
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Bioeng. Biotechnol. , 02 April 2025
Sec. Synthetic Biology
Volume 13 - 2025 | https://doi.org/10.3389/fbioe.2025.1567596
As interest in space exploration and in situ resource utilization grows, the potential to leverage synthetic biology and engineered microorganisms has garnered significant attention. Microorganisms provide a robust and efficient biological chassis to demonstrate the human blueprint for advancing space biology. However, progress toward these applications is hindered by the limited access to space-like environments and a lack of knowledge about how unique environmental factors affect relevant microbial systems. To address these issues, we evaluated the Salmonella Pathogenicity Island 1 (SPI-1) type Ⅲ secretion system (T3SS) as a protein production platform for space applications. Using a NASA-designed microgravity-simulating bioreactor system, we investigated the effects of simulated microgravity on cell growth, stress response, and protein secretion via SPI-1 T3SS. Our results demonstrated increased stress responses in cells grown under simulated microgravity. However, the SPI-1 T3SS maintained its ability to secrete proteins directly into the extracellular space in a single step under simulated microgravity, simplifying downstream purification processes. These findings suggest that the SPI-1 T3SS is a viable candidate for future space biology applications.
Humans have an instinctive curiosity to explore unknown territory, including space, which in turn prompts the development of innovations that improve daily life (Comstock and Lockney, 2007). In addition, advances from space exploration technologies are often uniquely suited for in situ resource utilization (ISRU) in terrestrial contexts where resources and/or infrastructure are limited. Synthetic biology, which is a multidisciplinary field combining biology and engineering, is considered one of the key technologies for developing biomanufacturing with limited resources and for space applications (Menezes et al., 2015; Brooks and Alper, 2021; Tang et al., 2023). In particular, synthetic biology can be used to systematically, and modularly, engineer microbial hosts, which is seen as a powerful solution to address the challenges of space exploration and ISRU (Menezes et al., 2015; Santomartino et al., 2023).
The application and transfer of current synthetic biology technologies to space are hindered by the unique environmental conditions encountered beyond Earth, for which there are currently limited data sets. Understanding the effects of space environments on biological systems is imperative for future applications. The International Space Station (ISS) has served as the primary location for conducting space biology research, but its limited accessibility and high operational costs restrict more extensive research. Ground-based microgravity simulators, such as rotary cell culture systems (RCCS) with High Aspect Ratio Vessels (HARVs), were designed by researchers at NASA Johnson Space Center to build low-shear environments to simulate this aspect of the space environment (Schwarz and Wolf, 1991; Schwarz et al., 1992). This ground-based microgravity simulator system has been widely used to study the effects of microgravity on microbial hosts (Nickerson et al., 2003; Anken, 2013; Acres et al., 2021). Multiple studies have reported that microorganisms, including Salmonella, respond to microgravity by altering physiological processes, gene expression, and stress responses (Nickerson et al., 2000; Tucker et al., 2007; Wilson et al., 2007; Kim and Rhee, 2016; Yang et al., 2016; Senatore et al., 2020; Su et al., 2021).
Proteins are essential macromolecules which serve diverse biological functions across all organisms and have become indispensable components in a variety of industrial applications, ranging from degradative enzymes to advanced materials and therapeutics (Whitford, 2005; Ferrer-Miralles et al., 2009; Overton, 2014). To meet the increasing industrial demand for these multifunctional biomolecules, microbial expression systems have emerged as the dominant platforms for recombinant protein production. These systems have been continually optimized to enable efficient protein synthesis on an industrial scale (Ferrer-Miralles et al., 2009; Overton, 2014; Chen et al., 2021; Tepap et al., 2023). Potential applications of proteins extend beyond terrestrial environments to space, especially supporting ISRU and long-duration missions. The versatility of proteins could address a diverse range of challenges in extraterrestrial environments, from the biomanufacturing of replacement or novel materials, such as biopolymers and biomaterials for radioprotection and habitation (Ouchen et al., 2014; Jemison and Olabisi, 2021) to the development of targeted medical treatments including personalized medicine and regenerative medicine for damages caused by radiation (Pavez Loriè et al., 2021). Furthermore, establishing microbial protein production platforms in space could provide adaptable and resource-efficient biomanufacturing facilities that are important for future space missions (Averesch et al., 2023; Santomartino et al., 2023).
Bacterial protein secretion systems are promising technology to enable the modular, on-demand production of such useful recombinant proteins. Harnessing native bacterial secretion mechanisms could simplify protein production in space by eliminating the need for complicated cell lysis and downstream purification steps that otherwise hinder protein production from adoption for resource-limited situations (Reed and Chen, 2013; Burdette et al., 2018). Among the microbial strains extensively studied in synthetic biology, Salmonella species have gained attention for its applications in medicine and pharmaceuticals (Yoon, 2018; Yan et al., 2023) due to their tractability and their high genetic similarity to Escherichia coli, which enables many tools developed in the canonical Gram-negative model system to be successfully transferred with relative ease. The Salmonella Pathogenicity Island 1 (SPI-1) type Ⅲ secretion system (T3SS) is a highly efficient tool for recombinant protein secretion (Metcalf et al., 2014; Azam et al., 2016; Metcalf et al., 2016; Song et al., 2017; Burdette et al., 2021; Liang et al., 2023; Burdette et al., 2024). The T3SS, utilized by many Gram-negative bacteria, comprises a needle-like structure known as the injectisome which is natively used to translocate effector proteins from the cytosol directly into eukaryotic hosts, but also functions in the absence of such hosts to secrete heterologous proteins appended to appropriate N-terminal signal sequences in one step to the extracellular space (Green and Mecsas, 2016; Burdette et al., 2018; Lou et al., 2019; Burdette et al., 2024). The system is able to secrete a wide variety of protein cargo, including antibody fragments, enzymes (Metcalf et al., 2016), material-forming proteins (Widmaier et al., 2009; Azam et al., 2016), and growth factors (Burdette et al., 2024), across a range of sizes (up to ∼95 kDa) (Widmaier et al., 2009), many of which are able to fold and function following secretion to the media in bacterial culture (Metcalf et al., 2016), at reported titers of up to 140 mg/L (Glasgow et al., 2017). Thus, the SPI-1 T3SS shows promise as a key component of a bacterial platform for recombinant protein production in resource-limited environments like space. While traditional intracellular protein expression systems require extensive and specialized downstream processing steps, the SPI-1 T3SS confers up to 80% purity without any additional purification steps, with further purity to >90% following as few as one additional unit operation (Azam et al., 2016). Despite these advantages, recombinant protein production through the SPI-1 T3SS remains subject to environmental sensitivity, which is attributed to its intricate regulatory network (Tartera and Metcalf, 1993; Ellermeier et al., 2005; Golubeva et al., 2012; Song et al., 2017; Lou et al., 2019). Thus, it is unknown how its function may be affected by conditions of relevance to space environmental conditions.
