
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Plant Sci. , 26 February 2025
Sec. Functional and Applied Plant Genomics
Volume 16 - 2025 | https://doi.org/10.3389/fpls.2025.1509472
This article is part of the Research Topic Innovative Molecular Strategies for Enhancing Plant Defense Against Biotic Stresses View all 9 articles
12-oxo-phytodienoic acid reductase (OPR) is one of the key enzymes in the octadecanoid pathway, and it controls the last step of jasmonic acid (JA) biosynthesis. Although multiple isoforms and functions of OPRs have been identified in various plants, no OPR genes have been identified, and their possible roles in grapevine development and defense mechanisms remain unknown. In this study, nine VvOPR genes were identified from grapevine genome and classified into two subfamilies. Systematic analyses of the physical and chemical properties, the expression and structure of the VvOPR genes, promoter elements, and chromosome locations were performed via bioinformatics and molecular biology methods. In addition, we described the characterization of the OPRI gene VvOPR1, which was synthesized via a PCR-based two-step DNA synthesis quantification reverse-transcription (PTDS) method. VvOPR1 expression is tissue-specific and induced by various stresses. The overexpression of VvOPR1 in Arabidopsis and rice (OT) significantly increased tolerance to Cu, Zn stress, and Cu, Zn stress-induced restriction of the germination rate, root/shoot length and fresh weight was significantly alleviated in OT. In OT, VvOPR1 enhanced the photosynthetic capacity, promoted ABA synthesis and the ABA-dependent stress response pathway, improved the antioxidation capacity by increasing the activities of ROS scavengers and the expression level of the related genes, while enhancing the accumulation of proline, AsA, GSH and reducing MDA and H2O2 levels. Moreover, VvOPR1 reduced Cu2+, Zn2+ accumulation and translocation. Together, we first systematically characterized the grapevine OPR gene family and reported that VvOPR1 responded to Cu, Zn stress in an ABA-dependent manner, and was quite independent of JA synthesis and signaling. All of the above results provide an important research basis and theoretical basis for further revealing the functions of VvOPR in grapevines in the future.
The 12-oxo-phytodienoic acid reductase (OPR) is a key enzyme that catalyzes the conversion of 12-oxophytodienoic acid (OPDA) to 12-oxo-phytodienoic acid (OPC-8:0), a reaction that is a key process in jasmonic acid (JA) biosynthesis (Liechti and Farmer, 2006; Schaller, 2001). OPR enzymes are classified as flavin mononucleotide (FMN)-dependent oxidoreductases and belong to the old yellow enzyme (OYE) family, which is well represented in the yeast genome (Liu et al., 2020).
On the basis of substrate specificity, OPRs are split into two subgroups in dicots, subgroupI (OPRI) (e.g, AtOPR1 and AtOPR2) and subgroupII members (OPRII) (e.g, AtOPR3) which is involved in JA biosynthesis (Stintzi and Browse, 2000). Phylogenetic analysis of OPR genes in rice revealed that OPRs in monocots can be divided into five groups: sub. I-V (Li et al., 2011). AtOPRs can be activated by wounding, pathogens, cadmium, and hormone signaling molecules, such as JA, abscisic acid (ABA), salicylic acid (SA), and ethylene (ET) (Fattorini et al., 2018; Zheng et al., 2016). Recently, nine IbOPR genes were identified in sweet potato, and IbOPR2 may play a crucial role in its response to salt stress by participating in JAs synthesis (Li et al., 2024). Seven OPR family genes (CaOPR1-7) were identified from the Capsicum annuum genome, the CaOPR6 was highly similar to AtOPR3, and it played important roles in response to abiotic and biotic stresses (Nie et al., 2022). Five OPR family genes were identified in watermelon, expression analysis revealed that ClOPR genes, except for ClOPR5, were highly expressed in the flower and fruits, furthermore, the findings suggested ClOPR2 and ClOPR4 involved in red-light-induced defense against root-knot nematode (Guang et al., 2021).
The biochemical and physiological functions of OPRs have been reported in monocots such as rice (Wu et al., 2024), cotton (Liu et al., 2020; Hu et al., 2018), tomato (Breithaupt et al., 2006), maize (Zhang et al., 2005), pea (Matsui et al., 2004), wheat (Mou et al., 2019) and tea (Xin et al., 2017). The overexpression of OsOPR1 improved the Cd tolerance of yeast cells by affecting the expression of antioxidant enzyme related genes and reducing Cd content in yeast cells (Wu et al., 2024). In upland cotton (Gossypium hirsutum), GhOPR3 can be phosphorylated by GhCPK33 at threonine-246 (Thr246) in peroxisomes, decreasing the protein level of GhOPR3, which consequently suppresses JA biosynthesis and reduces the resistance of cotton to Verticillium dahliae (Hu et al., 2018). In wheat (Triticum aestivum), the constitutive expression of TaOPR2 can rescue the male sterility phenotype of the AtOPR3 mutant, and these results suggest that TaOPR2 is involved in the biosynthesis of JA (Wang et al., 2016a). In tomato (Solanum lycopersicum), the SiOPR3 mutant plants exhibited increased susceptibility to Botrytis cinerea (Scalschi et al., 2015). In the tea (Camellia sinensis) plants, CsOPR3 plays an important role in JA biosynthesis and defense against herbivorous insects (Xin et al., 2017). In maize (Zea mays), ZmOPR7 and/or ZmOPR8 are highly induced by wounding or treatment with JA, ethylene and ABA (Zhang et al., 2005), and the double mutant ZmOPR7 ZmOPR8 exhibited delayed leaf senescence accompanied by reduced ethylene and ABA levels and a lack of anthocyanin pigmentation in brace roots (Yan et al., 2012). OPRI genes, which are not involved in the octadecanoid pathway (Schaller et al., 1998), are typically upregulated by pathogen invasion, wounding, and oxidative stress (Biesgen and Weiler, 1999; (Biesgen and Weiler, 1999; Strassner et al., 2002; Dong et al., 2013; Agrawal et al., 2003), events associated with ROS acceleration, so OPRIs are claimed to be concerned with antioxidant activity (Fitzpatrick et al., 2003; Trotter et al., 2006; Beynon et al., 2009). To date, the genome-wide identification of OPRs and the function of OPRI genes in grapevine have not been explicitly studied.
Grapevine (Vitis vinifera) is one of the most important economic crops in the world, and its growth and yield are restricted by various biotic and abiotic stresses in the filed. Cu, Zn are essential mineral elements for the normal growth and development of plants (Rehman et al., 2019; Adrees et al., 2015). However, long-term application of copper-based pesticides and fungicides (Manousaki et al., 2008; Nagajyoti et al., 2010), such as Bordeaux mixtures [with the Cu concentration at approximately 1.5 g/L] (Druart et al., 2012), which has been used intensively to control grapevine fungal diseases. Inorganic zinc (Zn) either alone or mixed with agrochemicals, such as fungicides against various diseases and organic fertilizers as sources or nutrients for grapevines is also often used in vineyards (Mirlean et al., 2007; Ramos and López-Acevedo, 2004). The accumulation of Cu, Zn in the vineyard soils has been increasing and often far beyond the required limits for normal grapevine growth (Yruela, 2005; Zalamena et al., 2015). Previous studies have explored the toxic effects of Cu, Zn on grapevines (Toselli et al., 2009; Juang et al., 2012; Miotto et al., 2014). However, the tolerance mechanism of the grapevines in response to Cu, Zn stress is not fully understood.
A genome-wide analysis of OPRs is essential for understanding the functions of the OPR gene family in grapevines and the possible roles of VvOPRs in development and defense against stress should be explored. In the present study, OPR family genes were firstly identified from the grapevine genome and nine VvOPRs which classified into two subfamilies were systematically analyzed. In addition, the OPRI gene VvOPR1 was synthesize by PTDS (PCR-based two-step DNA synthesis quantification reverse-transcription) method, the function and mechanism of the grapevine abiotic stress-inducible VvOPR1 gene was investigated via transgenic plants. Detailed analysis was carried out on the transgenic plants, and the results showed that the heterologous expression of VvOPR1 in plants improved tolerance to Cu, Zn stress. All these results could contribute to screening more potential functional genes to improve tolerance against abiotic stresses in grapevines.
Based on NCBI (http://blast.ncbi.nlm.nih.gov/Blast.cgi) database for the reported OPRs in Arabidopsis (Biesgen and Weiler, 1999; Schaller and Weiler, 1997), the BLASTP and TBLASTP programs were used to search the Grape Genome Browser (http://www.genoscope.cns.fr/externe/GenomeBrowser/Vitis/) and Database (http://genomes.cribi.unipd.it/grape/), TAIR (http://www.arabidopsis.org/) and PlantTFDB (http://rice.plantbiology.msu.edu/) and a local protein database of homologous OPR genes in plants was constructed and OPR family genes in grapevine were obtained. A pairwise comparison of nucleotide and amino acid sequence similarity (%) between VvOPR family members was developed. The physical and chemical properties of the VvOPR family members, including the number of amino acids, molecular weight (MW), and isoelectric point (theoretical pI) were calculated using the online ExPaSy tool (http://www.ExPASy.org). The subcellular localization of the VvOPR proteins was predicted via the TargetP (https://services.healthtech.dtu.dk/services/TargetP-2.0/) and WoLF PSORT (https://wolfpsort.hgc.jp/) online servers.
A paired comparison analysis between VvOPRs was developed via ClustalX software (2.1) (Larkin et al., 2007). OPRs from grapevine, rice and Arabidopsis were used to construct a phylogenetic tree via the neighbor-joining (NJ) method using MEGA software 6.0 (Tamura et al., 2013). The MEME program (http://meme-suite.org/tools/meme) was used to identify the conserved motifs of the VvOPR family members (Bailey et al., 2009). The gene structure display server program (GSDS 2.0) was used to illustrate the exon/intron organization of the VvOPR genes (Guo et al., 2007). The 2 kb upstream sequences of all of the VvOPR genes were considered as the promoters, and these sequences were extracted for the prediction of cis-regulatory elements via PlantCARE (Mou et al., 2019) Each position of the VvOPR gene on the grapevine chromosome was mapped via the software MapInspect.
The plants (grapevine ecotype Maincure Finger, Arabidopsis thaliana ecotype Columbia L. and rice ecotype Wuyun Jing) were stored in our laboratory. The seeds of Arabidopsis and rice were surface-sterilized with 75% ethanol for 1.5 min, followed by 0.5% calcium hypochlorite for 20 min, and then rinsed at least three times with sterile distilled water. The sterilized seeds were placed on Murashige and Skoog (MS) medium with 1% agar and stratified in the dark at 4°C for 48 h. Then these plates were transferred to a controlled environmental chamber at a light intensity of ~120 μmol photons m-2s-1 at 22°C, maintained on a 16/8 h day/night cycle, with 70% relative humidity, and grown horizontally and/or vertically. After the rice seeds germinated, the seedlings were transferred into Hoagland’s liquid medium and/or pots filled with a 9:3:1 mixture of vermiculite/peat moss/perlite under the above conditions.
