
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
BRIEF RESEARCH REPORT article
Front. Vet. Sci. , 05 February 2025
Sec. Veterinary Pharmacology and Toxicology
Volume 12 - 2025 | https://doi.org/10.3389/fvets.2025.1540132
 Zi Wang1,2,3†
Zi Wang1,2,3† Miao Sun1†
Miao Sun1† Yongqiang Wang2,4
Yongqiang Wang2,4 Jinchuan Shi1
Jinchuan Shi1 Wei Gao1
Wei Gao1 Dongxu Han1
Dongxu Han1 Fanjun Zeng1
Fanjun Zeng1 Sanren5
Sanren5 Liyin Du1,3*
Liyin Du1,3* Hongxia Ma6*
Hongxia Ma6* Kai Liu1,2,3*
Kai Liu1,2,3*Escherichia coli is a major pathogen responsible for calf diarrhea. However, it has developed resistance to many antimicrobial drugs for their inappropriate usage. The bacterial quorum sensing system transmits information between bacteria, it's important in regulating bacterial virulence, drug and acid resistance and so on. This system can found in Gram-negative bacteria and operates through acyl-homoserine lactone (AHL) signaling molecules. In this study, a type I quorum sensing AHL, N-Octanoyl-L-Homoserine lactone (C8), was added to E. coli growth medium to investigate its regulatory functions in drug resistance. After screening out the strains of E. coli that showed an obvious regulatory effect to the drug ofloxacin (OFX), transcriptomic sequencing was performed on the E. coli strains from the sub-inhibitory concentration group that concentration plus C8 group, and the control group. It shows that C8 significantly influenced resistance to OFX and the minimum inhibitory concentration of OFX in the tested strain was significantly increased. To Analyze transcriptome sequencing results identified 415 differentially expressed genes between the control and sub-inhibitory concentration groups, of which 201 were up-regulated and 214 were down. There were 125 differentially expressed genes between bacteria treated with a sub-inhibitory concentration of OFX and those treated with C8, of which 102 were up-regulated and 23 were down. Finally, It found that to adding the C8 significantly increased the resistance of tested bacteria to OFX. Data from transcriptome sequencing on differently expressed genes helps to explain how the type I quorum sensing system controls drug resistance in E. coli.
Escherichia coli is a widely distributed Gram-negative bacterium and is the most numerous facultative anaerobe inhabiting the intestines of warm-blooded animals (1, 2). Many E. coli strains are harmless or even beneficial to the host; however, some strains can be pathogenic to both humans and animals (3). At present, one of the more challenging diseases facing veterinarians and farmers today is colibacillosis, which is highly contagious and can cause severe diarrhea and septicemia in calves, with especially high morbidity and mortality rates in younger animals, resulting in significant financial loss to farmers and the cattle industry (4, 5).
It is common practice in the livestock industry to routinely administer large amounts of antimicrobials to prevent and treat infectious diseases, as well as to promote growth (6, 7). While antimicrobials are important in the prevention and treatment of bacterial infections, their misuse has led to increased drug resistance and the development of multidrug-resistant strains (8). This increased antimicrobial resistance (AMR) is becoming a concern for veterinarians and livestock producers (9). As previous studies show, cattle are an important source of AMR bacteria. The resistance in E. coli is especially concerning due to the risk it poses to human health via the food chain. E. coli strains are normally resident in the intestinal tract and bovine carriers are often asymptomatic. Consequently, the transfer of resistance from non-pathogenic to pathogenic strains can occur with ease in the same environment, resulting in the emergence of multidrug-resistant strains, as well as increasing the selection pressure for the clinical therapeutic use of antimicrobial drugs (10).
Bacterial cells can communicate with one another through the production and detection of signaling molecules, a mechanism known as quorum sensing (QS). This signaling is mediated by molecules known as autoinducers (AIs). Two types of bacterial cell-cell communication are recognized, namely, the AI-1 and AI-2 QS regulatory systems (11). The recent discovery of these microbial QS systems has provided new hope for investigating the regulatory mechanisms underlying the development of drug resistance and thus overcoming this resistance (12). It has been found that QS systems modulate various cellular processes in microorganisms, particularly in the regulation of drug efflux pumps and the formation of microbial biofilms (13, 14). The currently most studied QS pathways are mediated by acyl homoserine lactones (AHLs or AI-1) in Gram-negative bacteria (15). Bacteria produce and detect specific signaling molecules to coordinate their gene expression and physiological activities in relation to the bacterial cell density, promoting biofilm formation, the secretion of extracellular enzymes, synthesis of antimicrobial substance synthesis, competence, and virulence factor expression. This complex signaling network allows large populations of microbial cells to exhibit a multicellular behavioral pattern in response to changing environmental conditions (16–19).
This study aimed to investigate the impact of QS molecule C8 on the antibiotic resistance in E. coli strains that cause calf diarrhea to common antibiotics and to elucidate the underlying molecular mechanisms. In this study, OFX, enrofloxacin (ENR), doxycycline (DOX), florfenicol (FFC), and tetracycline (TET) were used to determine their minimum inhibitory concentration (MICs). In addition, 200 mg/L (1 × 10−3 mol/L) of the AHL, N-Octanoyl-L-Homoserine lactone (C8), a type I QS molecule, was added to the growth medium to identify drugs in which resistance was regulated by C8. The transcriptomes of three groups of bacteria, one treated with a sub-inhibitory drug concentration, another treated with a sub-inhibitory drug concentration plus C8, and a control group, were sequenced, and differentially expressed genes (DEGs) were identified. The expression of these genes was verified using RT-qPCR to determine the effects of C8 on drug-resistant E. coli causing diarrhea in calves. This study provides data supporting the research and development of new drugs and drug resistance inhibitors to counteract the development of bacterial resistance, thereby reducing the economic losses caused by E. coli diarrhea in calves.
Calf diarrhea E. coli was isolated from a large-scale beef cattle farm in Tongliao City and kept in the Laboratory of Preventive Veterinary Medicine, College of Animal Science and Technology, Inner Mongolia MINZU University.
