
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Vet. Sci. , 10 March 2025
Sec. Veterinary Epidemiology and Economics
Volume 12 - 2025 | https://doi.org/10.3389/fvets.2025.1527279
 Tefera Woldemariam1†
Tefera Woldemariam1† Temesgen Mohammed2,3*†
Temesgen Mohammed2,3*† Aboma Zewude2,3
Aboma Zewude2,3 Mahlet Chanyalew3
Mahlet Chanyalew3 Hazim O. Khalifa2,4
Hazim O. Khalifa2,4 Gezahegne Mamo5
Gezahegne Mamo5 Gobena Ameni2*
Gobena Ameni2*Introduction: The zoonotic transmission of tuberculosis (TB) from cattle to humans has long been recognized, while the reverse transmission from humans to animals has only recently been reported. The socioeconomic conditions in rural Ethiopia are conducive to the zoonotic and reverse zoonotic transmission of TB between cattle and humans. This study aimed to explore the transmission of the Mycobacterium tuberculosis complex between cattle and humans in Central Ethiopia.
Methods: To achieve this objective, a cross-sectional study was conducted involving 1,896 cattle slaughtered at two abattoirs and 392 human subjects suspected of tuberculosis who visited health institutions for treatment. Mycobacteriological culture and spoligotyping were used for the study. Additionally, the Spoligotype International Types and VNTR (variable number of DNA tandem repeats) International Types (SITVIT2) database and the online tool “Run TB-Lineage” were used to identify SIT and lineages of the isolates from both humans and cattle.
Results: Culture positivity was found in 26.3% (21/80) of the gross TB-suspicious tissue lesions in the lungs and lymph nodes (mandibular, retropharyngeal, cranial, and caudal mediastinal, as well as left and right bronchial, hepatic, and mesenteric lymph nodes) of cattle. Of the 21 cattle isolates, 12 (57.2%) were identified as M. bovis, while the remaining 9 (42.8%) were classified as M. tuberculosis. Similarly, only 22% (86/392) of the sputum samples from TB-suspicious humans were culture positive. These 86 human isolates included 81 M. tuberculosis, three M. africanum, and two M. bovis, as determined by spoligotyping. SIT50, SIT118, and SIT1318, which belong to the human species, were isolated from both humans and cattle. The two human M. bovis isolates exhibited the pattern of SB1443, which was not identified in cattle within this study area.
Conclusion: The zoonotic and reverse zoonotic transmissions of TB were confirmed in Ethiopia by isolating two M. bovis from humans and nine M. tuberculosis from cattle, which suggested a greater role for M. tuberculosis in cattle compared to M. bovis in humans.
Bovine tuberculosis (bTB) is a chronic disease affecting animals, characterized by the progressive development of granulomatous lesions or tubercles in the lymph nodes, lung tissue, and other organs. This disease is primarily caused by Mycobacterium bovis, a member of the Mycobacterium tuberculosis (M. tuberculosis) complex, and to a lesser extent by M. tuberculosis. The disease leads to significant economic losses for the agriculture sector and poses public health challenges by causing zoonotic TB in humans. According to the World Organization for Animal Health, 82 (44%) of the 188 countries and territories reported the presence of bTB between January 2017 and June 2018 (1). Thus, this disease occurs worldwide, although most developed countries have reduced or eliminated it from their cattle populations, except in limited areas where wild animals, as reservoirs of the infection, interfere with control programs. Conversely, the disease remains prevalent in many African countries and parts of Asian countries (1–4). The prevalence of bTB in animal populations in developing countries can be attributed to insufficient or ineffective control measures that address the socioeconomic issues related to the current test-and-slaughter approach (3).
The direct correlation between M. bovis infection in cattle and zoonotic TB in the human population has been well documented in many industrialized countries (3). Zoonotic TB, caused by M. bovis, is transmitted from animals to humans through the consumption of contaminated foods and the inhalation of cough droplets from infected animals during close physical contact. It also poses an occupational hazard for farmers, slaughterhouse workers, butchers, and veterinary professionals.
Epidemiological studies of TB in cattle and humans in Zambia (5) and Ethiopia (6) indicated associations between tuberculin-positive cattle and human TB. In Zambia, it was observed that households reporting a TB case within the last year were seven times more likely to own herds containing tuberculin-positive cattle. Similarly, tuberculin positivity was three times higher in cattle owned by farmers with active TB than in those owned by farmers without active TB in Ethiopia. These observations suggest potential transmission of mycobacteria between humans and cattle, which could involve the transmission of M. bovis from cattle to humans or the transmission of M. tuberculosis from humans to cattle. A few studies in Ethiopia reported the isolation of M. bovis from sputum and fine needle aspirates of human TB cases (7, 8). According to a previously published study (9), in countries where bTB is still common and pasteurization of milk is not practiced, an estimated 10–15% of human TB is caused by M. bovis, with infections believed to occur through drinking raw milk (10) and through inhalation in rural areas among people living in close contact with cattle (11).
The limited number of studies conducted in Ethiopia lacks extensive geographic coverage and representativeness. Geographic regions such as the East Shewa Zone of central Ethiopia, where dairy cattle and beef farming are commonly practiced, have not been considered. The East Shewa Zone is at risk for mycobacteria transmission between cattle and humans. Numerous butchers in the area slaughter cattle from beef farms and nearby rural areas to sell raw meat, making it well-known for this business. In addition to beef cattle, farmers in the surrounding towns and rural districts also keep dairy cows. A significant percentage of raw milk is supplied to consumers in the area. These common practices of consuming raw milk and raw meat could facilitate the transmission of M. bovis from cattle to humans. Furthermore, Ethiopia ranks seventh among the 20 countries with the highest burden of human TB, with an incidence rate of 165 cases per 100,000 population (12).
East Shewa Zone is one of the most densely populated areas of Ethiopia, characterized by a high rate of human mobility, as the main road to the Port of Djibouti passes through the Zone. Therefore, there are favorable conditions for the transmission of human TB, and there is a significant risk of M. tuberculosis transmission from humans to cattle due to close physical contact between them and their cattle. It is essential to investigate whether transmission of TB occurs between cattle and humans, given these conducive epidemiological conditions. The present study was conducted to examine the zoonotic transmission of the Mycobacterium tuberculosis complex between cattle and humans in central Ethiopia.
The study was conducted from 2018 to August 2020 in the towns of Adama and Bishoftu, located within the East Shewa Zone of Oromia Regional State in Central Ethiopia. Bishoftu and Adama are situated 47 km and 95 km southeast of Addis Ababa, respectively. These two towns are known for their beef production and the sale of beef meat. Cattle from the beef farms and rural areas surrounding these towns are slaughtered in local abattoirs, and raw meat consumption is quite common in the region. In addition to beef cattle, dairy cows are raised by farmers living in these towns and the surrounding rural districts. Consequently, a significant percentage of raw milk is supplied to consumers in these towns and nearby districts. Unfortunately, BTB is prevalent in the country’s dairy farms, which are not expected to be free from the disease.
A cross-sectional study design was used to evaluate the transmission of M. tuberculosis complex species between cattle and humans in Central Ethiopia.
Sample size determination was based on the availability of funds and logistical support for the project’s research activities. Since the main objective of this research was to evaluate the transmission of M. tuberculosis between cattle and humans, efforts were made to collect the maximum number of samples by examining a large number of cattle and gathering a significant number of sputum samples. No statistical calculations were used to estimate the sample size in either the human or cattle studies. Thus, the available funds and logistics facilitated the examination of 1,896 cattle at two abattoirs (1,266 at Adama and 630 at Bishoftu). Additionally, 392 sputum samples (274 from Adama and 118 from Bishoftu) were collected from the two towns.
The study subjects comprised cattle slaughtered at these two abattoirs and human TB patients who visited healthcare centers in the two towns during the study period. A postmortem examination for BTB was conducted on cattle from beef farms located in either of the two towns or the surrounding districts. Similarly, sputum samples from individuals living in these towns or the surrounding districts with connections to either dairy or beef cattle were used for bacteriological and molecular analysis.
