- 1Department of Medical Genetics, Shahid Beheshti University of Medical Sciences, Tehran, Iran
- 2Urology and Nephrology Research Center, Shahid Beheshti University of Medical Sciences, Tehran, Iran
Coronary artery disease (CAD) is the main reason of cardiovascular mortalities worldwide. This condition is resulted from atherosclerotic occlusion of coronary arteries. MicroRNAs (miRNAs) are implicated in the regulation of proliferation and apoptosis of endothelial cells, induction of immune responses and different stages of plaque formation. Up-regulation of miR-92a-3p, miR-206, miR-216a, miR-574-5p, miR-23a, miR-499, miR-451, miR-21, miR-146a, and a number of other miRNAs has been reported in CAD patients. In contrast, miR-20, miR-107, miR-330, miR-383-3p, miR-939, miR-4306, miR-181a-5p, miR-218, miR-376a-3p, and miR-3614 are among down-regulated miRNAs in CAD. Differential expression of miRNAs in CAD patients has been exploited to design diagnostic or prognostic panels for evaluation of CAD patients. We appraise the recent knowledge about the role of miRNAs in the development of diverse clinical subtypes of CAD.
Introduction
Coronary artery disease (CAD) is the principal source of cardiovascular mortalities worldwide (1). In 2020, it is expected that 11.1 million patients die as a results of CAD related complications (2). Clinically, CAD has different categories ranging from stable angina pectoris to acute coronary syndromes which comprises unstable angina (UA) and myocardial infarction (MI) (3). The majority of MI cases are resulted from the establishment of acute intraluminal coronary thrombus inside an epicardial coronary artery and the subsequent occlusion of the coronary artery (4, 5). The acute coronary thrombosis results in a sudden decrease in the blood flow and induction of necrosis in the myocardial region which is takes the blood supply from this coronary artery (6). Some other cardiovascular pathologies might be associated with CAD. For instance, acute MI might lead to defects in functioning myocytes resulting in myocardial fibrosis and left ventricle dilatation. Subsequent induction of neurohormonal responses and left ventricle remodeling results in progressive weakening of the residual viable myocardium (7). Moreover, ischemic conditions leads to upsurge of endogenous catecholamines in the myocardial interstitial fluid which in turn increases myocardial apoptosis and fibrosis (8). Dysregulation of several microRNAs (miRNAs) has been displayed in different categories of CAD, potentiating these transcripts as biomarkers of this devastating condition (9). miRNAs have been shown to modulate gene expression at post transcriptional level via destroying mRNA targets or by obstructing their translation (10). Since each miRNA is capable of regulating expression of several transcripts, it is estimated that more than half of protein-coding genes in the human genome are influenced by miRNAs (11). Therefore, miRNAs can affect numerous important biological and cellular function such as cell differentiation, proliferation, and cell death in the cardiovascular system (12). Understanding the role of miRNAs in the pathogenesis of CAD would lead to identification of appropriate therapies for this global health problem. We appraise the recent knowledge about the role of miRNAs in the development of diverse clinical subtypes of CAD.
miRNAs in CAD
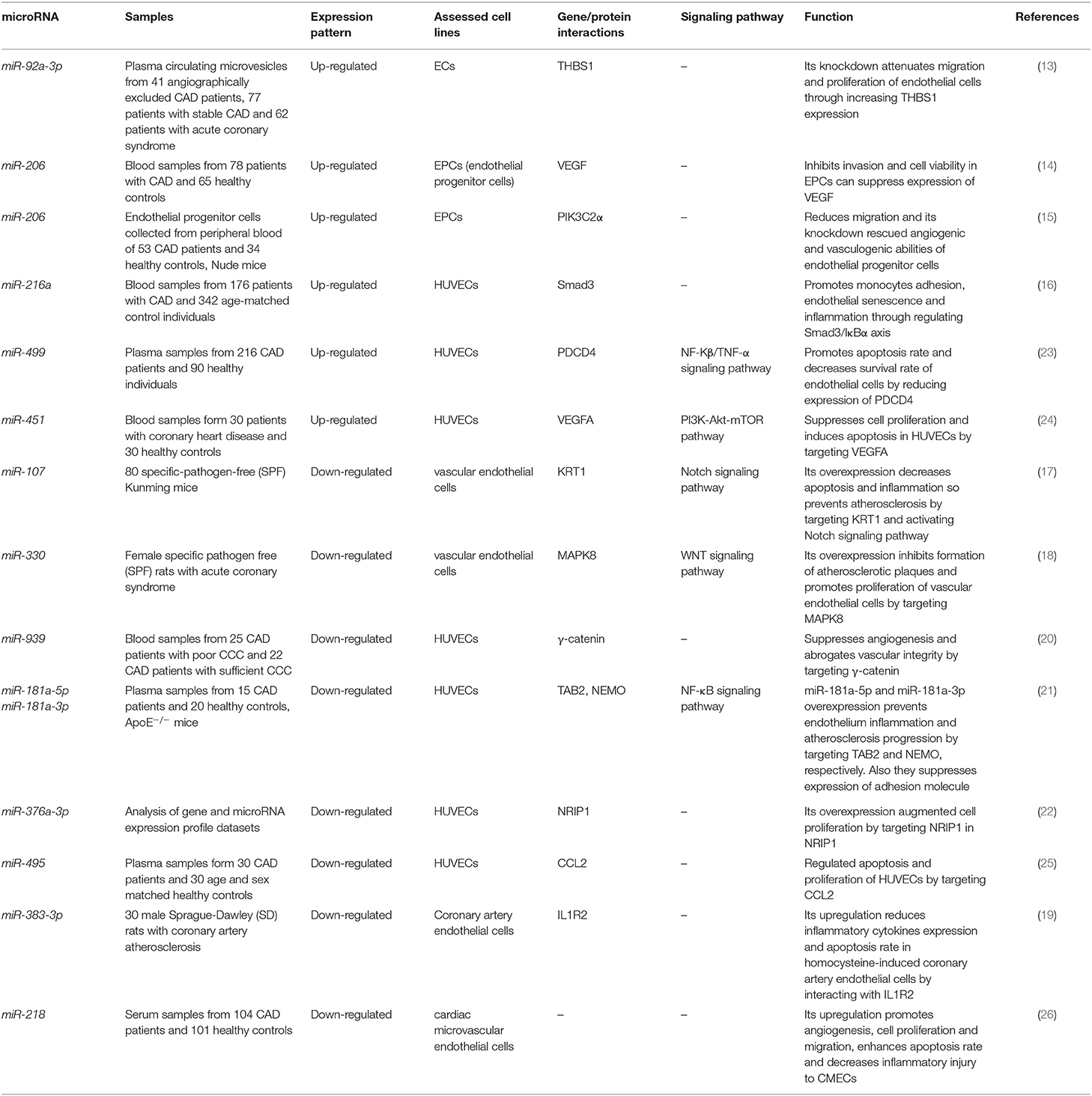
Function of miRNAs in CAD has been assessed in different cell types. Endothelial cells have been the mostly assessed cell type in this regard. Liu et al. have extracted circulating microvesicles (MVs) from plasma samples of CAD patients to assess signature of their miRNA constituents. Among miRNAs which were reported to regulate vascular performance, miR-92a-3p has been shown to be up-regulated in CAD cases compared with non-CAD individuals. MVs enclosing miR-92a-3p have been demonstrated to be mostly originated from endothelial cells. Treatment of these cells with oxidized LDL and IL-6 has resulted in up-regulation of miR-92a-3p levels in these cells and higher incorporation of this miRNA in MVs. Transport of these MVs to other endothelial cells has enhanced their migration and proliferation. miR-92a-3p exerts these functions through inhibition of expression of THBS1, the inhibitor of angiogenesis. Taken together, atherosclerosis enhances the incorporation of endothelial miR-92a-3p into MVs, which controls angiogenesis in recipient endothelial cells through a THBS1-associated route (13). Wang et al. have demonstrated up-regulation of miR-206 in endothelial progenitor cells as well as plasma samples gathered from CAD patients. However, expression levels of miR-206 have not been associated with clinicopathological characteristics of CAD patients. Functionally, miR-206 has been shown to inhibit the viability and invasion of endothelial progenitor cells in CAD patients, while enhancing apoptosis in these cells. miR-206 can also suppress expression of vascular endothelial growth factor (VEGF) (14). Moreover, this miRNA modulates endothelial progenitor cell functions through targeting the protein kinase PIK3C2α. This protein kinase has been shown to be down-regulated in endothelial progenitor cells of CAD patients. miR-206 silencing in these cells enhanced their angiogenic and vasculogenic capacities both in vitro and in an animal model of ischemia. Besides, miR-206 silencing enhanced activities of PIK3C2α, Akt, and endothelial nitric oxide synthase (15). miR-216a is another miRNA which is involved in endothelial aging and dysfunction through modulating expression of Smad3. Over-expression of miR-216a in human umbilical vein endothelial cells (HUVECs) has activated an untimely senescence-like feature in these cells which was accompanied by defects in proliferation and migration. The consequent suppression of Smad3 has resulted in enhancement of adhesion of these cells to monocytes, modulation of the destruction of NF-κB inhibitor alpha (IκBα) and stimulation of adhesion proteins. Levels of miR-216a has been shown to be elevated in the plasma samples of old CAD patients in association with higher susceptibility to CAD (16). Figure 1 shows the cascade of involvement of miR-216a in CAD.