In this study, we sought to investigate the effects of simulated microgravity on the SPI-1 T3SS as a potential protein production platform for space applications. We employed Salmonella enterica serovar Typhimurium ASTE13, a high-efficiency protein secretion strain (Song et al., 2017; Burdette et al., 2021; Liang et al., 2023). By comparing growth and stress responses under different gravitational conditions, we evaluated SPI-1 T3SS functionality through transcriptional activity and protein secretion experiments. Consistent with previous reports, we observed altered stress responses to simulated microgravity in this strain (Nickerson et al., 2003; Yang et al., 2016). Interestingly, the SPI-1 T3SS showed comparable secretion activity under both gravitational conditions, indicating its potential as a robust bio-device for future space missions.
Salmonella enterica serovar Typhimurium ASTE13 (ASTE13), a derivative of S. enterica serovar Typhimurium DW01 (Song et al., 2017; Burdette et al., 2021; Liang et al., 2023), or its derivatives were used in this study unless otherwise noted. All strains and plasmids used in this study are listed in Table 1.
ASTE13 was used to compare the growth and stress resistance at both gravitational conditions. A single colony of ASTE13 from a lysogeny broth (LB) agar plate was inoculated into LB-Lennox (L) media (10 g/L tryptone, 5 g/L yeast extract, and 5 g/L NaCl) and statically cultured overnight at 37°C. The overnight culture samples were diluted to an OD600 of 0.01 in LB-L and then transferred to the HARVs for the culture on the RCCS (10 mL autoclavable HARVs, RCCS-4D, Synthecon Inc., United States) at 37°C and 25 rpm in a humidified incubator.
After the 10-h cultivation in the HARVs, bacterial cultures were either kept untreated or exposed to one of the following three stress conditions. For the pH stress experiment, 1M of concentrated citrate buffer (pH 3.0) (Yang et al., 2016) was added to the culture, which was then held for 30 min at room temperature. For the heat stress experiment, cultures were placed in a water bath at 50°C for 30 min. For the bile salt stress experiment, 10% (w/v) of bile salt was added to the cell cultures for 30 min. Following the stress exposure, cell viabilities were analyzed using LIVE/DEAD staining followed by flow cytometry to compare the survival rate under stress conditions. The viability assay was carried out by following the manufacturer’s protocol for the LIVE/DEAD Bacterial Viability Kit (L7012, Thermo Fisher Scientific). Briefly, the cell cultures were adjusted by OD670of 0.03 and then diluted 100 times in PBS buffer. Then, 3 µL of the reagent mixture from the kit was added to each sample and incubated at room temperature for 15 min in the dark. After the incubation, the samples were transferred to a 96-well plate and analyzed by flow cytometry using an Attune NxT flow cytometer (Life Technologies) for fluorescence measurements. Data analysis was performed by FlowJo.
Autoclavable HARVs were selected due to their ability to be re-used, enabling experimental consistency and replications. Prior to all experiments using the RCCS with HARVs, the vessel membranes were pre-wet and the HARVs were checked for contamination issues by filling 70% of the vessel volume with culture media followed by rotating at 25 rpm and 37°C overnight. For bacterial culture experiments using the RCCS with HARVs, the media pre-occupied vessels were emptied and then the diluted bacterial culture was transferred to, and completely filled, the HARVs to avoid bubble formation. This required maintaining a larger volume of diluted culture than was needed for the HARVs. Before attaching the HARVs to the RCCS, the HARVs were checked for the presence of any bubbles. If present, the remaining bubbles were removed using the syringe attached to the HARVs. The RCCS was placed at 37°C in a humidified incubator and the rotation speed of HARVs was set at 25 rpm unless otherwise noted. End-point sampling was used for all experiments to prevent any issues related to microgravity disruption.
To assess promoter activity under different gravitational states, ASTE13 sipC:GFPmut2 was used. GFPmut2 acts as a reporter of PsicA promoter activity (Burdette et al., 2021). This strain was streaked from a frozen stock, and a single colony was inoculated into 5 mL of LB media with 0.4% (w/v) glucose and cultured overnight at 225 rpm and 37°C. The overnight culture was diluted 100-fold in LB-L and then transferred to the HARVs and loaded onto the RCCS. The cultures were grown at 37°C and 25 rpm. As noted above, there was a single HARV culture used for each time point. Thus for each time point, a vessel was removed from the RCCS and samples were withdrawn from the HARV. These were diluted to an OD600 of 0.01 in PBS with 1 mg/mL kanamycin and transferred to a 96-well plate, which was stored at 4°C overnight. The next day, the cells were analyzed by flow cytometry using the Attune NxT flow cytometer. Data was analyzed using FlowJo 10.5.3.
ASTE13 and ASTE13 ΔprgI strains were transformed with the plasmid harboring PsicA-DH (Widmaier et al., 2009), which encodes for the catalytic domain of the human intersectin-1 protein, to create the strains referred to herein as WT-DH and ΔprgI-DH, respectively (Table 1). A single colony was inoculated into 5 mL of LB-L media with chloramphenicol (34 μg/mL) and cultured overnight at 225 rpm and 37°C. The overnight culture was diluted 100-fold into LB-L with chloramphenicol (34 μg/mL) and transferred to HARVs. The strains were grown for 7 h in the HARVs at 37°C and 25 rpm unless noted otherwise. Cultivation for over 7 h in the HARVs leads to substantial cell lysis (data not shown). Therefore, all secretion-related experiments were completed within 7 h to minimize experimental complications from cellular lysis. The cultures were centrifuged at 4,000 x g for 10 min to separate the cells from the media. The supernatant is considered the secretion fraction. Sodium dodecyl sulfate-polyacrylamide electrophoresis (SDS-PAGE) samples for the secretion fractions were prepared by adding Laemmli buffer to the supernatant. These were compared to whole culture lysate samples, which were not subjected to centrifugation. Instead, for whole culture samples, the cell cultures were mixed directly with Laemmli buffer. All SDS-PAGE samples (secreted fraction and whole culture) were prepared by boiling at 95°C for 5 min immediately after sample preparation, which lyses the whole culture samples. The samples were loaded onto 12.5% SDS-PAGE gels and electrophoresed at 130 V for 70 min. After the separation of each sample by SDS-PAGE, proteins were transferred to a polyvinylidene difluoride (PVDF) membrane (Millipore) for Western blotting. Membranes were probed overnight at 4°C with mouse anti-FLAG (Sigma-Aldrich F3165) or anti-GroEL (Sigma-Aldrich G6532) antibodies according to the manufacturer’s protocol. Briefly, the antibodies were diluted in TBST buffer (2.42 g/L Tris base, 8.76 g/L of NaCl, 0.05% v/v Tween-20, pH 7.5) at ratios of 1:6666 and 1:10000, respectively. Chemiluminescent detection was carried out with secondary antibodies for 1 h at room temperature using goat anti-mouse IgG (Invitrogen 32430) or anti-rabbit IgG (Invitrogen 32460), each diluted 1:1000 in TBST buffer, following the manufacturer’s protocol. Western blot images were visualized using the West Pico PLUS chemiluminescent substrate (Thermo Scientific) followed by detection using a ChemiDoc XRS+ (Bio-Rad). The Western blot images, Figure 4 and Supplementary Figure 2, were cropped to exclude irrelevant regions while maintaining the integrity of this experimental data. Protein size markers were labeled in the same positions as in the original blot images. The original images were reviewed and verified during the revision process to ensure transparency. All relative protein quantities were calculated from OD600-normalized samples using ImageJ software and further adjusted to the average of the replicates of the specified normalization condition for relative comparison.