Cu/Zn treatments (Mikkelsen et al., 2012; Moravcová et al., 2018) were performed on grapevine, VvOPR1-overexpressing Arabidopsis and rice seedlings. Six-month-old grapevine seedlings were grown in Hoagland’s liquid medium supplemented with 10 μM CuSO4·5H2O or 0.1 mM ZnSO4·7H2O for 0, 3, 6, 9, 12 or 24 h. The whole grapevine plants from each treatment and/or the different organs (roots, stems and leaves) were sampled, frozen in liquid N2, and stored at -70 °C. The germination rate was determined when VvOPR1-overexpressing Arabidopsis seedlings were grown horizontally in MS medium supplemented with Cu (150 or 200 μM) or Zn (1.25 or 1.5 mM) for 2 weeks, respectively. The germination rate was calculated as the proportion of seedlings for which the radicle had begun to emerge from the seed coat. The morphological characteristics (root length and fresh weight) were determined at various Cu (50, 100 μM)/Zn (0.75, 0.87 mM) concentrations after VvOPR1-overexpressing Arabidopsis seedlings were oriented vertically in Petri dishes containing MS medium incubated for 14 days. A Cu/Zn solution of deionized water was added to Hoagland’s liquid medium to obtain concentrations of Cu 0, 200, 600 and 900 μM/Zn 0, 2.5, 5.0 and 7.5 mM, and two-week-old transgenic rice seedlings were used to detect Cu/Zn tolerance. Then, 4-week-old transgenic rice seedlings were subjected to Cu (750 μM), Zn (7.5 mM), Cu+Zn (750 μM+7.5 mM) treatment for 5 d, and immediately transferred to liquid nitrogen and then a freezer (-80 °C) until further physical and chemical analyses.
Three seedlings were selected randomly from each treatment, and frozen grapevine and rice tissues (0.2 g) were ground thoroughly in liquid nitrogen using a mortar and pestle, respectively. Grapevine RNA extraction was performed via the modified CTAB method (Wang et al., 2011), and total RNA from VvOPR1-overexpressing rice was extracted from the disrupted tissues using the extraction medium (Qiagen) according to the manufacturer’s instructions. The RNA concentration was measured using a NanoDrop 2000 (Thermo Fisher Scientific, Waltham, MA, USA) and RNA integrity was assessed using an RNA Nano 6000 Assay Kit in conjunction with an Agilent Bioanalyzer 2100 system (Agilent Technologies, Santa Clara, CA, USA). The genomic DNA was removed by incubating the RNA sample in gDNA wipeout buffer (Qiagen) at 42 °C for 2 min before complementary DNA (cDNA) synthesis. First-strand cDNA was synthesized from 5 μg of total RNA with the QuantiTect Reverse Transcription Kit (Qiagen) in a 20 μL reaction volume according to the manufacturer’s instructions and it was stored at -20 °C until it was used in real-time quantitative PCR assays.
Quantitative real-time PCR was performed with a 7500 Fast Real-Time PCR system (Applied Biosystems), using SYBR Green chemistry. PCRs were conducted in a final volume of 20 μL containing 2 μL of a 1:10 dilution of cDNA sample, 10 μL of iQ SYBR Green Supermix, and 0.5 μmol/L of each primer. The thermal cycling conditions included initial denaturation at 95 °C for 30 s, followed by 40 cycles of 95 °C for 5 s, and 65 °C for 15 s for annealing and extension. PCR efficiency was checked for all primers used for gene expression analyses (Supplementary Table S1). Only primers with higher amplification efficiency (>90%) were used in this experiment. The VvActin and OsUBQ5 genes were used as references in grapevine and rice seedlings, respectively. The relative expression levels of the target genes (antioxidative enzyme-encoding genes and ABA- and JA-related genes) were determined via the standard 2−ΔΔCT method of Livak and Schmittgen (2001). All samples were tested in duplicate for the reference genes as well as for the genes of interest.
The full-length of VvOPR1 (XP_002281119) gene was artificially biosynthesized via PTDS method (Xiong et al., 2004). The synthesized VvOPR1 was cloned into the TA cloning vector Simple pMD-18 (pMD18-T) and the high integrity of the construct was confirmed by sequencing. The coding sequence was subsequently excised from the pMD18-T vector and inserted into the pCAMBIA1301 vector under the control of the CaMV 35S promoter. The constructed plant expression vectors were transformed into Agrobacterium (strain EHA105) via the freeze-thaw procedure, and strains containing this vector were transformed into the rice cultivar Wuyun Jing according to the methods of Tang (Tang et al., 2017). VvOPR1-overexpressing Arabidopsis was constructed via the floral dip method (Zhang et al., 2006). Positive transformants were selected via hygromycin (50 μg/ml) resistance and confirmed by PCR using the specific primers. In addition, VvOPR1 was transformed into the Arabidopsis NCED1 mutant which is defective in ABA synthesis to construct VvOPR1/NCED1 lines.
Three VvOPR1-overexpressing A. thaliana seedlings were selected randomly from each treatment, and their morphological characteristics (germination rate, root length and fresh weight) were measured and compared. Four-week-old OT and WT rice seedlings grown in Hoagland’s liquid medium supplemented with Cu (750 μM), Zn (7.5 mM), Cu+Zn (750 μM+7.5 mM) for five days were sampled for physical and chemical analyses. The Chl and Car contents were estimated via the Lichtenthaler (1987) and Wellburn (1994) methods. The root activity and MDA was determined as described by Wang et al. (2016b). The content of H2O2 was determined via the method of Zhou et al. (2018) with slight modifications. The free proline content was determined according to the methods of Bates et al. (1973) and Song et al. (2011). The antioxidants and antioxidant enzymes in the AsA-GSH cycle were detected refer to previous studies (Zhou et al., 2018; Geng et al., 2018; Thounaojam et al., 2012).
Rice leaves were extracted for ABA (Lou and Baldwin, 2003) and JA (Hung and Kao, 2003) analysis following previous studies with slight modifications. The crude extract was centrifuged and passed through polyvinylpyrrolidone column and C18 cartridges to remove plant pigments and other nonpolar compounds. JA extraction was analyzed by GC-MS with a doubly labeled internal standard ([1,2-13C]-JA). The ABA eluates were then concentrated to dryness by vacuum evaporation and resuspended in Tris-buffered saline. Afterwards, ABA was subsequently determined spectrophotometrically at 405 nm with an ABA immunoassay detection kit (model PGR-1; Sigma-Aldrich, St. Louis MO, USA).
For Cu2+/Zn2+ determination, both the aerial and root parts of the Cu/Zn-treated OT and WT seedlings were harvested, carefully rinsed with distilled water, oven dried at 80 °C for two days and ground (Li et al., 2017). The dried plant parts were digested with HNO3 (11N) at 200°C for 10 h, and the digested samples were then diluted with HNO3 (0.1 N) until a clear liquid formed. The filtrate was analyzed via ICP-OES (GBC INTEGRAXL, Australia) and standard solutions from MERCK and Analytika Praha were used to determine the metal content. The Cu2+ content was determined at a wavelength of 324.754 nm, and the Zn2+ content was measured at 213.856 nm (Tanhan et al., 2007). The TF, TI and AR for Cu2+/Zn2+ were detected using the following formulas developed by previous studies (Shu et al., 2002; Khan et al., 2015) based on the dry weight of the seedlings (Supplementary Tables S1, S2).
The treatments of the experiments were arranged in a completely randomized block design, and three replicates were performed for each treatment. The data analysis was performed statistically using analysis of variance (ANOVA) via SPSS 17.0 and the data are presented as the means ± SEs (n=3). A statistically significant difference (p<0.05) was calculated by the least significant difference (LSD) test and is shown in the figures.
Based on NCBI database and literature, 121 homologous OPR genes were identified from 12 plant species representing six major plant lineages, including algae, moss, pteridophyta, gymnosperm, monocot and dicot (Table 1). Nine VvOPRs (OPRs from Vitis vinifera) were identified from grapevine and named VvOPR1~9. The putative VvOPR genes were predicted to encode proteins ranging from 340 to 398 amino acids in length, the molecular weight (Mw) and isoelectric point (pI) of the VvOPR proteins ranged from 37.63 to 43.71 KDa and from 5.45 to 8.23, respectively (Table 2). In terms of amino acid composition, positive amino acids accounted for the greatest percentage, followed by aliphatic amino acids and aromatic acids, and negative amino acids accounted for the lowest percentage (Table 2). Almost all of the VvOPR proteins were predicted to accumulate in the cytoplasm (Cyto). The chromosomal positions of the VvOPR genes revealed that OPR genes were distributed in clusters, the VvOPR genes were distributed across two chromosomes with different densities (Figure 1A), the putative VvOPR3 genes were mapped to chromosome 11, and the other eight VvOPRs were mapped to chromosome 18. VvOPR1~2 mapped to the top of chromosome 18, and VvOPR4~9 mapped to the bottom of chromosome 18.