The sensitivities of the bacterial isolate to five antimicrobial drugs, namely, OFX, ENR, DOX, FFC, and TET, were tested using the broth microdilution method, as described in the Clinical and Laboratory Standards Institute (CLSI) guidelines (20). Briefly, bacteria were grown in LB broth until reaching an OD600 of 0.5. Two hundred microliters of serial dilutions of the antimicrobial agents were placed in sterile 96-well polystyrene plates; the concentrations (wells 1–15) were 512, 256, 128, 64, 32, 16, 8, 4, 2, 1, 0.5, 0.25, 0.125, 0.0625, and 0.03125 μg/mL. Then, 100 μL of the bacterial suspension was added. The 16th well was used as a blank control (LB culture medium containing no antibiotics). Background blank and positive controls were included in each experiment. Experiments were performed with at least three independent replicates. The plates were incubated at 37°C for 24 h. The lowest concentration with no visible bacteria growth was considered the MIC (21, 22). The MIC of each antibiotic was classified as resistant (R), intermediate (I), or susceptible (S) based on CLSI breakpoints, or the National Antibiotic Resistance Monitoring System (NARMS) breakpoints for intestinal bacteria when the CLSI breakpoints were unavailable (23).
As described by Zhang et al. (24), 40 mg/L (2 × 10−4 mol/L), 60 mg/L (3 × 10−4 mol/L), 80 mg/L (4 × 10−4 mol/L), 100 mg/L (5 × 10−4 mol/L), and 200 mg/L (1 × 10−3 mol/L) of the AHL C8 were added for pre-experimentation, with a concentration of 200 mg/L (1 × 10−3 mol/L) ultimately selected for the subsequent experiments. After the addition of the antibiotics and bacterial suspension described in 2.2, 200 mg/L (1 × 10−3 mol/L) C8 was added, and the plates were incubated at 37°C for 24 h. The MIC results were observed.
Three groups of bacterial cells were established, namely, a group treated with a sub-inhibitory concentration of OFX, a second group treated with a sub-inhibitory concentration of OFX plus C8, and a control group. Specifically, Group 1 (TL-3) was treated with 16 μg/ml OFX, Group 2 (TL-4) was treated with 64 μg/ml OFX with the addition of 200 mg/L (1 × 10−3 mol/L) C8, and the control group (TL-2) had no OFX and C8. The bacteria were collected for total RNA extraction when they reached the exponential growth phase (OD600 = 0.5). Total RNA was extracted from the TL-2, TL-3, and TL-4 groups using a Bacteria RNA Extraction Kit (Tiangen Biochemical Technology (Beijing) Co., Ltd.), according to the manufacturer's instructions. The RNA samples were sent to Guangzhou Gene Denovo Biotechnology Co. for high-throughput sequencing (Illumina's Novaseq 6000 platform) and transcriptomic analysis. The raw reads were filtered to remove low-quality reads, including reads containing adapters, reads containing nucleotides with quality scores lower than 20, and reads with unknown nucleotides larger than 10%, before mapping to the ribosomal RNA (rRNA) database using Bowtie 2 version 2.2.8. The rRNA-mapped reads were removed, and the remaining reads were mapped to the reference genome. The reference genome sequence accession number was CP103295-CP103297. The prediction of the new transcripts was conducted using Rockhopper. Gene expression levels were normalized using the Fragments Per Kilobase of transcript per Million mapped reads (FPKM) method. The edgeR package (version 3.12.1; http://www.r-project.org) was used to identify DEGs across samples or groups using the criteria of a fold change ≥2 and a false discovery rate (FDR) < 0.05.
All DEGs were annotated using the Gene Ontology (GO) and the Kyoto Encyclopedia of Genes and Genomes (KEGG) databases. GO functional annotation was performed to describe the molecular functions, cellular components, and biological processes associated with the DEGs. KEGG enrichment analysis was used to identify the metabolic and signaling pathways associated with the DGEs, with significant enrichment determined by comparison to the annotation of reference transcripts (FDR ≤ 0.05). In addition, 9 DEGs (Supplementary Table 1) were selected for verification of their expression levels using quantitative reverse transcription-polymerase chain reaction (qRT-PCR). All qPCR reactions were performed in a total volume of 20 μL. The cycling parameters included an initial denaturation at 95°C for 30 s, followed by 40 cycles of 95°C for 10 s, and finally 60°C for 30 s. The 16S gene was used as an internal control and the changes in relative gene expression were calculated using with the 2−ΔΔCt method.
All sequences were deposited in the NCBI Sequence Read Archive and are publicly accessible under accession number PRJNA1163313. These files can be accessed via the following link: https://www.ncbi.nlm.nih.gov/bioproject/PRJNA1163313 (accessed on 20 September 2024).
Statistical analysis was performed using the GraphPad Prism 5 software package (Graph Software, San Diego, CA, USA). All data are expressed as the mean ± standard error based on three independent experiments. A p-value < 0.05 was considered statistically significant.
The MIC values of the five drugs, ENR, OFX, DOX, FFC, and TET, after treatment with 200 mg/L (1 × 10−3 mol/L) C8 are shown in Table 1. In the C8-treated group, the MIC values for OFX were increased from an initial 32–128 μg/ml, while those for ENR were increased from an initial 256–512 μg/ml, and the values for DOX, FFC, and TET remain unchanged.
The gene expression levels are calculated using the FPKM method. Due to the varying sequencing depths among different samples, the absolute expression levels of genes are amplified after the normalization of FPKM values. The intergroup comparison of gene expression levels is illustrated in the boxplot presented in Figure 1. Distribution plots of gene expression abundance are created by plotting the log10 (FPKM) values across different samples to compare the trends in expression changes (Figure 2). The expression density plots for each sample conform to a normal distribution, and the expression trends of biological replicates show consistency.