The primary outcome was the isolation of the same species of M. tuberculosis complex from both cattle and humans, suggesting possible TB transmission between the two. Other secondary outcomes included the percentage of cattle with TB lesions in their tissues and the percentage of suspicious TB lesions that tested positive via culture and spoligotyping. In the human study, the secondary outcome focused on the percentage of presumptive patients who tested positive through culture and spoligotyping. Diagnostic criteria for the cattle study included gross BTB lesions confirmed by culture and spoligotyping. For the human study, the diagnostic criteria included clinical examinations such as persistent coughing for more than 2 weeks, chest pain, hemoptysis, night sweats, and weight loss, followed by confirmation through sputum culturing and spoligotyping of the cultured isolates. Individuals showing signs and symptoms of TB, confirmed by sputum culturing, were treated with anti-TB drugs. The final confirmation of probable transmission of M. tuberculosis complex species between cattle and humans was achieved through spoligotyping.
The primary objective of the study was to confirm the transmission of the Mycobacterium tuberculosis complex between cattle and humans. Focus was placed on searching for BTB lesions in the tissues of cattle, followed by the collection of suspicious tissue lesions. The tissues with detected suspicious lesions were recorded. BTB suspicious tissue samples were collected in universal bottles containing normal saline and transported under cold chain conditions to the TB laboratory at the Aklilu Lemma Institute of Pathobiology, Addis Ababa University.
A total of 1,896 cattle were examined at two abattoirs, from which 80 BTB suspicious tissue lesions were collected from the lungs and lymph nodes (mandibular, retropharyngeal, cranial, and caudal mediastinal, left and right bronchial, hepatic, and mesenteric lymph nodes) for culturing. The number of animals recruited was determined by logistical availability and the convenience of the slaughterhouses. Human sample collection was conducted as part of routine diagnostic procedures for TB. Sputum samples were submitted to hospitals and health centers by 392 TB-suspicious cases. Consequently, 274 sputum samples were collected from Adama Town, while 118 sputum samples were collected from Bishoftu Town. The sputum samples were transported under cold chain conditions to the TB laboratory at the Aklilu Lemma Institute of Pathobiology, Addis Ababa University, for culturing.
A postmortem examination was conducted following the procedure described by (13). Each of the seven lobes of the lungs was thoroughly inspected and palpated for any suspicious gross TB lesions. Similarly, the mandibular, retropharyngeal, cranial and caudal mediastinal, left and right bronchial, hepatic, and mesenteric lymph nodes were sliced into 2 mm sections and subsequently examined for the presence of visible lesions, as per the protocol described earlier (14, 15). Tissues showing macroscopic lesions compatible with bTB were collected in a universal bottle with a 0.9% saline solution and transported to the laboratory under cold chain conditions. A total of 80 suspicious lesions were collected from the 1,896 cattle investigated and processed for culturing, as detailed below.
Samples from suspected TB lesions were further processed to isolate mycobacteria according to the protocols of the World Organization for Animal Health (16). Tissue specimens for culture were collected in sterile universal bottles containing 5 mL of a 0.9% saline solution. They were then transported in an icebox at a stable temperature of 4°C to the Aklilu Lemma Institute of Pathobiology. In the laboratory, the specimens were sectioned with sterile blades and homogenized using a mortar and pestle. The homogenate was decontaminated by adding an equal volume of 4% NaOH and centrifuging at 1,865 g for 15 min. The supernatant was discarded, and the sediment was neutralized with 1% (0.1 N) HCl, using phenol red as an indicator. Neutralization was considered to have been achieved when the solution’s color changed from purple to yellow. Subsequently, 0.1 mL of suspension from each sample was spread over a slant of Lowenstein-Jensen medium. Duplicate slants were used, one enriched with sodium pyruvate and the other with glycerol. Cultures were incubated aerobically at 37°C for at least eight weeks, with weekly observations of colony growth.
Sputum samples were collected and processed for culture following the WHO guidelines (17). Briefly, an equal volume of 4% NaOH was mixed with the sputum sample, and the mixture was centrifuged at 3,000 rpm for 15 min at room temperature. After decanting the supernatant, the sediment was neutralized with 2 N HCl using phenol red as an indicator. Neutralization was achieved when the solution changed color from purple to yellow. Thereafter, 100 μL of the suspension was inoculated onto two sterile LJ medium slopes, which were enriched with either pyruvate or glycerol. The inoculated media were then incubated at 37°C in a slanted position for one week and in an upright position for four to five weeks. Bacterial growth was monitored weekly until the 8th week of culture.
The colonies were harvested from the surface of the LJ medium and re-suspended in 200 μL of sterile double-distilled water. Subsequently, the colonies and water were mixed thoroughly. The mixture was then heated to 80°C for 50 min in a water bath to release mycobacterial DNA, which was used for subsequent spoligotype analysis.
Spoligotyping was performed on 107 M. tuberculosis complex isolates at the Aklilu Lemma Institute of Pathobiology, Addis Ababa University, following the standard operating procedure that was used by (18) and primarily developed by (19). The DNA released by heat-killing of the colonies was used as a template to amplify the direct repeat (DR) region of M. tuberculosis complex by polymerase chain reaction (PCR) using oligonucleotide primers derived from the DR sequence, RDa (5’GGTTTTGGGTTTGAACGAC3’) and RDb (5’CCGAGAGGGGACG GAAAC3’) primers (19). The total volume of the PCR reaction mixture was 25 μL, consisting of 12.5 μL of HotStarTaq Master Mix (Qiagen; providing a final concentration of 1.5 mM MgCl2 and 200 mM of each deoxyribonucleotide triphosphate), 2 μL of each primer (20 pmol each), 5 μL of a suspension of DNA template (approximately 10–50 ng), and 3.5 μL of Qiagen water. The mixture was heated for 15 min at 96°C and then subjected to 30 cycles of 1 min at 96°C, 1 min at 55°C, and 30 s at 72°C, followed by a final extension at 72°C for 10 min.
Immediately before running the spoligotyping, the PCR product was denatured in a thermocycler at 96°C for 10 min and then removed from the thermocycler and placed on ice to prevent further denaturation of the PCR products. The denatured PCR product was subsequently loaded onto a membrane covalently bonded with a set of 43 oligonucleotides, each corresponding to a unique spacer DNA sequence within the DR locus of the M. tuberculosis complex, and hybridized at 60oc for 1 h. After hybridization, the membrane was washed twice for 10 min in 2x SSPE (1x SSPE consists of 0. 18 M NaCl, 10 mM NaH2 PA4, and 1 mM EDTA [pH 7. 7]) with 0.5% sodium dodecyl sulfate at 60oC and then incubated in a 1: 4000 dilution of streptavidin peroxidase (Boehringer) for 1 h at 42°C. The membrane underwent two additional washes for 10 min in 2 x SSPE with 0.5% sodium dodecyl sulfate at 42°C and was rinsed with 2 x SSPE for 5 min at room temperature. Hybridized DNA was detected using the enhanced chemiluminescence method (Amersham, Biosciences, Amersham, UK) and exposed to X-ray film (Hyperfilm ECL, Amersham). A mixture of 10 mL of ECL reagent 1 and 10 mL of ECL reagent 2 was prepared and added to the membrane, which was rinsed in the solution for 5 min at room temperature. The membrane was then attached to a film in the darkroom, placed in a cassette, and incubated for 15 min at room temperature. After incubation, the film was removed and placed in a developer solution for 2 min. The film was subsequently rinsed with tap water for 15 s and placed in a fixer solution for 1 min. Finally, the film was dried and used for result interpretation. The presence of a spacer was indicated as a black square, while its absence was indicated as a white square on the film. Subsequently, the black squares were converted to 1, while the white squares were converted to 0 and transferred to the Spoligotype International Types-VNTR International Types (SITVIT) database for identifying the Spoligotype International Types (SIT) and the lineages of the isolates.