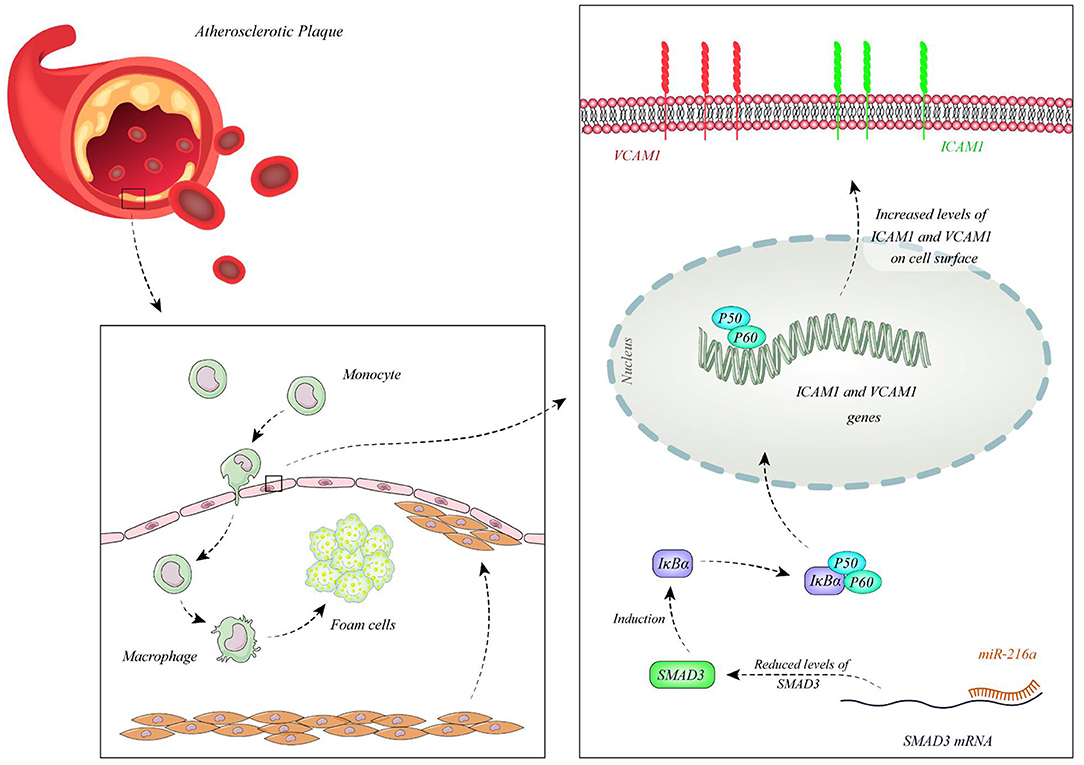
Figure 1. miR-216a is over-expressed in CAD patients. miR-216a attaches to 3' UTR of Smad3 and decreases its levels. Down-regulation of Smad3 leads to reduction of IκBα releasing NF-κB and enhancing its nuclear transport. Subsequent up-regulation of ICAM1 and VCAM1 enhances attachment of monocytes to endothelial cells promoting development of CAD (16).
Gao et al. have demonstrated high concentrations of lipids, atherosclerotic index, apoptotic index, and KRT1-positive expression while suppression of Notch signaling pathway in the atherosclerotic mice. miR-107 has been shown to bind with KRT1, thus reducing its expression. This miRNA has been down-regulated in animal models of CAD (17). Ren et al. have reported down-regulation of miR-330 in CAD group. Overexpression of miR-330 has been shown to inhibit atherosclerotic plaques creation whereas enhancing proliferation of vascular endothelial cells through modulating MAPK8 via the WNT signaling pathway (18). Lian et al. have shown down-regulation of miR-383-3p and up-regulation of IL1R2 in myocardial tissues of atherosclerotic animals. Forced over-expression of miR-383-3p has reduced expression of IL1R2, caspase-1, IL-1β, IL-6, and IL-18, ameliorated cell apoptosis in the coronary artery endothelial cells, while enhanced IL-10 levels, cell survival, and tube construction (19). Hou et al. have reported down-regulation of miR-939 in the blood of patients with adequate coronary collateral circulation compared with those having insufficient coronary collateral circulation. Up-regulation of miR-939 in HUVECs has remarkably suppressed proliferation, adhesion and tube construction, while increasing migration capacity of these cells. γ-catenin has been identified as a direct target of miR-939 (20). Expressions of both miR-181a-5p and miR-181a-3p have been lower in the aorta plaque and plasma of animal models of CAD. Up-regulation of these miRNAs considerably delays atherosclerotic plaque development in animals. These miRNAs have functional roles in the reduction of expression of pro-inflammatory proteins and diminishing the infiltration of macrophage, leukocyte and T cell into the atherosclerotic plaques through suppression of adhesion molecule expressions in HUVECs (21). miR-376a-3p has also been down-regulated in CAD samples. In vitro studies have shown the effects of miR-376a-3p silencing in the suppression of proliferation of HUVECs through modulating NRIP1 expression (22). Table 1 displays the functional roles of miRNAs in the development of CAD, based on the results of studies which have been conducted in endothelial cells.
