, 111–122. doi:10.1136/jmedgenet-2014-102757
Hu, Y. J., Zhu, S. H., Xu, R. Z., Wang, M. X., Chen, F. R., Zhang, Z. S., et al. (2022). Delta-catenin attenuates medulloblastoma cell invasion by targeting EMT pathway. Front. Genet. 13, 867872. doi:10.3389/fgene.2022.867872
Huang, F., Chen, J. Y., Lan, R. L., Wang, Z., Chen, R. Q., Lin, J. A., et al. (2019). Hypoxia induced δ-Catenin to enhance mice hepatocellular carcinoma progression via Wnt signaling. Exp. Cell Res. 374, 94–103. doi:10.1016/j.yexcr.2018.11.011
Huang, F., Chen, J. Y., Wang, Z., Lan, R. L., Fu, L. X., and Zhang, L. R. (2018). δ-Catenin promotes tumorigenesis and metastasis of lung adenocarcinoma. Oncol. Rep. 39, 809–817. doi:10.3892/or.2017.6140
Huang, F. Y., Chiu, P. M., Tam, K. F., Kwok, Y. K., Lau, E. T., Tang, M. H., et al. (2006). Semi-quantitative fluorescent PCR analysis identifies PRKAA1 on chromosome 5 as a potential candidate cancer gene of cervical cancer. Gynecol. Oncol. 103, 219–225. doi:10.1016/j.ygyno.2006.02.028
Huang, Y. T., Jiang, Z. J., Gao, X. Y., Luo, P., and Jiang, X. F. (2021). ARMC subfamily: structures, functions, evolutions, interactions, and diseases. Front. Mol. Biosci. 8, 791597. doi:10.3389/fmolb.2021.791597
Ide, N., Hata, Y., Deguchi, M., Hirao, K., Yao, I., and Takai, Y. (1999). Interaction of S-SCAM with neural plakophilin-related Armadillo-repeat protein/delta-catenin. Biochem. Bioph Res. Co. 256, 456–461. doi:10.1006/bbrc.1999.0364
Izawa, I., Nishizawa, M., Ohtakara, K., and Inagaki, M. (2002). Densin-180 interacts with delta-catenin/neural plakophilin-related armadillo repeat protein at synapses. J. Biol. Chem. 277, 5345–5350. doi:10.1074/jbc.M110052200
Jun, G., Moncaster, J. A., Koutras, C., Seshadri, S., Buros, J., McKee, A. C., et al. (2012). δ-Catenin is genetically and biologically associated with cortical cataract and future Alzheimer-related structural and functional brain changes. Plos One 7, e43728. doi:10.1371/journal.pone.0043728
Kang, H. J., Kim, K. T., Park, Y., Yoo, K. H., Kim, J. W., Lee, J. Y., et al. (2021). Genetic markers for depressive disorders with earlier age at onset. Prog. Neuro-Psychoph. 108, 110176. doi:10.1016/j.pnpbp.2020.110176
Keil, R., Schulz, J., and Hatzfeld, M. (2013). p0071/PKP4, a multifunctional protein coordinating cell adhesion with cytoskeletal organization. Biol. Chem. 394, 1005–1017. doi:10.1515/hsz-2013-0114
Kim, K., Sirota, A., Chen, Yh Y. H., Jones, S. B., Dudek, R., Lanford, G. W., et al. (2002). Dendrite-like process formation and cytoskeletal remodeling regulated by delta-catenin expression. Exp. Cell Res. 275, 171–184. doi:10.1006/excr.2002.5503