Proportions were used to indicate the number of cattle with TB lesions. Tissue lesions and sputum culture positivity were presented as proportions. In the molecular aspect of the study, SIT numbers and lineages of isolates were identified using the SITVIT1 Database and Run TB-Lineage. The results of spoligotyping were converted into octal and binary formats and then submitted into the query box, allowing the retrieval of the strain names from the database if the spoligotype pattern of the strain matched one already registered in the SPolDB4 database, accessible at http://www.pasteur-guadeloupe.fr:8081/SITVITDemo/ (SITVIT1 Database) (20). If the strain’s pattern had not been previously registered in the database, it was classified as an orphan. The lineages were also generated by entering binary and octal formats into the query box of the SITVIT1 Database and Run TB-Lineage.
Gross TB lesions were detected in 80 out of 1,896 (4.22%) of the cattle slaughtered at the Bishoftu and Adama abattoirs. These lesions were observed in various tissues of the lungs and lymph nodes, including the mandibular, retropharyngeal, cranial, and caudal mediastinal, left and right bronchial, hepatic, and mesenteric lymph nodes. They were more frequent and severe in the lungs and thoracic lymph nodes, followed by those in the head and neck. Accordingly, about 21 out of 80 (26.3%) of the suspected tissue lesions tested positive for mycobacterial growth.
Table 1 shows selected demographic characteristics of human TB-suspicious cases, which were used as a source of sputum from the two towns. The sputum culture positivity rate was 21.9% (86/392). Fifty culture-positive samples originated from Adama, while 36 were from Bishoftu. The culture positivity rate for samples collected in Adama Town was 18.2% (50/274), whereas it was 30.5% (36/118) for those collected in Bishoftu Town.

Table 1. Selected demographic characteristics of the study participant’s and their association with culture positivity.
Spoligotyping of 21 isolates from cattle yielded 18 different spoligotype patterns. Of the 21 isolates, 12 (57.2%) were confirmed to be M. bovis, while the remaining nine (42.8%) were M. tuberculosis. The isolates were grouped into 3 clustered strains, each containing 2 isolates (28.63%), and 15 (71.4%) single isolates (Table 2). Out of the 18 spoligotype patterns, 6 were registered in the international database, while the remaining 12 were not found in it. The 12 strains of M. bovis isolated from cattle were represented by 9 different spoligotype patterns. Of these 9 clusters of M. bovis, 7 were new and had not been previously reported in the M. bovis spoligotype database, while the remaining two belonged to spoligotypes SB1265 (two isolates) and SB1176 in the database M.bovis.org. The nine strains of M. tuberculosis isolated from cattle represented 9 different spoligotype patterns. Of these nine clusters, five clusters were not registered in the international spoligotype SITVIT2 database and were thus designated as orphan strains, while the remaining four clusters corresponded to spoligotypes SIT 50, 118, 117, and 1,318 in the SITVIT2 database, with the majority belonging to the Euro-American lineage.
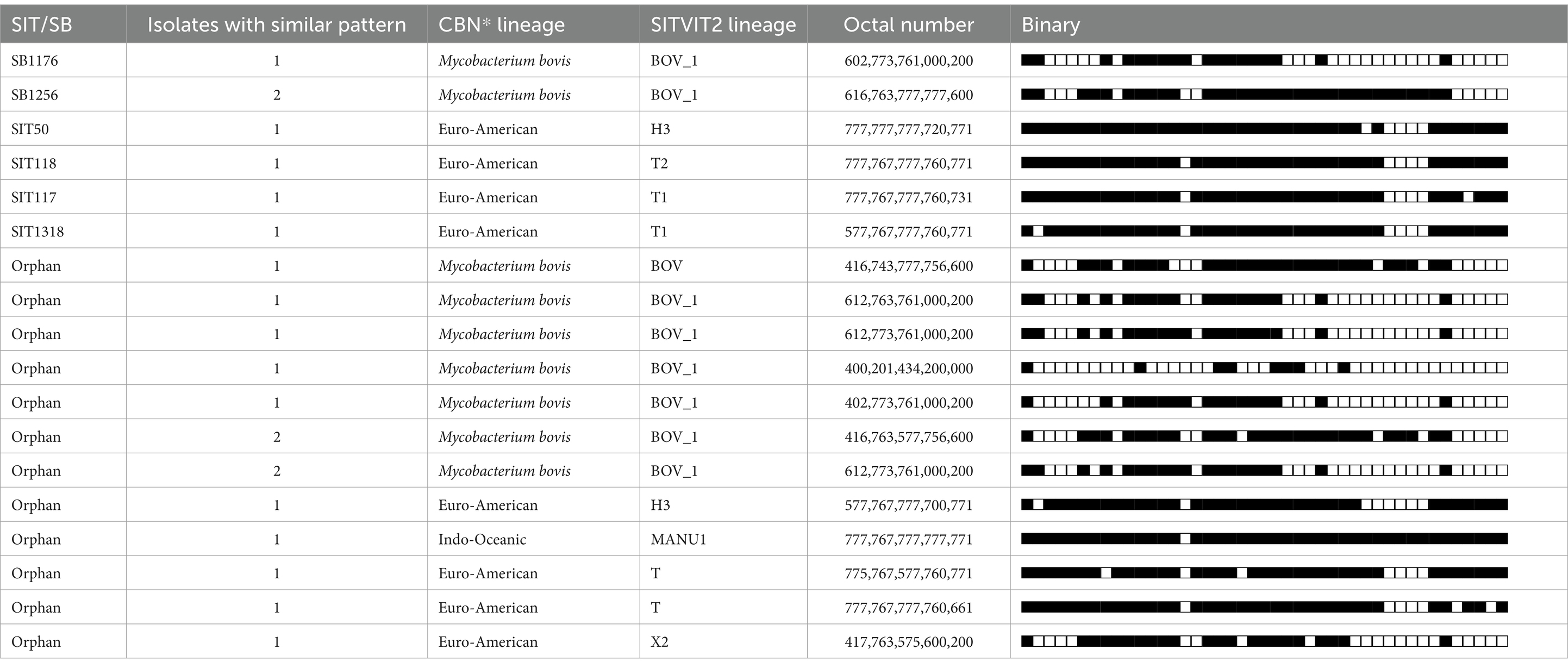
Table 2. Spoligotype patterns of M. bacterium tuberculosis complex species isolated from cattle in Central Ethiopia.
Spoligotyping of 86 isolates from humans yielded 35 different spoligotype patterns. The isolates were grouped into 14 clustered strains, comprising 65 (75.6%) isolates and 21 (24.4%) single strains (Table 3). The overall diversity among the isolates was 40.7%. Out of the 35 spoligotype patterns, 27 were recorded in the international database, while the remaining eight were not found in the database. The most frequently identified strains were SIT 149, SIT 53, and SIT 118, representing 18, 11, and 6 isolates, respectively, and belonging to the M. tuberculosis species. Classification of the spoligotype patterns using TB-insight RUN TB-Lineage showed that the Euro-American (E-A) lineages dominated, comprising 87.2%, while 3.5% of the isolates were classified as Central-Asian (CAS) lineages, another 3.5% as M. africanum lineages, and 1.2% as Indo-oceanic lineages. Two M. bovis isolates (2.3%) were identified by their characteristic spoligotype features, which include missing spacers 3, 9, 16, and 39–43. Among the unregistered patterns, four strains were attributed to the E-A lineage, three to M. africanum, one to CAS, and one to the Indo-oceanic lineage. The two M. bovis strains isolated from sputum had spoligotypes SB01443 in the M.bovis.org database.