All quantitative data are presented as the mean ± standard deviation (SD) of three or four biological replicates, as noted. Statistical significance was determined using the Student’s t-test via Excel’s data analysis tool, with significance levels indicated as * (p < 0.05), ** (p < 0.01), and *** (p < 0.001).
The RCCS is a bioreactor system that provides simulated microgravity and low-shear environments (<1 dyn/cm2) for microbial culture by rotating the culture vessels at speeds to avoid cell sedimentation (Nickerson et al., 2003; Anken, 2013; Acres et al., 2021). In order to investigate the impact of simulated microgravity on our S. enterica ASTE13 protein production strain, we needed to establish a protocol for bacterial culture and sampling in the system under simulated microgravity that was amenable to adaptation for Earth gravity operation as a control. Briefly, the RCCS is designed to be operated in the simulated microgravity (vertical) orientation, such that rotation of the HARVs is in the plane vertical to the ground and leads to perpetually falling cells within the culture (Figure 1). As a control orientation, we chose to run the system in Earth gravity (horizontal) orientation. The RCCS is not designed to operate in this condition, so we used a custom-fabricated support in order to hold the RCCS in this Earth gravity orientation, which allowed rotation in the plane horizontal with the ground. The RCCS with HARVs was placed in the humidified incubator to avoid media evaporation. We selected a 25 rpm rotation speed for the HARVs regardless of orientation, because this speed emulates microgravity environments in this system when operated with rotation in the vertical orientation (Nickerson et al., 2000; Wilson et al., 2002; Crabbé et al., 2010; Yang et al., 2016; Kim and Rhee, 2016; Kim and Rhee, 2021). Importantly, the RCCS with HARVs at this speed avoids fluid mixing as assessed with a dye experiment (Crabbé et al., 2010).
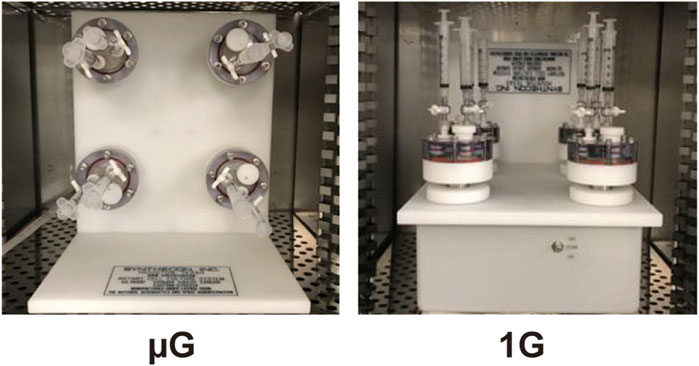

Figure 1. Setup of Rotary Cell Culture System (RCCS) with High Aspect Ratio Vessels (HARVs) to generate simulated microgravity (µG, left) and Earth gravity (1G, right) conditions.
Sampling from the HARVs during cell culture requires using a syringe and the sample port (Supplementary Figure 1), which must be done while rotating the vessels. The procedure is challenging in the microgravity orientation, and would alter vessel volume, potentially causing unknown effects on the simulated microgravity environment. Importantly, this process also does not yield homogeneous samples at Earth gravity due to cell sedimentation. To avoid these two critical issues, we opted to use only end-point sampling. The resulting procedure requires more time and samples (one HARV culture is used per time point), but we reasoned it should lead to more reproducible datasets.
Using the RCCS bioreactor culture system established above, we next set out to determine whether gravity alters cell growth and stress responses for ASTE13. We cultured the bacteria for 10 h in LB media in both microgravity and Earth gravity environments, taking time points throughout the culture period. Cell growth appears indistinguishable between the two conditions through the 4-h time points, but then begins to deviate. At 10 hours, a 36% growth enhancement (p < 0.001) was observed in the strain grown in the Earth gravity orientation as compared to the microgravity orientation (Figure 2A).
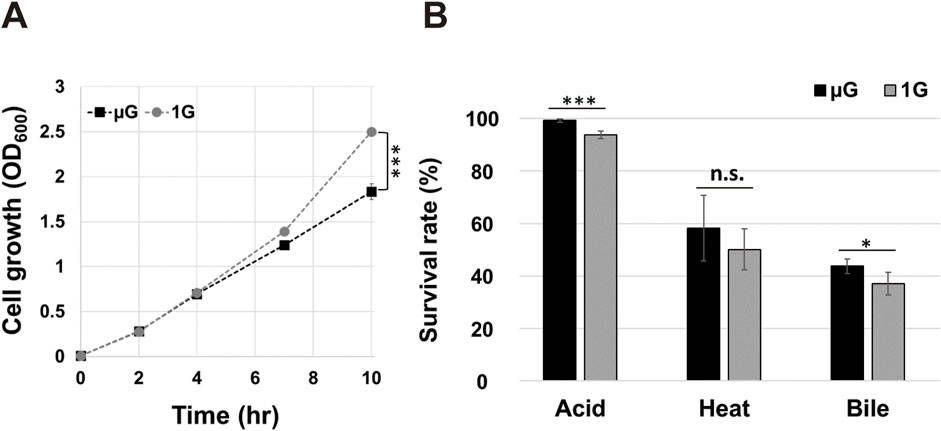
Figure 2. Comparison of Salmonella ASTE13 growth and stress resistance under simulated microgravity (µG) and Earth gravity (1G) conditions. (A) Growth of the Salmonella ASTE13 strain cultured under µG and 1G was measured by optical density at 600 nm (OD600) after end-point sampling. Data represent the mean of three biological replicates, with error bars displaying standard deviation. (B) Stress resistance of Salmonella ASTE13 strain cultured under µG and 1G. After 10 h of growth in HARVs with RCCS, the strain was exposed to acid stress (pH < 4), heat stress (55 °C), and bile stress (10% bile salts) for 30 min. Survival rates were determined using the LIVE/DEAD Cell Viability Assays. Data represent the mean of three biological replicates with error bars indicating standard deviation. A two-tailed Student’s t-test was performed to determine a statistically significant difference between two gravitational conditions. The significances were marked as *** (p < 0.001), * (p < 0.05), and n. s. (p > 0.05).
Salmonella stress tolerance is impacted by growth under microgravity as opposed to Earth gravity conditions, but the effects can vary depending on strain, nutrition, and other culture conditions. For example, S. enterica x3339 and x4973 exhibited enhanced stress tolerance to low pH, heat, and osmotic shock when grown under simulated microgravity compared to Earth gravity (Nickerson et al., 2000; Wilson et al., 2002), while S. enterica D23580 decreases in stress tolerance to either low pH or bile stress following a simulated microgravity exposure (Yang et al., 2016). Likewise, studies on other microbial strains have reported varying stress responses to environmental factors such as low pH, low and high temperature under different gravitational conditions (Crabbé et al., 2010; Kim and Rhee, 2016; Sheet et al., 2020; Kim and Rhee, 2021). Therefore, we sought to examine the stress responses of our S. enterica serovar Typhimurium ASTE13 strain, specifically targeting low pH, heat, and bile salt as stress stimulators─conditions known to elicit marked stress responses in Salmonella (Humphrey, 2004; Prieto et al., 2004). We grew ASTE13 under both simulated microgravity and Earth gravity conditions, and then exposed each sample to environmental stressors. Stress resistance was assessed by determining survival rates using a viability assay after 30 min of exposure to each stressor. The cells grown at simulated microgravity exhibited higher survival rates for both acid stress (p < 0.001) and bile salts (p < 0.05) compared to those grown at Earth gravity (Figure 2B) while survival for heat stress was not significantly different across gravity environments.