Koutras, C., Lessard, C. B., and Lévesque, G. (2011). A nuclear function for the presenilin 1 neuronal partner NPRAP/δ-catenin. J. Alzheimers Dis. 27, 307–316. doi:10.3233/JAD-2011-110536
Koutras, C., and Lévesque, G. (2011). Identification of novel NPRAP/δ-catenin-interacting proteins and the direct association of NPRAP with dynamin 2. PLoS One 6, e25379. doi:10.1371/journal.pone.0025379
Lauritano, D., Lucchese, A., Di Stasio, D., Della Vella, F., Cura, F., Palmieri, A., et al. (2019). Molecular aspects of drug-induced gingival overgrowth: an in vitro study on amlodipine and gingival fibroblasts. Int. J. Mol. Sci. 20, 2047. doi:10.3390/ijms20082047
Lee, N., Park, S. J., Haddad, G., Kim, D. K., Park, S. M., Park, S. K., et al. (2016). Interactomic analysis of REST/NRSF and implications of its functional links with the transcription suppressor TRIM28 during neuronal differentiation. Sci. Rep-Uk 6, 39049. doi:10.1038/srep39049
Li, Q., Cai, D., Huang, H., Zhang, H., Bai, R., Zhao, X., et al. (2021). Phosphoproteomic profiling of the hippocampus of offspring rats exposed to prenatal stress. Brain Behav. 11, e2233. doi:10.1002/brb3.2233
Li, Y. J., Goh, L., Khor, C. C., Fan, Q., Yu, M., Han, S., et al. (2011a). Genome-wide association studies reveal genetic variants in CTNND2 for high myopia in Singapore Chinese. Ophthalmology 118, 368–375. doi:10.1016/j.ophtha.2010.06.016
Li, Y. J., Goh, L. A., Khor, C. C., Fan, Q. A., Yu, M. A., Han, S. Y., et al. (2011b). Genome-wide association studies reveal genetic variants in CTNND2 for high myopia in Singapore Chinese. Ophthalmology 118, 368–375. doi:10.1016/j.ophtha.2010.06.016
Liang, Y. X., Feng, Y. C., Wu, W. L., Chang, C. L., Chen, D. A., Chen, S. C., et al. (2020). microRNA-218-5p plays a protective role in eosinophilic airway inflammation via targeting δ-catenin, a novel catenin in asthma. Clin. Exp. Allergy 50, 29–40. doi:10.1111/cea.13498
Ligon, C., Seong, E., Schroeder, E. J., DeKorver, N. W., Yuan, L., Chaudoin, T. R., et al. (2020). δ-Catenin engages the autophagy pathway to sculpt the developing dendritic arbor. J. Biol. Chem. 295, 10988–11001. doi:10.1074/jbc.RA120.013058
Liu, H., Song, H., Jiang, Y., Jiang, Y., Zhang, F., Liu, Y., et al. (2021b). A single-step genome wide association study on body size traits using imputation-based whole-genome sequence data in yorkshire pigs. Front. Genet. 12, 629049. doi:10.3389/fgene.2021.629049
Liu, J., and Zhang, H. X. (2014). Polymorphism in the 11q24.1 genomic region is associated with myopia: a comprehensive genetic study in Chinese and Japanese populations. Mol. Vis. 20, 352–358.