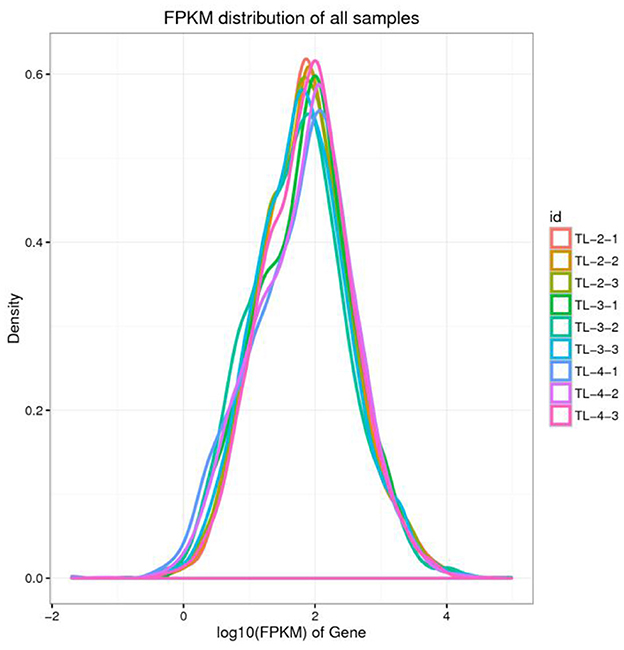
Figure 2. Density map of gene expression values of different samples. The abscissa is log10 (FPKM), and the ordinate is the density of genes.
Differentially expressed genes were identified in edgeR software using the criteria |log2FC| > 1 and FDR < 0.05. The TL-2 vs. TL-3 comparison identified 415 DGEs (P < 0.05), including 201 upregulated and 214 down-regulated genes. As shown in Table 2, the DGEs showing the most significant up-regulation were yiaG, gadA, rplW, yebG, cspI, and narG, while the most significantly down-regulated DGEs were yibA, gadE, argG, ydiE, artJ, htpG, and vapB. The TL-3 vs. TL-4 comparison identified 125 DGEs (P < 0.05), including 102 upregulated and 23 downregulated genes. As shown in Table 3, the most significantly up-regulated genes were dnaN, recF, recB, ycaQ, fimC, fimI, and fimA, while the most significantly down-regulated genes were yibA, gadE, argG, ydiE, artJ, htpG, and vapB. Volcano plots (Figure 3) and heatmaps (Figure 4) illustrate the identified DEGs.
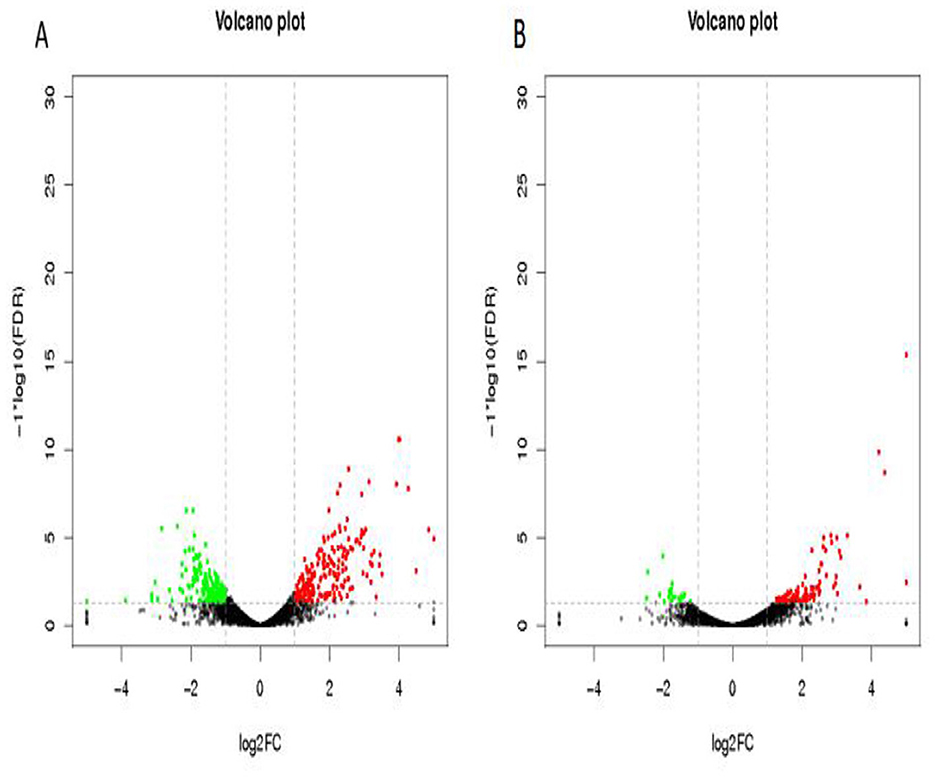
Figure 3. Volcano plots of differentially expressed genes. (A) TL-2 vs. TL-3 volcano plot, (B) TL-3 vs. TL-4 volcano plot. Black indicates genes with no significant changes, green indicates down-regulated genes, and red indicates up-regulated genes.