Slight environmental changes can alter the SPI-1 T3SS transcriptional activity, in turn affecting protein secretion titer and efficiency (Tartera and Metcalf, 1993; Burdette et al., 2021; Liang et al., 2023). Given the distinct impact of gravity on stress responses, we wished to investigate whether gravity influences the transcriptional activity of SPI-1 T3SS. For this purpose, we used a variation of the ASTE13 strain harboring a GFP reporter encoded in place of the downstream SPI-1 effector sipC (sipC:GFPmut2), which reports on the activity of the sic promoter (Burdette et al., 2021). The sic promoter is activated when the T3SS needle is constructed and operational (Darwin and Miller, 2000). SipC, known as a needle tip complex protein, is located within the sic operon, and its expression is controlled by the sic promoter (Lou et al., 2019). The sic promoter plays a critical role in the secretion-activated network of SPI-1 T3SS operon, driving the expression of genes involved in protein secretion (Widmaier et al., 2009). The promoter is the final promoter to be activated in the secretion activation cascade. Given that our goal is to establish active secretion through the SPI-1 T3SS under space conditions, we reasoned that this promoter is an excellent proxy for the full activation of the T3SS. Using the strain harboring the GFP reporter, we examined sic promoter activity by monitoring GFP fluorescence on a per-cell basis using flow cytometry over time from 0 to 6 h (Figure 3). An active SPI-1 T3SS population was detected in 3-h samples for both gravity environments (Figures 3A, B). This population increases in both size of the “sic active” population (Figure 3B) and the fluorescence intensity of this fraction of the population (Figure 3C). Importantly, T3SS activation under simulated microgravity was not significantly different (p > 0.05) to that observed at Earth gravity at all time points. These results are encouraging for the potential use of the SPI-1 T3SS as a protein production platform in space.
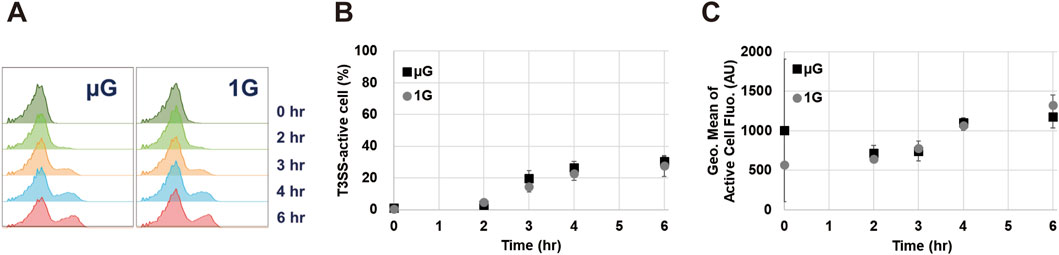
Figure 3. Comparison of SPI-1 T3SS transcriptional activity under simulated microgravity (µG) and Earth gravity (1G) conditions. (A) Representative histograms showing the GFP signal detected via flow cytometry in sipC:GFPmut2 indicating transcriptional activity from sicA promoter. Histograms are representative of three to four biological replicates. (B) percentage of cells expressing GFP signal, and (C) geometric mean fluorescence intensity of the GFP signal representing T3SS activity. Data in B and C shows the mean value of three to four biological replicates with error bars indicating standard deviation. Measured values for simulated microgravity were not significantly different (p > 0.05) to those observed at Earth gravity at any time point.
We next directly evaluated the protein secretion capabilities of the SPI-1 T3SS in our lab-developed ASTE13 strain using the ground-based bioreactor system. To assess protein secretion into the extracellular environment, we transformed ASTE13 with an export plasmid, which harbors a gene encoding the DH protein fused to an N-terminal SptP secretion sequence and a C-terminal FLAG tag, all under the control of the sic promoter (Widmaier et al., 2009). The DH protein is the 24 kDa catalytic domain of the human intersectin-1 protein that has long served as a model secretion substrate for the SPI-1 system, with SptP as well as other N-terminal secretion tags (Widmaier et al., 2009; Metcalf et al., 2014; Burdette et al., 2021; Liang et al., 2023). Using the DH protein, therefore, enables the assessment of secretion efficiency in the context of the established SPI-1 secretion model and in comparison to prior studies. Concurrently, to help assess whether DH protein detected in the supernatant is the result of SPI-1 T3SS-mediated protein secretion, we used another strain, ASTE13 ΔprgI, to serve as a negative control (Metcalf et al., 2014; Burdette et al., 2024). This strain contains a deletion of the gene encoding the needle-forming protein PrgI, a key SPI-1 T3SS component. In the absence of PrgI, the needle structure is not formed, and the strain cannot secrete proteins via SPI-1 T3SS (Metcalf et al., 2014; Metcalf et al., 2016; Burdette et al., 2024). We used Western blotting against the FLAG tag to assess protein expression and secretion for both ASTE13 and our negative control ASTE13 ΔprgI strain. To assess protein expression, we examined protein content of the entire culture (lysed bacterial cells plus the media), while to assess protein secretion, we examined the media of the culture alone by pelleting the bacterial cells. Under the simulated microgravity condition, DH protein expression levels were similar in both strains. However, the media of the ASTE13 strain exhibited intense protein bands while the non-secreting negative control strain had no detectable protein bands at the size expected for the DH fusion under the simulated microgravity condition (Figure 4; Supplementary Figure 2). From the data, we concluded that the DH protein is secreted in a T3SS-dependent manner under simulated microgravity.
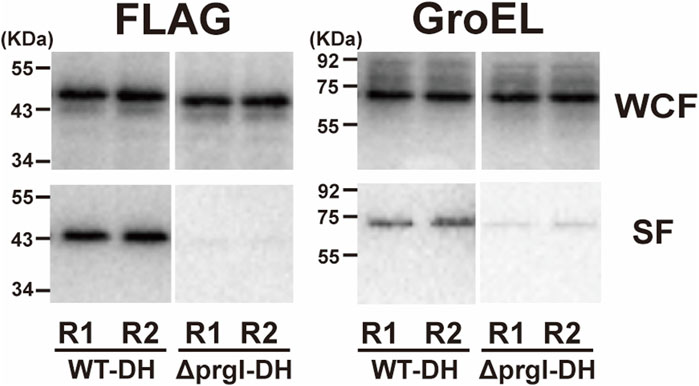
Figure 4. Evaluation of DH protein secretion through SPI-1 T3SS in simulated microgravity (µG) conditions. The WT-DH and ΔprgI-DH strains were cultured for 7 h at 25 rpm in HARVs with RCCS. Protein secretion was analyzed by Western blotting, and two biological replicates (marked as R1 and R2) from WT-DH and ΔprgI-DH strain were loaded on SDS-PAGE gel. Loading volumes were determined by OD600. SF and WCF represent the secreted fraction and whole culture lysate fraction, respectively.