Liu, J., Zhang, R., Sun, L., Zheng, Y., Chen, S., Chen, S. L., et al. (2021a). Genotype-phenotype correlation and interaction of 4q25, 15q14 and MIPEP variants with myopia in southern Chinese population. Br. J. Ophthalmol. 105, 869–877. doi:10.1136/bjophthalmol-2019-314782
Lu, B. Y., Jiang, D., Wang, P. F., Gao, Y., Sun, W. M., Xiao, X. S., et al. (2011). Replication study supports CTNND2 as a susceptibility gene for high myopia. Invest Ophth Vis. Sci. 52, 8258–8261. doi:10.1167/iovs.11-7914
Lu, J. P., Zhang, J. A., Kim, K., Case, T. C., Matusik, R. J., Chen, Y. H., et al. (2010). Human homolog of Drosophila Hairy and enhancer of split 1, Hes1, negatively regulates δ-catenin (CTNND2) expression in cooperation with E2F1 in prostate cancer. Mol. Cancer 9, 304. doi:10.1186/1476-4598-9-304
Lu, L., Gao, Y., Zhang, Z., Cao, Q., Zhang, X. N., Zou, J. H., et al. (2015). Kdm2a/b lysine demethylases regulate canonical Wnt signaling by modulating the stability of nuclear β-catenin. Dev. Cell 33, 660–674. doi:10.1016/j.devcel.2015.04.006
Lu, Q. (2010). δ-Catenin dysregulation in cancer: interactions with E-cadherin and beyond. J. Pathol. 222, 119–123. doi:10.1002/path.2755
Lu, Q., Aguilar, B. J., Li, M., Jiang, Y., and Chen, Y. H. (2016). Genetic alterations of δ-catenin/NPRAP/Neurojungin (CTNND2): functional implications in complex human diseases. Hum. Genet. 135, 1107–1116. doi:10.1007/s00439-016-1705-3
Lu, Q., Paredes, M., Medina, M., Zhou, J. H., Cavallo, R., Peifer, M., et al. (1999). delta-catenin, an adhesive junction-associated protein which promotes cell scattering. J. Cell Biol. 144, 519–532. doi:10.1083/jcb.144.3.519
Luckert, K., Götschel, F., Sorger, P. K., Hecht, A., Joos, T. O., and Pötz, O. (2011). Snapshots of protein dynamics and post-translational modifications in one experiment-β-catenin and its functions. Mol. Cell Proteomics 10, M110.007377. doi:10.1074/mcp.M110.007377
Ma, J., Zhao, D., Wu, Y., Xu, C., and Zhang, F. (2015). Cyclic stretch induced gene expression of extracellular matrix and adhesion molecules in human periodontal ligament cells. Arch. Oral Biol. 60, 447–455. doi:10.1016/j.archoralbio.2014.11.019
Mackinnon, M. J., Ndila, C., Uyoga, S., Macharia, A., Snow, R. W., Band, G., et al. (2016). Environmental correlation analysis for genes associated with protection against malaria. Mol. Biol. Evol. 33, 1188–1204. doi:10.1093/molbev/msw004
Maity, A. K., Stone, T. C., Ward, V., Webster, A. P., Yang, Z., Hogan, A., et al. (2022). Novel epigenetic network biomarkers for early detection of esophageal cancer. Clin. Epigenetics 14, 23. doi:10.1186/s13148-022-01243-5
Medina, M., Marinescu, R. C., Overhauser, J., and Kosik, K. S. (2000). Hemizygosity of δ-catenin (CTNND2) is associated with severe mental retardation in cri-du-chat syndrome. Genomics 63, 157–164. doi:10.1006/geno.1999.6090
Miller, D. E., Squire, A., and Bennett, J. T. (2020). A child with autism, behavioral issues, and dysmorphic features found to have a tandem duplication within CTNND2 by mate-pair sequencing. Am. J. Med. Genet. A 182, 543–547. doi:10.1002/ajmg.a.61442
Moncaster, J. A., Moir, R. D., Burton, M. A., Chadwick, O., Minaeva, O., Alvarez, V. E., et al. (2022). Alzheimer's disease amyloid-β pathology in the lens of the eye. Exp. Eye Res. 221, 108974. doi:10.1016/j.exer.2022.108974
