
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Bioeng. Biotechnol. , 20 March 2025
Sec. Biosensors and Biomolecular Electronics
Volume 13 - 2025 | https://doi.org/10.3389/fbioe.2025.1499164
This article is part of the Research Topic Sensing Technologies for Sustainable Healthcare View all 6 articles
Introduction: Testosterone is a steroid hormone, which plays a pivotal role in regulating metabolism and protein synthesis in the body. The detection of testosterone is of paramount importance for diagnostic purposes in clinical settings, as well as for monitoring athletes’ physiological parameters and ensuring the integrity of sports competitions.
Methods: Herein, we present a phage display-derived biosensing platform through genetic engineering of the TS77 antibody variable domains. The variable region genes of the heavy and light chains from TS77 antibody were cloned into the pDong1 plasmid and displayed on the phage surfaces through phage display technology. Subsequently, a novel non-competitive open-sandwich ELISA (OS-ELISA) was developed for testosterone detection, leveraging the antigen-induced interaction changes in antibody variable regions.
Results: OS-ELISA based on anti L-chain antibody achieved a limit of detection (LOD) of 2.71 nM and a half-maximal effective concentration (EC50) of 0.22 μM for testosterone detection. Furthermore, the enhanced OS-ELISA platform incorporating purified maltose binding protein fused with VL (MBP-VL) and VH phage demonstrated a LOD of 1.07 pM and a wide working range from 1 pM to 10 mM.
Discussion: The OS-ELISA developed in this study exhibits high sensitivity and a broad dynamic range for testosterone quantification, showing significant potential for clinical diagnostics and athlete monitoring applications.
Testosterone (Ide, 2023), a steroidal hormone, plays a critical role in regulating physiological processes and promoting protein synthesis in the body. It is essential for maintaining muscle strength and mass, preserving bone density and strength, improving metabolism, and enhancing overall physical performance. Deviations in testosterone levels have been consistently associated with a range of pathological conditions, such as Polycystic Ovary Syndrome (PCOS) (Hahn et al., 2007), Ovarian Steroid Cell Tumor (OSCT) (Mizoguchi et al., 2014), and Adrenal Cortex Hyperplasia (Ciumas et al., 2009). Furthermore, the detection of testosterone is of vital significance in the physiological assessment of elite athletes. Firstly, testosterone levels serve as a biomarker reflecting an athlete’s athletic capabilities and anabolic metabolism (Wang et al., 2023). Secondly, owing to its potential for performance enhancement, testosterone has been proscribed as a prohibited substance in sports competitions (You et al., 2011). Therefore, the detection of testosterone is of paramount importance for diagnostic purposes in clinical settings, as well as for monitoring athlete performance and ensuring the integrity of sports competitions.
Current methodologies for testosterone detection primarily encompass two categories: those reliant on large-scale equipment and those based on antibody-mediated immunological detection. Within the former, notable examples include the Gas Chromatography-Tandem Mass Spectrometry (GC-MS/MS) approach employed by institutions such as Ghent University in Belgium (Thienpont et al., 1994) and the German Society for Clinical Chemistry and Laboratory Medicine (Siekmann, 1979), as well as the Liquid Chromatography-Tandem Mass Spectrometry (LC-MS/MS) method utilized by organizations like the Health Sciences Authority (Chen et al., 2019), Center for Disease Control and Prevention (Botelho et al., 2013), and National Institute of Standards and Technology (Tai et al., 2007). These methods, while precise, often require skilled operators and complex preprocessing steps.
Alternatively, immunoassay that exploit antigen-antibody interactions, such as the Indirect Competitive Enzyme-Linked Immunosorbent Assay (ic-ELISA) (Jiang et al., 2011) offer simplicity in operation. However, due to the inherent limitations of the competitive format, achieving marked signal changes in the presence of trace analytes necessitates minimizing the amounts of both antibodies and competitive haptens, which leads to the loss in assay sensitivity.
Open Sandwich Enzyme-linked Immunosorbent Assay (OS-ELISA) (Lim et al., 2007) is a novel immunoassay based on the interaction of variable regions of the heavy chain (VH) and light chain (VL) of the antibody. The principle of OS-ELISA is that the interaction between VH and VL is modulated by the presence of an antigen. VH and VL barely bind to each other in the absence of antigen. When antigen is present, they bind to the antigen to form a trimeric structure, and the signal gradually intensifies as the antigen concentration increases. Based on the principle, a non-competitive immunoassay is established for rapid detection of analytes of interest. OS-ELISA has as already proven successful in the detection of a variety of haptens, including environmental hazards and food additives, such as Microcystin-LR (MCLR) (Chen et al., 2020), Clenbuterol (CLEN) (Cong et al., 2019), Tenuazonic acid (TeA) (Liang et al., 2021) and so on.
In this study, we have developed a novel noncompetitive immunoassay strategy for testosterone. This approach utilizes antigen-driven affinity enhancement, enabling rapid quantification of testosterone. By incorporating the advantages of OS-ELISA, our technique promises to offer a highly sensitive and efficient means for testosterone detection, contributing to both medical diagnostics and athletic performance monitoring.