Lai et al. have reported over-expression of miR-574-5p in the serum samples and vascular smooth muscle cells (VSMCs) of CAD patients. Up-regulation of miR-574-5p has enhanced cell proliferation and suppressed apoptotic processes in VSMCs through targeting ZDHHC14 (27). Down-regulation of miR-146a has been demonstrated to attenuate apoptosis of vascular smooth muscle cells. Autologous injection of endothelial stem cells in a rat model of acute myocardial infarction has led to downregulation of miR-146a levels, reduction of apoptosis in the myocardial cells and decrease in infarct area. Such effects have been accompanied by up-regulation of VEGF (28). Expression of miR-93 has been increased in ventricle tissues and blood samples of mice model of MI. Moreover, miR-93 has been shown to be released from cardiomyocytes cultured in hypoxic conditions. miR-93 suppresses apoptotic processes and guards cardiomyocytes from ischemia/reperfusion damage. miR-93 silencing has deteriorated cardiac remodeling in these animal models. Thus, miR-93 over-expression and release from cardiomyocytes has been regarded as an adaptive mechanism following MI to attenuate cardiac remodeling and heart failure (29). miR-448 has been shown to be over-expressed in vascular smooth muscle cells (VSMCs) obtained from atherosclerotic plaques of coronary artery compared with those obtained from normal arteries. Expression of this miRNA is induced by PDGF-bb, a growth factor that enhances proliferation of VSMCs. MEF2C has been recognized as a direct target of miR-448 in VSMCs, though its down-regulation miR-448 enhances VSMCs migration (30). Table 2 shows the list of CAD-related miRNAs whose function has been assessed in myocardial cells or vascular smooth muscle cells.
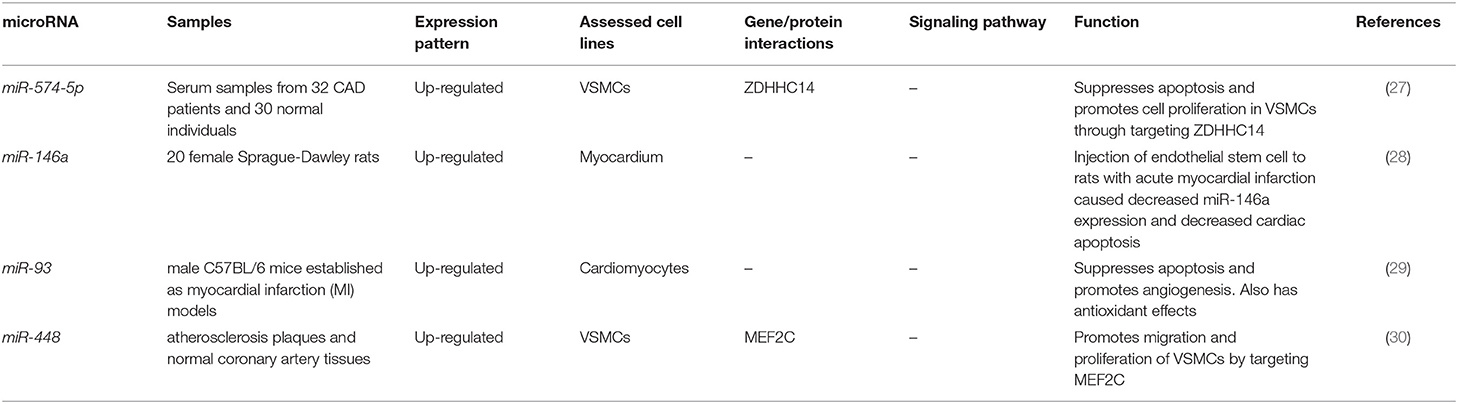
Table 2. CAD-related miRNAs whose function has been assessed in myocardial cells or vascular smooth muscle cells.
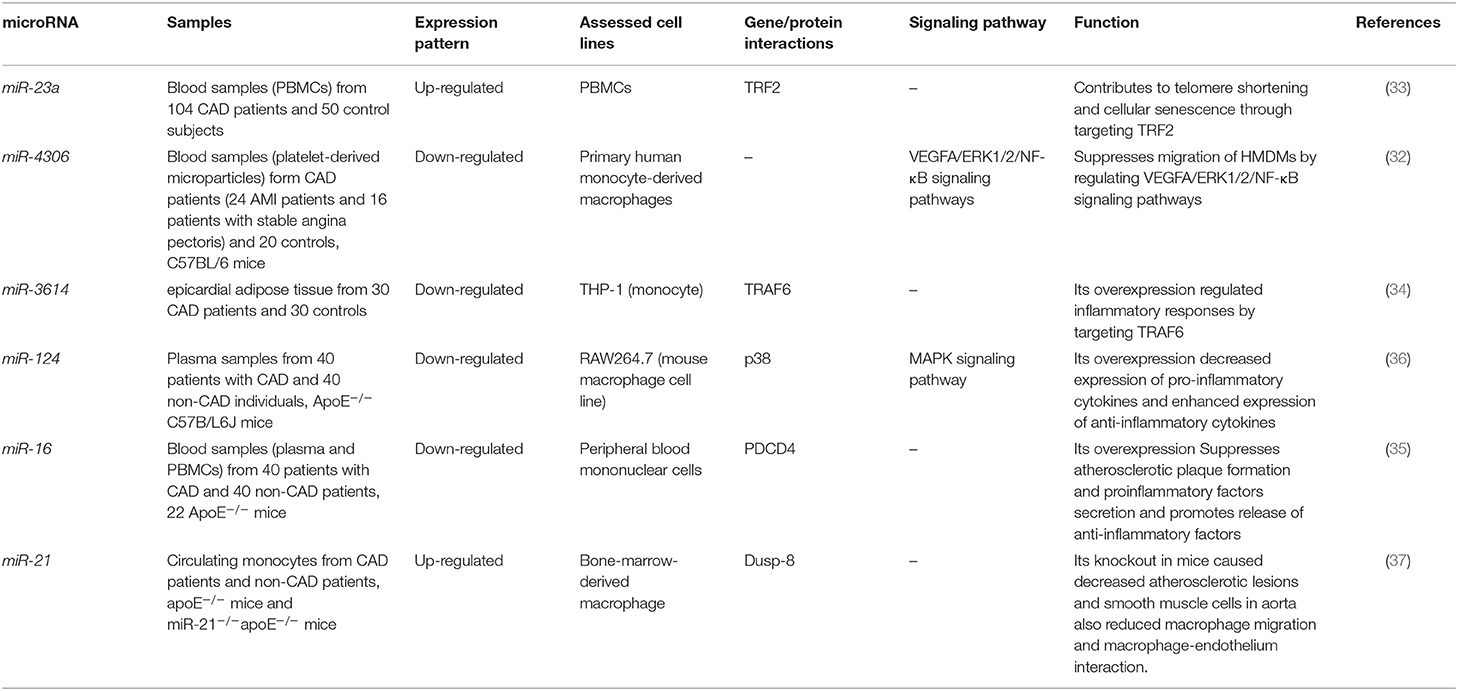
Wang et al. have reported down-regulation of miR-20 in animal models of CAD in association with over-expression of VEGF and PTEN. Levels of miR-20a have been up-regulated following exercise in CAD animals. Up-regulation of miR-20a has reduced levels of ET-1, TxA2, ANGII, PTEN and enhanced levels of eNOS, PGI2, and VEGF. miR-20a exerts its functions through binding with the 3′UTR of PTEN, thus enhancing cell survival and proliferation via induction of the PI3K/Akt signaling (31). Expression of miR-4306 has been decreased in platelets and platelet-originated microparticles of CAD patients. Plasma miRNA-4306 has been mostly fractionated with microparticles rather than Argonaute2 complexes or HDL. These microparticles have the ability to transfer miR-4306 into human monocyte-derived macrophages, thus suppressing their migration and decreasing the quantity of macrophages in cardiac tissue in mouse model of MI. Mechanistically, miR-4306 binds with VEGFA to suppress ERK/NF-κB signaling (32). Expression of miR-23a has been higher in the peripheral blood mononuclear cells (PBMCs) of CAD patients compared with control subjects parallel with down-regulation of TRF2 levels. Aggressive lipid lowering therapy has reduced miR-23a, enhanced TRF2 expression and attenuated telomere erosion through this route (33). Expression of miR-3614 has been decreased by lipopolysaccharide (LPS) in macrophages, while LPS-associated inflammatory damage can be attenuated by up-regulation of miR-3614. This miRNA has been shown to target TRAF6 and suppress phosphorylation of kinases in the MAPK and NF-κB cascades Therefore, miR-3614/TRAF6/MAPK/NF-κB cascade can suppress devastating inflammatory responses (34). Animal studies have shown the role of miR-16 in reduction of development of atherosclerotic plaques and suppression of accretion of inflammatory factors while enhancement of release of anti-inflammatory factors. Mechanistically, miR-16 exerts these effects through downregulation of PDCD4 and activation of p38 and ERK1/2, while inactivation of JNK pathway (35). Table 3 demonstrates the relevance of miRNAs with the pathogenesis of CAD through summarizing the results of studies which reported function of miRNAs in macrophages/monocytes.