Nishizawa, D., Nagashima, M., Kasai, S., Hasegawa, J., Nakayama, K., Ebata, Y., et al. (2022). Associations between the C3orf20 rs12496846 polymorphism and both postoperative analgesia after orthognathic and abdominal surgeries and C3orf20 gene expression in the brain. Pharmaceutics 14, 727. doi:10.3390/pharmaceutics14040727
Nivard, M. G., Mbarek, H., Hottenga, J. J., Smit, J. H., Jansen, R., Penninx, B. W., et al. (2014). Further confirmation of the association between anxiety and CTNND2: replication in humans. Genes Brain Behav. 13, 195–201. doi:10.1111/gbb.12095
Nopparat, J., Zhang, J., Lu, J. P., Chen, Y. H., Zheng, D., Neufer, P. D., et al. (2015). δ-Catenin, a Wnt/β-catenin modulator, reveals inducible mutagenesis promoting cancer cell survival adaptation and metabolic reprogramming. Oncogene 34, 1542–1552. doi:10.1038/onc.2014.89
Oh, J. H., Lee, S., Thor, M., Rosenstein, B. S., Tannenbaum, A., Kerns, S., et al. (2023). Predicting the germline dependence of hematuria risk in prostate cancer radiotherapy patients. Radiother. Oncol. 185, 109723. doi:10.1016/j.radonc.2023.109723
Paffenholz, R., and Franke, W. W. (1997). Identification and localization of a neurally expressed member of the plakoglobin/armadillo multigene family. Differentiation 61, 293–304. doi:10.1046/j.1432-0436.1997.6150293.x
Pauly, M., Krumbiegel, M., Trumpp, S., Braig, S., Rupprecht, T., Kraus, C., et al. (2024). Severe manifestation of Rauch-Azzarello syndrome associated with biallelic deletion of CTNND2. Clin. Genet. 106, 180–186. doi:10.1111/cge.14532
Peifer, M., McCrea, P. D., Green, K. J., Wieschaus, E., and Gumbiner, B. M. (1992). The vertebrate adhesive junction proteins beta-catenin and plakoglobin and the Drosophila segment polarity gene armadillo form a multigene family with similar properties. J. Cell Biol. 118, 681–691. doi:10.1083/jcb.118.3.681
Qu, J., Cheng, X., Liu, M., and Zhang, Q. (2024). CTNND2 gene expression in melanoma tissues and its effects on the malignant biological functions of melanoma cells. Transl. Cancer Res. 13, 6347–6363. doi:10.21037/tcr-24-2159
Ramakrishna, K., Nalla, L. V., Naresh, D., Venkateswarlu, K., Viswanadh, M. K., Nalluri, B. N., et al. (2023). WNT-Β catenin signaling as a potential therapeutic target for neurodegenerative diseases: current status and future perspective. Diseases 11, 89. doi:10.3390/diseases11030089
Sardina, J. M., Walters, A. R., Singh, K. E., Owen, R. X., and Kimonis, V. E. (2014). Amelioration of the typical cognitive phenotype in a patient with the 5pter deletion associated with Cri-du-chat syndrome in addition to a partial duplication of CTNND2. Am. J. Med. Genet. A 164, 1761–1764. doi:10.1002/ajmg.a.36494
Shen, J., Wang, Y., Liu, Y., Lan, J., Long, S., Li, Y., et al. (2025). Behavioral Abnormalities, Cognitive Impairments, Synaptic Deficits, and Gene Replacement Therapy in a CRISPR Engineered Rat Model of 5p15.2 Deletion Associated With Cri du Chat Syndrome. Adv. Sci. (Weinh), e2415224. doi:10.1002/advs.202415224
Sheth, F., Gohel, N., Liehr, T., Akinde, O., Desai, M., Adeteye, O., et al. (2012). Gain of Chromosome 4qter and Loss of 5pter: An Unusual Case with Features of Cri du Chat Syndrome. Case Rep. Genet. 2012, 153405–153414. doi:10.1155/2012/153405