The TS77 sequence was designed with reference to previously reported sequences (Valjakka et al., 2002). The genes and primers utilized in the experiments were synthesized by Sangon Biotechnology (Shanghai, China). The primers used are summarized in Table 1. Escherichia coli DH5α used for gene cloning and plasmid amplification, was purchased from Tsingke Biotech (Beijing, China). Escherichia coli TG-1 used for phage display was obtained from GE Healthcare (Tokyo, Japan). Escherichia coli BL21-Gold (DE3) pLysS used for expressing the antibody fusion protein, was purchased from Agilent Technologies (La Jolla, California, United States). The restriction endonucleases were sourced from New England Biolabs (Beverly, Massachusetts, United States). Ligation High Ver.2 used for DNA ligation and KOD-Plus-Neo used for PCR, were purchased from TOYOBO (Osaka, Japan). Testosterone was purchased from Aladdin Biochemical Technology (Shanghai, China). Additionally, all other chemicals and reagents were sourced from Vazyme (Nanjing, China) or Sangon Biotech (Shanghai, China).
The single chain variable fragment (scFv) gene of TS77 was synthesized and cloned into the pUC57 (Shanghai Sangon Biotech Co., Ltd.). This plasmid was then utilized as a template for the amplification of the scFv gene via polymerase chain reaction (PCR) using KOD Plus Neo and the primers M13+ and M13-. Subsequently, the PCR products were analyzed using agarose gel electrophoresis, and the desired fragments were recovered using a gel extraction kit.
The scFv gene, following its recovery, was subjected to restriction digestion along with the pDong1 (blank) plasmid using NcoI and XhoI at 37°C for 4 h. The digestion products were analyzed via agarose gel electrophoresis, and the target fragments were purified using a gel extraction kit. Subsequently, the digested plasmid and gene fragment were ligated using Ligation High Vers.2 at 16°C for 2 h. The ligation mixture was then evenly spread onto LBA solid medium (10 g/L tryptone, 5 g/L yeast extract, 5 g/L NaCl, 15 g/L agar, 100 μg/mL ampicillin) for bacterial transformation.
After verifying the clones using PCR with M13Rv and pHENseq primers, the positive clones were chosen, and their plasmids were isolated for sequencing. The positive plasmids containing the scFv gene were further processed with SalI and NotI at 37°C for 4 h. The digested products were analyzed, purified, and subsequently ligated again. The ligation mixture was once more spread onto LBA solid medium, and after PCR analysis with primers pDong1VLback and pDong1CKfor, positive clones were picked, their plasmids extracted, and subjected to sequencing to confirm the integrity of the inserted gene. The plasmid with the correct sequence was named pDong1/Fab-TS77.
The plasmid pDong1/Fab-TS77 was transformed into TG-1 competent cells and spread onto 2YTAG solid medium (16 g/L tryptone, 10 g/L yeast extract, 5 g/L NaCl, 15 g/L agar, 100 μg/mL ampicillin and 1% glucose). Single colonies were selected and cultivated in 2YTAG liquid medium (16 g/L tryptone, 10 g/L yeast extract, 5 g/L NaCl, 100 μg/mL ampicillin and 1% glucose) until the OD600 reached 0.2. Subsequently, 1010 KM13 helper phages (Dong et al., 2009) were added, and the mixture was incubated at 37°C for 1 h. Following incubation, the cells were centrifuged at 5,500 g for 10 min, and the pellet was resuspended in 4 mL of 2YTAGK medium (2YT medium containing 100 μg/mL ampicillin, 50 μg/mL kanamycin, and 0.1% glucose). The culture was then grown at 30°C with shaking at 250 rpm for 20 h.
After cultivation, the cells were centrifuged at 3,300 g for 30 min, and the supernatant was collected. To precipitate the phage, A fifth of the total volume of PEG/NaCl solution (20% polyethylene glycol 6,000, 2.5 M NaCl) was added to the supernatant, followed by incubation on ice for 1 h. Subsequently, the mixture was centrifuged at 5,500 g for 30 min at 4°C. The supernatant was discarded, and the phage pellet was resuspended in 200 μL of phosphate-buffered saline (PBS). To eliminate any remaining cellular debris, the suspension was centrifuged at 11,600 g for 10 min, and the supernatant containing the displaying antigen-binding fragment (Fab) phages was collected.
To eliminate the CH1 from the pDong1/Fab-TS77, the plasmid was digested with SgrAI at 37°C for 4 h. Subsequently, the target fragment was recovered using a gel extraction kit and subsequently ligated to itself using Ligation High Ver.2 at 16°C for 2 h. The ligation product was then transformed into TG-1 competent cells, which were evenly spread on 2YTAG solid medium and incubated at 37°C overnight. After confirmation of positive clones through colony PCR with primers M13Rv and pHENseq and sequencing, the positive clone was subjected to expanded culture, and plasmid extraction was performed to obtain pDong1/OS-TS77. The plasmid was transformed into TG-1 competent cells for phage display, with steps similar to those described above. After culture, the supernatant which contained the VH phage and L chains was collected.
For the implementation of OS-ELISA, 100 μL/well of 2 μg/mL rabbit anti-human kappa chain antibody was coated onto microplates and incubated overnight at 4°C. The plates were then blocked with MPBS (PBS containing 2% skim milk) at 25°C for 2 h, followed by 3 washes with PBST (PBS containing 1% Tween 20). Subsequently, a 100 μL mixture of phage solution (containing 1010 phages) and antigen dilutions at various concentrations in equal volumes was added and incubated at 25°C for 1 h. After 5 washes with PBST, 0.2 μg/mL of HRP-conjugated anti-M13 antibody was added and incubated at 25°C for 1 h. Following another 5 washes with PBST, the TMBZ solution (100 μg/mL 3, 3′, 5, 5′-tetramethylbenzidine and 0.02 μL/mL H2O2 in 100 mM NaOAc, pH 6.0) was added for color development, and 50 μL/well of 10% H2SO4 was used to stopped. After the reaction was terminated, the absorbance values at 450 nm and 630 nm were measured, and the data were analyzed to construct a standard curve.