Diagnostic/Prognostic Significance of miRNAs in CAD
Altered levels of miRNA in the circulation of CAD patients potentiates their usage as biomarkers in this condition. Zhong et al. have demonstrated differential expressions of tens of miRNAs in patients with UA or ST-segment elevation MI compared with normal controls. Receiver operating characteristics (ROC) curves have revealed miR-142-3p and miR-17-5p as possible markers for diagnosis of these two classes of CAD. Moreover, differential expressed miRNAs have been correlated with the pathological events during the course of CAD (38). Vahed et al. have reported down-regulation of miR-21 in the PBMCs of patients with insignificant coronary artery stenosis compared with CAD patients or healthy subjects. Levels of this miRNA have been negatively correlated with the PTEN. Moreover, they reported a gradual elevation miR-25 expression from healthy subjects to those with insignificant coronary artery stenosis and CAD patients. Expression levels of miR-21 and miR-25 in the PBMCs could differentiate three groups of study participants (39). Yao et al. have demonstrated the capacity of miRNAs in distinguishing CAD patients with heart failure (HF) from those without HF. Among the most significantly dysregulated miRNAs between these two groups of patients have miR-221, miR-19b-5p, and miR-25-5p. Combination of expression levels of these miRNAs in PBMCs and hypertension have been significantly correlated with higher risk of HF risk in CAD patients (40). Another miRNA with promising results in diagnostic approaches is miR-122-5p which could differentiate unstable CAD patients from healthy controls with accuracy of 0.9, yet its accuracy in differentiation of stable patients from controls was not appropriate (41). A brief review of studies which demonstrated this function is presented in Table 4.
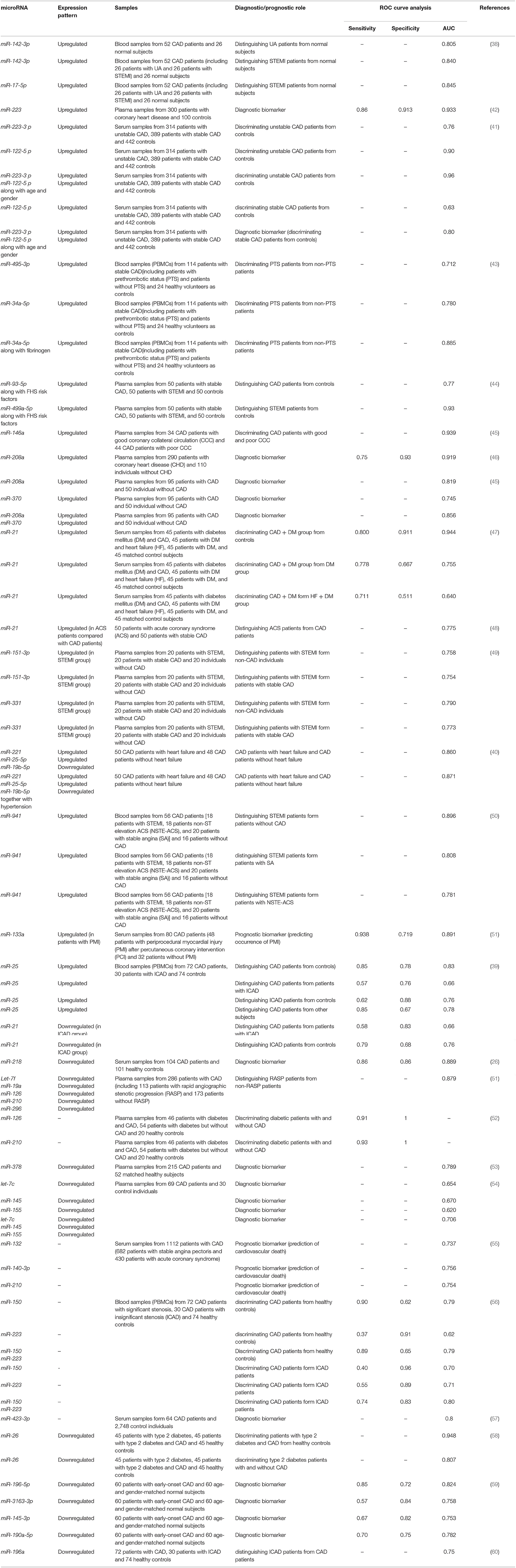
Table 4. Diagnostic/prognostic significance of miRNAs in CAD (UA, unstable angina; STEMI: ST-segment elevation myocardial infarction).