We next set out to explore the secretion efficiency of SPI-1 T3SS under simulated microgravity and Earth gravity conditions. In both gravity conditions, similar levels of DH protein expression were observed (Supplementary Figure 3). The strains grown at Earth gravity exhibited 2.3-fold (p < 0.001) more intense DH protein bands in the secreted fraction than those grown under simulated microgravity (Supplementary Figure 3). However, intracellular GroEL protein bands were also approximately 1.5-fold (p < 0.05) higher in the secreted fraction under Earth gravity conditions. In contrast, the GroEL protein bands under simulated microgravity were detectable but displayed relatively low intensity. Taken together, our results show there is measurable cell lysis in the HARV format, precluding comparison of secretion efficiency across gravity conditions (Supplementary Figure 3).
With the expansion of the space economy, interest in space exploration and opportunities continue to grow (Corrado et al., 2023). However, astronauts face health and safety challenges during missions, particularly due to the resource-limited environments (Braddock, 2017; Douglas et al., 2020; Douglas et al., 2021). Therefore, there is an increasing need for robust systems that provide essential goods such as medicine, food, and energy through ISRU (Jemison and Olabisi, 2021; Berliner et al., 2024). The advancements in biotechnology have led to the development of microorganisms as efficient cell factories for producing useful chemicals and proteins owing to their rapid growth and ease of genetic manipulation (Menezes et al., 2015; Nielsen and Keasling, 2016; Sanchez-Garcia et al., 2016; Santomartino et al., 2023). In principle, by maintaining bacterial stocks and a DNA synthesizer, a variety of bioproducts could be produced on demand from the same reagents. Despite this potential, space environments impose unique constraints on microbial systems, impacting their physiological, metabolic, and genetic behavior compared to what may be observed on Earth. To fully leverage microorganisms for space applications, we must understand how space conditions affect biological systems. Several prior studies suggested that simulated microgravity can enhance protein production (Xiang et al., 2010; Huangfu et al., 2020; Chen et al., 2021). Here, we examined a potential method for facile protein purification to utilize this increased protein production in a space or resource-limited environment.
For this purpose, we assessed S. enterica serovar Typhimurium for its potential as a protein production bio-factory. Since S. enterica includes pathogenic strains, its beneficial applications in biotechnology are often overlooked. However, some S. enterica strains have been successfully utilized as vaccines (e.g., Ty21a and Vi CPS) (MacLennan et al., 2014) and are also considered a promising vehicle for the delivery of therapeutic proteins for anti-cancer drugs and immunotherapies, with approval from the U.S. Food and Drug Administration (Badie et al., 2021; Aganja et al., 2022). In particular, we explored the robustness of the SPI-1 T3SS, an efficient protein secretion system found in S. enterica, using ground-based, microgravity-simulating bioreactors. The SPI-1 T3SS offers a simplified and efficient method for extracellular recombinant protein production bypassing the need for complex downstream processing. However, its performance under microgravity has not been thoroughly evaluated. Nickerson and coworkers reported increased virulence of S. enterica grown in microgravity and simulated microgravity environments (Nickerson et al., 2000). Because the T3SS encoded by SPI-1 is natively an important factor for S. enterica virulence in pathogenic strains (Green and Mecsas, 2016; Burdette et al., 2018; Lou et al., 2019), we reasoned that the SPI-1 T3SS may maintain or even increase in activity under microgravity conditions, and therefore be useful as a platform for producing recombinant proteins such as therapeutic agents and enzymes in space via ISRU.
Our laboratory strain S. enterica ASTE13 provides high growth and protein secretion and was engineered for industrial protein production applications. Our results indicate that ASTE13 responds to microgravity by exhibiting altered growth and stress responses (Figure 1). The reduced growth observed in this study may be attributed to a stress response triggered by simulated microgravity. Previous research has demonstrated that microgravity can act as a stress stimulus in S. enterica (Nickerson et al., 2000; Wilson et al., 2002), resulting in altered cellular responses. Similarly, a recent study in E. coli reported not only growth reduction under simulated microgravity condition but also the upregulation of stress-related genes compared to the Earth gravity condition (Yim et al., 2020). When we compared the stress tolerance under both gravitational conditions, the heat stress resistance of ASTE13 under simulated microgravity was not as significant as the acid and bile-induced stresses. Salmonella employ a range of distinct mechanisms to counteract diverse stressors (Shen and Fang, 2012). Acid and bile stress resistance frequently involves the activity of efflux pumps or modifications to the outer membrane (Gunn, 2000; Schumacher et al., 2023), while heat shock resistance primarily relies on the synthesis of heat shock or chaperone proteins (Roncarati and Scarlato, 2017). The activation of stress responses under simulated microgravity may have enhanced resistance to acid and bile stress but did not sufficiently prepare the cells for heat shock resistance. This indicates that ASTE13 grown under simulated microgravity might not have been physiologically adapted to tolerate heat stress as readily as acid and bile-induced stresses. The increased stress tolerance observed in ASTE13 under simulated microgravity can be considered a desirable feature for its use in space environments.
Importantly, the microgravity condition does not appear to negatively impact the functionality of SPI-1 T3SS. Both protein expression levels and transcriptional activity remained comparable to those observed at Earth gravity (Figure 3; Supplementary Figure 3). Furthermore, our experiment with the ASTE13 ΔprgI confirmed that S. enterica could efficiently secrete proteins through the SPI-1 T3SS even under microgravity conditions (Figure 4; Supplementary Figure 2). While this is useful in our ASTE13 strain for protein production, it raises questions with respect to how virulent strains may behave. However, we note that evolution optimized secretion levels for infection, so if the observed increases in ASTE13 extend to more pathogenic strains, this still may not translate to increased virulence. Additional studies are warranted to explore the impact of microgravity on pathogenicity.
The RCCS with HARVs as a microgravity simulator may not be ideal for cultures requiring extended incubation times due to the potential of biofilm formation (Diaz et al., 2022) and the cell lysis we observed in this study. Since protein secretion via the T3SS of ASTE13 has not given rise to extensive cell lysis when grown with shaking in the standard laboratory environment, and the observed effects were less under the microgravity condition than the Earth gravity condition, we conclude that something related to the HARV format (e.g., changes in oxygen delivery) is responsible for inducing lysis. For all bacterial culture studies using the HARV system, it will be imperative to understand and address cell lysis in advance. However, since the S. enterica T3SS protein production platform requires only short culture durations, this microgravity-simulating bioreactor system provides an efficient test platform for studying microgravity impacts on the T3SS. To further reduce lysis, future studies on this system could further optimize the culture conditions in microgravity-simulating bioreactor systems or engineer ASTE13 to enhance its robustness, e.g., via adaptive laboratory evolution using growth in the HARV.