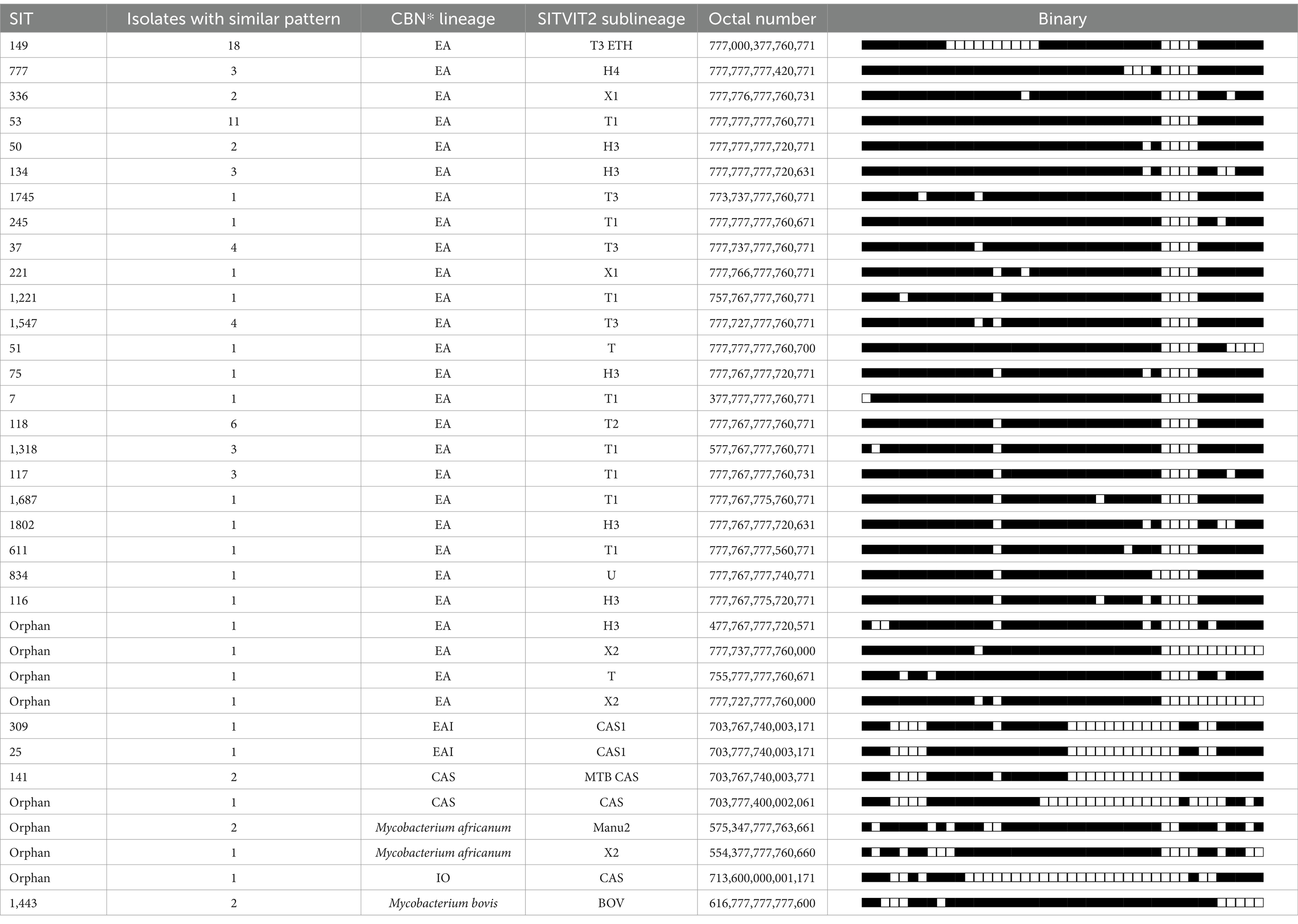
Table 3. Spoligotype patterns of Mycobacterium tuberculosis complex species isolated from humans in Central Ethiopia.
In the present study, M. tuberculosis complex species were isolated from 86 human TB patients at Bishoftu and Adama Referral Hospital and Adama Health Center who visited these health institutions. Additionally, 21 MTBC species were isolated from suspected tuberculosis lesions in cattle slaughtered at the Adama municipal abattoir and Bishoftu ELFORA export abattoir. The isolates were identified at the strain and lineage levels based on spoligotyping. Analyzing strain diversity and comparing it with international databases is key to understanding the global distribution of M. tuberculosis strains. In this context, we described the strain diversity of M. tuberculosis complex among clinical isolates from pulmonary TB patients and from tuberculosis lesions in cattle slaughtered in central Ethiopia, and we compared these strains with those in the SITVIT2 database.
In the present study, culture positivity in primary culture media from animals was found in 26.3% (21/80) of tissue lesions suspected of TB. However, the isolation percentage of the M. tuberculosis complex from TB-suspected lesions was low in this study, similar to results from earlier studies (18, 21, 22). This low percentage of M. tuberculosis complex isolation in zebu cattle in Ethiopia could be associated with various factors. One potential factor is the containment of the M. tuberculosis complex within granulomas, which are usually calcified and could lead to the death of M. tuberculosis, resulting in a low yield from culturing these lesions. Over the last 2 decades, most lesions detected in zebu cattle have been localized to one anatomical site and calcified (14, 15, 22, 23). This low isolation rate of mycobacteria may also stem from reduced culture sensitivity due to prolonged storage at field sites, freeze–thaw cycles during transportation, contamination of tissue samples, and the overgrowth of M. bovis by environmental mycobacteria (24). Furthermore, the presence of caseous and/or calcified lesions, as well as lesions resembling tuberculous lesions, may not always indicate mycobacterial origin; they can be caused by other intracellular organisms or parasites, or viable mycobacteria may not be present in calcified lesions (13, 25).
In the present study, 21 animal isolates showed evidence of M. tuberculosis complex species, with 12 identified as M. bovis and nine as M. tuberculosis. Further characterization of the M. bovis strain through spoligotyping revealed the presence of 12 isolates grouped into nine clusters of spoligotype patterns. Of the nine clusters of M. bovis, seven were novel and had not been previously reported in the M. bovis spoligotype database, while the remaining two clusters corresponded to spoligotype SB1265 (two isolates) and SB1176 in the M.bovis.org database. Similar to this study, the spoligotype SB1265 has also been reported earlier (26). The isolation of SB1176 (M. bovis) indicates its dominance as a major cause of BTB in Ethiopian cattle (18, 23, 26–28). The spoligotype SB1176 carries a specific spoligotype feature (spacers 4–7 missing) typical of members of a clonal complex identified as M. bovis African 2 (Af2), which has so far only been found in East Africa (29).
The nine M. tuberculosis strains isolated from TB-suspected lesions in cattle belong to eight Euro-American lineages (SIT 50, 118, 117, and 1,318) and one Indo-Oceanic lineage. M. tuberculosis infection has been reported in a wide range of domestic and wildlife animal species, most frequently in those living in close, prolonged contact with humans (30–33).
Similar to the present study, the isolation of M. tuberculosis from TB-suspected lesions was reported in cattle in Ethiopia by others (18, 22, 34, 35). Moreover, M. tuberculosis has been sporadically isolated from cattle in other countries, as reported by (36). Animal attendants with active TB in the respiratory tract, urinary tract, or gastrointestinal tract also represent an active source of M. tuberculosis for animals, spreading the bacillus via sputum, urine, or feces (33). The isolation of M. tuberculosis from bovine tissues likely supports the possible transmission of M. tuberculosis from humans to cattle.
The isolation of M. tuberculosis from bovine tissues raises concerns about potential human-to-cattle transmission, although M. tuberculosis is primarily considered a human pathogen. Unlike M. bovis, the main causative agent of bovine tuberculosis with a well-established zoonotic cycle, M. tuberculosis typically lacks a known reservoir in cattle. However, previous studies have documented cases of M. tuberculosis infection in cattle, suggesting that close contact with infected humans, particularly in environments with poor biosecurity measures, may facilitate transmission (37, 38). Human-to-animal transmission of M. tuberculosis has been reported in other animals, indicating that cross-species transmission is possible under certain conditions (39, 40).
Possible routes of transmission to cattle include direct aerosol exposure in confined environments, ingestion of contaminated feed or water, and contamination during handling and milking. While such occurrences are rare, they highlight the need for improved surveillance of bovine tuberculosis cases that do not fit the classical M. bovis infection profile. Given the zoonotic potential of tuberculosis, distinguishing between M. tuberculosis and M. bovis infections in cattle is critical for understanding transmission dynamics and implementing appropriate control measures. Further molecular epidemiological studies are needed to confirm the origins of M. tuberculosis in cattle and to assess the risk of reverse zoonosis in agricultural settings.