To facilitate the fusion expression of maltose-binding protein (MBP) with the TS77 VL domain, the plasmid pMAL-VL (TS77) was developed based on the pMAL-VL (3A8) plasmid (Chen et al., 2020). The pMAL-VL (3A8) plasmid and the TS77 scFv gene were individually amplified and subsequently digested using SalI and NotI restriction enzymes. The digested plasmid and gene fragments were then ligated using Ligation High Ver.2, followed by transformation into DH5α competent cells and even spreading on LBA solid medium. Positive clones were identified through PCR verification using MalE and M13F primers, and selected clones underwent plasmid extraction and sequencing.
The plasmid was then transformed into BL21 (DE3) pLysS competent cells and evenly spread on LBA solid medium. After overnight incubation, a single clone was picked and inoculated into 4 mL LBA liquid medium for cultivation for 12–16 h, which was then transferred to 300 mL LBA liquid medium for expanded growth. When the OD600 reached 0.6, isopropyl β-D-1-thiogalactopyranoside (IPTG) with a final concentration of 1 mM was added and then cultured at 25°C with shaking at 250 rpm for 21 h. After a centrifugation at 8,000 g for 20 min at 4°C, the bacterial pellet was resuspended, and the cells were lysed by sonication to obtain a soluble protein solution. This solution was subsequently purified using Ni-NTA Sefinose Resin (Shanghai Sango Biotechnology, Co. Ltd.). To assess the concentration and purity of the purified protein, 10 μL of each sample was analyzed by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and stained with Coomassie Blue R250.
After phage display of pDong1/OS-TS77, a supernatant containing VH phage and L chain was mixed with a fifth of the total volume of PEG/NaCl and incubated in ice for 1 h, followed by centrifugation at 5,500 g for 1 h at 4°C, and the supernatant was removed. The pellet was resuspended in PBS and centrifuged again, yielding a final supernatant enriched with VH phage.
For the assay, 100 μL/well of 2 μg/mL MBP-VL was coated onto microplates and incubated overnight at 4°C. 2% MPBS was used to block the wells at 25°C for 2 h, followed by 3 washes with PBST. A mixture of 1010 cfu of VH phage solution and a series of concentrations of antigen was added to the wells and incubated at 25°C for 1 h. After washing 5 times with PBST, 0.2 μg/mL of HRP-conjugated anti-M13 antibody was added and incubated at 25°C for 1 h. Following another 5 washes with PBST, the assay was performed as those described above.
Microplates were coated with 100 μL/well of 2 μg/mL MBP-VL and incubated overnight at 4°C. 2% MPBS was used to block the wells at 25°C for 2 h. After washing 3 times with PBST, 1010 cfu of VH phage solution was mixed with different concentrations of testosterone, estradiol, and cortisol. The mixture was added to the wells and incubated at 25°C for 1 h. Then the steps of OS-ELISA was performed as those described above. The data obtained were analyzed to compare the responses generated by testosterone against those of estradiol and cortisol, allowing for the assessment of the specificity of the MBP-VL based enhanced OS-ELISA towards testosterone detection. The results were used to plot standard curves for each hormone, enabling a direct comparison of the sensitivity and specificity of the assay towards testosterone versus potential cross-reactive hormones such as estradiol and cortisol.
To evaluate the binding affinity and interaction patterns between TS77 and testosterone, we used Autodock Vina 1.2.2 (Zuo et al., 2005), a computer-aided protein-ligand docking software. The molecular structure of testosterone was retrieved from the PubChem compound database (https://pubchem.ncbi.nlm.nih.gov/). The variable region sequence of the TS77 antibody was predicted using Swiss-Model (http://swissmodel.expasy.org/). Initially, the antibody and small molecule files were prepared by converting them to the PDBQT format, excluding all water molecules, and appending polar hydrogen atoms. The grid box was centered to encompass the domain of each protein structure while accommodating the movement of the free molecule. The docking pocket was configured as a 30 Å × 30 Å×30 Å cubic box with a grid spacing of 0.05 nm. The molecular docking was performed using Autodock Vina 1.2.2 (http://autodock.scripps.edu/), and the results were utilized for model visualization.
The data obtained from OS-ELISA were analyzed using Prism 9 (GraphPad Software, San Diego, CA, United States). The dose–response curves were fitted with a four-parameter equation as follows:
where:
y is the response at concentration x,
a is the maximum response (in arbitrary units, a.u.),
b is the slope factor (dimensionless),
c is the half-maximal effective concentration (in picoMolar, pM),
d is the minimum response (in arbitrary units, a.u.).
The EC50 was obtained from Nonlin fit results of dose–response curves. The LOD for each assay was obtained by adding the estimated antigen concentration that showed the mean blank value to 3 times the standard deviation for the OS-ELISA.
To validate the antigen-binding activity of the TS77 Fab expressed via phage display, we constructed the pDong1/Fab-TS77 Figure 1A. The Fd was fused with the phage pIII protein for expression, and the SgrAI site was added to both ends of the CH1 domain, while the L chain was expressed independently. Through PCR amplification of the TS77 scFv, distinct bands were observed between 750 and 1,000 bp after agarose gel electrophoresis, indicating successful amplification of the TS77 scFv Figure 1B. Following digestion with restriction endonucleases, the VH and VL genes were obtained. These were subsequently cloned into the pDong1-blank vector, resulting in the acquisition of pDong1/Fab-TS77.