Overall, our findings suggest that the SPI-1 T3SS is still active in microgravity, making it a promising platform for protein production in space. To further refine this system for space applications, we propose integrating systems biology approaches to identify the engineering targets that could optimize the SPI-1 T3SS for efficient protein secretion machinery in space. Additionally, it would be useful to evaluate the versatility of Salmonella T3SS under the two distinct gravitational conditions by introducing a range of distinct protein classes as secretion substrates, such as enzymes, potential receptor-binding proteins, and material-forming proteins. Such experiments will serve to further optimize the capacity of the system, particularly with regard to the folding, functionality, and activity of secreted proteins. Finally, a complete biosafety evaluation of S. enterica ASTE13 is warranted given its potential as a protein producer and its close evolutionary relationship to more pathogenic strains. Should such studies reveal potential for latent pathogenicity, strain engineering could be used to address such issues. Engineering of ASTE13 to limit pathogenicity and enhance protein production in a standard laboratory environment as well as under simulated microgravity will inform engineering strategies for future space missions.
The original contributions presented in the study are included in the article/Supplementary Material, further inquiries can be directed to the corresponding author.
M-KK: Conceptualization, Funding acquisition, Investigation, Formal Analysis, Methodology, Validation, Visualization, Writing–original draft, Writing–review and editing. JB: Conceptualization, Methodology, Writing–review and editing. DT-E: Conceptualization, Funding acquisition, Methodology, Validation, Supervision, Writing–original draft, Writing–review and editing.
The author(s) declare that financial support was received for the research and/or publication of this article. This study was funded by the National Research Foundation of Korea (Grant NRF 2021R1A5A8029490) awarded to M-KK, and National Science Foundation (Grant BBE 1706125) awarded to DT-E.
The authors wish to thank members of the Tullman-Ercek lab, including Dr. Nolan Kennedy, Dr. Lisa Burdette, Dr. Julie Liang, and Dr. Han Teng Wong, for their helpful comments on the work described in this manuscript. We would also like to thank Dr. Jiseon Yang from Arizona State University for her valuable comments on handling the HARV system.
Author JB was employed by Interplanetary Exploration Institute Ltd. DTE has a financial interest in Opera Bioscience, which is commercializing bacterial protein production and secretion. DTE is an inventor on a US patent application related to this work. DTE's conflict of interest is reviewed and managed by Northwestern University in accordance with its established conflict-of-interest policies.
The remaining author declares that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fbioe.2025.1567596/full#supplementary-material
Acres, J. M., Youngapelian, M. J., and Nadeau, J. (2021). The influence of spaceflight and simulated microgravity on bacterial motility and chemotaxis. npj Microgravity 7 (1), 7. doi:10.1038/s41526-021-00135-x
Aganja, R. P., Sivasankar, C., Senevirathne, A., and Lee, J. H. (2022). Salmonella as a promising curative tool against cancer. Pharmaceutics 14 (10), 2100. doi:10.3390/pharmaceutics14102100
Anken, R. (2013). Simulation of microgravity for studies in gravitational biology: principles, devices and applications. Curr. Biotechnol. 2 (3), 192–200. doi:10.2174/22115501113029990012
Averesch, N. J. H., Berliner, A. J., Nangle, S. N., Zezulka, S., Vengerova, G. L., Ho, D., et al. (2023). Microbial biomanufacturing for space-exploration-what to take and when to make. Nat. Commun. 14 (1), 2311. doi:10.1038/s41467-023-37910-1
Azam, A., Li, C., Metcalf, K. J., and Tullman-Ercek, D. (2016). Type III secretion as a generalizable strategy for the production of full-length biopolymer-forming proteins. Biotechnol. Bioeng. 113 (11), 2313–2320. doi:10.1002/bit.25656
Badie, F., Ghandali, M., Tabatabaei, S. A., Safari, M., Khorshidi, A., Shayestehpour, M., et al. (2021). Use of Salmonella bacteria in cancer therapy: direct, drug delivery and combination approaches. Front. Oncol. 11, 624759. doi:10.3389/fonc.2021.624759
Berliner, A. J., Zezulka, S., Hutchinson, G. A., Bertoldo, S., Cockell, C. S., and Arkin, A. P. (2024). Domains of life sciences in spacefaring: what, where, and how to get involved. npj Microgravity 10 (1), 12. doi:10.1038/s41526-024-00354-y
Braddock, M. (2017). Ergonomic challenges for astronauts during space travel and the need for space medicine. J. Ergon. 7 (221), 2. doi:10.4172/2165-7556.1000221
Brooks, S. M., and Alper, H. S. (2021). Applications, challenges, and needs for employing synthetic biology beyond the lab. Nat. Commun. 12 (1), 1390. doi:10.1038/s41467-021-21740-0
Burdette, L. A., Leach, S. A., Kennedy, N., Ikwuagwu, B. C., Summers, J. S., and Tullman-Ercek, D. (2024). Characterization and engineering of the type 3 secretion system needle monomer from Salmonella through the construction and screening of a comprehensive mutagenesis library. Msphere 9, e0036724. doi:10.1128/msphere.00367-24
Burdette, L. A., Leach, S. A., Wong, H. T., and Tullman-Ercek, D. (2018). Developing Gram-negative bacteria for the secretion of heterologous proteins. Microb. Cell Fact. 17 (1), 196. doi:10.1186/s12934-018-1041-5
Burdette, L. A., Wong, H. T., and Tullman-Ercek, D. (2021). An optimized growth medium for increased recombinant protein secretion titer via the type III secretion system. Microb. Cell Fact. 20 (1), 44. doi:10.1186/s12934-021-01536-z
Chen, X., Li, C., and Liu, H. (2021). Enhanced recombinant protein production under special environmental stress. Front. Microbio 12, 630814. doi:10.3389/fmicb.2021.630814
Comstock, D., and Lockney, D. (2007). “NASA's legacy of technology transfer and prospects for future benefits,” in AIAA SPACE 2007 conference & exposition.
Corrado, L., Cropper, M., and Rao, A. (2023). Space exploration and economic growth: New issues and horizons. Proc. Natl. Acad. Sci. U. S. A. 120, e2221341120. doi:10.1073/pnas.2221341120
Crabbé, A., Pycke, B., Van Houdt, R., Monsieurs, P., Nickerson, C., Leys, N., et al. (2010). Response of Pseudomonas aeruginosa PAO1 to low shear modeled microgravity involves AlgU regulation. Environ. Microbiol. 12 (6), 1545–1564. doi:10.1111/j.1462-2920.2010.02184.x
Darwin, K. H., and Miller, V. L. (2000). The putative invasion protein chaperone SicA acts together with InvF to activate the expression of Salmonella typhimurium virulence genes. Mol. Microbiol. 35 (4), 949–960. doi:10.1046/j.1365-2958.2000.01772.x
Diaz, A. M., Li, W., Irwin, T., Orourke, A., Khodadad, C., Hummerick, M., et al. (2022). “Investigation into space effects on biofilm growth using simulated microgravity,” in 51st International conference on environmental systems.