Out of 392 positive sputum samples cultured, 86 (21.9%) were culture-positive. The culture positivity rate of sputum samples collected from TB-suspected human cases, such as those who have coughed for more than two weeks and experienced night sweats, was low. Similarly, an earlier study in Ethiopia reported a low percentage of M. tuberculosis isolation from the sputum of TB patients (41). This study indicated that 83.2% (247/297) of patients suspected of having pulmonary TB had a negative smear result for acid-fast bacilli. The likelihood of isolating the M. tuberculosis complex from smear-negative TB cases was low. The culture positivity of sputum is directly related to the presence of bacilli, which can also be affected by the nature of the granuloma. If the granuloma is dense and solid, the chances of bacilli excretion in the sputum are minimal, leading to a low yield from sputum culture. Additionally, if the sputum is not collected correctly or if its storage and processing for culture are not conducted according to optimal procedures, the yield of sputum culture will be low.
Spoligotyping of the 86 human isolates revealed 35 distinct spoligotype patterns, representing 40.7% of genotype diversity. The diversity of the spoligotype strains observed in this study was consistent with the 41.9% reported by others (42, 43), but higher than the percentages reported in earlier studies conducted in Ethiopia (44–46). This low diversity of spoligotype strains of M. tuberculosis could suggest that a limited number of lineages are responsible for the disease in the study area despite significant migrations of various infected ethnic groups to Adama City and its surroundings from other regions of the country.
A total of 65 human isolates were grouped into 14 clusters, resulting in an overall clustering percentage of 75.6%. The clustering rate observed in the present study was comparable to a national survey (47) and a study performed in the Afar pastoral region (42). However, the clustering rate reported in this study is higher than that found in earlier research (43, 48, 49). The observed differences in clustering rates might be related to variations in sanitation, housing, and population density. Additionally, a high level of strain clustering could suggest recent and ongoing TB transmission in the study area (50–52). Consequently, poor sanitation, poor housing, and high population density (overcrowding) facilitate TB transmission, thereby increasing the clustering rate, while adequate sanitation, proper housing, and low population density reduce TB transmission, resulting in a decreased clustering rate.
In the present study, the most commonly identified strains were SIT 149, SIT 53, and SIT 118, in order of decreasing frequency. Similar to this study, SIT149 was predominantly isolated in previous research conducted in the Afar Pastoral Region of Ethiopia (42). In accordance with the present study, SIT 149 (T3-ETH) is frequently isolated in Ethiopia and among Ethiopian immigrants in Denmark (20). A previous study in Ethiopia showed that SIT149 was the most frequently clustered strain among MDR-TB isolates (53). The second most common M. tuberculosis strain found in this study was SIT53, which has also been reported earlier in the SITVIT database as the most common type in Ethiopia by other researchers (54).
The strains of the M. tuberculosis complex isolated in the present study belonged to the M. tuberculosis species lineages: Euro-American, Indo-Oceanic, East-African-Indian, or the species M. bovis or M. africanum species. The dominant lineage was the Euro-American lineage, constituting 87.2% of the isolates. The Euro-American lineage is the most prevalent in the world (55) and in Ethiopia, ranging from 32.5% in Gambella near the South Sudan border to 86.8% (48) in central Ethiopia (56). It can be hypothesized that the Euro-American lineage may have been introduced to Ethiopia by Europeans during the Italian invasion. Screening of the SITVIT2 database also identified 3.5% (3/86) of the isolates as members of M. africanum. This finding is in line with the findings reported by (49). M. africanum has been recognized as a significant cause of human TB in West African countries (57, 58).
Among the 86 human isolates, 2 (2.3%) isolates were identified as M. bovis. The two human M. bovis isolates had identical spoligo patterns, suggesting that they originated from the same sources. The spoligotype is identical to SB1443 in the www.mbovis.org database. Additionally, the M. bovis isolates from human pulmonary TB patients matched the spoligotype of animal isolates (SB1443), which were previously identified in cattle from Nigeria (59). This finding suggests cattle-to-human transmission of M. bovis, as cattle are the primary hosts of this pathogen.
Humans can become infected with M. bovis through the ingestion of contaminated food from cattle or by inhaling particles from infected cattle. This finding is also supported by the findings of other studies (7), which indicate the involvement of M. bovis in TB, presumably linked to contact with infected cattle and the consumption of raw milk and meat (2). Overall, the contribution of M. bovis to TB in central Ethiopia was low (2.3%), similar to other studies conducted in Ethiopia.
In the present study, M. bovis was isolated from human TB patients, while on the other hand, M. tuberculosis was isolated from TB lesions in cattle. Thus, this study indicates that TB is a significant example of zoonosis and reverse zoonosis. Similarly, earlier studies reported the isolation of M. bovis from human TB patients in Ethiopia, although the percentage of isolation was low (27, 42, 49, 60, 61). Additionally, M. tuberculosis was isolated from TB lesions in cattle in Ethiopia and other countries (18, 36, 61–64). Moreover, M. tuberculosis was also isolated from various species of domestic and zoo animals (65–67). Although such studies are generally scarce in Africa, the isolation of M. tuberculosis has been reported from animals in a few African countries (68, 69). The possible sources of infection for animals with M. tuberculosis may include the sputum, urine, and feces of humans, arising from active cases of human TB in the respiratory tract, urinary tract, or gastrointestinal tract, respectively (33).
The present studies and earlier Ethiopian studies reported a higher percentage of M. tuberculosis (reverse zoonosis) isolation in cattle tissues than the percentage of M. bovis isolation in humans. This observation could suggest that reverse zoonosis is more important than zoonosis in terms of TB transmission in Ethiopia. However, people infected with M. tuberculosis may be more likely to seek medical attention, leading to an underestimation of the prevalence of M. bovis infection in the population.
The prevalence of bTB in indigenous cattle, which constitute approximately 98% of the Ethiopian cattle population, is low, resulting in a minimal chance of bTB transmission from cattle to humans. Conversely, Ethiopia is one of the 10 countries worldwide with a high burden of human TB (12), which increases the likelihood of M. tuberculosis transmission from humans to cattle due to the socio-cultural connections Ethiopian farmers maintain with their cattle. However, contrary to Ethiopian reports, studies from other African countries have indicated a higher percentage of M. bovis isolation from humans than the percentage of M. tuberculosis isolation from animals (7, 70–72).
This study indicated that both M. tuberculosis and M. bovis strains are circulating among humans and cattle in central Ethiopia. This finding highlights the challenges of TB control in the study area. Therefore, a One Health approach incorporating multidisciplinary control and preventive measures should be considered by the departments of public health, veterinary services, and environmental health in the study area.
The datasets presented in this study can be found in online repositories. The names of the repository/repositories and accession number(s) can be found in the article/Supplementary material.
The studies involving human participants were approved by the Institutional Review Board (IRB) of the Oromia Health Bureau (Reference No. IRB/BEFO/HBTFH/1–8/3647). These studies were conducted according to local legislation and institutional requirements. Written informed consent for participation in this study was obtained from the participants’ legal guardians or next of kin. The animal study was approved by the Oromia Health Bureau’s Institutional Review Board (IRB) (Reference No. IRB/BEFO/HBTFH/1–8/3647) and was conducted in compliance with local legislation and institutional requirements.
TW: Writing – original draft, Writing – review & editing. TM: Writing – original draft, Writing – review & editing. AZ: Conceptualization, Data curation, Formal analysis, Methodology, Writing – review & editing. MC: Writing – review & editing. HK: Writing – review & editing. GM: Writing – review & editing. GA: Writing – original draft, Writing – review & editing.
The author(s) declare that financial support was received for the research, authorship, and/or publication of this article. The study was financially supported by the Office of the Vice President for Research and Graduate Studies at Addis Ababa University (AAU) for the thematic research project “Mycobacterial Infections in Animals and Their Livestock,” with Account Number 15000018800006223. The AAU monitored the implementation of this research project through annual scientific and financial reports. Additionally, the final scientific and financial reports were submitted to the AAU. The APC of the manuscript is covered by United Arab Emirates University.