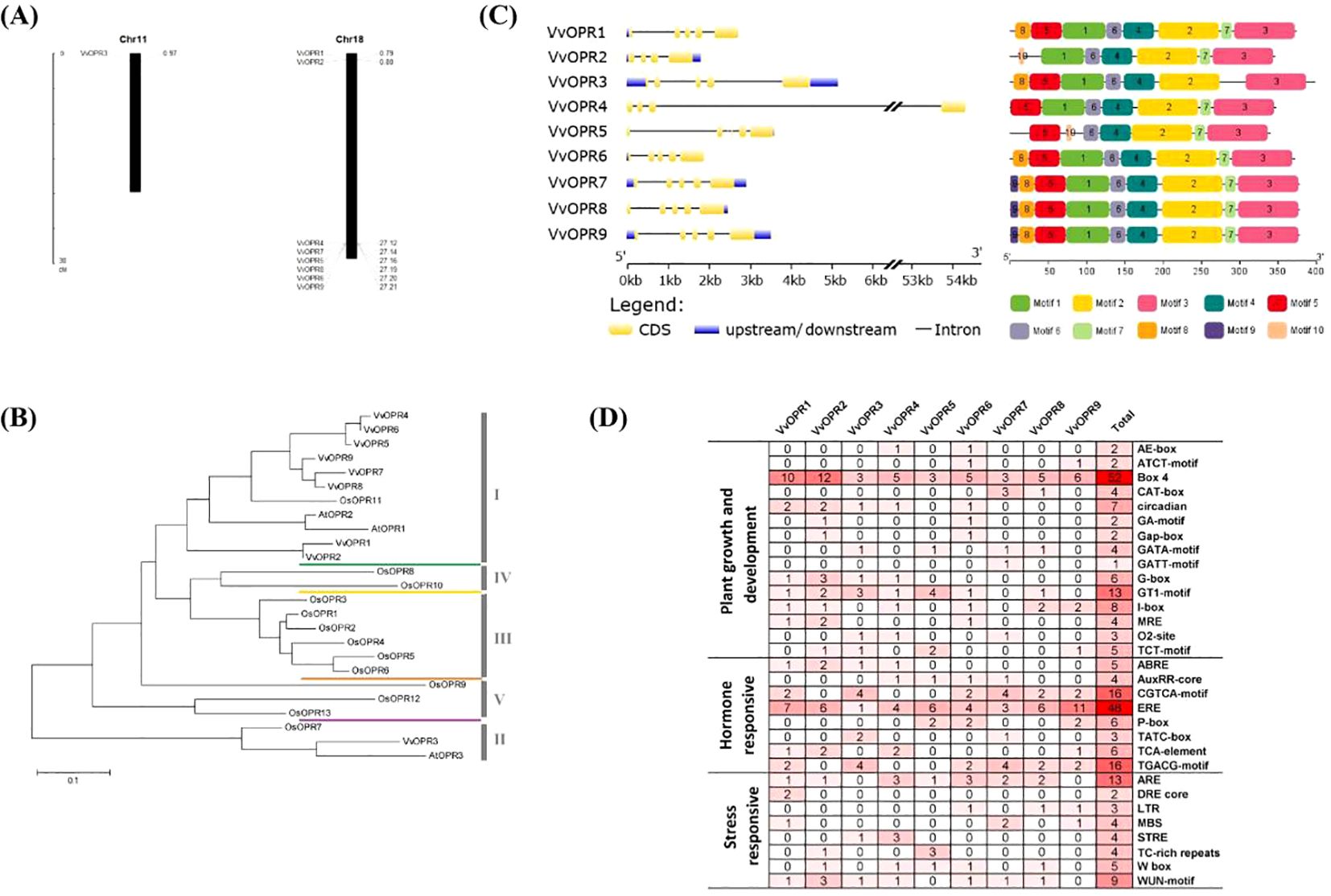
Figure 1. Genome-wide Analysis of VvOPR Family Genes. (A) The chromosomal positions of the OPR genes in grapevine. (B) The phylogenetic tree of OPR proteins in grapevine, Arabidopsis and rice. (C) Gene structure and conserved motifs of VvOPR family members. (D) Frequency and function of cis-regulatory elements (CREs) in the promoter regions of VvOPR genes. 1: Plant growth and development; 2: Hormone responsive; 3: Stress responsive.
To evaluate the evolutionary relationships within the OPR gene family in grapevine, a rooted maximum-likelihood (ML) phylogenetic tree with 25 OPR genes from monocots (rice) and dicots (Arabidopsis and grapevine) (Figure 1B) was established. Phylogenetic analysis revealed that the OPR gene family can be subdivided into five well-conserved subfamilies, and these subfamilies are numbered sub.I to V. All OPR genes from grapevine, Arabidopsis and several OPRs from rice were grouped into sub.I and II, indicating that all OPR genes from angiosperms shared a common ancestor before the divergence between dicots and monocots. Several OPRs from rice were grouped into sub.III to V, showing that lineage-specific expansion and divergence events occurred in monocots after divergence from dicots and sub.III to V were generated exclusively in monocots. To explore different selective constraints on duplicated VvOPR genes, the Ka (synonymous nucleotide substitution), Ks (non-synonymous nucleotide substitution) and Ka/Ks ratios for each pair of duplicated VvOPR genes were calculated, and seven duplicated pairs of paralogous genes were identified in the nine VvOPRs. The results showed that the ratios of Ka/Ks for two duplicated pairs were <1, ranging from 0.1986 to 0.6999 (Table 3). These results revealed that the gene duplication events occurred in these genes and that the functions of the duplicated genes did not diverge during genome evolution after the duplication events.
To further detect the origin and evolution of the VvOPR family genes, the nucleotide and amino acid sequences of the nine VvOPRs were blasted via pairwise alignment, respectively (Table 4). The results revealed that the level of nucleotide and amino acid sequence identity among the VvOPRs varied from 28.3% to 98.1% and 44.4% to 97.4%, respectively. Five pairs of VvOPRs showed extremely high homology (the level of sequence identity between VvOPRs was above 90%), as follows: VvOPR4/VvOPR6 > VvOPR1/VvOPR2 > VvOPR7/VvOPR8 > VvOPR8/VvOPR9 > VvOPR7/VvOPR9. The level of nucleotide/amino acid identity between VvOPR3 and the other eight VvOPRs varied from 28.3% to 36.7%/44.4% to 51.9%. These results indicated that VvOPR3 may have undergone rapid evolution and functioned differently from the other eight VvOPR genes.
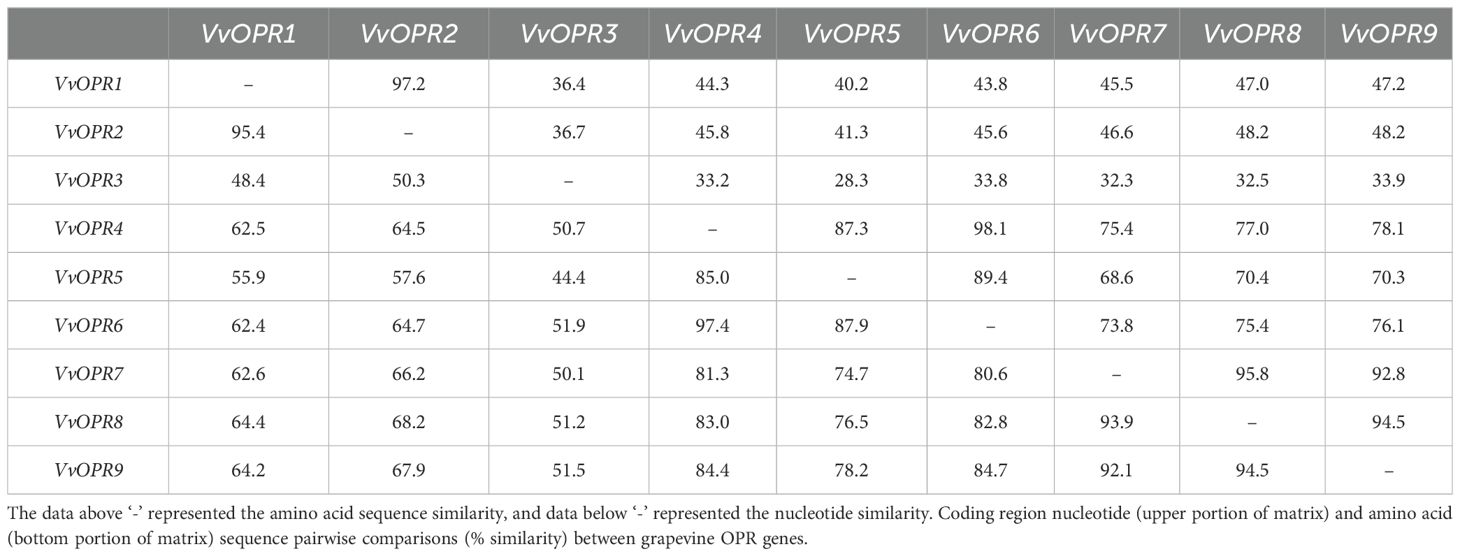
Table 4. Pairwise comparison of nucleotide and amino acid sequence identity (%) between OPR family members in grapevine.
An exon-intron analysis was performed by comparing the predicted coding sequence (CDS) with the sequence of the VvOPR genes. As shown in Figure 1C, the VvOPR family members presented a highly conserved exon-intron structure, which contained 4~6 exons and 3~5 introns. VvOPR4 shared the longest intron sequence, an untranslated region of 53 kb. In addition, the nine VvOPR proteins shared at least four conserved motifs (motifs 6, 4, 2 and 3), which showed the same alignment. VvOPR7, VvOPR8 and VvOPR9 were completely consistent and motif 9 is characteristic of the three VvOPR proteins. Motif 7 was only deleted in VvOPR3. Phylogenetic analysis revealed that VvOPR members in the same branch were structurally conserved, indicating that motif 7 may play an important role in the evolution and function of VvOPRs. Taken together, the fact that the VvOPR proteins not only share very similar intron/exon structures but also contain common motifs supports their close evolutionary relationships and membership in the same subfamily. All of these results suggested that the strategies of VvOPR genes classification were relevant and reliable.
To further understand the biological regulation of the VvOPR genes, the cis-regulatory elements (CREs) in the upstream of these genes were identified via the PlantCare database. The cis-elements included plant growth and development elements, hormone responsive elements, and environmental stress responsive elements and so on (Figure 1D). The frequency of these elements in the regulatory region of each corresponding gene, as well as their overall frequency in family members, is very diverse. Most CREs involved in plant growth and development, such as Box 4 (52 times), GT1-motif (13 times) and I-box (8 times) are related to light-responsive elements. Hormone-responsive elements include those involved in ethylene response (ERE) and methyl jasmonate (MeJA). Classic plant hormones such as SA, ETH and MeJA are involved in the regulation and integration of plant immune responses against pests and pathogens.
Different tissues (roots, stems and leaves) of six-month-old grapevine (Maincure Finger) seedlings were sampled, and real-time quantitative PCR (qRT-PCR) was used to examine the VvOPR1 expression profile. The results revealed that various expression levels of VvOPR1 could be detected in all of the tissues tested, and the expression level of VvOPR1 was the highest in the roots and the lowest in the leaves (Figure 2A). As shown in Figure 2B, under Cu, Zn stress, VvOPR1 expression rapidly decreased throughout the 3 h period, and the expression of VvOPR1 was higher under Zn stress than under Cu stress. Then, VvOPR1 expression increased and peaked at 6 h, and the expression of VvOPR1 level was higher under Cu stress than under Zn stress from 6 h to 12 h, the plateau lasted from 9 h to 12 h testing period. Exposure to excess Cu, Zn stress could stimulate the overproduction of reactive oxygen species (ROS) and abscisic acid (ABA), the former impairs antioxidant defense systems and the latter triggers downstream stress-responsive pathways (Mittler et al., 2011). Here, the expression level of VvOPR1 almost mirrored the same trend after exposure to 10 mM H2O2 and 200 μM ABA, the expression level of VvOPR1 increased gradually from 0 h to 12 h under the treatment conditions (Figures 2C, D). All these results suggest that the expression pattern of VvOPR1 is tissue-specific and that its expression can be induced by various stresses.