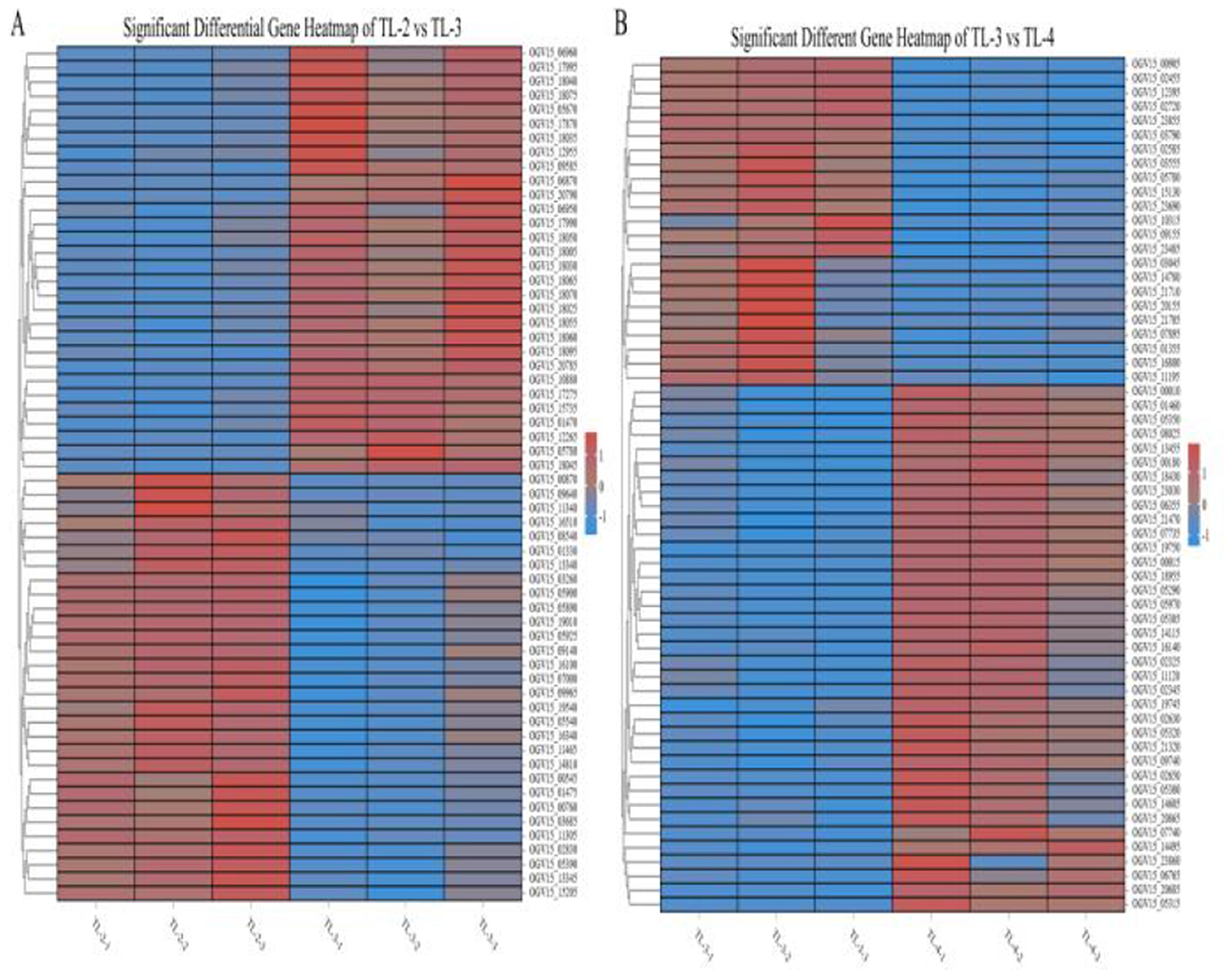
Figure 4. Heatmaps of partial differentially expressed genes. (A) Genes showing significant changes in the TL-2 vs. TL-3 comparison. (B) Genes showing significant changes in the TL-3 vs. TL-4 comparison.).
To compare the functions of the DEGs between the different groups, the up- and downregulated DGEs identified via edgeR were analyzed using GO, with a threshold value P < 0.05. The GO results were classified into cellular component (CC), molecular function (MF), and biological process (BP). The DEGs in the TL-2 vs. TL-3 comparison were classified into 13 MFs, 18 CCs, and 22 BPs, while those in the TL-3 vs. TL-4 comparison include 11 MFs, 15 CCs, and 19 BPs (Figures 5A, B). Categorizing the GO enrichment results in Level 2 GO terms showed that most of the DEGs were significantly enriched in metabolic, cellular, and single-organism processes in the BP category, cellular and cell membrane fractions in the CC category, and transporter activity, catalytic activity, and molecular binding in MFs.
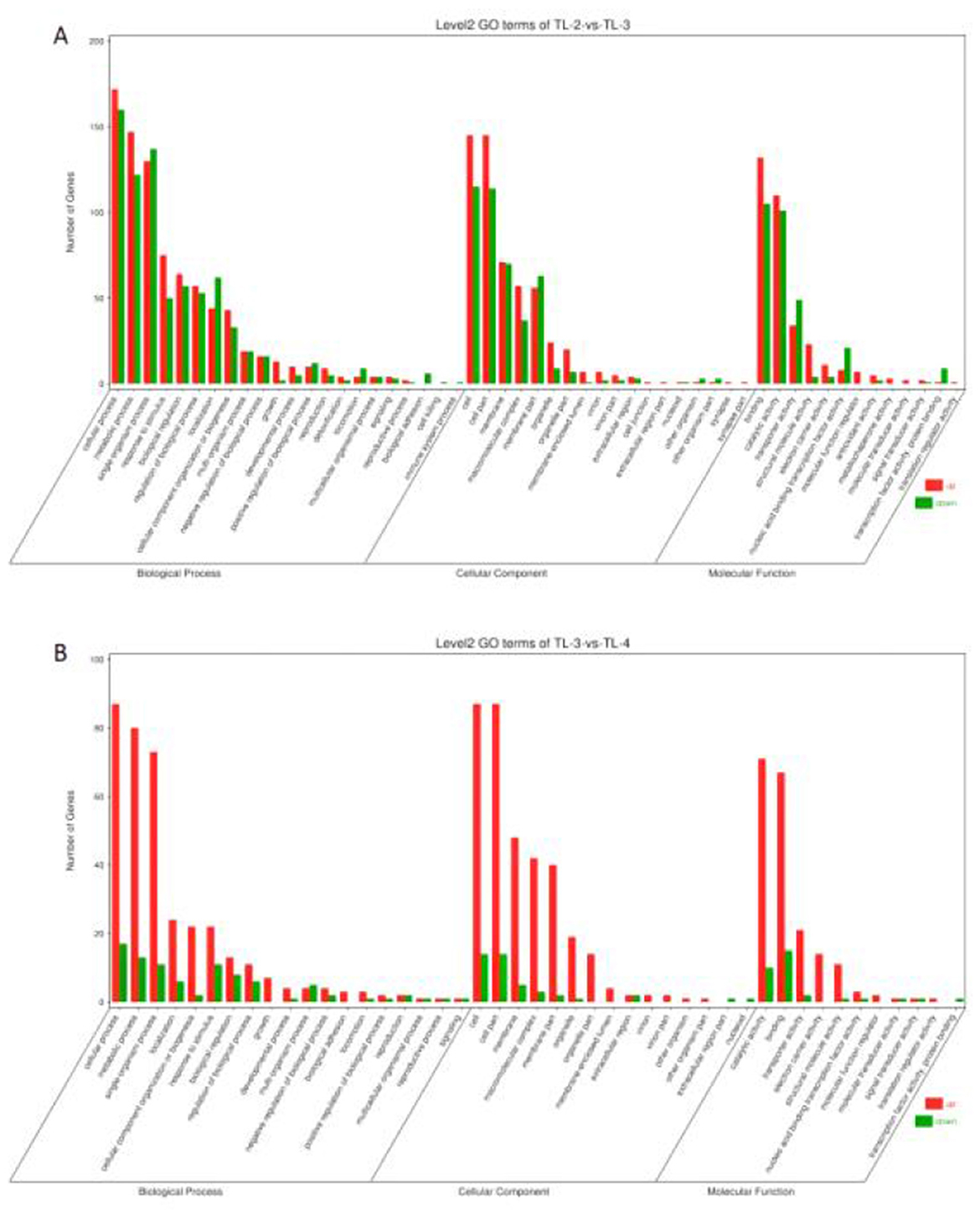
Figure 5. Gene Ontology enrichment analysis of differentially expressed genes. (A) TL-2 vs. TL-3 enrichment analysis. (B) TL-3 vs. TL-4 enrichment analysis.).