miRNA Polymorphisms and Copy Number Variations in CAD
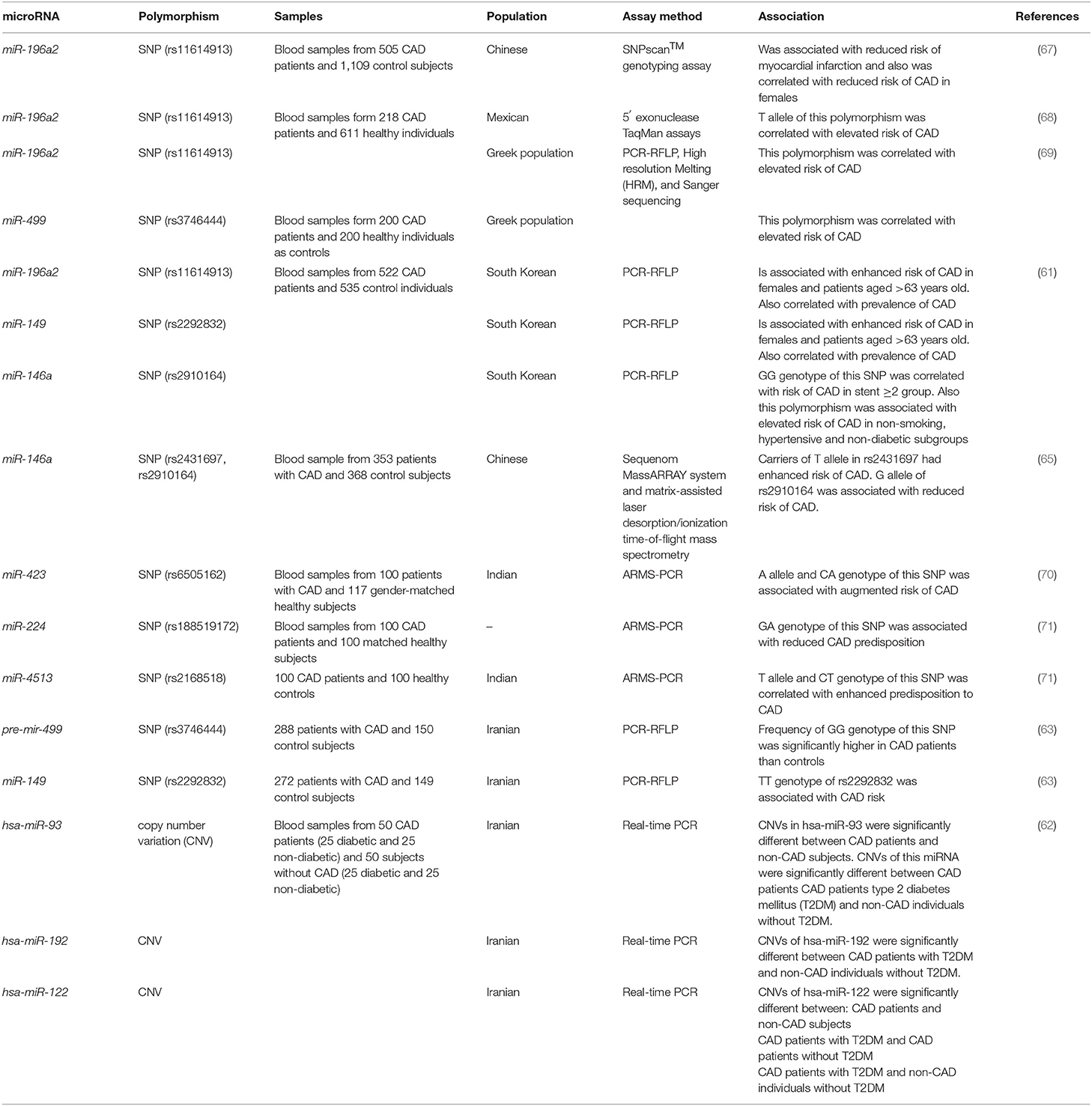
Both single nucleotide polymorphisms (SNPs) and copy number variations (CNVs) within miRNA coding genes have been associated with risk of CAD. Sung et al. have examined the relation between miR-146a, miR-149, miR-196a2, and miR-499 SNPs and CAD in a Korean population. They have reported association between the miR-149 rs2292832 and miR-196a2 rs11614913 SNPs and this disorder. Notably, the miR-146a rs2910164 GG genotype has been more prevalent among CAD patients with more than two stents. Moreover, combination of miR-146a G, miR-149 T, miR-196a2 C, and mIR-499 G alleles has been considerably associated with CAD occurrence. Certain SNPs have been reported to increase susceptibility to CAD in different subclasses of study participants such as non-smokers, hypertensive and non-diabetic individuals (61). Sohrabifar et al. have evaluated the presence of CNVs of hsa-miR-93, hsa-miR-122, hsa-miR-192 in CAD patients with or without type 2 diabetes mellitus. They have reported remarkable differences in the distribution of CNVs of hsa-miR-93 between CAD and non-CAD as well as between diabetic CAD and diabetic non-CAD individuals. In addition, hsa-miR-122 CNVs have been differently distributed among three subgroups (62). The rs2292832 miR-149 has been associated with risk of CAD in Iranian population. However, this SNP does not either affect the secondary structure of pre-miR-149 or the stability of the miRNA hairpin structure (63). As this SNP is located outside the sequence of mature miR-149, it has been proposed that it might affect the maturation process and therefore decrease expression of miR-149 (64). T allele of rs2431697 in miR-146a has been associated with higher risk of CAD (65). In addition, the rs2910164 within this miRNA affects risk of CAD (61). This SNP resides in the precursor of miR-146a and results in down-regulation of levels of mature miR-146a (66). Table 5 reviews the investigations which appraised the role of SNPs/CNVs in conferring risk of CAD.
Conclusions and Perspectives
Aberrant expression of miRNAs in CAD patients has been recognized through high throughput sequencing methods in addition to candidate gene assays. An example of the former type of assays has been conducted through investigation of Gene Expression Omnibus (GEO) database showing frequent differential expression of 150 genes and 5 miRNAs (22). Luciferase reporter assays have shown the functional interactions between a number of miRNAs and mRNAs (24, 72). miRNAs can regulate development of CAD through different mechanisms such as modulation of angiogenesis [miR-92a-3p (13), miR-939 (20), and miR-206 (14)], inflammatory responses [miR-181a-5p, miR-181a-3p (21), miR-216a (16), and miR-383-3p (19)], leukocyte adhesion [miR-21 (37) and miR-25 (39)] and modulation of activity of VSMCs [miR-574-5p (27)]. Notably, a number of miRNAs influence different aspects of this process or different targets in a certain process. For instance, miR-206 regulated expressions of VEGF, PIK3C2α, Akt, and endothelial nitric oxide synthase, all of them being involved in the angiogenic processes. NF-Kβ/TNF-α, PI3K-Akt-mTOR, WNT, and VEGFA/ERK1/2/NF-κB are among signaling pathways which are regulated by miRNAs in the context of CAD.
In addition to dysregulation of expression of miRNAs in endothelial cells and VSMCs, microvesicles originated from these cells have been shown to contain abnormal levels of miRNAs, thus these particles can broaden the extent of miRNAs effects on diverse cells. The presence of miRNAs in the circulation of CAD patients endowed them the ability to predict disease course and distinguish CAD patients from healthy subjects. Both plasma and PBMC levels of miRNAs could be used as diagnostic markers for CAD. Most importantly, miRNAs signature can predict the occurrence of CAD-related complications such as HF. Their ability in distinguishing UA from MI is another promising result of recent investigations, potentiating them as accurate diagnostic marker for stratifying patients who need urgent interventions. However, a major limitation of application of miRNAs as diagnostic or prognostic markers in CAD is the influence of other age-related factors on their expression. Identification of CAD-specific miRNAs whose expressions are not affected by patients' health condition is a major issue in this regard. Longitudinal assessment of miRNA profile in relation with health status of CAD patients and measurement of possible confounding parameters would help in identification of markers for clinical application.
Finally, several SNPs and CNVs within miRNA coding genes have been associated with risk of CAD, providing further evidence for crucial partake of miRNAs in the pathogenesis of CAD. Most notably, some genotypes of these SNPs have been associated with risk of CAD in patients with specific lifestyles or habits (61, 62), demonstrating the possible interaction between these genetic variants and environmental factors. However, the impact of these SNPs on CAD-related biological processes such as cell adhesion, inflammation, proliferation or apoptosis has not been appraised in vitro. Conduction of these types of studies would pave the way for design of targeted therapeutic interventions in CAD. Taken together, miRNAs participate in different aspects of CAD pathogenesis and could be used as specific/sensitive markers for this condition. The therapeutic application of miRNAs in CAD should be judged in upcoming studies.
Author Contributions
MT and SG-F wrote the draft and revised it. MG designed the tables and collected the data. All authors contributed to the article and approved the submitted version.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Abbreviations
ARMS, Amplification-Refractory Mutation System; ANGII, angiotensinogen; TAB2, binding protein 2, CNVs, Copy Number Variations; CCL2, C-C motif chemokine ligand 2; copy number variations, ; CAD, Coronary artery disease; EPCs, endothelial progenitor cells; ET-1, endothelin 1;GEO, Gene Expression Omnibus; HF, heart failure; HRM, High resolution Melting; HUVECs, human umbilical vein endothelial cells; ICAM1, intercellular adhesion molecule 1; KRT1, keratin 1; LDL, low-density lipoprotein; LPS, lipopolysaccharide; miRNAs, MicroRNAs; MI, myocardial infarction; MVs, microvesicles; MAPK8, mitogen-activated protein kinase 8; MEF2C, myocyte enhancer factor 2C; MEF2C, myocyte enhancer factor 2C; IκBα, NF-κB inhibitor alpha; NRIP1, nuclear receptor interacting protein 1; eNOS, nitric oxide synthase 3; PBMCs, peripheral blood mononuclear cells; PIK3C2α, phosphatidylinositol-4-phosphate 3-kinase catalytic subunit type 2 alpha; PDCD4, programmed cell death 4; RFLP, Restriction fragment length polymorphism; STEMI, ST-segment elevation myocardial infarction; SNPs, single nucleotide polymorphisms; SMAD3, SMAD family member 3; SPF, specific-pathogen-free; THBS1, thrombospondin 1; MAP3K7, TGF-beta activated kinase 1; TxA2, thromboxane A2 TxA2; TRF2, telomeric repeat binding factor 2; TRAF6, TNF receptor associated factor 6; UA, unstable angina; VEGF, vascular endothelial growth factor; VSMCs, vascular smooth muscle cells; VCAM1, vascular cell adhesion molecule 1; ZDHHC14, zinc finger DHHC-type palmitoyltransferase 14.