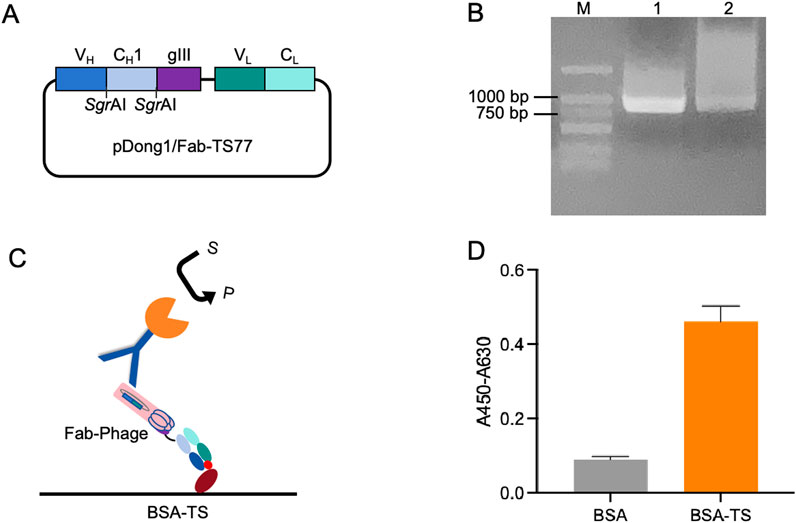
Figure 1. Phage display of TS77 antigen-binding fragment. (A) Schematic of the pDong1/Fab-TS77. (B) Agarose gel electrophoresis analysis of TS77 scFv gene. M: DNA marker. (C) Schematic of the ELISA. (D) ELISA analysis of the antigen binding activity of phage-displayed Fab. Error bars represent standard deviation (SD) of triplicate samples.
After phage display, the TS77 Fab phage was obtained with a titer of 1013 cfu/mL. To verify the activity of the phage-displayed TS77 Fab, we coated microplates with BSA and BSA-TS separately and identified its antigen-binding activity using ELISA Figure 1C. As shown in Figure 1D, the absorbance of the BSA-TS was significantly higher than that of the negative control BSA, indicating that the phage-displayed TS77 Fab exhibited good binding activity towards testosterone.
Subsequently, we removed the SgrAI restriction sites from both ends of CH1 to obtain the pDong1/OS-TS77 plasmid Figure 2A. Following phage display, the VH was presented on the phage surface, while the VL remained free in the culture medium due to the loss of the CH1-CL dimerization. Utilizing an Anti-L chain antibody, we directly employed the culture medium containing both the L chain and VH phage for testosterone detection. The schematic of the OS-ELISA is depicted in Figure 2B. The OS-ELISA results showed an increase in absorbance with rising testosterone concentrations, indicating a dose-dependent relationship Figure 2C. The limit of detection (LOD) was calculated to be 2.71 nM, and the half-maximal effective concentration (EC50) was 0.22 μM. These results demonstrate that the OS-ELISA method is capable of detecting testosterone effectively.
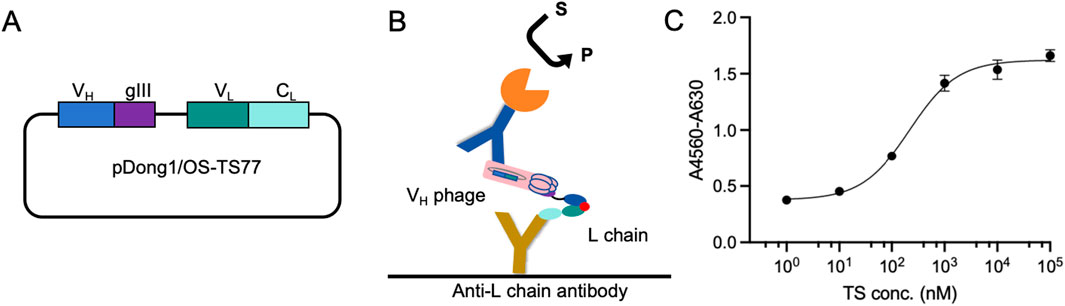
Figure 2. Open sandwich phage ELISA. (A) Schematic of the pDong1/OS-TS77. (B) Schematic of the OS-ELISA. (C) Dose-response curve for the detection of testosterone concentration. Error bars represent standard deviation (SD) of triplicate samples.
To further enhance the detection sensitivity and range, we fused the VL with MBP to obtain MBP-VL. Fusion with MBP enhances the stability, expression level, and coating efficiency of VL, while mitigating signal loss in multi-step binding processes during OS-ELISA. The TS77 VL was cloned into the pMAL-blank vector to generate pMAL-TS77VL Figure 3A. The plasmid was expressed in BL21 (DE3) pLysS. The purity of the expressed MBP-VL was confirmed by SDS-PAGE, which revealed a distinct protein band near 55–75 kDa Figure 3B, consistent with the predicted size of 60.7 kDa for MBP-VL, indicating successful expression.