The authors acknowledge the study participants, animal owners, and the administrators of the abattoirs and healthcare provider centers (hospitals and health centers) for their willingness to support the study.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Gen AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
The Supplementary material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fvets.2025.1527279/full#supplementary-material
Supplementary IMAGE 1 | Spoligotype pattern of mycobacteria isolated from humans. The profile is based on presence or absence of 43 spacers. No 1 and 43 show the patterns of positive control for M. tuberculosis; No 2 shows patterns of a negative control; No 3 show a positive control for Mycobacterium bovis. Patterns other than No 1, 2, 23 and 43 show the patterns of mycobacteria isolated from humans. Black squares show the presence of spacers while white dots show the absence of spacers.
Supplementary ImAGE 2 | Spoligotype pattern of mycobacteria isolated from humans. The profile is based on presence or absence of 43 spacers. No 1 and 43 show the patterns of positive control for M. tuberculosis; No 2 shows patterns of a negative control; No 3 show a positive control for Mycobacterium bovis. Patterns other than No 1, 2, 23 and 43 show the patterns of mycobacteria isolated from humans. Black squares show the presence of spacers while white dots show the absence of spacers.
Supplementary IMAGE 3 | Spoligotype pattern of mycobacteria isolated from animal and humans. The profile is based on presence or absence of 43 spacers. No 1 and 43 show the patterns of positive control for M. tuberculosis; No 2 shows patterns of a negative control; No 3 show a positive control for Mycobacterium bovis. Patterns other than No 1, 2, 23 and 43 show the patterns of mycobacteria isolated from humans. Black squares show the presence of spacers while white dots show the absence of spacers.
BTB, Bovine tuberculosis; CAS, Central Asian Strain; EA, Euro-American; EAI, East-African-Indian; IO, Indio-Oceanic; LJ, Lowenstein-Jensen; SITVIT, Spoligotype International Types-VNTR International Types; APHA, Plant and Animal Health Agency; UAE, United Arab Emirates; VNTR, Variable Number of DNA Tandem Repeats.
1. OIE. Bovine Tuberculosis. World Organization for Animal Health, OIE, Protecting Animals, Preserving Our Future. (2020). Available at: https://www.woah.org/ (Accessed June 22, 2024).
2. Ayele, WY. Bovine tuberculosis: an old disease but a new threat to Africa. Int J Tuberc Lung Dis. (2004) 8:924–37.
3. Cosivi, O, Grange, JM, Daborn, CJ, Raviglione, MC, Fujikura, T, Cousins, D, et al. Zoonotic tuberculosis due to Mycobacterium bovis in developing countries. Emerg Infect Dis. (1998) 4:59–70. doi: 10.3201/eid0401.980108
4. Jebara, KB. Ten years’ work on the world organisation for animal health (OIE) worldwide animal disease notification system. Prev Vet Med. (2012) 107:149–59. doi: 10.1016/j.prevetmed.2012.08.008
5. Cook, AJ, Tuchili, LM, Buve, A, Foster, SD, Godfrey-Fausett, P, Pandey, GS, et al. Human and bovine tuberculosis in the Monze District of Zambia--a cross-sectional study. Br Vet J. (1996) 152:37–46. doi: 10.1016/s0007-1935(96)80084-4
6. Regassa, GM, and Ameni, G. Bovine tuberculosis is more prevalent in cattle owned by farmers with active tuberculosis in Central Ethiopia. Vet J. (2008) 178:119–25. doi: 10.1016/j.tvjl.2007.06.019
7. Firdessa, R, Berg, S, Hailu, E, Schelling, E, Gumi, B, Erenso, G, et al. Mycobacterial lineages causing pulmonary and extrapulmonary tuberculosis, Ethiopia. Emerg Infect Dis. (2013) 19:460–3. doi: 10.3201/eid1903.120256
8. Kidane, D, Olobo, JO, Habte, A, Negesse, Y, Aseffa, A, Abate, G, et al. Identification of the causative organism of tuberculous lymphadenitis in Ethiopia by PCR. J Clin Microbiol. (2002) 40:4230–4. doi: 10.1128/JCM.40.11.4230-4234.2002
9. Ashford, DA, Whitney, E, Raghunathan, P, and Cosivi, O. Epidemiology of selected mycobacteria that infect humans and other animals. Rev Sci Tech. (2001) 20:325–37. doi: 10.20506/rst.20.1.1266
10. Kazwala, RR, Daborn, CJ, Kusiluka, LJ, Jiwa, SF, Sharp, JM, and Kambarage, DM. Isolation of Mycobacterium species from raw milk of pastoral cattle of the southern highlands of Tanzania. Trop Anim Health Prod. (1998) 30:233–9. doi: 10.1023/a:1005075112393
11. Yates, MD, Pozniak, A, and Grange, JM. Isolation of mycobacteria from patients seropositive for the human immunodeficiency virus (HIV) in south East England: 1984-92. Thorax. (1993) 48:990–5. doi: 10.1136/thx.48.10.990
12. WHO. Global tuberculosis report 2020. (2020). Available online at: https://www.who.int/publications-detail-redirect/9789240013131 (Accessed January 20, 2024).
13. Corner, LA. Post mortem diagnosis of Mycobacterium bovis infection in cattle. Vet Microbiol. (1994) 40:53–63. doi: 10.1016/0378-1135(94)90046-9
14. Ameni, G, Aseffa, A, Engers, H, Young, D, Hewinson, G, and Vordermeier, M. Cattle husbandry in Ethiopia is a predominant factor affecting the pathology of bovine tuberculosis and gamma interferon responses to mycobacterial antigens. Clin Vaccine Immunol. (2006) 13:1030–6. doi: 10.1128/CVI.00134-06
15. Vordermeier, M, Ameni, G, and Glass, EJ. Cytokine responses of Holstein and Sahiwal zebu derived monocytes after mycobacterial infection. Trop Anim Health Prod. (2012) 44:651–5. doi: 10.1007/s11250-011-9950-x
16. OIE. Bovine tuberculosis. In Manual of Diagnostic Tests and Vaccines for Terrestrial Animals. Part 2, Section 2.3. Chapter 2.3.3. World Organization for Animal Health. (2010). Available at: https://www.woah.org/ (Accessed June 22, 2024).
17. WHO. Laboratory Services in Tuberculosis Control: Culture. Part III, WHO, Geneva, Switzerland. (1998). Available at: https://iris.who.int/ (Accessed June 22, 2024).
18. Berg, S, Firdessa, R, Habtamu, M, Gadisa, E, Mengistu, A, Yamuah, L, et al. The burden of mycobacterial disease in ethiopian cattle: implications for public health. PLoS One. (2009) 4:e5068. doi: 10.1371/journal.pone.0005068
19. Kamerbeek, J, Schouls, L, Kolk, A, van Agterveld, M, van Soolingen, D, Kuijper, S, et al. Simultaneous detection and strain differentiation of Mycobacterium tuberculosis for diagnosis and epidemiology. J Clin Microbiol. (1997) 35:907–14. doi: 10.1128/jcm.35.4.907-914.1997
20. Brudey, K, Driscoll, JR, Rigouts, L, Prodinger, WM, Gori, A, Al-Hajoj, SA, et al. Mycobacterium tuberculosis complex genetic diversity: mining the fourth international spoligotyping database (SpolDB4) for classification, population genetics and epidemiology. BMC Microbiol. (2006) 6:23. doi: 10.1186/1471-2180-6-23
21. Ameni, G, Ameni, K, and Tibbo, M. Bovine tuberculosis: prevalence and risk factor assessment in cattle and cattle owners in Wuchale-Jida district Central Ethiopia (2003) 1:1–4.