References
1. Roth GA, Johnson C, Abajobir A, Abd-Allah F, Abera SF, Abyu G, et al. Global, regional, and national burden of cardiovascular diseases for 10 causes, 1990 to 2015. J Am Coll Cardiol. (2017) 70:1–25. doi: 10.1016/j.jacc.2017.04.052
2. Mathers CD, Loncar D. Projections of global mortality and burden of disease from 2002 to 2030. PLoS Med. (2006) 3:e442. doi: 10.1371/journal.pmed.0030442
3. Spiekerman RE, Brandenburg JT, Achor RW, Edwards JE. The spectrum of coronary heart disease in a community of 30,000: A clinicopathologic study. Circulation. (1962) 25:57–65. doi: 10.1161/01.CIR.25.1.57
4. Davies MJ, Thomas AC. Plaque fissuring–the cause of acute myocardial infarction, sudden ischaemic death, and crescendo angina. Br Heart J. (1985) 53:363. doi: 10.1136/hrt.53.4.363
5. Ghafouri-Fard S, Gholipour M, Taheri M. The emerging role of long non-coding RNAs and circular RNAs in coronary artery disease. Front Cardiovas Med. (2021) 8:42. doi: 10.3389/fcvm.2021.632393
6. Ambrose JA, Singh M. Pathophysiology of coronary artery disease leading to acute coronary syndromes. F1000Prime Rep. (2015) 7:08. doi: 10.12703/P7-08
7. Sutton MGSJ, Sharpe N. Left ventricular remodeling after myocardial infarction: pathophysiology and therapy. Circulation. (2000) 101:2981–8. doi: 10.1161/01.CIR.101.25.2981
8. Gheorghiade M, Sopko G, De Luca L, Velazquez EJ, Parker JD, Binkley PF, et al. Navigating the crossroads of coronary artery disease and heart failure. Circulation. (2006) 114:1202–13. doi: 10.1161/CIRCULATIONAHA.106.623199
9. Melak T, Baynes HW. Circulating microRNAs as possible biomarkers for coronary artery disease: a narrative review. EJIFCC. (2019) 30:179–94.
10. Pasquinelli AE. MicroRNAs and their targets: recognition, regulation and an emerging reciprocal relationship. Nat Rev Genet. (2012) 13:271–82. doi: 10.1038/nrg3162
11. Bajan S, Hutvagner G. Regulation of miRNA processing and miRNA mediated gene repression in cancer. Microrna. (2014) 3:10–7. doi: 10.2174/2211536602666140110234046
12. Landskroner-Eiger S, Moneke I, Sessa WC. miRNAs as modulators of angiogenesis. Cold Spring Harbor Perspect Med. (2013) 3:a006643. doi: 10.1101/cshperspect.a006643
13. Liu Y, Li Q, Hosen MR, Zietzer A, Flender A, Levermann P, et al. Atherosclerotic conditions promote the packaging of functional microRNA-92a-3p into endothelial microvesicles. Circul Res. (2019) 124:575–87. doi: 10.1161/CIRCRESAHA.118.314010
14. Wang M, Ji Y, Cai S, Ding W. MiR-206 suppresses the progression of coronary artery disease by modulating vascular endothelial growth factor (VEGF) expression. Med Sci Monit. (2016) 22:5011. doi: 10.12659/MSM.898883
15. Tang Y, Zhang Y, Chen Y, Xiang Y, Xie Y. Role of the micro RNA, miR-206, and its target PIK 3C2α in endothelial progenitor cell function–potential link with coronary artery disease. FEBS J. (2015) 282:3758–72. doi: 10.1111/febs.13372
16. Yang S, Mi X, Chen Y, Feng C, Hou Z, Hui R, et al. MicroRNA-216a induces endothelial senescence and inflammation via Smad3/IκBα pathway. J Cell Mol Med. (2018) 22:2739–49. doi: 10.1111/jcmm.13567
17. Gao ZF, Ji XL, Gu J, Wang XY, Ding L, Zhang H. microRNA-107 protects against inflammation and endoplasmic reticulum stress of vascular endothelial cells via KRT1-dependent Notch signaling pathway in a mouse model of coronary atherosclerosis. J Cell Physiol. (2019) 234:12029–41. doi: 10.1002/jcp.27864
18. Ren J, Ma R, Zhang ZB, Li Y, Lei P, Men JL. Effects of microRNA-330 on vulnerable atherosclerotic plaques formation and vascular endothelial cell proliferation through the WNT signaling pathway in acute coronary syndrome. J Cell Biochem. (2018) 119:4514–27. doi: 10.1002/jcb.26584
19. Lian Z, Lv FF, Yu J, Wang JW. The anti-inflammatory effect of microRNA-383-3p interacting with IL1R2 against homocysteine-induced endothelial injury in rat coronary arteries. J Cell Biochem. (2018) 119:6684–94. doi: 10.1002/jcb.26854
20. Hou S, Fang M, Zhu Q, Liu Y, Liu L, Li X. MicroRNA-939 governs vascular integrity and angiogenesis through targeting γ-catenin in endothelial cells. Biochem Biophys Res Commun. (2017) 484:27–33. doi: 10.1016/j.bbrc.2017.01.085
21. Su Y, Yuan J, Zhang F, Lei Q, Zhang T, Li K, et al. MicroRNA-181a-5p and microRNA-181a-3p cooperatively restrict vascular inflammation and atherosclerosis. Cell Death Dis. (2019) 10:1–15. doi: 10.1038/s41419-019-1599-9
22. Du L, Xu Z, Wang X, Liu F. Integrated bioinformatics analysis identifies microRNA-376a-3p as a new microRNA biomarker in patient with coronary artery disease. Am J Transl Res. (2020) 12:633–48.