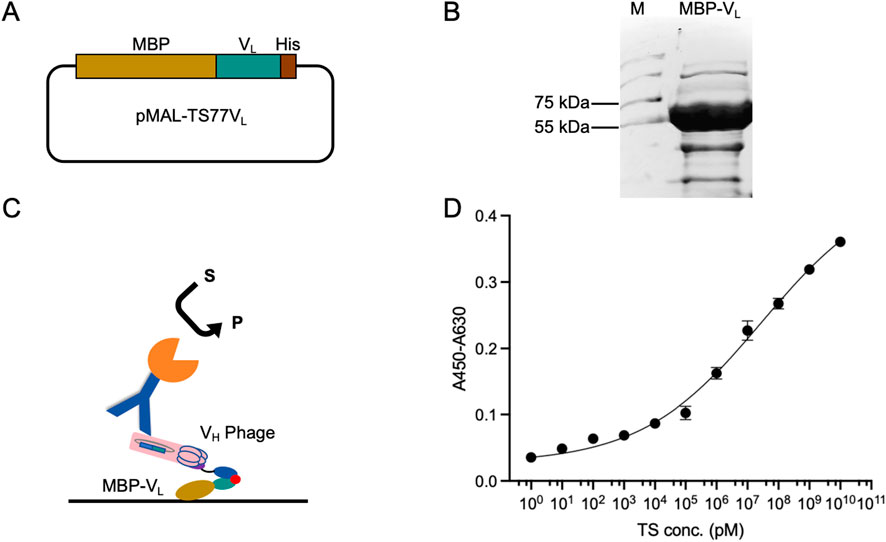
Figure 3. Enhanced OS-ELISA using purified phage displayed VH and MBP-VL fusion protein. (A) Schematic of the pMAL-TS77VL. (B) Schematic of the enhanced OS-ELISA. (C) SDS-PAGE analysis of MBP-VL with Coomassie Blue staining. (D) Dose-response curve for the detection of testosterone concentration. Error bars represent standard deviation (SD) of triplicate samples.
OS-ELISA for testosterone detection was performed using MBP-VL and VH phage Figure 3C. As shown in Figure 3D, a dose-dependent relationship was observed between the absorbance and testosterone concentration, with significant improvements in sensitivity and detection range. And the wide detection range is from 1 pM to 10 mM. Besides, the LOD was lowered to 1.07 pM, and the EC50 was 28.7 μM. This enhanced OS-ELISA exhibited a 2000-fold decrease in LOD, resulting in a substantial improvement in sensitivity. In comparison, commercial testosterone ELISA kits typically have LOD ranging from 100 to 200 pM, suggesting that the enhanced OS-ELISA based on MBP-VL and VH phage possesses ultra-high sensitivity.
To further validate the specificity of the OS-ELISA for testosterone detection, we performed simultaneous detection of estradiol (Waifalkar et al., 2022) and cortisol (Hameed et al., 2013), which has a similar structure to testosterone Figure 4A, using the enhanced OS-ELISA. The absorbance of the reaction system was measured in the presence of a series of concentrations of these analytes. As shown in Figure 4B, the OS-ELISA exhibited distinct differences in response to the different analytes. With increasing concentrations of testosterone, the absorbance displayed a dose-dependent increase. In contrast, no significant increase in absorbance was observed with increasing concentrations of estradiol or cortisol. These results suggest that the OS-ELISA detection method exhibits strong specificity towards testosterone.
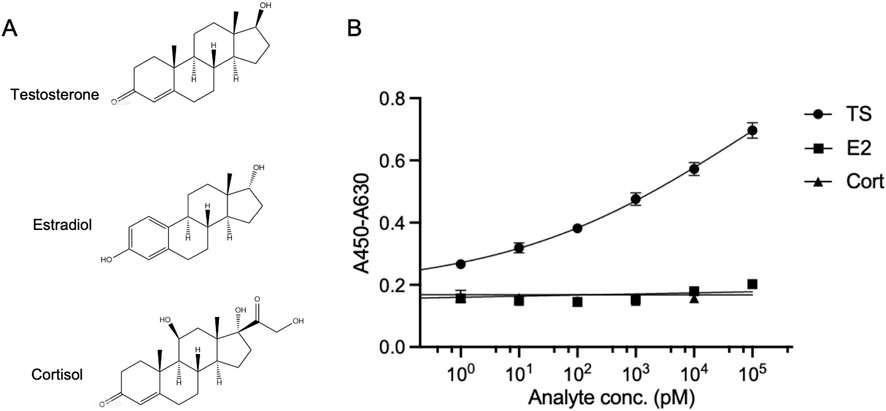
Figure 4. Cross-reactivity of OS-ELISA with structural analogues. (A) Structures of testosterone, estradiol and cortisol. (B) Dose-response curve of the analyte detection. Error bars represent standard deviation (SD) of triplicate samples.
To analyze the binding site of TS77 with testosterone, molecular docking analysis was performed. The binding position and interactions between the TS77 scFv and testosterone were obtained using AutoDock Vina v.1.2.2. The binding free energy was calculated to be −8.275, indicating a highly stable binding. Notably, the GLY-108 of VH and the LYS-196 of VL were identified as key residues forming visible hydrogen bonds with testosterone Figure 5. This suggests that the presence of testosterone facilitates the formation of a more stable trimer consisting of these three components, which is crucial for the successful establishment of the OS-ELISA.
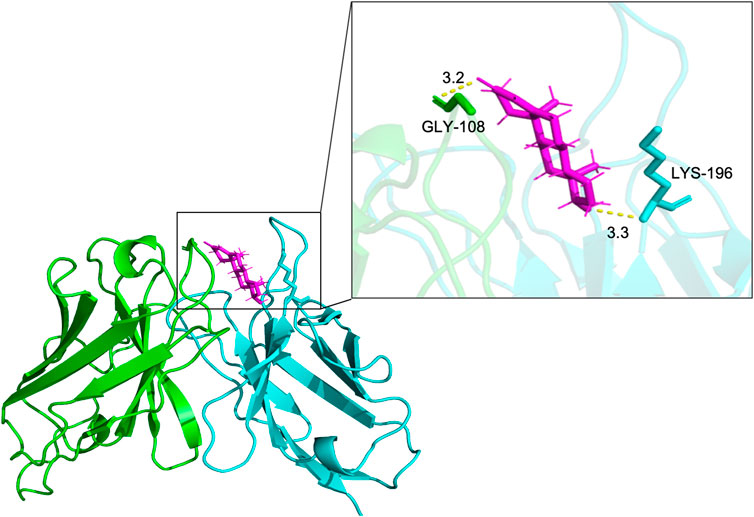
Figure 5. The docking results of antibody V region and testosterone. Key amino acids in VH and VL were presented by different colors of sticks. The results were presented in form of PyMOL. Hydrogen bonds are shown as yellow dotted lines.