Shimizu, T., Ishida, J., Kurozumi, K., Ichikawa, T., Otani, Y., Oka, T., et al. (2019). δ-Catenin promotes bevacizumab-induced glioma invasion. Mol. Cancer Ther. 18, 812–822. doi:10.1158/1535-7163.MCT-18-0138
Simats, A., Ramiro, L., García-Berrocoso, T., Briansó, F., Gonzalo, R., Martín, L., et al. (2020). A mouse brain-based multi-omics integrative approach reveals potential blood biomarkers for ischemic stroke. Mol. Cell Proteomics 19, 1921–1936. doi:10.1074/mcp.RA120.002283
Singh, V., Singh, A. P., Sharma, I., Singh, L. C., Sharma, J., Borthakar, B. B., et al. (2019). Epigenetic deregulations of Wnt/β-catenin and transforming growth factor beta-Smad pathways in esophageal cancer: outcome of DNA methylation. J. Cancer Res. Ther. 15, 192–203. doi:10.4103/jcrt.JCRT_634_17
Singh, V., Singh, L. C., Vasudevan, M., Chattopadhyay, I., Borthakar, B. B., Rai, A. K., et al. (2015). Esophageal cancer epigenomics and integrome analysis of genome-wide methylation and expression in high risk northeast Indian population. Omics 19, 688–699. doi:10.1089/omi.2015.0121
Srinivasalu, N., Lu, C., Pan, M., Reinach, P. S., Wen, Y., Hu, Y., et al. (2018). Role of cyclic adenosine monophosphate in myopic scleral remodeling in Guinea pigs: a microarray analysis. Invest Ophthalmol. Vis. Sci. 59, 4318–4325. doi:10.1167/iovs.17-224685
Stern, E. P., Guerra, S. G., Chinque, H., Acquaah, V., González-Serna, D., Ponticos, M., et al. (2020). Analysis of anti-RNA polymerase III antibody-positive systemic sclerosis and altered GPATCH2L and CTNND2 expression in scleroderma renal crisis. J. Rheumatol. 47, 1668–1677. doi:10.3899/jrheum.190945
Tuncay, I. O., Parmalee, N. L., Khalil, R., Kaur, K., Kumar, A., Jimale, M., et al. (2022). Analysis of recent shared ancestry in a familial cohort identifies coding and noncoding autism spectrum disorder variants. Npj Genom Med. 7, 13. doi:10.1038/s41525-022-00284-2
Turner, T. N., Sharma, K., Oh, E. C., Liu, Y. F. P., Collins, R. L., Sosa, M. X., et al. (2015). Loss of δ-catenin function in severe autism. Nature 520, 51–56. doi:10.1038/nature14186
van Rootselaar, A. F., Groffen, A. J., de Vries, B., Callenbach, P. M. C., Santen, G. W. E., Koelewijn, S., et al. (2017). δ-Catenin (CTNND2) missense mutation in familial cortical myoclonic tremor and epilepsy. Neurology 89, 2341–2350. doi:10.1212/WNL.0000000000004709
Vaz, R., Edwards, S., Dueñas-Rey, A., Hofmeister, W., and Lindstrand, A. (2023). Loss of ctnnd2b affects neuronal differentiation and behavior in zebrafish. Front. Neurosci. 17, 1205653. doi:10.3389/fnins.2023.1205653
Vershinin, Z., Feldman, M., Chen, A., and Levy, D. (2016). PAK4 methylation by SETD6 promotes the activation of the wnt/β-catenin pathway. J. Biol. Chem. 291, 6786–6795. doi:10.1074/jbc.M115.697292
Vrijenhoek, T., Buizer-Voskamp, J. E., van der Stelt, I., Strengman, E., Sabatti, C., Geurts van Kessel, A., et al. (2008). Recurrent CNVs disrupt three candidate genes in schizophrenia patients. Am. J. Hum. Genet. 83, 504–510. doi:10.1016/j.ajhg.2008.09.011
Wang, L., Xu, M., Wang, Y., Wang, F., Deng, J., Wang, X., et al. (2024a). Melatonin improves synapse development by PI3K/Akt signaling in a mouse model of autism spectrum disorder. Neural Regen. Res. 19, 1618–1624. doi:10.4103/1673-5374.387973