23. Liang S, Gong X, Zhang G, Huang G, Lu Y, Li Y. The lncRNA XIST interacts with miR-140/miR-124/iASPP axis to promote pancreatic carcinoma growth. Oncotarget. (2017) 8:113701. doi: 10.18632/oncotarget.22555
24. Lin J, Jiang J, Zhou R, Li X, Ye J. MicroRNA-451b participates in coronary heart disease by targeting VEGFA. Open Med. (2019) 15:1–7. doi: 10.1515/med-2020-0001
25. Liu D, Zhang X-l, Yan C-h, Li Y, Tian X-x, Zhu N, et al. MicroRNA-495 regulates the proliferation and apoptosis of human umbilical vein endothelial cells by targeting chemokine CCL2. Thrombosis Res. (2015) 135:146–54. doi: 10.1016/j.thromres.2014.10.027
26. Gao W, Cui H, Li Q, Zhong H, Yu J, Li P, et al. Upregulation of microRNA-218 reduces cardiac microvascular endothelial cells injury induced by coronary artery disease through the inhibition of HMGB1. J Cell Physiol. (2020) 235:3079–95. doi: 10.1002/jcp.29214
27. Lai Z, Lin P, Weng X, Su J, Chen Y, He Y, et al. MicroRNA-574-5p promotes cell growth of vascular smooth muscle cells in the progression of coronary artery disease. Biomed Pharmacotherap. (2018) 97:162–7. doi: 10.1016/j.biopha.2017.10.062
28. Fang Y, Chen S, Liu Z, Ai W, He X, Wang L, et al. Endothelial stem cells attenuate cardiac apoptosis via downregulating cardiac microRNA-146a in a rat model of coronary heart disease. Exp Therap Med. (2018) 16:4246–52. doi: 10.3892/etm.2018.6702
29. Li K, Lin T, Chen L, Wang N. MicroRNA-93 elevation after myocardial infarction is cardiac protective. Med Hypotheses. (2017) 106:23–5. doi: 10.1016/j.mehy.2017.07.003
30. Zhang R, Sui L, Hong X, Yang M, Li W. MiR-448 promotes vascular smooth muscle cell proliferation and migration in through directly targeting MEF2C. Environ Sci Pollut Res Int. (2017) 24:22294–300. doi: 10.1007/s11356-017-9771-1
31. Wang D, Wang Y, Ma J, Wang W, Sun B, Zheng T, et al. MicroRNA-20a participates in the aerobic exercise-based prevention of coronary artery disease by targeting PTEN. Biomed Pharmacotherap. (2017) 95:756–63. doi: 10.1016/j.biopha.2017.08.086
32. Yang Y, Luo H, Liu S, Zhang R, Zhu X, Liu M, et al. Platelet microparticles-containing miR-4306 inhibits human monocyte-derived macrophages migration through VEGFA/ERK1/2/NF-κB signaling pathways. Clin Exp Hypertens. (2019) 41:481–91. doi: 10.1080/10641963.2018.1510941
33. Satoh M, Nasu T, Takahashi Y, Osaki T, Hitomi S, Morino Y, et al. Expression of miR-23a induces telomere shortening and is associated with poor clinical outcomes in patients with coronary artery disease. Clin Sci. (2017) 131:2007–17. doi: 10.1042/CS20170242
34. Guo T, Wang J, Cheng G, Huang H. miR-590-5p may regulate colorectal cancer cell viability and migration by targeting PDCD4. Exp Ther Med. (2020) 20:55. doi: 10.3892/etm.2020.9183
35. Zhou Y, Cheng X, Wan Y, Chen T, Zhou Q, Wang Z, et al. MicroRNA-421 inhibits apoptosis by downregulating Caspase-3 in human colorectal cancer. Cancer Manag Res. (2020) 12:7579–87. doi: 10.2147/CMAR.S255787
36. Liang X, Wang L, Wang M, Liu Z, Liu X, Zhang B, et al. MicroRNA-124 inhibits macrophage cell apoptosis via targeting p38/MAPK signaling pathway in atherosclerosis development. Aging. (2020) 12:13005. doi: 10.18632/aging.103387
37. Bai F, Yu Z, Gao X, Gong J, Fan L, Liu F. Simvastatin induces breast cancer cell death through oxidative stress up-regulating miR-140-5p. Aging. (2019) 11:3198. doi: 10.18632/aging.101974
38. Zhong Z, Hou J, Zhang Q, Zhong W, Li B, Li C, et al. Circulating microRNA expression profiling and bioinformatics analysis of dysregulated microRNAs of patients with coronary artery disease. Medicine. (2018) 97:e11428. doi: 10.1097/MD.0000000000011428
39. Vahed SZ, Aghaee-Bakhtiari SH, Daraei A, Saadatian Z, Kafil HS, Yousefi B, et al. Expression pattern of miR-21, miR-25 and PTEN in peripheral blood mononuclear cells of patients with significant or insignificant coronary stenosis. Gene. (2019) 698:170–8. doi: 10.1016/j.gene.2019.02.074
40. Yao Y, Song T, Xiong G, Wu Z, Li Q, Xia H, et al. Combination of peripheral blood mononuclear cell miR-19b-5p, miR-221, miR-25-5p, and hypertension correlates with an increased heart failure risk in coronary heart disease patients. Anatolian J Cardiol. (2018) 20:100. doi: 10.14744/AnatolJCardiol.2018.43255
41. Singh S, de Ronde MW, Kok MG, Beijk MA, De Winter RJ, van der Wal AC, et al. MiR-223-3p and miR-122-5p as circulating biomarkers for plaque instability. Open Heart. (2020) 7:e001223. doi: 10.1136/openhrt-2019-001223
42. Guo J-F, Zhang Y, Zheng Q-X, Zhang Y, Zhou H-H, Cui L-M. Association between elevated plasma microRNA-223 content and severity of coronary heart disease. Scand J Clin Lab Invest. (2018) 78:373–8. doi: 10.1080/00365513.2018.1480059
43. Gao J, Liu J, Zhang Y, Guan B, Qu H, Chai H, et al. PBMCs-Derived microRNA signature as a prethrombotic status discriminator in stable coronary artery disease. Thromb Haemostasis. (2020) 120:121–31. doi: 10.1055/s-0039-1700518
44. John F, Neylon A, McGorrian C, Blake GJ. miRNA-93-5p and other miRNAs as predictors of coronary artery disease and STEMI. Int J Cardiol. (2016) 224:310–6. doi: 10.1016/j.ijcard.2016.09.016
45. Wang J, Yan Y, Song D, Liu B. Reduced plasma miR-146a is a predictor of poor coronary collateral circulation in patients with coronary artery disease. BioMed Res Int. (2016) 2016:4285942. doi: 10.1155/2016/4285942
46. Zhang Y, Li H-H, Yang R, Yang B-J, Gao Z-Y. Association between circulating microRNA-208a and severity of coronary heart disease. Scand J Clin Lab Invest. (2017) 77:379–84. doi: 10.1080/00365513.2017.1328740
47. Al-Hayali MA, Sozer V, Durmus S, Erdenen F, Altunoglu E, Gelisgen R, et al. Clinical value of circulating microribonucleic acids miR-1 and miR-21 in evaluating the diagnosis of acute heart failure in asymptomatic type 2 diabetic patients. Biomolecules. (2019) 9:193. doi: 10.3390/biom9050193
48. Darabi F, Aghaei M, Movahedian A, Pourmoghadas A, Sarrafzadegan N. The role of serum levels of microRNA-21 and matrix metalloproteinase-9 in patients with acute coronary syndrome. Mol Cell Biochem. (2016) 422:51–60. doi: 10.1007/s11010-016-2805-z
49. Horváth M, Horváthová V, Hájek P, Štěchovský C, Honěk J, Šenolt L, et al. MicroRNA-331 and microRNA-151-3p as biomarkers in patients with ST-segment elevation myocardial infarction. Sci Rep. (2020) 10:5845. doi: 10.1038/s41598-020-62835-w