22. Aylate, A. Bovine tuberculosis: prevalence and diagnostic efficacy of routine meat inspection procedure in Woldiya municipality abattoir north Wollo zone, Ethiopia. Trop Anim Health Prod. (2013) 45:855–64. doi: 10.1007/s11250-012-0298-7
23. Ameni, G, Aseffa, A, Engers, H, Young, D, Gordon, S, Hewinson, G, et al. High prevalence and increased severity of pathology of bovine tuberculosis in Holsteins compared to zebu breeds under field cattle husbandry in Central Ethiopia. Clin Vaccine Immunol. (2007) 14:1356–61. doi: 10.1128/CVI.00205-07
24. Cleaveland, S, Shaw, DJ, Mfinanga, SG, Shirima, G, Kazwala, RR, Eblate, E, et al. Mycobacterium bovis in rural Tanzania: risk factors for infection in human and cattle populations. Tuberculosis (Edinb). (2007) 87:30–43. doi: 10.1016/j.tube.2006.03.001
25. Diguimbaye-Djaibé, C, Hilty, M, Ngandolo, R, Mahamat, HH, Pfyffer, GE, Baggi, F, et al. Mycobacterium bovis isolates from tuberculous lesions in Chadian zebu carcasses. Emerg Infect Dis. (2006) 12:769–71. doi: 10.3201/eid1205.050691
26. Biffa, D, Skjerve, E, Oloya, J, Bogale, A, Abebe, F, Dahle, U, et al. Molecular characterization of Mycobacterium bovis isolates from Ethiopian cattle. BMC Vet Res. (2010) 6:28. doi: 10.1186/1746-6148-6-28
27. Firdessa, R, Tschopp, R, Wubete, A, Sombo, M, Hailu, E, Erenso, G, et al. High prevalence of bovine tuberculosis in dairy cattle in Central Ethiopia: implications for the dairy industry and public health. PLoS One. (2012) 7:e52851. doi: 10.1371/journal.pone.0052851
28. Mekibeb, A, Fulasa, TT, Firdessa, R, and Hailu, E. Prevalence study on bovine tuberculosis and molecular characterization of its causative agents in cattle slaughtered at Addis Ababa municipal abattoir, Central Ethiopia. Trop Anim Health Prod. (2013) 45:763–9. doi: 10.1007/s11250-012-0287-x
29. Berg, S, Garcia-Pelayo, MC, Müller, B, Hailu, E, Asiimwe, B, Kremer, K, et al. African 2, a clonal complex of Mycobacterium bovis epidemiologically important in East Africa. J Bacteriol. (2011) 193:670–8. doi: 10.1128/JB.00750-10
30. Alfonso, R, Romero, RE, Diaz, A, Calderon, MN, Urdaneta, G, Arce, J, et al. Isolation and identification of mycobacteria in New World primates maintained in captivity. Vet Microbiol. (2004) 98:285–95. doi: 10.1016/j.vetmic.2003.10.023
31. Montali, RJ, Mikota, SK, and Cheng, LI. Mycobacterium tuberculosis in zoo and wildlife species. Rev Sci Tech. (2001) 20:291–303. doi: 10.20506/rst.20.1.1268
32. Pavlik, I, Yayo Ayele, W, Parmova, I, Melicharek, I, Hanzlikova, M, Svejnochova, M, et al. Mycobacterium tuberculosis in animal and human populations in six central European countries during 1990-1999. Vet Med. (2003) 48:83–9. doi: 10.17221/5754-VETMED
33. Thoen, CO, Karlson, AG, and Himes, EM. Mycobacterial infections in animals. Rev Infect Dis. (1981) 3:960–72. doi: 10.1093/clinids/3.5.960
34. Ameni, G, Desta, F, and Firdessa, R. Molecular typing of Mycobacterium bovis isolated from tuberculosis lesions of cattle in north eastern Ethiopia. Vet Rec. (2010) 167:138–41. doi: 10.1136/vr.b4881
35. Tsegaye, W, Aseffa, A, Mache, A, Mengistu, Y, Berg, S, and Ameni, G. Conventional and molecular epidemiology of bovine tuberculosis in dairy farms in Addis Ababa City, the Capital of Ethiopia. Int J Applied Res Vet Med. (2010) 8:143–51.
36. Ocepek, M, Pate, M, Zolnir-Dovc, M, and Poljak, M. Transmission of Mycobacterium tuberculosis from human to cattle. J Clin Microbiol. (2005) 43:3555–7. doi: 10.1128/JCM.43.7.3555-3557.2005
37. Okeke, LA, Cadmus, S, Okeke, IO, Muhammad, M, Awoloh, O, Dairo, D, et al. Prevalence and risk factors of Mycobacterium tuberculosis complex infection in slaughtered cattle at Jos south abattoir, plateau state, Nigeria. Pan Afr Med J. (2014) 18 Suppl 1:7. doi: 10.11694/pamj.supp.2014.18.1.3841
38. Ghebremariam, MK, Michel, AL, Nielen, M, Vernooij, JCM, and Rutten, VPMG. Farm-level risk factors associated with bovine tuberculosis in the dairy sector in Eritrea. Transbound Emerg Dis. (2018) 65:105–13. doi: 10.1111/tbed.12622
39. Stephens, N, Vogelnest, L, Lowbridge, C, Christensen, A, Marks, GB, Sintchenko, V, et al. Transmission of Mycobacterium tuberculosis from an Asian elephant (Elephas maximus) to a chimpanzee (Pan troglodytes) and humans in an Australian zoo. Epidemiol Infect. (2013) 141:1488–97. doi: 10.1017/S095026881300068X
40. Borham, M, Oreiby, A, El-Gedawy, A, Hegazy, Y, Khalifa, HO, Al-Gaabary, M, et al. Review on bovine tuberculosis: an emerging disease associated with multidrug-resistant Mycobacterium species. Pathogens. (2022) 11:715. doi: 10.3390/pathogens11070715
41. Desta, K, Asrat, D, Lemma, E, Gebeyehu, M, and Feleke, B. Prevalence of smear negative pulmonary tuberculosis among patients visiting St. Peter’s Tuberculosis Specialized Hospital, Addis Ababa, Ethiopia. Ethiop Med J. (2009) 47:17–24.
42. Belay, M, Ameni, G, Bjune, G, Couvin, D, Rastogi, N, and Abebe, F. Strain diversity of Mycobacterium tuberculosis isolates from pulmonary tuberculosis patients in Afar pastoral region of Ethiopia. Biomed Res Int. (2014) 2014:238532. doi: 10.1155/2014/238532
43. Disassa, H, Tafess, K, Worku, A, and Ameni, G. A preliminary study on molecular characterization of Mycobacterium tuberculosis in Benishangul Gumuz region, Western Ethiopia. BMRJ. (2015) 10:1–10. doi: 10.9734/BMRJ/2015/20032
44. Debebe, T, Admassu, A, Mamo, G, and Ameni, G. Molecular characterization of Mycobacterium tuberculosis isolated from pulmonary tuberculosis patients in Felege Hiwot referral hospital, Northwest Ethiopia. J Microbiol Immunol Infect. (2014) 47:333–8. doi: 10.1016/j.jmii.2013.03.012
45. Diriba, B, Berkessa, T, Mamo, G, Tedla, Y, and Ameni, G. Spoligotyping of multidrug-resistant Mycobacterium tuberculosis isolates in Ethiopia. Int J Tuberc Lung Dis. (2013) 17:246–50. doi: 10.5588/ijtld.12.0195
46. Yimer, SA, Hailu, E, Derese, Y, Bjune, GA, and Holm-Hansen, C. Spoligotyping of Mycobacterium tuberculosis isolates among pulmonary tuberculosis patients in Amhara region, Ethiopia. APMIS. (2013) 121:878–85. doi: 10.1111/apm.12046
47. Getahun, M, Ameni, G, Kebede, A, Yaregal, Z, Hailu, E, Medihn, G, et al. Molecular typing and drug sensitivity testing of Mycobacterium tuberculosis isolated by a community-based survey in Ethiopia. BMC Public Health. (2015) 15:751. doi: 10.1186/s12889-015-2105-7
48. Asebe, G, Adane, W, Aboma, Z, Tafess, K, and Ameni, G. Molecular characterization of Mycobacterium tuberculosis complex in Gambella region, Southwest Ethiopia. Afr J Microbiol Res. (2016) 10:1086–93. doi: 10.5897/AJMR2015.7415
49. Nuru, A, Mamo, G, Teshome, L, Zewdie, A, Medhin, G, Pieper, R, et al. Bovine tuberculosis and its risk factors among dairy cattle herds in and around Bahir Dar City, Northwest Ethiopia. Ethiop Vet J. (2015) 19:27. doi: 10.4314/evj.v19i2.3
50. Barnes, PF, Yang, Z, Preston-Martin, S, Pogoda, JM, Jones, BE, Otaya, M, et al. Patterns of tuberculosis transmission in Central Los Angeles. JAMA. (1997) 278:1159–63. doi: 10.1001/jama.1997.03550140051039
51. Easterbrook, PJ, Gibson, A, Murad, S, Lamprecht, D, Ives, N, Ferguson, A, et al. High rates of clustering of strains causing tuberculosis in Harare, Zimbabwe: a molecular epidemiological study. J Clin Microbiol. (2004) 42:4536–44. doi: 10.1128/JCM.42.10.4536-4544.2004
52. Small, PM, Hopewell, PC, Singh, SP, Paz, A, Parsonnet, J, Ruston, DC, et al. The epidemiology of tuberculosis in San Francisco -- a population-based study using conventional and molecular methods. N Engl J Med. (1994) 330:1703–9. doi: 10.1056/NEJM199406163302402
53. Agonafir, M, Lemma, E, Wolde-Meskel, D, Goshu, S, Santhanam, A, Girmachew, F, et al. Phenotypic and genotypic analysis of multidrug-resistant tuberculosis in Ethiopia. Int J Tuberc Lung Dis. (2010) 14:1259–65.