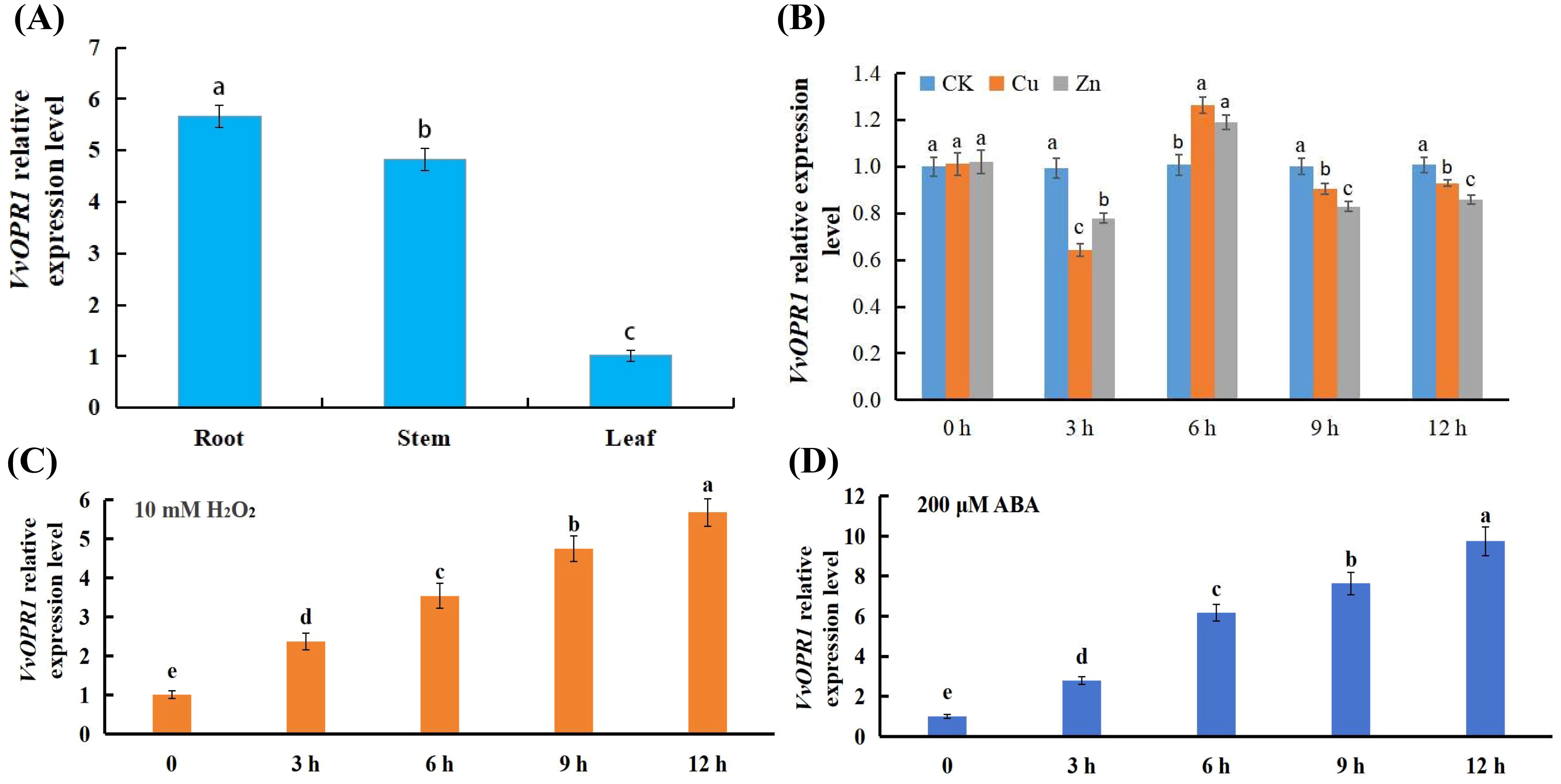
Figure 2. Expression analysis of the VvOPR1 gene and VvOPR1 is induced by various abiotic stresses in grapevine leaves. (A) The qRT-PCR analysis of the expression level of VvOPR1 in different tissues of grapevine during seedling stage. (B-D) The qRT-PCR analysis of VvOPR1 expression in grapevine leaves subjected to (B) Cu, Zn; (C) 10 mM H2O2; (D) 200 μM ABA stress, respectively. Different lowercase letters indicate significant differences (P<0.05) in the expression level between the different tissues of grapevine (A) and between different time points subjected to various stress treatments, respectively (B-D).
VvOPR1 was synthesized via the PTDS method and transferred into both Arabidopsis and rice (Figure 3A). Several independent lines of transgenic seedlings (T1 generation) were subsequently generated. After selecting for hygromycin resistance, PCR was performed (T3 generation) (Figure 3B), the putative transgenic plants were confirmed, and several transgenic T3 plant lines (OT-1, OT-2 and OT-3) were chosen for further experiments. The phenotypes of the transgenic plants (T3 generation) were similar to those of the wild-type (WT) plants both on MS plates and in soil pots. These findings indicated that overexpression of VvOPR1 in Arabidopsis and rice caused no visible morphological changes.
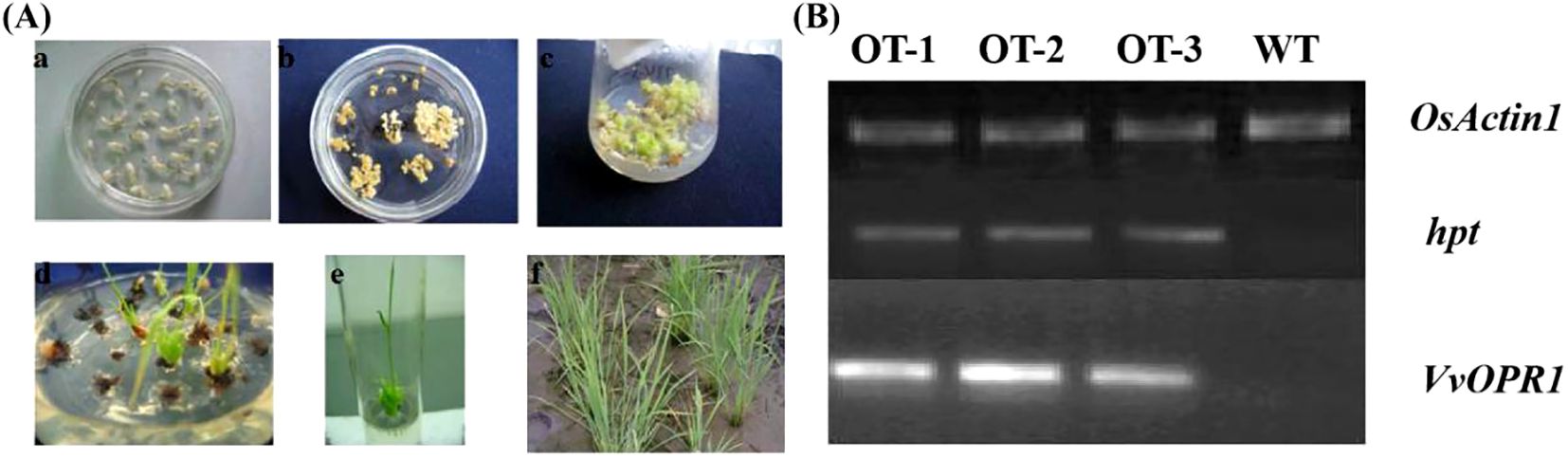

Figure 3. The transformation and VvOPR1 PCR in rice seedlings. (A) The procedure of Agrobacterium-mediated genetic transformation of rice calli, including: (a) Calli induction; (b) First screening; (c) The second screening; (d) Differentiation; (e) Taking root; (f) Transplantation. (B) RT-PCR analysis of the VvOPR1 gene fragments from the transgenic (OT-1, OT-2, and OT-3) and wild-type (WT) plants, OsActin1 gene was used as a reference.
The performance of WT and OT Arabidopsis seedlings under both normal and Cu and Zn stress conditions was compared to investigate whether the overexpression of VvOPR1 could potentiate the tolerance of OTs to Cu, Zn stress (Figures 4A-E). No significant difference in growth performance was detected between the two types of seedlings in the absence of stress, but all seedlings from each type were obviously suppressed when they were grown for three weeks in the medium containing Cu, Zn, and the suppression was more severe in the WT seedlings than in the OT seedlings (Figures 4A-E). The germination rate of OT Arabidopsis seeds was nearly twofold higher compared with that of WT seeds on medium supplemented with 150, 200 μM Cu. A similar pattern was observed for OT seeds under 1.2 mM Zn stress condition (Figures 4A, B). Exposure to Zn stress, compared with that of WT seedlings, the root length of OT seedlings increased by 32% to 46%, whereas the fresh weight increased by 56% to 80% (Figures 4C, D, E). Under Cu stress, the root length increased by 30% in the 50 μM Cu treatment, however, the root length increased from 50% to 78% in the OT lines compared with that in the WT plants under the 150 μM Cu treatment (Figures 4C, D), whereas the fresh weight increased by from 20% to 45% (Figures 4C, E).