Testosterone, as a steroid hormone, plays a pivotal role in physiological regulation and protein synthesis, making it a key reference in disease diagnosis and functional assessment of athletes (Mulhall et al., 2018). Conventional instrumental methods for testosterone detection primarily include GC-MS/MS, LC-MS/MS, among others. These methodologies exhibit high sensitivity, along with a broad linear range. However, they necessitate specialized technical expertise from operators and impose stringent requirements on experimental instruments and conditions.
In this study, we have developed a non-competitive immunoassay method specifically for testosterone, which differentiates itself from traditional immunoassays that employ full-length antibody. The OS-ELISA is a detection method utilizing recombinant antibody fragments, VH and VL domains (Ueda et al., 1996). This approach circumvents the limitation of epitope quantity and is capable of detecting haptens with molecular weights less than 1,000 Da (Suzuki et al., 2000). The efficacy of ic-ELISA is often dictated by the affinity of the antibodies used, thereby restricting its sensitivity and detection range. Conversely, OS-ELISA offers non-competitive detection of small molecules, with enhanced sensitivity and an expanded detection range (Gonzalez-Techera et al., 2007).
The success of this detection method hinges critically on the enhancement of the affinity between VH and VL in the presence of the antigen, thereby necessitating a low inherent affinity between VH and VL (Islam et al., 2010). The pDong1 phage display system (Dong et al., 2009) employed in this study not only display antibody fragments on the phage surface but also facilitates the assessment of VH-VL interactions. After validating the antigen-binding activity of the Fab fragment through phage display, we utilized the SgrAI sites flanking the CH1 domain in pDong1/Fab-TS77 to excise CH1. Since Fd is fused to the phage pIII protein, while the L chain exists as a separate open reading frame, excision of CH1 disrupts their dimerization, allowing the light chain to exist independently in solution. Subsequently, the results of OS-ELISA can be used to evaluate the interaction between VH and VL. It also provides reference for the development of another immunoassay, Quenchbody (Q-body) (Dong and Ueda, 2021).
MBP exhibits excellent solubilizing properties and is frequently incorporated into prokaryotic expression systems as a solubilizing tag. In this study, the fusion of VL with MBP enhanced the sensitivity of OS-ELISA. Firstly, in the light chain antibody detection system, the ratio of VH to VL is fixed and cannot be altered. By preparing MBP-VL separately, the sensitivity can be improved through optimized adjustment of the VH-to-VL ratio. Secondly, the fusion with MBP increased the expression level of VL, facilitating subsequent modifications and transformations. The use of MBP-VL not only augmented the quantity of VL but also, due to its higher molecular weight, exposed more VL-binding sites during the coating process, thereby enhancing the sensitivity of the assay.
Molecular docking was performed to simulate the binding position and interaction sites of the TS77 variable region with testosterone. As expected, testosterone forms hydrogen bonds with amino acids on both VH and VL. When VH and VL bind to testosterone, a stable trimer is formed among the three components, reinforcing the potential of this approach for highly sensitive and specific detection.
A comparison of OS-ELISA with previously published systems for testosterone detection is shown in Table 2. Compared to the traditional ic-ELISA method, OS-ELISA exhibits a broader detection range and higher sensitivity. It was reported that the reference range of serum total testosterone in male adults showed to be 2.01–7.50 ng/mL (6.97–26.00 nM) (Iwamoto et al., 2004). In this study, OS-ELISA based on anti L chain antibody achieved a LOD of 2.71 nM and an EC50 of 0.22 μM. Furthermore, the enhanced ELISA with a detection range from 1 pM to 10 mM utilizing MBP-VL achieved an even lower LOD of 1.07 pM, and an EC50 of 28.7 μM, significantly outperforming commercial ic-ELISA kits in terms of sensitivity. Additionally, OS-ELISA demonstrated high specificity when compared against two structurally similar hormones. This methodology has the potential to offer a novel approach for the prevention, rapid diagnosis, prognosis of testosterone-related diseases, as well as the monitoring of physical condition and doping detection in elite athletes.
In this study, we present the development of a novel non-competitive OS-ELISA for the sensitive and specific detection of testosterone. This innovative approach exploits the enhanced interaction between antigen-driven antibody VH and VL, offering a significant advancement in immunoassay technology. Initially, a foundational OS-ELISA platform was established using an expression mixture of VH and dissociated VL derived from phage display. Building on this, we further optimized the assay by incorporating VH and MBP-VL, which markedly improved both sensitivity and specificity. The resulting OS-ELISA demonstrates exceptional performance, positioning it as a highly reliable and practical analytical tool for testosterone detection. With its robust design and superior performance, this method holds great promise for applications in clinical monitoring and in vitro diagnostics, offering an alternative to conventional immunoassays.
The original contributions presented in the study are included in the article/supplementary material, further inquiries can be directed to the corresponding author.
GZ: Data curation, Formal Analysis, Investigation, Software, Writing–original draft, Writing–review and editing. SZ: Writing–review and editing. YZ: Writing–review and editing. NJ: Writing–review and editing. LZ: Methodology, Software, Writing–review and editing. JD: Funding acquisition, Methodology, Project administration, Resources, Writing–review and editing.