Douglas, G. L., Wheeler, R. M., and Fritsche, R. F. (2021). Sustaining astronauts: resource limitations, technology needs, and parallels between spaceflight food systems and those on Earth. Sustainability 13 (16), 9424. doi:10.3390/su13169424
Douglas, G. L., Zwart, S. R., and Smith, S. M. (2020). Space food for thought: challenges and considerations for food and nutrition on exploration missions. J. Nutr. 150 (9), 2242–2244. doi:10.1093/jn/nxaa188
Ellermeier, C. D., Ellermeier, J. R., and Slauch, J. M. (2005). HilD, HilC and RtsA constitute a feed forward loop that controls expression of the SPI1 type three secretion system regulator hilA in Salmonella enterica serovar Typhimurium. Mol. Microbiol. 57 (3), 691–705. doi:10.1111/j.1365-2958.2005.04737.x
Ferrer-Miralles, N., Domingo-Espín, J., Corchero, J. L., Vázquez, E., and Villaverde, A. (2009). Microbial factories for recombinant pharmaceuticals. Microb. Cell Fact. 8 (1), 17. doi:10.1186/1475-2859-8-17
Glasgow, A. A., Wong, H. T., and Tullman-Ercek, D. (2017). A secretion-Amplification role for Salmonella enterica Translocon protein SipD. ACS Synth. Biol. 6 (6), 1006–1015. doi:10.1021/acssynbio.6b00335
Golubeva, Y. A., Sadik, A. Y., Ellermeier, J. R., and Slauch, J. M. (2012). Integrating global regulatory Input into the Salmonella pathogenicity island 1 type III secretion system. Genetics 190 (1), 79–90. doi:10.1534/genetics.111.132779
Green, E. R., and Mecsas, J. (2016). Bacterial secretion systems: an Overview. Microbiol. Spectr. 4 (1), 213–239. doi:10.1128/9781555819286.ch8
Gunn, J. S. (2000). Mechanisms of bacterial resistance and response to bile. Microbes Infect. 2 (8), 907–913. doi:10.1016/s1286-4579(00)00392-0
Huangfu, J., Kim, H. S., Xu, K., Ning, X., Qin, L., Li, J., et al. (2020). Omics analysis reveals the mechanism of enhanced recombinant protein production under simulated microgravity. Front. Bioeng. Biotechnol. 8, 30. doi:10.3389/fbioe.2020.00030
Humphrey, T. (2004). Salmonella, stress responses and food safety. Nat. Rev. Microbiol. 2 (6), 504–509. doi:10.1038/nrmicro907
Jemison, M., and Olabisi, R. (2021). Biomaterials for human space exploration: a review of their untapped potential. Acta Biomater. 128, 77–99. doi:10.1016/j.actbio.2021.04.033
Kim, H., and Rhee, M. (2016). Influence of low-shear modeled microgravity on heat resistance, membrane fatty acid composition, and heat stress-related gene expression in Escherichia coli O157: H7 ATCC 35150, ATCC 43889, ATCC 43890, and ATCC 43895. Appl. Environ. Microbiol. 82 (10), 2893–2901. doi:10.1128/aem.00050-16
Kim, H. W., and Rhee, M. S. (2021). Low-shear modeled microgravity impacts the acid stress response and post-thermal stress behavior of acid-resistant, adaptable, and sensitive Escherichia coli O157:H7 strains. Food control. 121, 107603. doi:10.1016/j.foodcont.2020.107603
Liang, J. M., Burdette, L. A., Wong, H. T., and Tullman-Ercek, D. (2023). Construction of a constitutively active type III secretion system for heterologous protein secretion. Appl. Microbiol. Biotechnol. 107 (5), 1785–1800. doi:10.1007/s00253-023-12411-9
Lou, L., Zhang, P., Piao, R., and Wang, Y. (2019). Salmonella pathogenicity island 1 (SPI-1) and its complex regulatory network. Front. Cell Infect. Microbio 9, 270. doi:10.3389/fcimb.2019.00270
MacLennan, C. A., Martin, L. B., and Micoli, F. (2014). Vaccines against invasive Salmonella disease: current status and future directions. Hum. Vaccin Immunother. 10 (6), 1478–1493. doi:10.4161/hv.29054
Menezes, A. A., Cumbers, J., Hogan, J. A., and Arkin, A. P. (2015). Towards synthetic biological approaches to resource utilization on space missions. J. R. Soc. Interface 12 (102), 20140715. doi:10.1098/rsif.2014.0715
Menezes, A. A., Montague, M. G., Cumbers, J., Hogan, J. A., and Arkin, A. P. (2015). Grand challenges in space synthetic biology. J. R. Soc. Interface 12 (113), 20150803. doi:10.1098/rsif.2015.0803
Metcalf, K. J., Bevington, J. L., Rosales, S. L., Burdette, L. A., Valdivia, E., and Tullman-Ercek, D. (2016). Proteins adopt functionally active conformations after type III secretion. Microb. Cell Fact. 15 (1), 213. doi:10.1186/s12934-016-0606-4
Metcalf, K. J., Finnerty, C., Azam, A., Valdivia, E., and Tullman-Ercek, D. (2014). Using transcriptional control to increase titers of secreted heterologous proteins by the type III secretion system. Appl. Enviro Microbiol. 80 (19), 5927–5934. doi:10.1128/aem.01330-14
Nickerson, C. A., Ott, C. M., Mister, S. J., Morrow, B. J., Burns-Keliher, L., and Pierson, D. L. (2000). Microgravity as a novel environmental signal affecting Salmonella enterica serovar Typhimurium virulence. Infect. Immun. 68 (6), 3147–3152. doi:10.1128/iai.68.6.3147-3152.2000
Nickerson, C. A., Ott, C. M., Wilson, J. W., Ramamurthy, R., LeBlanc, C. L., zu Bentrup, K. H., et al. (2003). Low-shear modeled microgravity: a global environmental regulatory signal affecting bacterial gene expression, physiology, and pathogenesis. J. Microbiol. Methods 54 (1), 1–11. doi:10.1016/s0167-7012(03)00018-6
Nielsen, J., and Keasling, J. D. (2016). Engineering cellular metabolism. Cell 164 (6), 1185–1197. doi:10.1016/j.cell.2016.02.004
Ouchen, F., Yaney, P., Rau, I., Kajzar, F., and Grote, J. (2014). Biopolymers suitable for space environments. Proc. SPIE 9226, 92260T. doi:10.1117/12.2070254
Overton, T. W. (2014). Recombinant protein production in bacterial hosts. Drug Discov. Today 19 (5), 590–601. doi:10.1016/j.drudis.2013.11.008
Pavez Loriè, E., Baatout, S., Choukér, A., Buchheim, J.-I., Baselet, B., Dello Russo, C., et al. (2021). The future of personalized medicine in space: from observations to countermeasures. Front. Bioeng. Biotechnol. 9, 739747. doi:10.3389/fbioe.2021.739747
Prieto, A. I., Ramos-Morales, F., and Casadesús, J. (2004). Bile-induced DNA damage in Salmonella enterica. Genetics 168 (4), 1787–1794. doi:10.1534/genetics.104.031062
Reed, B., and Chen, R. (2013). Biotechnological applications of bacterial protein secretion: from therapeutics to biofuel production. Res. Microbiol. 164 (6), 675–682. doi:10.1016/j.resmic.2013.03.006
Roncarati, D., and Scarlato, V. (2017). Regulation of heat-shock genes in bacteria: from signal sensing to gene expression output. FEMS Microbiol. Rev. 41 (4), 549–574. doi:10.1093/femsre/fux015
Sanchez-Garcia, L., Martín, L., Mangues, R., Ferrer-Miralles, N., Vázquez, E., and Villaverde, A. (2016). Recombinant pharmaceuticals from microbial cells: a 2015 update. Microb. Cell Fact. 15 (1), 33. doi:10.1186/s12934-016-0437-3
Santomartino, R., Averesch, N. J. H., Bhuiyan, M., Cockell, C. S., Colangelo, J., Gumulya, Y., et al. (2023). Toward sustainable space exploration: a roadmap for harnessing the power of microorganisms. Nat. Commun. 14 (1), 1391. doi:10.1038/s41467-023-37070-2
Schumacher, K., Brameyer, S., and Jung, K. (2023). Bacterial acid stress response: from cellular changes to antibiotic tolerance and phenotypic heterogeneity. Curr. Opin. Microbiol. 75, 102367. doi:10.1016/j.mib.2023.102367
Schwarz, R. P., Goodwin, T. J., and Wolf, D. A. (1992). Cell culture for three-dimensional modeling in rotating-wall vessels: an application of simulated microgravity. J. Tissue Cult. Methods 14 (2), 51–57. doi:10.1007/bf01404744
Schwarz, R. P., and Wolf, D. A. (1991). Rotating bio-reactor cell culture apparatus. United States. Patent no US4988623A.