To investigate which pathways may affect drug resistance in E. coli, all DEGs were further mapped using the KEGG database. The results are shown in Figure 6, with higher RichFactors indicating higher degrees of enrichment. The number of DEGs in the pathways is indicated by the size of the dots, with larger the dots representing greater numbers; dot colors indicate the q-value corrected by multiple hypothesis testing, with values ranging from 0 to 1, and the smaller the value, the more significant the enrichment. The results showed that the DEGs were mainly enriched in pathways associated with arginine, ribosomes, nitrogen metabolism, biosynthesis, and proline and arginine metabolism.

Figure 6. KEGG enrichment analysis of differentially expressed genes. (A) TL-2 vs. TL-3 comparison. (B) TL-3 vs. TL-4 comparison.).
To verify the reliability of the transcriptomic results, 9 DEGs were randomly selected for RT-qPCR verification (ycaQ, nrdD, fimI, nrdE, gadE, rpsS, fimC, rpsJ, and maIF). Measurement of the expression levels of these nine genes showed that the expression was consistent with the transcriptome sequencing results (Figures 7A, B), indicating that the transcriptome sequencing results were reliable.
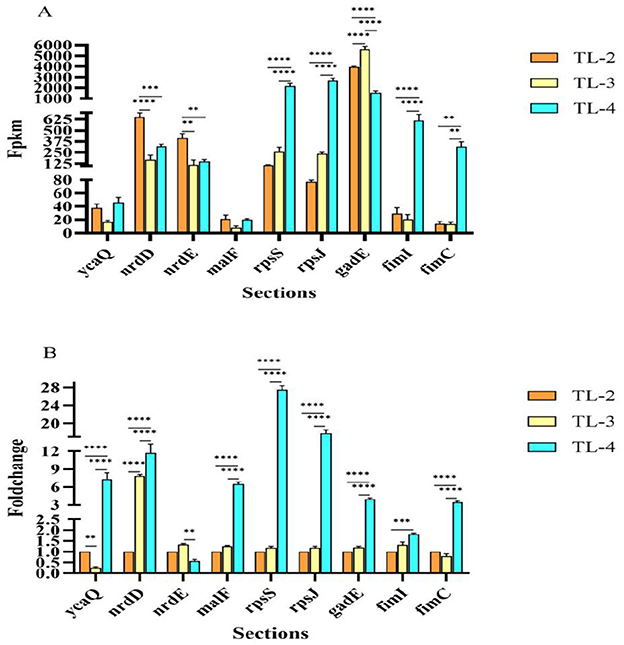
Figure 7. RT-qPCR verification of differentially expressed gene expression levels. (A) RT-qPCR verification of DGE FPKM values. (B) RT-qPCR verification of DGE fold changes). **, ***, ****represent significant differences at the 0.05 level, respectively.
Neonatal calf diarrhea (NCD) caused by primary or secondary infections of E. coli has become one of the major diseases in the global cattle industry (25, 26). At present, the treatment of E. coli mainly depends on antibiotics. However, due to the extensive use of antibiotics in clinical treatment, E. coli is prone to drug resistance. Quinolones are antimicrobial agents used to treat pathogenic E. coli infections. In recent years, multidrug-resistant strains of E. coli have appeared, and E. coli has caused huge economic losses to the cattle industry worldwide. Ibrahim et al. discovered that 80.5% of the 41 E. coli strains were resistant to quinolones (27). E. coli is regarded as a reservoir of antibiotic resistance determinants. This bacterium readily develops antibiotic resistance and is capable of transferring antibiotic resistance factors to other pathogenic microbes in gastrointestinal tracts (28).
The OFX exert their actions by inhibiting bacterial nucleic acid synthesis through disrupting the enzymes topoisomerase IV and DNA gyrase, and by causing breakage of bacterial chromosomes (29–31). In this study, we found that the expression of some genes involved in DNA synthesis, such as nrdD and nrdE, decreased significantly after the addition of sub-inhibitory concentration of OFX, which was consistent with the antibacterial mechanism of OFX (32). In addition, the expression of malF gene was also significantly decreased. The malF gene is mainly involved in the decomposition and synthesis of sugar-containing compounds and the glycosylation of antibiotics in organisms (33). Therefore, it is speculated that OFX can also control the synthesis of membrane structures such as bacterial cell wall, cell membrane and biofilm by regulating the expression of malF gene, thereby inhibiting bacterial growth.
There are extensive studies on the regulation of bacterial virulence, acid resistance and biofilm formation by type I QS. Zhang et al. studied the mechanism of type I QS signal molecules to enhance the acid resistance of enterohemorrhagic E. coli 0157: H7. It was confirmed that the QS signal molecule AHL regulates the expression of a variety of acid resistant related genes by regulating the ribosome regulatory factor RMF to enhance its acid resistance, also the rmf gene also affects the survival ability of E. coli under various stress conditions (34). Studies on the regulation of bacterial virulence-related genes by type I QS have confirmed that the signaling molecules can enhance the stability of bacteria after colonization, both through the enhancement of biofilm formation ability to resist bacterial resistance to environmental stresses and enhance bacterial adhesion to colonized tissues, as well as regulating the expression of other adhesion factors, such as flagellum, bacteriophage, and so on (35). The regulation of bacterial biofilm by type I QS system not only affects the secretion of virulence factors, but is also very important for bacterial drug resistance. At present, there are relatively few studies on the regulation of bacterial resistance by type I QS. In this study, the addition of AHL signaling molecule C8 found that it had a regulatory effect on the resistance of multiple drugs in the tested strains, of which the regulatory effect on OFX was the most obvious. Transcriptome sequencing was performed on each treatment group at sub-inhibitory concentration, and functional annotation analysis was performed after obtaining DEGs. It was found that there may be two ways for AHL signaling molecules to regulate the resistance of the tested strains to OFX:
Firstly, it is involved in the regulation of bacterial SOS response, after adding AHL signaling molecules, the expression of some key genes of SOS response changed significantly, such as recA, recN, recX, etc. In addition, the expression of ycaQ, a newly discovered gene encoding glycosylase, also increased significantly in recent years, and the ycaQ protein can cut the interstrand DNA crosslinks (36). Therefore, it is hypothesized that the AHL signaling molecule can regulate the expression of the ycaQ gene, which cleaves the cross-links between OFX and DNA, and works in conjunction with the SOS response to protect the bacterial DNA and thus increase resistance to OFX.