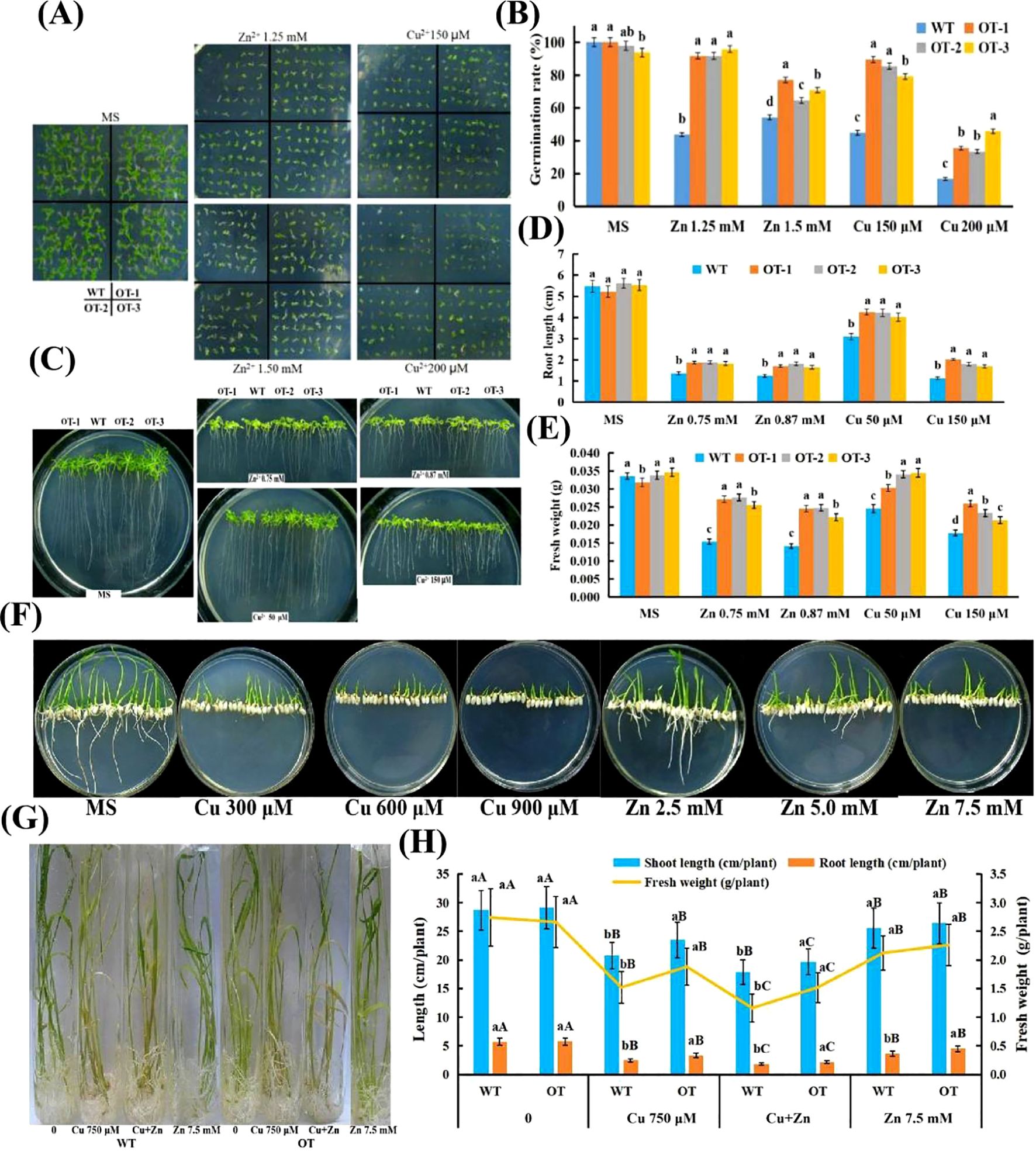
Figure 4. Performance of WT and OT Arabidopsis/rice under normal and Cu, Zn stress conditions. (A) Left panel, seedlings reared under normal conditions for three weeks; right panel, seedlings exposed to 125, 150 mM Zn and 150, 200 μM Cu for three weeks, respectively. (B) The germination rates of plants after grown horizontally on the MS medium with different Cu, Zn concentrations for four weeks. (C) VvOPR1 over-expressing and WT plants grown vertically in MS medium with or without Cu, Zn for three weeks, respectively. (D) Comparison of root length between OT Arabidopsis linesand WT seedlings presented in panel (C). (E) Comparison of fresh weight between OT lines and WT seedlings presented in panel (C). (F) Visual aspects of Cu/Zn-toxicity on rice seedlings. WT and OT rice seedlings grown vertically for two weeks on MS medium with different Cu, Zn concentrations. (G) Visual aspects of Cu, Cu+Zn and Zn-toxicity on rice seedlings. The four-week-old WT and OT rice seedlings grown in Hoagland’s liquid medium with 750 μM Cu, 750 μM Cu+7.5 mM Zn and 7.5 mM Zn for five days. (H) Effects of Cu, Cu+Zn and Zn on shoot length, root length and fresh weight of WT and OT rice seedlings. Each data point is the mean value ± SD of three replicates. The statistical significance was determined by Duncan’s multiple comparison tests. Different lowercase letters indicate significant differences (P<0.05) between WT and OT rice seedlings in the same treatment, and different capital letters indicate significant difference (P<0.05)among different treatments in WT and OT, respectively.
The effects of Cu, Zn stress on OT rice seedlings grown in MS medium and Hoagland’s liquid medium were detected (Figures 4F–H). When treated with Cu, WT rice showed neither any roots nor any cotyledons, while several cotyledons were detected in OT grown in MS medium (Figure 4F). When Zn stress was imposed on the rice seedlings, all of the WT presented no any roots and much fewer cotyledons than the OT did, whereas all of the Zn-treated OT seedlings showed several leaves and roots that became shorter with increasing Zn concentration in the MS medium (Figure 4F). After exposure to Hoagland’s liquid medium containing Cu, Zn for 5 days, several morphological disturbances were detected in the OT and WT. Visible toxicity symptoms such as chlorosis, necrosis and rolling of the leaves were more obvious upon Cu and Cu+Zn impositions (Figure 4G), and the roots of the Cu- and Cu+Zn-stressed seedlings became reddish, and their health deteriorated compared with that of the control seedlings (Figure 4G). Both the OT and WT seedlings exhibited a decrease in shoot length, root length and fresh weight significantly when they were exposed to 750 μM Cu solution (Figure 4G); to some degree, the greatest reduction in shoot length, root length and fresh weight occurred in the seedlings subjected to Cu+Zn stress (Figures 4G, H). Compared with those grown under Cu, Cu+Zn conditions, the Zn-stressed seedlings presented fewer toxic effects and retained a similar leaf color (Figures 4G, H). Together, compared with the WT seedlings, the OT presented significantly higher shoot length, root length and fresh weight under stress conditions, indicating that the tolerance of the OT seedlings to Cu, Zn stress was greater than that of the WT seedlings.
Chlorophyll fluorescence has been routinely used to monitor the photosynthetic performance of plants noninvasively and to screen plants for tolerance to environmental stresses (Baker and Rosenqvist, 2004). To determine whether the chlorophyll content of OT is affected by Cu, Zn stress, the chlorophyll (Chla) and carotenoid (Car) contents were detected. As shown in Table 5, there was no significant difference in the Chla, Chlb, Chla/Chlb or Car contents between WT and OT under normal conditions. In contrast, following the Cu, Zn and Cu+Zn treatments, the Chla, Chlb, Chla/Chlb and Car contents decreased in both the WT and OT seedlings compared with those under the control conditions, but the Chla, Chlb and Chla/Chlb values in the leaves of the OT were significantly higher than those in the WT, whereas the Car content in the OT was lower than that in the WT. The Chla, Chlb and Car contents in the leaves of the rice seedlings treated with 7.5 mM Zn were greater than those in the rice lines treated with 0.75 mM Cu and 0.75 mM Cu+7.5 mM Zn. Cu, Zn damage to the OT plants was evaluated both as photosystem II (PSII) stability and as injury to the whole plant. Fv/Fm (variable fluorescence/maximal fluorescence) was used to estimate the quantum yield of PSII photochemistry. The F0 (minimal fluorescence), Fm and Fv of WT and OT plants were measured, and no significant difference was detected between WT and OT plants under control condition. The values of OT were higher than those of WT seedlings after treatment with 0.75 mM Cu, 7.5 mM Zn and 0.75 mM Cu+7.5 mM Zn, respectively (Table 6). In particular, the combined Cu, Zn treatment markedly inhibited PSII, as indicated by a decrease in the Fv/Fm value. Together, these results are consistent with the hypothesis that plants overexpressing VvOPR1 are more tolerant of photoinhibition than are WT seedlings.
The roots were affected the most under heavy metal stress (Figure 4), and the OT and WT seedlings experienced different degrees of damage when exposed to Cu, Zn stress (Figure 5). Compared with that of the control group, the root activity of the OT-treated group decreased by 14.5%/30.8% when it was subjected to Zn/Cu stress, and there were significant differences between the two genotypes. However, the root activity reduced to the maximum level when these seedlings were subjected to Cu+Zn stress, and only the activity of the OT-2 line was significantly different from that of the WT line (Figure 5). Malondialdehyde (MDA) and H2O2 were induced by Cu, Zn stress (Figure 5), and the MDA content was remarkably reduced by 21.2%, 18.9% and 12.13%, respectively, compared with that of the WT with Zn, Cu and Cu+Zn treatment (Figure 5), whereas the H2O2 content increased by 69%, 134% and 173% respectively, under Zn, Cu and Cu+Zn stress compared with that the CK (Figure 5). In addition, the proline content was significantly higher in the OT (increased by 24.14%, 23.05% and 26.58%) than that in the WT under the stressed conditions (Cu, Zn and Cu+Zn stress, respectively) (Figure 5).
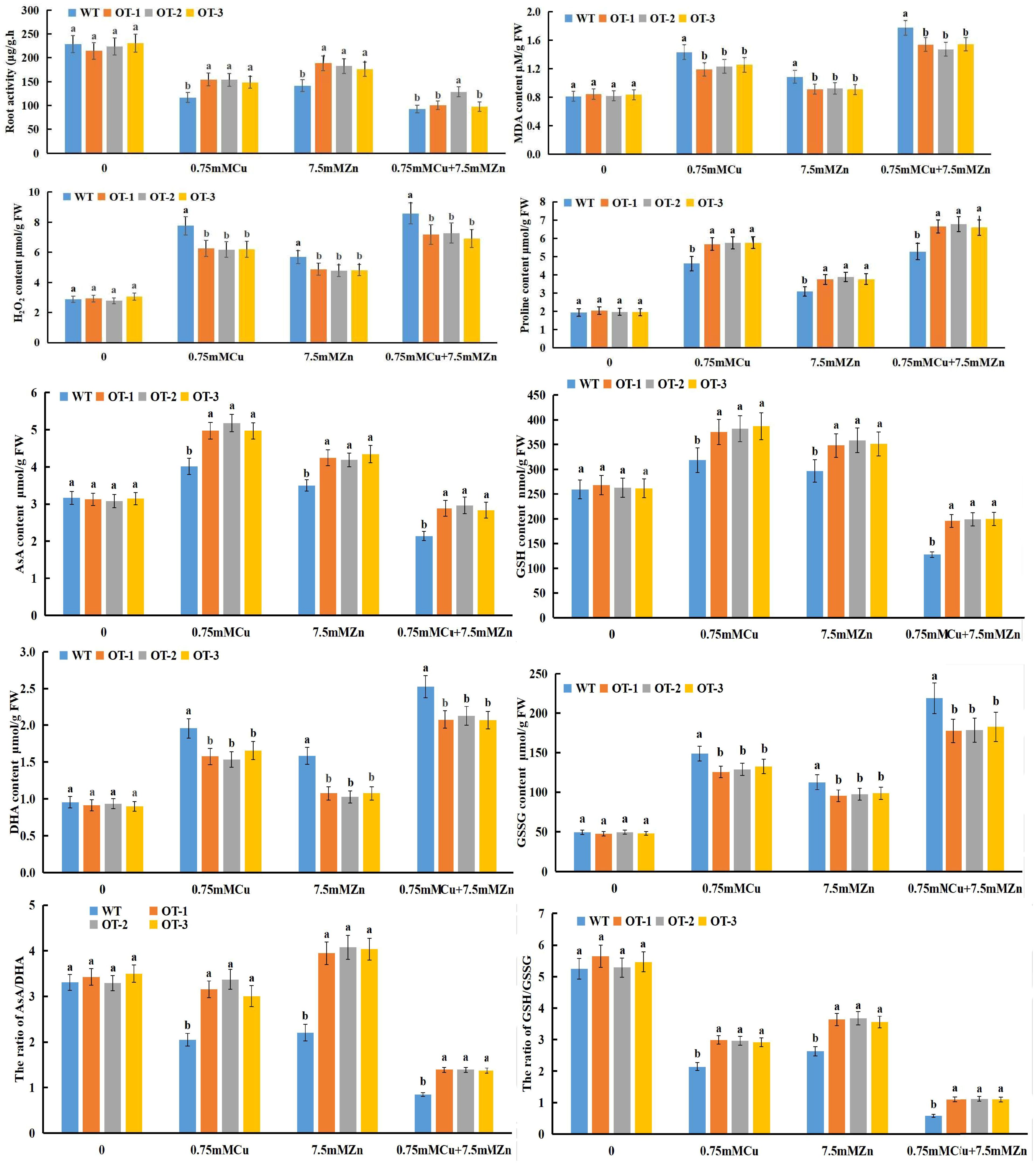
Figure 5. Quantitative analysis of various physiological indexes, including root activity, MDA, H2O2, Proline, AsA, GSH, DHA, GSSG content and AsA/DHA, GSH/GSSG ratio in WT and OT rice under normal and Cu, Zn stress conditions. Different letters indicate significant differences (p<0.05) between the OT and WT plants under the same conditions.