Wang, L. Y., Xu, M., Wang, Y., Wang, F. F., Deng, J., Wang, X. Y., et al. (2024b). Melatonin improves synapse development by PI3K/Akt signaling in a mouse model of autism spectrum disorder. Neural Regen. Res. 19, 1618–1624. doi:10.4103/1673-5374.387973
Wang, M. H., Dong, Q. Z., Zhang, D., and Wang, Y. J. (2011). Expression of delta-catenin is associated with progression of human astrocytoma. Bmc Cancer 11, 514. doi:10.1186/1471-2407-11-514
Wang, S., Deng, J., Wang, Y., Wang, L., Shao, C., Tang, H., et al. (2021a). Rictor is involved in Ctnnd2 deletion-induced impairment of spatial learning and memory but not autism-like behaviors. Front. Bioscience-Landmark 26, 335–346. doi:10.52586/4947
Wang, T., Chen, Y. H., Hong, H., Zeng, Y., Zhang, J., Lu, J. P., et al. (2009). Increased nucleotide polymorphic changes in the 5'-untranslated region of delta-catenin (CTNND2) gene in prostate cancer. Oncogene 28, 555–564. doi:10.1038/onc.2008.399
Wang, X., Xu, M., Xu, Q., Yang, F., Tang, H., Shao, C., et al. (2021b). Rictor is involved in Ctnnd2 deletion-induced impairment of spatial learning and memory but not autism-like behaviors. Front. Biosci. Landmark Ed. 26, 335–346. doi:10.52586/4947
Wang, Y. Y., Zhang, L., Chen, Y. T., Li, M., Ha, M. W., and Li, S. B. (2020). Screening and identification of biomarkers associated with the diagnosis and prognosis of lung adenocarcinoma. J. Clin. Lab. Anal. 34, e23450. doi:10.1002/jcla.23450
Wen, K., Zhang, L., Cai, Y., Teng, H., Liang, J., Yue, Y., et al. (2023). Identification and characterization of extrachromosomal circular DNA in patients with high myopia and cataract. Epigenetics 18, 2192324. doi:10.1080/15592294.2023.2192324
Workalemahu, T., Enquobahrie, D. A., Gelaye, B., Sanchez, S. E., Garcia, P. J., Tekola-Ayele, F., et al. (2018). Genetic variations and risk of placental abruption: a genome-wide association study and meta-analysis of genome-wide association studies. Placenta 66, 8–16. doi:10.1016/j.placenta.2018.04.008
Xie, B., Wu, T., Hong, D. G., and Lu, Z. (2024). Comprehensive landscape of junctional genes and their association with overall survival of patients with lung adenocarcinoma. Front. Mol. Biosci. 11, 1380384. doi:10.3389/fmolb.2024.1380384
Xu, M., Wang, L. Y., Wang, Y., Deng, J., Wang, X. Y., Wang, F. F., et al. (2023b). Melatonin ameliorates sleep-wake disturbances and autism-like behaviors in the Ctnnd2 knock out mouse model of autism spectrum disorders. Genes Brain Behav. 22, e12852. doi:10.1111/gbb.12852
Xu, M., Wang, X., Wang, L., Wang, S., Deng, J., Wang, Y., et al. (2023a). Effects of chronic sleep restriction on the neuro-phenotypes of Ctnnd2 knockout mice. Brain Behav. 13, e3075. doi:10.1002/brb3.3075
Yang, X., Yao, B. W., Niu, Y. S., Chen, T. X., Mo, H. Y., Wang, L., et al. (2019). Hypoxia-induced lncRNA EIF3J-AS1 accelerates hepatocellular carcinoma progression via targeting miR-122-5p/CTNND2 axis. Biochem. Bioph Res. Co. 518, 239–245. doi:10.1016/j.bbrc.2019.08.039
Yin, H., Li, D., Wang, Y., and Zhu, Q. (2019). Whole-genome resequencing analysis of Pengxian Yellow Chicken to identify genome-wide SNPs and signatures of selection. 3 Biotech. 9, 383. doi:10.1007/s13205-019-1902-6
Yu, Z. Q., Zhou, J. Q., Chen, X. L., Zhou, X. T., Sun, X. H., and Chu, R. Y. (2012). Polymorphisms in the CTNND2 gene and 11q24.1 genomic region are associated with pathological myopia in a Chinese population. Ophthalmologica. 228, 123–129. doi:10.1159/000338188