Secondly, it is involved in the regulation of the bacterial CpxAR two-component system. The CpxAR two-component system is an important regulatory system in E. coli, which can regulate the formation of bacterial biofilms, the expression of multidrug efflux pump genes and so on, so it has a regulatory effect on the resistance of E. coli. The CpxAR two-component system is mainly composed of CpxA, CpxR, and CpxP. CpxA is a histidine protein kinase located in the inner membrane of the cell, CpxR is a response protein located in the cytoplasm, and CpxP is a response regulatory protein located in the periplasm (37). In this study, we found that after the addition of AHL signaling molecules, the expression of CpxP genes differed significantly, and the fimbria adhesin genes fimA, fimC, and fimL were significantly up-regulated, and some of the ABC efflux pump-related genes and the acid-tolerant gene, gadE, were also differentially expressed. Fimbria adhesin are an important virulence factor and are required for biofilm formation, and deletion of the fim gene significantly reduces the ability of the bacterial biofilm to form (38). The gadE gene belongs to the regulatory protein of the LuxR family. Schwan et al. have shown that the gadE gene may indirectly inhibit the transcription of the fim gene, thereby affecting the formation of biofilm (39). In this study, we found that the expression of gadE gene decreased significantly after the addition of AHL signaling molecules, while at the same time the expression of fimbria adhesin genes fimA, fimC, and fimL was significantly up-regulated, which is in line with the results of Schwan WR et al. Based on the above research results, it is speculated that AHL signaling molecules can affect the resistance of the tested bacteria to OFX by regulating the CpxAR two-component system.
Biofilms are highly resistant to immune killing and removal, as well as antibiotic treatment. Because biofilms have certain physical and chemical effects that make the internal bacteria less susceptible to antibiotics and thus more resistant to antibiotics (40, 41). After regulating the expression of fimbriae adhesin genes, the ability of biofilm formation can be enhanced, and the drug resistance of bacteria in biofilm can be greatly improved. In addition, studies have confirmed that the type I QS system can affect the bacterial multidrug resistance (MDR) system, and the QS and MDR regulatory systems may share signal molecule regulatory pathways (42). The multidrug efflux pumps used in the MDR regulatory system can simultaneously efflux QS signals. The mechanism of these efflux pumps to output drug molecules is similar to that of QS signaling molecules. High concentrations of antibiotics can induce more efflux pump genes to be overexpressed, and these molecules are recognized by TetR family regulators, such as the classical ABC efflux pump system in E. coli. The QS signal may have antibacterial activity, and antibiotics also play the role of signal molecules. The antibiotic efflux pump is also the output of the QS signal. Therefore, there may be a certain evolutionary relationship and biological correlation between QS and MDR regulatory systems (43).
In this study, we found that the AHL signaling molecule C8 enhanced the drug resistance of the strains by adding 200 mg/L (1 × 10−3 mol/L) of AHL signaling molecule C8 to the subject strains, with a significant increase in resistance to OFX. The DEGs were obtained by transcriptome sequencing. Through the functional annotation analysis of the DEGs, it was speculated that there may be two ways for AHL signaling molecules to regulate the resistance of the tested strains to OFX: Firstly, it is involved in regulating the SOS response of bacteria, as well as regulating the expression of ycaQ, a gene encoding a glycosylase, to protect bacterial DNA and thereby increase resistance to OFX. Secondly, it is involved in the regulation of the bacterial CpxAR two-component system, which regulates the formation of bacterial biofilm and thus affects the resistance of the tested strain to OFX.
The datasets presented in this study can be found in online repositories. The names of the repository/repositories and accession number(s) can be found below: https://www.ncbi.nlm.nih.gov/, PRJNA1163313.
The animal study was approved by Laboratory Animal Ethics Committee of Inner Mongolia Minzu University College of Animal Science and Technology. The study was conducted in accordance with the local legislation and institutional requirements.
ZW: Writing – original draft, Writing – review & editing. MS: Writing – original draft, Writing – review & editing. YW: Data curation, Validation, Writing – review & editing. JS: Data curation, Formal analysis, Writing – review & editing. WG: Investigation, Writing – review & editing. DH: Formal analysis, Writing – review & editing. FZ: Software, Writing – review & editing. S: Supervision, Writing – review & editing. LD: Project administration, Supervision, Writing – review & editing. HM: Funding acquisition, Project administration, Supervision, Writing – review & editing. KL: Funding acquisition, Project administration, Supervision, Writing – review & editing.
The author(s) declare financial support was received for the research, authorship, and/or publication of this article. This study was supported by the National Natural Science Foundation of China (U23A20242, China), the Inner Mongolia Autonomous Region Colleges and Universities Basic Scientific Research Operating Expenses Program (GXKY22097, GXKY22082, Inner Mongolia, China), the Inner Mongolia Autonomous Region Central Guiding Local Development Fund Project (2022ZY0149, Inner Mongolia, China), the Inner Mongolia Xilin Gol League Science and Technology Project (202213, Inner Mongolia, China).