50. Bai R, Yang Q, Xi R, Li L, Shi D, Chen K. miR-941 as a promising biomarker for acute coronary syndrome. BMC Cardiovasc Disord. (2017) 17:227. doi: 10.1186/s12872-017-0653-8
51. Dai R, Liu Y, Zhou Y, Xiong X, Zhou W, Li W, et al. Potential of circulating pro-angiogenic microRNA expressions as biomarkers for rapid angiographic stenotic progression and restenosis risks in coronary artery disease patients underwent percutaneous coronary intervention. J Clin Lab Anal. (2020) 34:e23013. doi: 10.1002/jcla.23013
52. Amr K, Abdelmawgoud H, Ali Z, Shehata S, Raslan H. Potential value of circulating microRNA-126 and microRNA-210 as biomarkers for type 2 diabetes with coronary artery disease. Br J Biomed Sci. (2018) 75:82–7. doi: 10.1080/09674845.2017.1402404
53. Wang Y, Luo X, Liu Y, Han G, Sun D. Long noncoding RNA RMRP promotes proliferation and invasion via targeting miR-1-3p in non–small-cell lung cancer. J Cell Biochem. (2019) 120:15170–81. doi: 10.1002/jcb.28779
54. Faccini J, Ruidavets J-B, Cordelier P, Martins F, Maoret J-J, Bongard V, et al. Circulating miR-155, miR-145 and let-7c as diagnostic biomarkers of the coronary artery disease. Sci Rep. (2017) 7:42916. doi: 10.1038/srep42916
55. Karakas M, Schulte C, Appelbaum S, Ojeda F, Lackner KJ, Münzel T, et al. Circulating microRNAs strongly predict cardiovascular death in patients with coronary artery disease—results from the large AtheroGene study. Eur Heart J. (2017) 38:516–23. doi: 10.1093/eurheartj/ehw250
56. Saadatian Z, Nariman-Saleh-Fam Z, Bastami M, Mansoori Y, Khaheshi I, Parsa SA, et al. Dysregulated expression of STAT1, miR-150, and miR-223 in peripheral blood mononuclear cells of coronary artery disease patients with significant or insignificant stenosis. J Cell Biochem. (2019) 120:19810–24. doi: 10.1002/jcb.29286
57. Li P, Cai J-X, Han F, Wang J, Zhou J-J, Shen K-W, et al. Expression and significance of miR-654-5p and miR-376b-3p in patients with colon cancer. World J Gastrointestinal Oncol. (2020) 12:492. doi: 10.4251/wjgo.v12.i4.492
58. Al-Kafaji G, Al-Mahroos G, Abdulla Al-Muhtaresh H, Sabry MA, Abdul Razzak R, Salem AH. Circulating endothelium-enriched microRNA-126 as a potential biomarker for coronary artery disease in type 2 diabetes mellitus patients. Biomarkers. (2017) 22:268–78. doi: 10.1080/1354750X.2016.1204004
59. Ying D, Yang SH, Sha L, Cui CJ, Zhang Y, Zhu CG, et al. Circulating microRNAs as novel diagnostic biomarkers for very early-onset ( ≤ 40 years) coronary artery disease. Biomedical and Environmental Sciences. (2016) 29:545–54. doi: 10.3967/bes2016.073
60. Saadatian Z, Nariman-Saleh-Fam Z, Khaheshi I, Mansoori Y, Daraei A, Ghaderian SMH, et al. Peripheral blood mononuclear cells expression levels of miR-196a and miR-100 in coronary artery disease patients. Immunol Invest. (2020) 1–11. doi: 10.1080/08820139.2020.1791177. [Epub ahead of print].
61. Sung JH, Kim SH, Yang WI, Kim WJ, Moon JY, Kim IJ, et al. miRNA polymorphisms (miR-146a, miR-149, miR-196a2 and miR-499) are associated with the risk of coronary artery disease. Mol Med Rep. (2016) 14:2328–42. doi: 10.3892/mmr.2016.5495
62. Sohrabifar N, Ghaderian SMH, Vakili H, Ghaedi H, Rouhani B, Jafari H, et al. MicroRNA-copy number variations in coronary artery disease patients with or without type 2 diabetes mellitus. Arch Physiol Biochem. (2019) 1–7. doi: 10.1080/13813455.2019.1651340. [Epub ahead of print].
63. Ghaffarzadeh M, Ghaedi H, Alipoor B, Omrani MD, Kazerouni F, Shanaki M, et al. Association of miR-149 (RS2292832) variant with the risk of coronary artery disease. J Med Biochem. (2017) 36:251–8. doi: 10.1515/jomb-2017-0005
64. Wei WJ, Lu ZW, Li DS, Wang Y, Zhu YX, Wang ZY, et al. Association of the miR-149 Rs2292832 polymorphism with papillary thyroid cancer risk and clinicopathologic characteristics in a Chinese population. Int J Mol Sci. (2014) 15:20968–81. doi: 10.3390/ijms151120968
65. Wang Y, Wang X, Li Z, Chen L, Zhou L, Li C, et al. Two single nucleotide polymorphisms (rs2431697 and rs2910164) of miR-146a are associated with risk of coronary artery disease. Int J Environ Res Public Health. (2017) 14:514. doi: 10.3390/ijerph14050514
66. Ramkaran P, Khan S, Phulukdaree A, Moodley D, Chuturgoon AA. miR-146a polymorphism influences levels of miR-146a, IRAK-1, and TRAF-6 in young patients with coronary artery disease. Cell Biochem Biophys. (2014) 68:259–66. doi: 10.1007/s12013-013-9704-7
67. Qiu H, Chen Z, Lv L, Tang W, Hu R. Associations between microRNA polymorphisms and development of coronary artery disease: a case–control study. DNA Cell Biol. (2020) 39:25–36. doi: 10.1089/dna.2019.4963
68. Fragoso JM, Ramírez-Bello J, Martínez-Ríos MA, Peña-Duque MA, Posadas-Sánchez R, Delgadillo-Rodríguez H, et al. miR-196a2 (rs11614913) polymorphism is associated with coronary artery disease, but not with in-stent coronary restenosis. Inflamm Res. (2019) 68:215–21. doi: 10.1007/s00011-018-1206-z
69. Agiannitopoulos K, Samara P, Papadopoulou M, Efthymiadou A, Papadopoulou E, Tsaousis GN, et al. miRNA polymorphisms and risk of premature coronary artery disease. Hellenic J Cardiol. (2020) doi: 10.1016/j.hjc.2020.01.005 (in press).
70. Mir R, Jha CK, Elfaki I, Rehman S, Javid J, Khullar N, et al. MicroRNA-224 (rs188519172 A> G) gene variability is associated with a decreased susceptibility to coronary artery disease: A case-control study. MicroRNA. (2019) 8:198–205. doi: 10.2174/2211536608666181211153859
71. Mir R, Elfaki I, Javid J, Rehman S, Khullar N, Banu S, et al. Incidence of MicroR-4513C/T gene variability in coronary artery disease-a case-control study. Endocr Metab Immune Disord Drug Targets. (2019) 19:1216–23. doi: 10.2174/1871530319666190417111940
Keywords: coronary artery disease, miRNA, expression, biomarkers, myocardial infarction
Citation: Ghafouri-Fard S, Gholipour M and Taheri M (2021) Role of MicroRNAs in the Pathogenesis of Coronary Artery Disease. Front. Cardiovasc. Med. 8:632392. doi: 10.3389/fcvm.2021.632392
Received: 23 November 2020; Accepted: 18 March 2021;
Published: 12 April 2021.
Edited by:
Laiyuan Wang, Chinese Academy of Medical Sciences and Peking Union Medical College, ChinaReviewed by:
Zhi Xin Shan, Guangdong Provincial People's Hospital, ChinaChen Gao, UCLA, United States
Copyright © 2021 Ghafouri-Fard, Gholipour and Taheri. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Mohammad Taheri, bW9oYW1tYWRfODIzQHlhaG9vLmNvbQ==
 Soudeh Ghafouri-Fard
Soudeh Ghafouri-Fard Mahdi Gholipour
Mahdi Gholipour Mohammad Taheri
Mohammad Taheri