Yuan, L., Seong, E., Beuscher, J. L., and Arikkath, J. (2015). δ-Catenin regulates spine architecture via cadherin and PDZ-dependent interactions. J. Biol. Chem. 290, 10947–10957. doi:10.1074/jbc.M114.632679
Yue, Y., Tao, J., An, D., and Shi, L. (2023). Exploring the role of tumor stemness and the potential of stemness-related risk model in the prognosis of intrahepatic cholangiocarcinoma. Front. Genet. 13, 1089405. doi:10.3389/fgene.2022.1089405
Zhang, B., Willing, M., Grange, D. K., Shinawi, M., Manwaring, L., Vineyard, M., et al. (2016). Multigenerational autosomal dominant inheritance of 5p chromosomal deletions. Am. J. Med. Genet. A 170, 583–593. doi:10.1002/ajmg.a.37445
Zhang, P., Schaefer-Klein, J., Cheville, J. C., Vasmatzis, G., and Kovtun, I. V. (2018b). Frequently rearranged and overexpressed δ-catenin is responsible for low sensitivity of prostate cancer cells to androgen receptor and β-catenin antagonists. Oncotarget 9, 24428–24442. doi:10.18632/oncotarget.25319
Zhang, X. L., Ding, H. H., Xu, T., Liu, M., Ma, C., Wu, S. L., et al. (2018a). Palmitoylation of δ-catenin promotes kinesin-mediated membrane trafficking of Nav 1.6 in sensory neurons to promote neuropathic pain. Sci. Signal 11, eaar4394. doi:10.1126/scisignal.aar4394
Zhang, Y. L., Zhang, R. D., Ding, X., and Ai, K. X. (2018c). EFNB2 acts as the target of miR-557 to facilitate cell proliferation, migration and invasion in pancreatic ductal adenocarcinoma by bioinformatics analysis and verification. Am. J. Transl. Res. 10, 3514–3528.
Zhao, Y. X., Gao, G. X., Zhou, Y., Guo, C. X., Li, B., El-Ashram, S., et al. (2022). Genome-wide association studies uncover genes associated with litter traits in the pig. Animal 16, 100672. doi:10.1016/j.animal.2022.100672
Zheng, M., Simon, R., Mirlacher, M., Maurer, R., Gasser, T., Forster, T., et al. (2004). TRIO amplification and abundant mRNA expression is associated with invasive tumor growth and rapid tumor cell proliferation in urinary bladder cancer. Am. J. Pathol. 165, 63–69. doi:10.1016/S0002-9440(10)63275-0
Zhou, Y., Qiu, L., Sterpka, A., Wang, H., Chu, F., and Chen, X. (2019). Comparative phosphoproteomic profiling of type III adenylyl cyclase knockout and control, male, and female mice. Front. Cell Neurosci. 13, 34. doi:10.3389/fncel.2019.00034
Zhu, R. F., Gao, C., Feng, Q. Q., Guan, H. T., Wu, J. J., Samant, H., et al. (2022). Ferroptosis-related genes with post-transcriptional regulation mechanisms in hepatocellular carcinoma determined by bioinformatics and experimental validation. Ann. Transl. Med. 10, 1390. doi:10.21037/atm-22-5750
Keywords: cancer, CTNND2, δ-catenin, neurodevelopment, neuro-diseases
Citation: Zhang Y, Xie K and Jiang T (2025) Roles and regulation of δ-catenin in tumorigenesis and neuronal diseases. Front. Cell Dev. Biol. 13:1559059. doi: 10.3389/fcell.2025.1559059
Received: 16 January 2025; Accepted: 18 March 2025;
Published: 27 March 2025.
Edited by:
Brian D. Adams, Brain Institute of America, United StatesReviewed by:
Udayan Bhattacharya, NewYork-Presbyterian, United StatesCopyright © 2025 Zhang, Xie and Jiang. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Tingting Jiang, dHRqaWFuZ0BzY3V0LmVkdS5jbg==; Keping Xie, c2N1dG1lZGljaW5lQHNjdXQuZWR1LmNu
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.