The response of the ascorbate-glutathione (AsA-GSH) cycle to Cu, Zn stress in the OT is characterized in Figure 5. Compared with that under the control conditions, the AsA content of the OT/WT increased by 34.41%/17.72% and 19.96%/10.05% during Cu, Zn stress, respectively, whereas it decreased by 18.45%/34.42% when it was subjected to Cu+Zn stress. Compared with the WT, all of the OTs presented higher AsA level under the three stress conditions, and the greatest increase AsA content was detected in the Cu+Zn treatment, reached 21.78% (Figure 5). As represented in Figure 5, GSH and AsA contents had similar results, decreasing during Cu+Zn exposure, and increasing with Cu or Zn individual treatment. As shown in Figure 5, the Dehydroascorbate (DHA), Glutathione disulfide (GSSG) contents of the OT and WT seedlings increased under Cu, Zn and Cu+Zn stressful conditions, and the greatest increase was found under the Cu+Zn treatment. However, compared with the WT seedlings, the OT seedlings presented significantly lower DHA and GSSG contents under all three stress conditions. The changes in the AsA, GSH, DHA and GSSG contents of the OT and WT seedlings were conductive to reducing the AsA/DHA and GSH/GSSG ratios (Figure 5) after exposure to Cu, Zn single or combined stresses, except for the increased ratio of AsA/DHA in the OT seedlings under Zn stress compared with those under normal conditions.
The activities of antioxidative enzymes such as superoxide dismutase (SOD, EC1.15.1.1), peroxidase (POD, EC1.11.1.7), catalase (CAT, EC1.11.1.6), glutathioneperoxidase (GPX), ascorbate peroxidase (APX), glutathione reductase (GR), monodehydroascorbate reductase (MDHAR) and dehydroascorbate reductase (DHAR) were measured in the OTs and WTs exposed to Cu, Zn and Cu+Zn stress conditions (Figure 6A). In both the WT and OT rice seedlings, the activities of SOD and POD in response to the Cu, Zn and Cu+Zn treatments improved compared with those in the control, and significantly greater activities were detected in the OT than in the WT (Figure 6A) among the three stress treatments. Notably, there was a clear reduction in the CAT activity of the OT and WT plants under all three stress conditions (Figure 6A). This might be because the removal of H2O2 is carried out by APX and GPX, so an increase in CAT activity is not required. Compared with those of the control, the same increasing tendencies were observed for GPX, APX, GR, MDHAR and DHAR activities in both the OT and WT seedlings exposed to Cu, Zn stress, whereas a decreasing tendency was detected under Cu+Zn stress condition (Figure 6A). In conclusion, all of the results revealed greater activities of antioxidant enzymes in the OT lines than in the WT. These findings suggested that OT strengthened the ability of the plants to scavenge ROS and maintain ROS homeostasis by increasing the activities of antioxidant enzymes.
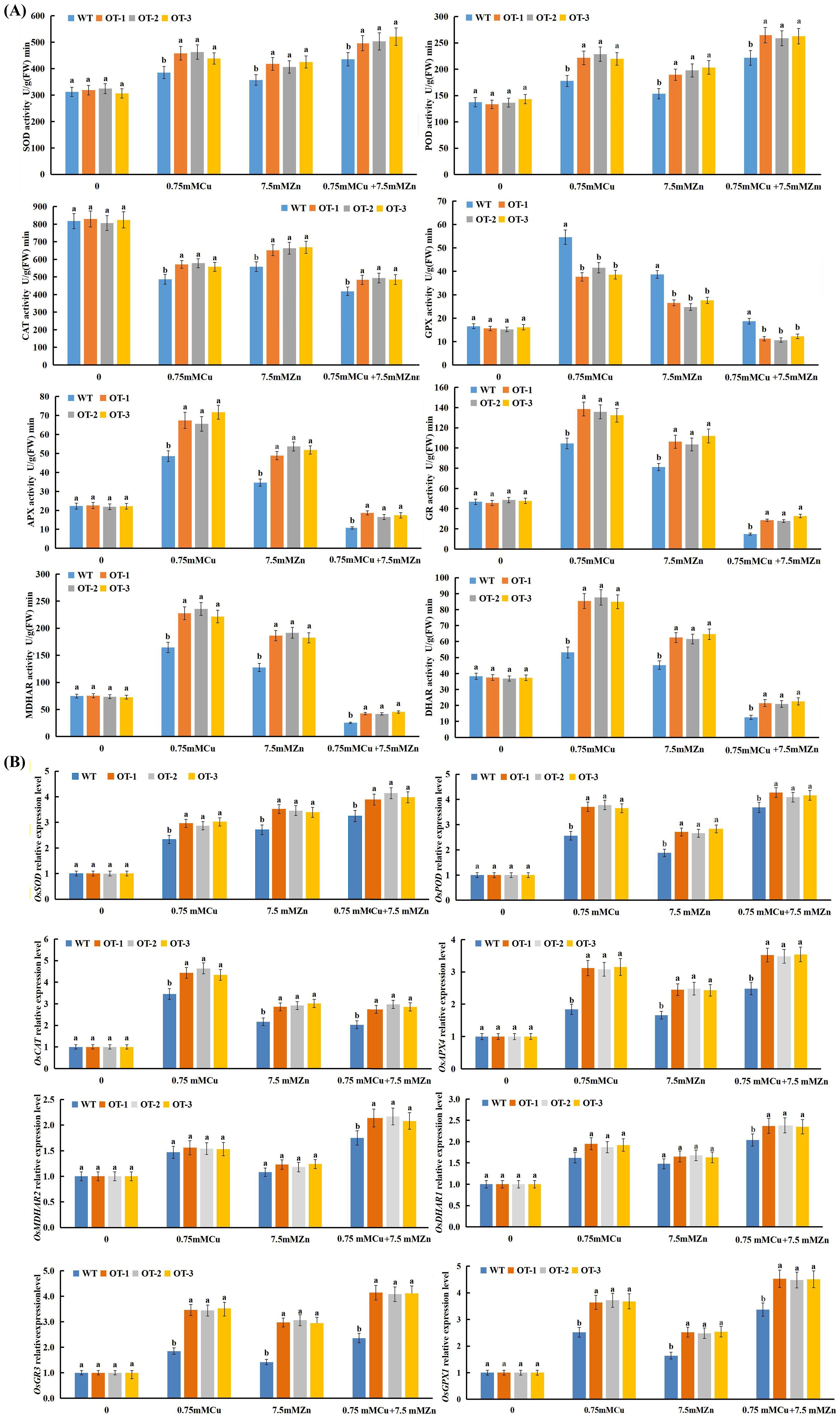
Figure 6. Effect of Cu, Zn on the activities of antioxidative enzymes (A) and on the expression level of antioxidative enzyme-encoding genes (B). Data presented are mean ± SE (n=3), different letters indicate significant differences (p<0.05) between the OT and WT plants under the same conditions.
To evaluate the regulatory role of VvOPR1 in gene expression, through which it could improve tolerance to Cu, Zn stress, several antioxidative enzyme-encoding genes were selected for qRT-PCR analysis (Figure 6B). The expression of all of the genes showed no significant difference between the WT and OT seedlings under controlled conditions. Conversely, the expression levels of all antioxidative enzyme-encoding genes were significantly greater in the OT and WT seedlings than in the control seedlings when they were exposed to Cu, Zn, Cu+Zn stress. The expression levels of OsSOD, OsPOD, OsCAT, OsAPX4, OsGR3 and OsGPX1 in the OTs were significantly higher than those in the WT seedlings under either Cu, Zn or Cu+Zn stress conditions (Figure 6B). OsSOD and OsPOD showed the highest expression levels under Cu+Zn stress condition (Figure 6B), while the highest expression level of OsCAT was observed in the Cu-treated seedlings (Figure 6B). The expression pattern of antioxidative enzyme-encoding genes involved in the AsA-GSH cycle (OsAPX4, OsMDHAR2, OsDHAR1, OsGR3 and OsGPX1) was similar to that of OsPOD when exposed to Cu, Zn, Cu+Zn stresses (Figure 6B), and the highest expression of these genes was also induced by Cu+Zn stress. Compared with those in the control seedlings, the expression levels of OsMDHAR2 and OsGR3 were higher in the OT and WT seedlings under Cu, Zn stress conditions, but no significant differences were detected between the OT and WT seedlings (Figure 6B).
To detect whether the overexpression of VvOPR1 would induce ABA or JA in OT under Cu, Zn stress conditions, the endogenous ABA and JA levels in OT and WT seedlings were determined. Under control conditions, no significant difference in ABA or JA content was detected between the OT and WT seedlings, whereas the levels of ABA and JA were elevated in both plant types after exposure to Cu, Zn, Cu+Zn stress, and the highest elevation was detected in the Cu+Zn treatment (Figure 7A). Specifically, significant differences in ABA levels were detected between the OT and WT under all stressed conditions, whereas no significant differences in JA levels were detected under the same stress conditions.