The author(s) declare that financial support was received for the research and/or publication of this article. This work was partly funded by the Shandong Provincial Natural Science Foundation, China (grant numbers: ZR2023MH021 and ZR2024QC347). This work was also supported in part by the World Research Hub (WRH) Program of the International Research Frontiers Initiative, Tokyo Institute of Technology, Japan.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Botelho, J. C., Shacklady, C., Cooper, H. C., Tai, S. S., Van Uytfanghe, K., Thienpont, L. M., et al. (2013). Isotope-dilution liquid chromatography-tandem mass spectrometry candidate reference method for total testosterone in human serum. Clin. Chem. 59 (2), 372–380. doi:10.1373/clinchem.2012.190934
Bulut, U., Sanli, S., Cevher, S. C., Cirpan, A., Donmez, S., and Timur, S. (2020). A biosensor platform based on amine functionalized conjugated benzenediamine-benzodithiophene polymer for testosterone analysis. J. Appl. Polym. Sci. 137 (43). doi:10.1002/app.49332
Cánovas, R., Daems, E., Campos, R., Schellinck, S., Madder, A., Martins, J. C., et al. (2022). Novel electrochemiluminescent assay for the aptamer-based detection of testosterone. Talanta 239, 123121. doi:10.1016/j.talanta.2021.123121
Chen, L. M., Tan, R. Y., Zhou, Y. M., Zhang, L. Q., Zhang, S. S., Li, X. Y., et al. (2020). Development of an Open sandwich ELISA for the detection of microcystin-LR. Microchem. J. 158, 105325. doi:10.1016/j.microc.2020.105325
Chen, Y., Teo, H. L., Liu, H., Loh, T. P., Liu, Q., Teo, T. L., et al. (2019). Simple and accurate candidate reference measurement procedure for total testosterone in human serum by one-step liquid-liquid extraction coupled with isotope dilution mass spectrometry. Anal. Bioanal. Chem. 411 (28), 7519–7528. doi:10.1007/s00216-019-02152-y
Ciumas, C., Linden Hirschberg, A., and Savic, I. (2009). High fetal testosterone and sexually dimorphic cerebral networks in females. Cereb. Cortex 19 (5), 1167–1174. doi:10.1093/cercor/bhn160
Cong, Y., Dong, H., Wei, X., Zhang, L., Bai, J., Wu, J., et al. (2019). A novel murine antibody and an open sandwich immunoassay for the detection of clenbuterol. Ecotoxicol. Environ. Saf. 182, 109473. doi:10.1016/j.ecoenv.2019.109473
Dong, J., Ihara, M., and Ueda, H. (2009). Antibody Fab display system that can perform open-sandwich ELISA. Anal. Biochem. 386 (1), 36–44. doi:10.1016/j.ab.2008.11.045
Dong, J., and Ueda, H. (2021). Recent advances in Quenchbody, a fluorescent immunosensor. Sensors (Basel) 21 (4), 1223. doi:10.3390/s21041223
Gonzalez-Techera, A., Kim, H. J., Gee, S. J., Last, J. A., Hammock, B. D., and Gonzalez-Sapienza, G. (2007). Polyclonal antibody-based noncompetitive immunoassay for small analytes developed with short peptide loops isolated from phage libraries. Anal. Chem. 79 (23), 9191–9196. doi:10.1021/ac7016713
Hahn, S., Kuehnel, W., Tan, S., Kramer, K., Schmidt, M., Roesler, S., et al. (2007). Diagnostic value of calculated testosterone indices in the assessment of polycystic ovary syndrome. Clin. Chem. Lab. Med. 45 (2), 202–207. doi:10.1515/CCLM.2007.031
Hameed, N., Yedinak, C. G., Brzana, J., Gultekin, S. H., Coppa, N. D., Dogan, A., et al. (2013). Remission rate after transsphenoidal surgery in patients with pathologically confirmed Cushing's disease, the role of cortisol, ACTH assessment and immediate reoperation: a large single center experience. Pituitary 16 (4), 452–458. doi:10.1007/s11102-012-0455-z
Hu, B., Li, J., Mou, L., Liu, Y., Deng, J., Qian, W., et al. (2017). An automated and portable microfluidic chemiluminescence immunoassay for quantitative detection of biomarkers. Lab. Chip 17 (13), 2225–2234. doi:10.1039/c7lc00249a
Ide, H. (2023). The impact of testosterone in men's health. Endocr. J. 70 (7), 655–662. doi:10.1507/endocrj.EJ22-0604
Islam, K. N., Ihara, M., Dong, J., Kasagi, N., Mori, T., and Ueda, H. (2010). Micro open-sandwich ELISA to rapidly evaluate thyroid hormone concentration from serum samples. Bioanalysis 2 (10), 1683–1687. doi:10.4155/bio.10.125
Iwamoto, T., Yanase, T., Koh, E., Horie, H., Baba, K., Namiki, M., et al. (2004). Reference ranges of total serum and free testosterone in Japanese male adults. Nihon Hinyokika Gakkai Zasshi 95 (6), 751–760. doi:10.5980/jpnjurol1989.95.751
Jiang, J., Wang, Z., Zhang, H., Zhang, X., Liu, X., and Wang, S. (2011). Monoclonal antibody-based ELISA and colloidal gold immunoassay for detecting 19-nortestosterone residue in animal tissues. J. Agric. Food Chem. 59 (18), 9763–9769. doi:10.1021/jf2012437