We would like to thank the Laboratory of Pharmacology and Toxicology, School of Animal Medicine, Jilin Agricultural University for help with the experiments.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Gen AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fvets.2025.1540132/full#supplementary-material
1. Umpiérrez A, Ernst D, Fernández M, Oliver M, Casaux ML, Caffarena RD, et al. Virulence genes of Escherichia coli in diarrheic and healthy calves. Rev Argent Microbiol. (2021) 53:34–8. doi: 10.1016/j.ram.2020.04.004
2. Arimizu Y, Kirino Y, Sato MP, Uno K, Sato T, Gotoh Y, et al. Large-scale genome analysis of bovine commensal Escherichia coli reveals that bovine-adapted E. coli lineages are serving as evolutionary sources of the emergence of human intestinal pathogenic strains. Genome Res. (2019) 29:1495–505. doi: 10.1101/gr.249268.119
3. Callaway TR, Elder RO, Keen JE, Anderson RC, Nisbet DJ. Forage feeding to reduce preharvest Escherichia coli populations in cattle, a review. J Dairy Sci. (2003) 86:852–60. doi: 10.3168/jds.S0022-0302(03)73668-6
4. Carter HSM, Renaud DL, Steele MA, Fischer-Tlustos AJ, Costa JHC. A narrative review on the unexplored potential of colostrum as a preventative treatment and therapy for diarrhea in neonatal dairy calves. Animals. (2021) 11:2221. doi: 10.3390/ani11082221
5. Kolenda R, Burdukiewicz M, Schierack P. A systematic review and meta-analysis of the epidemiology of pathogenic Escherichia coli of calves and the role of calves as reservoirs for human pathogenic E. coli. Front Cell Infect Microbiol. (2015) 5:23. doi: 10.3389/fcimb.2015.00023
6. Ueno Y, Suzuki K, Takamura Y, Hoshinoo K, Takamatsu D, Katsuda K. Antimicrobial resistance and associated genetic background of Histophilus somni isolated from clinically affected and healthy cattle. Front Vet Sci. (2022) 9:1040266. doi: 10.3389/fvets.2022.1040266
7. Jia Y, Mao W, Liu B, Zhang S, Cao J, Xu X. Study on the drug resistance and pathogenicity of Escherichia coli isolated from calf diarrhea and the distribution of virulence genes and antimicrobial resistance genes. Front Microbiol. (2022) 13:992111. doi: 10.3389/fmicb.2022.992111
8. Su J-H, Zhu Y-H, Ren T-Y, Guo L, Yang G-Y, Jiao L-G, et al. Distribution and antimicrobial resistance of Salmonella isolated from pigs with diarrhea in China. Microorganisms. (2018) 6:117. doi: 10.3390/microorganisms6040117
9. Morris C, Wickramasingha D, Abdelfattah EM, Pereira RV, Okello E, Maier G. Prevalence of antimicrobial resistance in fecal Escherichia coli and Enterococcus spp. isolates from beef cow-calf operations in northern California and associations with farm practices. Front Microbiol. (2023) 14:1086203. doi: 10.3389/fmicb.2023.1086203
10. Tabaran A, Soulageon V, Chirila F, Reget OL, Mihaiu M, Borzan M, et al. Pathogenic E. coli from cattle as a reservoir of resistance genes to various groups of antibiotics. Antibiotics. (2022) 11:404. doi: 10.3390/antibiotics11030404
11. Pacheco T, Gomes AÉI, Siqueira NMG, Assoni L, Darrieux M, Venter H, et al. SdiA, a quorum-sensing regulator, suppresses fimbriae expression, biofilm formation, and quorum-sensing signaling molecules production in Klebsiella pneumoniae. Front Microbiol. (2021) 12:597735. doi: 10.3389/fmicb.2021.597735
12. Zhao X, Yu Z, Ding T. Quorum-sensing regulation of antimicrobial resistance in bacteria. Microorganisms. (2020) 8:425. doi: 10.3390/microorganisms8030425
13. Wei C, Zhao X. Induction of viable but nonculturable Escherichia coli o157:h7 by low temperature and its resuscitation. Front Microbiol. (2018) 9:2728. doi: 10.3389/fmicb.2018.02728
14. Bäuerle T, Fischer A, Speck T, Bechinger C. Self-organization of active particles by quorum sensing rules. Nat Commun. (2018) 9:3232. doi: 10.1038/s41467-018-05675-7
15. Coquant G, Grill JP, Seksik P. Impact of N-acyl-homoserine lactones, quorum sensing molecules, on gut immunity. Front Immunol. (2020) 11:1827. doi: 10.3389/fimmu.2020.01827
16. Kwan JC, Meickle T, Ladwa D, Teplitski M, Paul V, Luesch H. Lyngbyoic acid, a “tagged” fatty acid from a marine cyanobacterium, disrupts quorum sensing in Pseudomonas aeruginosa. Mol Biosyst. (2011) 7:1205–16. doi: 10.1039/c0mb00180e
17. Chen L, Ku L, Li M. Editorial: frontiers in bacterial quorum sensing research. Front Cell Infect Microbiol. (2022) 12:999388. doi: 10.3389/fcimb.2022.999388
18. Huedo P, Coves X, Daura X, Gibert I, Yero D. Quorum sensing signaling and quenching in the multidrug-resistant pathogen Stenotrophomonas maltophilia. Front Cell Infect Microbiol. (2018) 8:122. doi: 10.3389/fcimb.2018.00122
19. Zeng X, Zou Y, Zheng J, Qiu S, Liu L, Wei C. Quorum sensing-mediated microbial interactions: mechanisms, applications, challenges and perspectives. Microbiol Res. (2023) 273:127414. doi: 10.1016/j.micres.2023.127414
20. CLSI. Performance Standards for Antimicrobial Disk and Dilution Susceptibility Tests for Bacteria Isolated from Animals. Wayne, PA: Clinical Laboratory Standard Institution [CLSI] (2015).