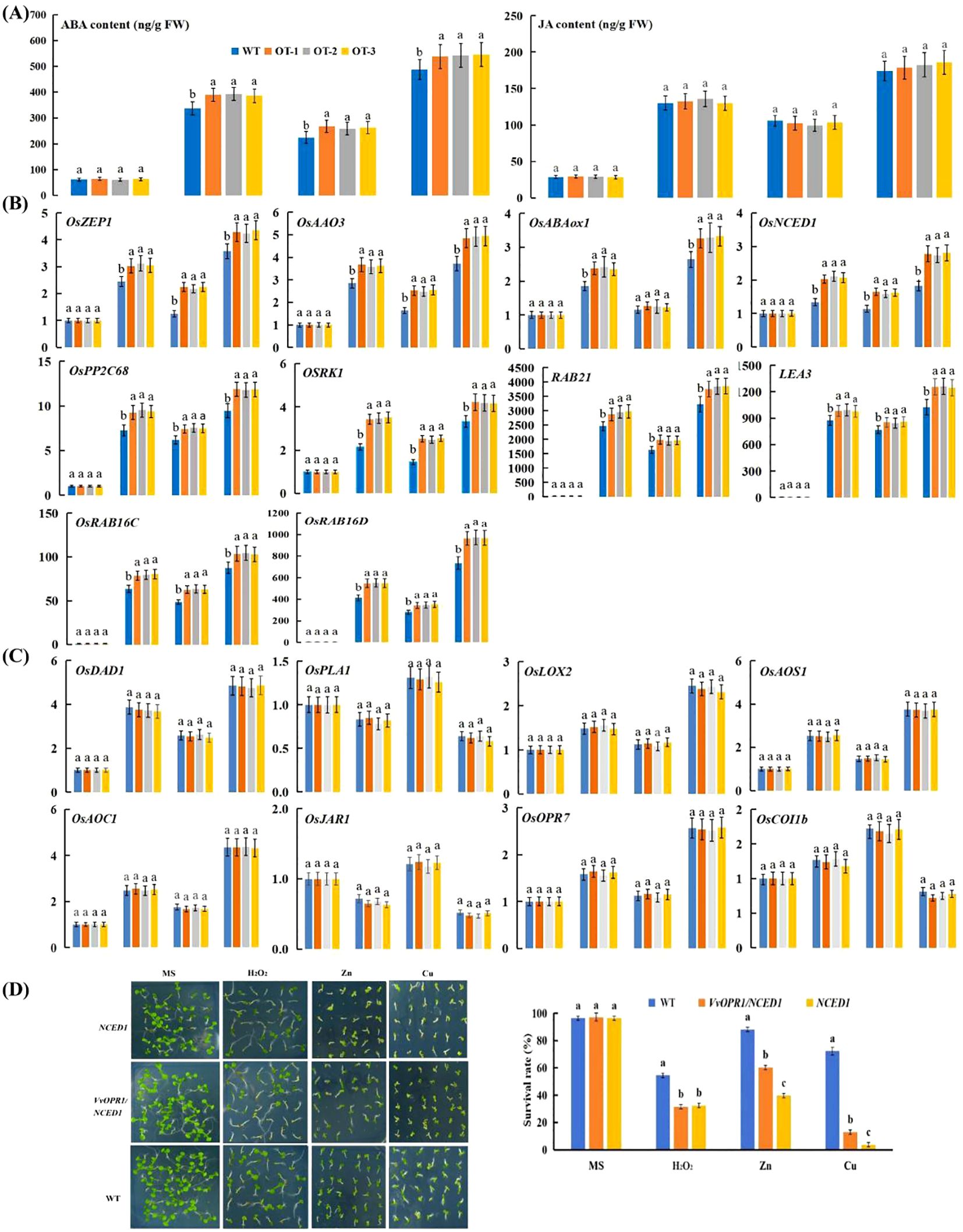
Figure 7. ABA and JA accumulation and the relative expression of ABA and JA- related genes under Cu, Zn stress conditions for five days. (A) ABA and JA content in normal and Cu, Zn-treated seedlings. (B) The qRT-PCR analysis of the expression of ABA-related genes. (C) The qRT-PCR analysis of the expression of JA-related gens. (D) The performance of WT, VvOPR1/NCED1 and NCED1 mutant Arabidopsis seedlings exposure to 1 mM H2O2, 500 μM Zn and 75 μM Cu stress for two weeks, respectively and the survival rate of seedlings. Each index represents an average of three replicates, and values are means ± standard deviation (SD); and different letters indicate significant differences (p<0.05) in the same index. WT: the wild-type Arabidopsis seedlings, NCED1: Arabidopsis NCED1 mutant, VvOPR1/NCED1: Arabidopsis NCED1 mutant lines heterologously expressing VvOPR1.
To further elucidate the molecular mechanism underlying the resistance of OT to Cu, Zn stress, the expression levels of ABA biosynthetic genes (OsNCED1, OsZEP1 and OsAAO3), catabolic genes (OsABA8ox1), signaling genes (OsPP2C68 and OSRK1) and responsive genes (RAB21, LEA3, RAB16C and RAB16D) and JA-related (OsDAD1, OsPLA1, OsLOX2, OsAOS1, OsAOC, OsOPR7, OsJAR1 and OsCOI1b) genes were assayed. The expression levels of ABA-related genes (except for the expression of OsABA8ox1 under Zn stress conditions) were significantly higher in the OT lines than in the WT seedlings under Cu, Zn stress conditions (Figure 7B). As shown in Figure 7C, the expression level of OsCOI1b in the OT and WT elevated after they were exposed to Cu, Zn stress, but reduced when they were subjected to Cu+Zn stress. The OsPLA1 and OsJAR1 genes were downregulated under Cu, Cu+Zn stress conditions but upregulated with Zn treatment. Other JA-related genes, such as OsDAD1, OsLOX2, OsAOS1, OsAOC and OsOPR7 exhibited similar expression patterns: Cu, Zn and Cu+Zn stress all elevated the expression of these genes, with the smallest increase occurring in Zn-treated seedlings, then the expression of these genes increased in the Cu-treated seedlings and peaked under Cu+Zn stress conditions.
To further detect the role of ABA in VvOPR1-enhanced Cu, Zn stress, VvOPR1 was transformed into the Arabidopsis NCED1 mutant, which is defective in ABA synthesis to construct VvOPR1/NCED1 lines (Figure 7D). Similar results were detected in the WT, VvOPR1/NCED1 and NCED1 lines under nonstressed conditions. The survival rate of the NCED1 mutant was seriously restricted by Cu, Zn stress, and the survival rate was lower by more than 60% compared with that of WT seedlings subjected to Cu stress (Figure 7D). VvOPR1 overexpression significantly increased the survival rate of seedlings subjected to Cu, Zn stress, and the survival rate of VvOPR1/NCED1 line was 12.96% to 3.7%, 60.19% to 39.81% to those of NCED1 line under Cu, Zn stress, respectively (Figure 7D). However, no significant difference was detected between the NCED1 mutant and VvOPR1/NCED1 line under H2O2 stress.
After exposure to 0.75 mM Cu and 7.5 mM Zn for 5 days, the contents of heavy metals drastically increased, and the concentrations of Cu2+/Zn2+ were significantly lower in the OT than in the WT (Tables 7, 8). Compared with those in the control seedlings, the Cu2+ content in the 0.75 mM Cu-treated WT and OT seedlings was nearly 2444- and 2364-fold greater, respectively, than that in the control; whereas the Cu2+ content was 2190- and 206-fold greater, respectively under Cu+Zn stress condition. A great increase in Zn2+ content was also detected in the Zn-treated seedlings, compared with the control, 849- and 788-fold increases in Zn2+ content were detected in the WT and OT seedlings under 7.5 mM Zn stress condition; 724- and 679-fold increases in Zn2+ content were detected in the WT and OT seedlings under Cu+Zn stress condition.
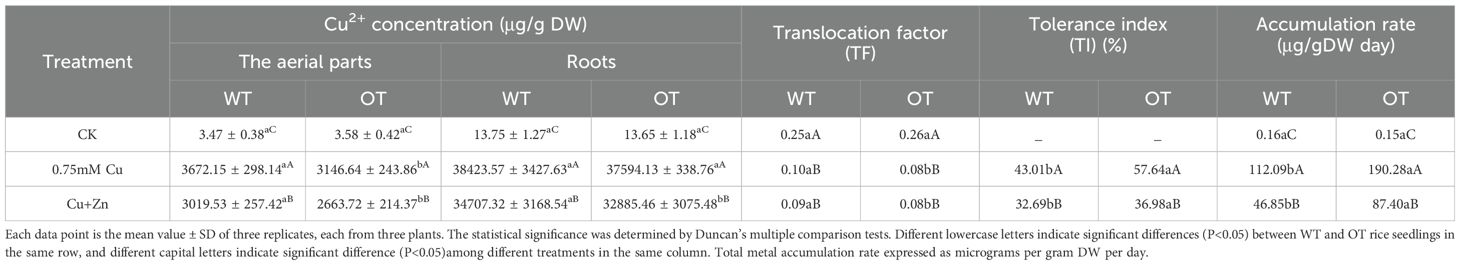
Table 7. Uptake and concentration of Cu2+ in the aerial parts and roots of OT and WT rice seedlings after five days treatment.
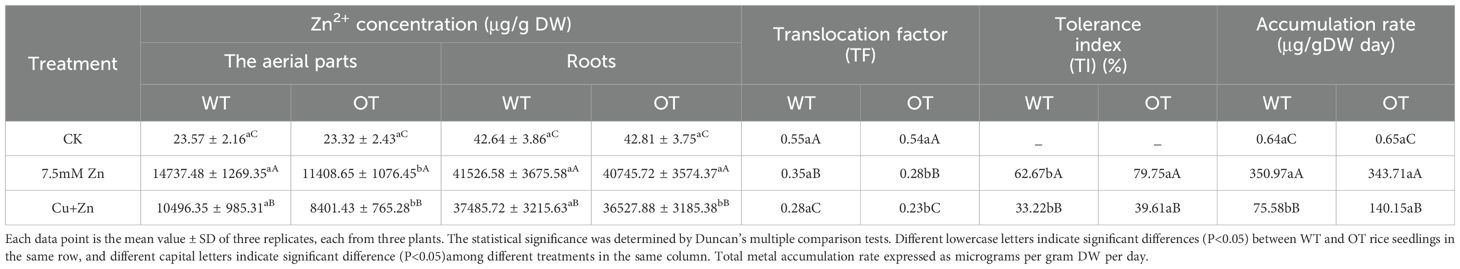
Table 8. Uptake and concentration of Zn2+ in the aerial parts and roots of OT and WT rice seedlings after five days treatment.
The distribution of Cu2+/Zn2+ taken up was not homogeneous, and a much higher proportion remained in the roots than was transported into the aerial parts. The proportion of Cu2+ in the roots versus the aerial parts was 79.85% versus 20.15%, 91.28% versus 8.72% and 92% versus 8% in WT under the control, Cu and Cu+Zn stress conditions, respectively; 79.22% versus 20.78%, 92.28% versus 7.72% and 92.51% versus 7.49% in the OT under the same conditions. In contrast to the Cu2+ distribution, a higher Zn2+ proportion was detected in the aerial parts, ranging from 18.7% (the proportion of Zn2+ in the OT treated with Cu+Zn stress) to 35.6% (the proportion of Zn2+ in the WT under control conditions).
In terms of the translocation factor (TF), under Cu, Zn stress conditions, both the OT and WT lines presented the reduced TFs compared with those under the control conditions. As a consequence of VvOPR1 overexpression, the distribution of Cu2+, Zn2+ in rice seedlings was altered, reducing the proportion of Cu2+, Zn2+ accumulation in the aerial parts of the OT and WT seedlings under Cu, Zn stress. TF values of Cu2+, Zn2+ in the OT were lower than those in the WT, but the TF value of Cu2+ was far below the TF value of Zn2+ (Tables 7, 8). The tolerance index (TI) of OT, which is based on root length, was higher than that of WT under all three treatment conditions, indicating that the sensitivity of WT to Cu, Zn stress was greater than that of OT. When exposed to 0.75 mM Cu and 0.75 mM Cu+7.5 mM Zn, the accumulation rates (ARs) of Cu2+ in the OT lines were 1.7- and 1.87-fold higher than those in the WT, respectively. While AR in OT was 0.98 and 1.85 fold for Zn2+ compared to WT when these seedlings were treated with 7.5 mM Zn or 0.75 mM Cu+7.5 mM Zn, respectively.