Li, G., Zhu, M., Ma, L., Yan, J., Lu, X., Shen, Y., et al. (2016). Generation of small single domain nanobody binders for sensitive detection of testosterone by electrochemical impedance spectroscopy. ACS Appl. Mater Interfaces 8 (22), 13830–13839. doi:10.1021/acsami.6b04658
Liang, Y., Wang, Y., Wang, F., Li, J., Wang, C., Dong, J., et al. (2021). An enhanced open sandwich immunoassay by molecular evolution for noncompetitive detection of Alternaria mycotoxin tenuazonic acid. Food Chem. 361, 130103. doi:10.1016/j.foodchem.2021.130103
Lim, S. L., Ichinose, H., Shinoda, T., and Ueda, H. (2007). Noncompetitive detection of low molecular weight peptides by open sandwich immunoassay. Anal. Chem. 79 (16), 6193–6200. doi:10.1021/ac070653z
Mizoguchi, M., Minami, S., Yamamoto, M., Tanizaki, Y., Kobayashi, A., and Ino, K. (2014). Ovarian steroid cell tumor, not otherwise specified, producing testosterone. J. Obstet. Gynaecol. Res. 40 (9), 2081–2085. doi:10.1111/jog.12467
Mulhall, J. P., Trost, L. W., Brannigan, R. E., Kurtz, E. G., Redmon, J. B., Chiles, K. A., et al. (2018). Evaluation and management of testosterone deficiency: AUA guideline. J. Urol. 200 (2), 423–432. doi:10.1016/j.juro.2018.03.115
Siekmann, L. (1979). Determination of steroid hormones by the use of isotope dilution--mass spectrometry: a definitive method in clinical chemistry. J. Steroid Biochem. 11 (1A), 117–123. doi:10.1016/0022-4731(79)90285-1
Suzuki, C., Ueda, H., Mahoney, W., and Nagamune, T. (2000). Open sandwich enzyme-linked immunosorbent assay for the quantitation of small haptens. Anal. Biochem. 286 (2), 238–246. doi:10.1006/abio.2000.4800
Tai, S. S., Xu, B., Welch, M. J., and Phinney, K. W. (2007). Development and evaluation of a candidate reference measurement procedure for the determination of testosterone in human serum using isotope dilution liquid chromatography/tandem mass spectrometry. Anal. Bioanal. Chem. 388 (5-6), 1087–1094. doi:10.1007/s00216-007-1355-3
Thienpont, L. M., De Brabandere, V. I., Stockl, D., and De Leenheer, A. P. (1994). Use of cyclodextrins for prepurification of progesterone and testosterone from human serum prior to determination with isotope dilution gas chromatography/mass spectrometry. Anal. Chem. 66 (22), 4116–4119. doi:10.1021/ac00094a041
Ueda, H., Tsumoto, K., Kubota, K., Suzuki, E., Nagamune, T., Nishimura, H., et al. (1996). Open sandwich ELISA: a novel immunoassay based on the interchain interaction of antibody variable region. Nat. Biotechnol. 14 (13), 1714–1718. doi:10.1038/nbt1296-1714
Valjakka, J., Hemminki, A., Niemi, S., Soderlund, H., Takkinen, K., and Rouvinen, J. (2002). Crystal structure of an in vitro affinity- and specificity-matured anti-testosterone Fab in complex with testosterone. J. Biol. Chem. 277 (46), 44021–44027. doi:10.1074/jbc.M208392200
Waifalkar, P. P., Noh, D., Derashri, P., Barage, S., and Oh, E. (2022). Role of estradiol hormone in human life and electrochemical aptasensing of 17β-estradiol: a review. Biosens. (Basel) 12 (12), 1117. doi:10.3390/bios12121117
Wang, H., Li, J., Zhang, X., Hu, B., Liu, Y., Zhang, L., et al. (2016). A microfluidic indirect competitive immunoassay for multiple and sensitive detection of testosterone in serum and urine. Analyst 141 (3), 815–819. doi:10.1039/c5an01835h
Wang, X., Huang, L., Zhao, D., Qiu, J., and Ma, Y. (2023). Research on the changes in catecholamine and other indicators during different training phases in female volleyball players. Altern. Ther. Health Med. 30, 192–201.
Wang, Y., Gay, G. D., Botelho, J. C., Caudill, S. P., and Vesper, H. W. (2014). Total testosterone quantitative measurement in serum by LC-MS/MS. Clin. Chim. Acta 436, 263–267. doi:10.1016/j.cca.2014.06.009
You, Y., Uboh, C. E., Soma, L. R., Guan, F., Li, X., Liu, Y., et al. (2011). Simultaneous separation and determination of 16 testosterone and nandrolone esters in equine plasma using ultra high performance liquid chromatography-tandem mass spectrometry for doping control. J. Chromatogr. A 1218 (26), 3982–3993. doi:10.1016/j.chroma.2011.04.087
Keywords: OS-ELISA, testosterone, phage display, rapid detection, antibody
Citation: Zhao G, Zhang S, Zou Y, Jia N, Zheng L and Dong J (2025) Detection of testosterone based on the interaction between variable regions of antibody heavy chain and light chain. Front. Bioeng. Biotechnol. 13:1499164. doi: 10.3389/fbioe.2025.1499164
Received: 20 September 2024; Accepted: 04 March 2025;
Published: 20 March 2025.
Edited by:
Wing Cheung Mak, The Chinese University of Hong Kong, ChinaReviewed by:
Wei Zhang, Swansea University, United KingdomCopyright © 2025 Zhao, Zhang, Zou, Jia, Zheng and Dong. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Jinhua Dong, amhkb25nQHVocnMuZWR1LmNu
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.