Senatore, G., Mastroleo, F., Leys, N., and Mauriello, G. (2020). Growth of Lactobacillus reuteri DSM17938 under two simulated microgravity systems: changes in reuterin production, gastrointestinal passage resistance, and stress genes expression response. Astrobiology 20 (1), 1–14. doi:10.1089/ast.2019.2082
Sheet, S., Yesupatham, S., Ghosh, K., Choi, M. S., Shim, K. S., and Lee, Y. S. (2020). Modulatory effect of low-shear modeled microgravity on stress resistance, membrane lipid composition, virulence, and relevant gene expression in the food-borne pathogen Listeria monocytogenes. Enzyme Microb. Technol. 133, 109440. doi:10.1016/j.enzmictec.2019.109440
Shen, S., and Fang, F. C. (2012). Integrated stress responses in Salmonella. Int. J. Food Microbiol. 152 (3), 75–81. doi:10.1016/j.ijfoodmicro.2011.04.017
Song, M., Sukovich, D. J., Ciccarelli, L., Mayr, J., Fernandez-Rodriguez, J., Mirsky, E. A., et al. (2017). Control of type III protein secretion using a minimal genetic system. Nat. Commun. 8 (1), 14737. doi:10.1038/ncomms14737
Su, X., Guo, Y., Fang, T., Jiang, X., Wang, D., Li, D., et al. (2021). Effects of simulated microgravity on the physiology of Stenotrophomonas maltophilia and multiomic analysis. Front. Microbiol. 12, 701265. doi:10.3389/fmicb.2021.701265
Tang, C., Wang, L., Zang, L., Wang, Q., Qi, D., and Dai, Z. (2023). On-demand biomanufacturing through synthetic biology approach. Mater Today Bio 18, 100518. doi:10.1016/j.mtbio.2022.100518
Tartera, C., and Metcalf, E. S. (1993). Osmolarity and growth phase overlap in regulation of Salmonella typhi adherence to and invasion of human intestinal cells. Infect. Immun. 61 (7), 3084–3089. doi:10.1128/iai.61.7.3084-3089.1993
Tepap, C. Z., Anissi, J., and Bounou, S. (2023). Recent strategies to achieve high production yield of recombinant protein: a review. J. Cell. Biotechnol. 9 (1), 25–37. doi:10.3233/jcb-220084
Tucker, D. L., Ott, C. M., Huff, S., Fofanov, Y., Pierson, D. L., Willson, R. C., et al. (2007). Characterization of Escherichia coli MG1655 grown in a low-shear modeled microgravity environment. BMC Microbiol. 7, 15–16. doi:10.1186/1471-2180-7-15
Widmaier, D. M., Tullman-Ercek, D., Mirsky, E. A., Hill, R., Govindarajan, S., Minshull, J., et al. (2009). Engineering the Salmonella type III secretion system to export spider silk monomers. Mol. Syst. Biol. 5, 309. doi:10.1038/msb.2009.62
Wilson, J. W., Ott, C. M., Ramamurthy, R., Porwollik, S., McClelland, M., Pierson, D. L., et al. (2002). Low-shear modeled microgravity alters the Salmonella enterica Serovar Typhimurium stress response in an RpoS-independent manner. Appl. Environ. Microbiol. 68 (11), 5408–5416. doi:10.1128/aem.68.11.5408-5416.2002
Wilson, J. W., Ott, C. M., zu Bentrup, K. H., Ramamurthy, R., Quick, L., Porwollik, S., et al. (2007). Space flight alters bacterial gene expression and virulence and reveals a role for global regulator Hfq. Proc. Natl. Acad. Sci. U. S. A. 104 (41), 16299–16304. doi:10.1073/pnas.0707155104
Xiang, L., Qi, F., Dai, D., Li, C., and Jiang, Y. (2010). Simulated microgravity affects growth of Escherichia coli and recombinant β-D-glucuronidase production. Appl. Biochem. Biotechnol. 162, 654–661. doi:10.1007/s12010-009-8836-0
Yan, X., Liu, X., Zhao, C., and Chen, G.-Q. (2023). Applications of synthetic biology in medical and pharmaceutical fields. Signal Transduct. Target Ther. 8 (1), 199. doi:10.1038/s41392-023-01440-5
Yang, J., Barrila, J., Roland, K. L., Ott, C. M., and Nickerson, C. A. (2016). Physiological fluid shear alters the virulence potential of invasive multidrug-resistant non-typhoidal Salmonella Typhimurium D23580. npj Microgravity 2 (1), 16021–16023. doi:10.1038/npjmgrav.2016.21
Yim, J., Cho, S. W., Kim, B., Park, S., Han, Y. H., and Seo, S. W. (2020). Transcriptional Profiling of the Probiotic Escherichia coli Nissle 1917 strain under simulated microgravity. Int. J. Mol. Sci. 21 (8), 2666. doi:10.3390/ijms21082666
Keywords: microgravity, Salmonella, type 3 secretion system (T3SS), protein secretion, synthetic biology
Citation: Kang M-K, Bevington J and Tullman-Ercek D (2025) Evaluation of the Salmonella type 3 secretion system (T3SS) as part of a protein production platform for space biology applications. Front. Bioeng. Biotechnol. 13:1567596. doi: 10.3389/fbioe.2025.1567596
Received: 27 January 2025; Accepted: 18 March 2025;
Published: 02 April 2025.
Edited by:
Cheemeng Tan, University of California, Davis, United StatesReviewed by:
Shaobo Yang, Dana–Farber Cancer Institute, United StatesCopyright © 2025 Kang, Bevington and Tullman-Ercek. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Danielle Tullman-Ercek, ZXJjZWtAbm9ydGh3ZXN0ZXJuLmVkdQ==
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.