21. Zhang S, Shu Y, Wang Y, Zhong Z, Wang M, Jia R, et al. High rate of multidrug resistance and integrons in Escherichia coli isolates from diseased ducks in select regions of China. Poult Sci. (2023) 102:102956. doi: 10.1016/j.psj.2023.102956
22. Gao L, Ma X. Transcriptome analysis of Acinetobacter baumannii in rapid response to subinhibitory concentration of minocycline. Int J Environ Res Public Health. (2022) 19:16095. doi: 10.3390/ijerph192316095
23. Wang Z, Sun M, Guo S, Wang Y, Meng L, Shi J, et al. Detection of drug resistance in Escherichia coli from calves with diarrhea in the Tongliao region: an analysis of multidrug-resistant strains. Front Vet Sci. (2024) 11:1466690. doi: 10.3389/fvets.2024.1466690
24. Zhang Y, Ma Q, Su B, Chen R, Lin J, Lin Z, et al. A study on the role that quorum sensing play in antibiotic-resistant plasmid conjugative transfer in Escherichia coli. Ecotoxicology. (2018) 27:209–16. doi: 10.1007/s10646-017-1886-0
25. Wang D, Gao H, Zhao L, Lv C, Dou W, Zhang X, et al. Detection of the dominant pathogens in diarrheal calves of Ningxia, China in 2021-2022. Front Vet Sci. (2023) 10:1155061. doi: 10.3389/fvets.2023.1155061
26. Brunauer M, Roch FF, Conrady B. Prevalence of worldwide neonatal calf diarrhoea caused by Bovine Rotavirus in combination with Bovine Coronavirus, Escherichia coli K99 and Cryptosporidium spp.: a meta-analysis. Animals. (2021) 11:1014. doi: 10.3390/ani11041014
27. Ibrahim D, Awad A, Younis G. Prevalence and characterization of quinolone-resistant Escherichia coli isolated from retail raw beef and poultry meat in Egypt. J Adv Vet Anim Res. (2023) 10:490–99. doi: 10.5455/javar.2023.j702
28. Gu X, Ma X, Wu Q, Tao Q, Chai Y, Zhou X, et al. Isolation, identification, molecular typing, and drug resistance of Escherichia coli from infected cattle and sheep in Xinjiang, China. Vet Med Sci. (2023) 9:1359–68. doi: 10.1002/vms3.1101
29. Pham TDM, Ziora ZM, Blaskovich MAT. Quinolone antibiotics. Medchemcomm. (2019) 10:1719–39. doi: 10.1039/C9MD00120D
30. Yang F, Zhang S, Shang X, Wang L, Li H, Wang X. Characteristics of quinolone-resistant Escherichia coli isolated from bovine mastitis in China. J Dairy Sci. (2018) 101:6244–52. doi: 10.3168/jds.2017-14156
31. Bush NG, Diez-Santos I, Abbott LR, Maxwell A. Quinolones: mechanism, lethality and their contributions to antibiotic resistance. Molecules. (2020) 25:5662. doi: 10.3390/molecules25235662
32. Torrents E, Buist G, Liu A, Eliasson R, Kok J, Gibert I, et al. The anaerobic (class III) ribonucleotide reductase from Lactococcus lactis. Catalytic properties and allosteric regulation of the pure enzyme system. J Biol Chem. (2000) 275:2463–71. doi: 10.1074/jbc.275.4.2463
33. Yakovlieva L, Walvoort MTC. Processivity in bacterial glycosyltransferases. ACS Chem Biol. (2020) 15:3–16. doi: 10.1021/acschembio.9b00619
34. Zhang X, Li Z, Pang S, Jiang B, Yang Y, Duan Q, et al. The impact of cell structure, metabolism and group behavior for the survival of bacteria under stress conditions. Arch Microbiol. (2021) 203:431–41. doi: 10.1007/s00203-020-02050-3
35. Peña-Díaz J, Woodward SE, Creus-Cuadros A, Serapio-Palacios A, Ortiz-Jiménez S, Deng W, et al. Quorum sensing modulates bacterial virulence and colonization dynamics of the gastrointestinal pathogen Citrobacter rodentium. Gut Microbes. (2023) 15:2267189. doi: 10.1080/19490976.2023.2267189
36. Bradley NP, Washburn LA, Christov PP, Watanabe CMH, Eichman BF. Escherichia coli YcaQ is a DNA glycosylase that unhooks DNA interstrand crosslinks. Nucleic Acids Res. (2020) 48:7005–17. doi: 10.1093/nar/gkaa346
37. Tierney AR, Rather PN. Roles of two-component regulatory systems in antibiotic resistance. Future Microbiol. (2019) 14:533–52. doi: 10.2217/fmb-2019-0002
38. Berne C, Ellison CK, Ducret A, Brun YV. Bacterial adhesion at the single-cell level. Nat Rev Microbiol. (2018) 16:616–27. doi: 10.1038/s41579-018-0057-5
39. Schwan WR, Luedtke J, Engelbrecht K, Mollinger J, Wheaton A, Foster JW, et al. Regulation of Escherichia coli fim gene transcription by GadE and other acid tolerance gene products. Microbiology. (2022) 168:001149. doi: 10.1099/mic.0.001149
40. Davies D. Understanding biofilm resistance to antibacterial agents. Nat Rev Drug Discov. (2003) 2:114–22. doi: 10.1038/nrd1008
41. Mah TF, O'Toole GA. Mechanisms of biofilm resistance to antimicrobial agents. Trends Microbiol. (2001) 9:34–9. doi: 10.1016/S0966-842X(00)01913-2
42. Yang S, Lopez CR, Zechiedrich EL. Quorum sensing and multidrug transporters in Escherichia coli. Proc Natl Acad Sci USA. (2006) 103:2386–91. doi: 10.1073/pnas.0502890102
Keywords: Escherichia coli, drug resistance, quorum-sensing molecule, transcriptomic sequencing, differentially expressed genes
Citation: Wang Z, Sun M, Wang Y, Shi J, Gao W, Han D, Zeng F, Sanren, Du L, Ma H and Liu K (2025) Regulation of ofloxacin resistance in Escherichia coli strains causing calf diarrhea by quorum-sensing acyl-homoserine lactone signaling molecules. Front. Vet. Sci. 12:1540132. doi: 10.3389/fvets.2025.1540132
Received: 05 December 2024; Accepted: 07 January 2025;
Published: 05 February 2025.
Edited by:
Zhigang Zhang, Northeast Agricultural University, ChinaReviewed by:
Wei Yang, Heilongjiang Bayi Agricultural University, ChinaCopyright © 2025 Wang, Sun, Wang, Shi, Gao, Han, Zeng, Sanren, Du, Ma and Liu. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Liyin Du, ZHVsaXlpbjIwMTZAMTYzLmNvbQ==; Hongxia Ma, bWFob25neGlhQGpsYXUuZWR1LmNu; Kai Liu, bGl1a2FpNzIxMDI2QDE2My5jb20=
†These authors have contributed equally to this work and share first authorship
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.