54. Bruchfeld, J, Aderaye, G, Palme, IB, Bjorvatn, B, Ghebremichael, S, Hoffner, S, et al. Molecular epidemiology and drug resistance of Mycobacterium tuberculosis isolates from Ethiopian pulmonary tuberculosis patients with and without human immunodeficiency virus infection. J Clin Microbiol. (2002) 40:1636–43. doi: 10.1128/JCM.40.5.1636-1643.2002
55. Reed, MB, Pichler, VK, McIntosh, F, Mattia, A, Fallow, A, Masala, S, et al. Major Mycobacterium tuberculosis lineages associate with patient country of origin. J Clin Microbiol. (2009) 47:1119–28. doi: 10.1128/JCM.02142-08
56. Bedewi, Z, Worku, A, Mekonnen, Y, Yimer, G, Medhin, G, Mamo, G, et al. Molecular typing of Mycobacterium tuberculosis complex isolated from pulmonary tuberculosis patients in Central Ethiopia. BMC Infect Dis. (2017) 17:184. doi: 10.1186/s12879-017-2267-2
57. de Jong, BC, Hill, PC, Brookes, RH, Gagneux, S, Jeffries, DJ, Otu, JK, et al. Mycobacterium africanum elicits an attenuated T cell response to early secreted antigenic target, 6 kDa, in patients with tuberculosis and their household contacts. J Infect Dis. (2006) 193:1279–86. doi: 10.1086/502977
58. Homolka, S, Post, E, Oberhauser, B, George, AG, Westman, L, Dafae, F, et al. High genetic diversity among Mycobacterium tuberculosis complex strains from Sierra Leone. BMC Microbiol. (2008) 8:103. doi: 10.1186/1471-2180-8-103
59. Müller, B, Hilty, M, Berg, S, Garcia-Pelayo, MC, Dale, J, Boschiroli, ML, et al. African 1, an epidemiologically important clonal complex of Mycobacterium bovis dominant in Mali, Nigeria, Cameroon, and Chad. J Bacteriol. (2009) 191:1951–60. doi: 10.1128/JB.01590-08
60. Zewdie, O, Mihret, A, Ameni, G, Worku, A, Gemechu, T, and Abebe, T. Molecular typing of mycobacteria isolated from tuberculous lymphadenitis cases in Addis Ababa, Ethiopia. Int J Tuberc Lung Dis. (2016) 20:1529–34. doi: 10.5588/ijtld.15.1023
61. Gumi, B, Schelling, E, Berg, S, Firdessa, R, Erenso, G, Mekonnen, W, et al. Zoonotic transmission of tuberculosis between pastoralists and their livestock in south-East Ethiopia. EcoHealth. (2012) 9:139–49. doi: 10.1007/s10393-012-0754-x
62. Ameni, G, Tadesse, K, Hailu, E, Deresse, Y, Medhin, G, Aseffa, A, et al. Transmission of Mycobacterium tuberculosis between farmers and cattle in Central Ethiopia. PLoS One. (2013) 8:e76891. doi: 10.1371/journal.pone.0076891
63. Ameni, G, Vordermeier, M, Firdessa, R, Aseffa, A, Hewinson, G, Gordon, SV, et al. Mycobacterium tuberculosis infection in grazing cattle in Central Ethiopia. Vet J. (2011) 188:359–61. doi: 10.1016/j.tvjl.2010.05.005
64. Prasad, HK, Singhal, A, Mishra, A, Shah, NP, Katoch, VM, Thakral, SS, et al. Bovine tuberculosis in India: potential basis for zoonosis. Tuberculosis (Edinb). (2005) 85:421–8. doi: 10.1016/j.tube.2005.08.005
65. Lyashchenko, KP, Greenwald, R, Esfandiari, J, Olsen, JH, Ball, R, Dumonceaux, G, et al. Tuberculosis in elephants: antibody responses to defined antigens of Mycobacterium tuberculosis, potential for early diagnosis, and monitoring of treatment. Clin Vaccine Immunol. (2006) 13:722–32. doi: 10.1128/CVI.00133-06
66. Mamo, G, Bayleyegn, G, Sisay Tessema, T, Legesse, M, Medhin, G, Bjune, G, et al. Pathology of camel tuberculosis and molecular characterization of its causative agents in pastoral regions of Ethiopia. PLoS One. (2011) 6:e15862. doi: 10.1371/journal.pone.0015862
67. Mamo, G, Kassaye, A, Ali, MS, and Ameni, G. A cross sectional study of camel tuberculosis in Ethiopia. Bull Anim Hlth Prod Afr. (2009) 57:13–20. doi: 10.4314/bahpa.v57i1.44047
68. Boulahbal, F, Benelmouffok, A, and Brahimi, K. Role of Mycobacterium tuberculosis in bovine tuberculosis. Arch Inst Pasteur Alger. (1978) 53:155–64.
69. Sulieman, MS, and Hamid, ME. Identification of acid fast bacteria from caseous lesions in cattle in Sudan. J Vet Med B Infect Dis Vet Public Health. (2002) 49:415–8. doi: 10.1046/j.1439-0450.2002.00565.x
70. Addo, K, Owusu-Darko, K, Yeboah-Manu, D, Caulley, P, Minamikawa, M, Bonsu, F, et al. Mycobacterial species causing pulmonary tuberculosis at the korle bu teaching hospital, Accra, Ghana. Ghana Med J. (2007) 41:52–7. doi: 10.4314/gmj.v41i2.55293
71. Idigbe, EO, Anyiwo, CE, and Onwujekwe, DI. Human pulmonary infections with bovine and atypical mycobacteria in Lagos, Nigeria. J Trop Med Hyg. (1986) 89:143–8.
Keywords: Ethiopia, reverse zoonosis, tuberculosis, zoonosis, Central Ethiopia
Citation: Woldemariam T, Mohammed T, Zewude A, Chanyalew M, Khalifa HO, Mamo G and Ameni G (2025) Zoonotic transmission of the Mycobacterium tuberculosis complex between cattle and humans in Central Ethiopia. Front. Vet. Sci. 12:1527279. doi: 10.3389/fvets.2025.1527279
Received: 14 November 2024; Accepted: 21 February 2025;
Published: 10 March 2025.
Edited by:
Moh A. Alkhamis, Kuwait University, KuwaitReviewed by:
Max Maurin, Centre Hospitalier Universitaire de Grenoble, FranceCopyright © 2025 Woldemariam, Mohammed, Zewude, Chanyalew, Khalifa, Mamo and Ameni. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Temesgen Mohammed, dGVtZXNnZW5fc0B1YWV1LmFjLmFl; Gobena Ameni, Z29iZW5hLmFtZW5pQHVhZXUuYWMuYWU=
†These authors have contributed equally to this work and share first authorship
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.