OPRs are multigene families that can be identified from various plants, and comparative genomics approaches have been used to analyze OPR gene families in different plant species (Liu et al., 2020; Stintzi and Browse, 2000; Li et al., 2011). However, no comprehensive studies have focused on the grapevine OPR family, and this study was the first to systematically investigate the VvOPR family.
In this study, nine VvOPRs were identified in grapevine. The rooted maximum-likelihood phylogenetic tree revealed two OPR subfamilies in grapevine following the classification system (Montanini et al., 2007), and all of the VvOPRs except VvOPR3 were clustered with sub.I (Figure 1B). VvOPR genes usually contain 4~6 exons and 3~5 introns according to the VvOPR exon/intron structure analyses (Figure 1C), and these results were similar to the gene structures of ZmOPR and TaOPR (Zhang et al., 2005; Mou et al., 2019). A combination of the phylogenetic (Figure 1B) and genetic structure analysis (Figure 1C) revealed that most of the VvOPR genes within a subfamily showed a similar exon/intron structures. VvOPR4 in sub.I contained the most introns (Figure 1C), indicating that the intron loss events occurred during the structural evolution of the OPR gene family (Li et al., 2009). As shown by the diversity of VvOPR protein-conserved motifs, the number and the position of VvOPR motifs in each subfamily are clearly conserved (Figure 1C), with VvOPR1 in sub.I lacking motif 9 and 10; analogously, the motif loss events were also detected in sub.II, and VvOPR3 lacks motif 7, and the absence of this motif is crucial for the secondary structure of proteins (Xin et al., 2017). In general, paralogous genes may have new biological functions relative to their ancestor genes (Guo et al., 2008), so we concluded that VvOPR1 might function in conjunction with the associated homologous genes AtOPR1,2.
The number of OPR members in the VvOPR family differed from that in other representative plants (Table 1), possibly because grapevine has undergone whole-genome duplications during its evolutionary history. Gene duplication is one of the primary driving forces in the evolution of genomes (Moore and Purugganan, 2004; Cannon et al., 2004), hence, it is highly possible that the gene duplication led to VvOPR gene family expansion, and that new genes contributed to the new structures and new biological functions. Therefore, we deduce that VvOPR1 might develop some new functions that are different from AtOPR1,2.
The expression of OPRs in dicots and monocots was found to be tissue specific (Liu et al., 2020; Stintzi and Browse, 2000; Li et al., 2011; Biesgen and Weiler, 1999; Mou et al., 2019; Xin et al., 2017; Creelman and Mullet, 1995). Here, VvOPR1 showed the highest expression level in roots, followed by stems and leaves, and the expression profiles of VvOPR1 under abiotic stress treatments (Cu, Zn, H2O2 and ABA stresses) (Figure 5) were much different from those of GhOPR (Liu et al., 2020), ZmOPR (Zhang et al., 2005) and TaOPR (Wang et al., 2016a; Dong et al., 2013). Therefore, based on the specific stress inductive patterns of VvOPR1 suggest that VvOPR1 is involved in the response to Cu, Zn stress.
Research on OPRs participating in development and response to various abiotic and biotic stresses in some plants has been conducted thoroughly (Bosch et al., 2014; Dave et al., 2011). In this study, the growth retardation was observed in the OT and WT seedlings under Cu, Zn stress, however, the OT seedlings showed obvious phenotypic changes compared with the WT seedlings, manifested as higher generation rates, longer shoot and root lengths and higher fresh weights (Figure 4). The growth reduction raised with increasing levels of Cu, Zn, which induced toxicity at elevated concentrations, and the impact on root elongation was greater than that on shoot growth (Figure 4); similar results have been reported for durum wheat (Michaud et al., 2007; Ghavri and Singh, 2012; Bibi et al., 2006; Shu et al., 2002). In Phaseolus vulgaris, Cu toxicity disrupted the capacity of cells to remove oxidatively damaged proteins by inhibiting the ubiquitin proteasome pathway in embryonic stages thus inhibiting seed germination (Karmous et al., 2014). Seed germination which is very sensitive to the external medium is regulated by changes in the cellular redox status (Nanda and Agrawal, 2016), and the addition of metal further aggravates the microenvironment, causing damage to proteins and leading to a reduction in germination (El-Maarouf-Bouteau and Bailly, 2008). In this study, the relatively high germination rate of OT under Cu, Zn stress conditions might have been due to decreased accumulation of oxidatively damaged proteins. The Cu, Zn-induced growth reduction has also been observed in Spirodela polyrhiza (Upadhyaya and Panda, 2010), Withania somnifera (Khatun et al., 2008) and Sorghum bicolor (Soudek et al., 2014).
ABA is a vital component of the abiotic stress response, and increasing ABA synthesis and/or limiting ABA catabolism frequently occurs in plant in response to abiotic stress (Jakab et al., 2005). In response to abiotic stresses, the ABA content dramatically increases in transgenic plants overexpressing some ABA biosynthesis-related genes to cope with the stress (Yue et al., 2012; Hwang et al., 2010). Similar expression levels of ABA biosynthetic (OsNCED1, OsZEP1 and OsAAO3) and catabolic (OsABA8ox1) genes were detected in the OT and WT seedlings under normal conditions, which indicated that VvOPR1 alone may not be sufficient to regulate ABA biosynthesis. However, the higher expression levels of these genes in the OT seedlings exposed to Cu, Zn stress clearly indicated that the possible role of VvOPR1 in Cu, Zn stress-induced ABA-biosynthesis, along with additional factors. In this study, although the expression levels of both ABA biosynthetic (OsNCED1, OsZEP1 and OsAAO3) and catabolic (OsABA8ox1) genes were elevated, the speed of ABA synthesis may be much faster than that of catabolism, so the ABA content was elevated in Cu-, Zn-treated seedlings, suggesting its involvement in the induction of protective mechanisms against excess Cu, Zn toxicity. This finding is in agreement with a previous study in which ABA was found to increase under Cu (Vishwakarma et al., 2017), Zn stress (Wang et al., 2014). The ABA content was significantly higher in the OT lines than in the WT, indicating the possible role of VvOPR1 induced ABA accumulation in response to Cu, Zn stress and an acceleration in ABA-dependent pathway caused by VvOPR1 overexpression is initiated from a burst in ABA synthesis.
ABA triggers a signaling cascade that regulates a suite of abiotic stress responsive genes (Nakashima et al., 2009). Here, the expression levels of the early ABA signaling genes (OsPP2C68 and OSRK1) and late ABA-responsive genes (RAB21, OsLEA3, RAB16C and RAB16D) in Cu-, Zn-treated seedlings were obviously elevated compared with those in the control condition, and the higher expression level in OT (Figure 7B) provides firm evidence for a positive role of VvOPR1 in stress-responsive ABA signaling. Overall the results suggested that the overexpression of VvOPR1 induced ABA synthesis and enhanced ABA content in OT seedlings under Cu, Zn stress conditions, thus, leading to increased expression levels of ABA signaling and responsive genes. However, further study is necessary in order to better understand how component genes involved in ABA signal transduction mediate Cu, Zn stress through the induction of gene expression induction.
Excess Cu, Zn often cause the generation of ROS, and excess ROS in the cell are associated with extensive lipid peroxidation (Deng et al., 2010; Olmos et al., 2003; Cho et al., 2012), which generates a range of toxic breakdown products, such as α, β unsaturated aldehydes (Trotter et al., 2006; Esterbauer, 1993). Here, MDA accumulated in WT and OT seedlings during Cu, Zn stress, albeit to a lesser extent in the OT lines than in the WT, significantly (Figure 5). High lipid peroxidation in rice seedlings might be the result of ROS-induced oxidative stress or increased lipoxygenase activity caused by Cu, Zn stress. Similar results have also been reported in Triticum aestivum (Panda et al., 2003) in response to Zn and in Brassica juncea (Radic et al., 2010) in response to Cu. These results indicate that higher protection against oxidative damage in OT might be due to less ROS generation, less peroxidation, and the ability of VvOPR1 to alleviate ROS damage.
In addition to counteracting the toxicity of ROS-induced lipid peroxidation, the direct neutralization of ROS has been proposed as a component of stress tolerance (Mittler, 2002). The OYZ family is believed to protect the cell against the damaging effects of lipid peroxidation products, and the function of OYE in yeast also appears to be to reduce the level of ROS present (Fitzpatrick et al., 2003). In this study, compared with those in WT seedlings, the reduced H2O2 in Cu, Zn-treated OT may be attributed to the increased levels of antioxidative enzymes involved in scavenging this oxidant (Thounaojam et al., 2012). Previous studies have shown that CAT has a high capacity but low affinity, whereas POD has a high affinity for H2O2 (Mittler et al., 2002). The decrease in CAT activity indicated that CAT was not required for the elimination of H2O2 in Cu-, Zn-stressed rice plants and that POD is the major effective H2O2-scavenging enzyme for reducing H2O2 in OT cells under Cu, Zn stress. In addition, APX has a greater affinity for H2O2 than dose CAT and it may play a more critical role in the regulation of ROS (Nanda and Agrawal, 2016), and increased APX activity (Figure 6A) under Cu stress has been previously reported (Tewari et al., 2006). Higher AsA (Figure 5) and increased SOD activity (Figure 6A) are associated with the detoxification of superoxide radicals to H2O2 which may be responsible for the increased activity of APX under Cu, Zn stress conditions (Li et al., 2013; Lukatkin et al., 2014). This result is in accordance with the findings of Drazkiewicz et al. (2003), where the ASH-GSH cycle plays an important role in reducing the toxic effect of Cu. Thus VvOPR1 overexpression increased the content of several low-molecular weight nonenzymatic compounds (Figure 5), increased the activities of several ROS scavenging enzymes (Figure 6A) and the expression of their encoding genes (Figure 6B) and there is some evidence that VvOPR1 promotes the efficiency of ROS scavenging to affect the removal of ROS and alleviate their deleterious effects.
ABA can increase the transcription and activity of ROS network genes, and defects in this network can also disrupt the expression of ABA and stress responsive genes (Miao et al., 2006). Moreover, the VvOPR1/NCED1 Arabidopsis mutant did not rescue the sensitivity of the NCED1 mutant to H2O2 (Figure 7D). Hence, it could be concluded that the enhanced efficiency of ROS scavenging promoted by VvOPR1 expression may be, at least in part, mediated by an acceleration of ABA synthesis and an upregulation of relevant signaling pathways.
In this work the behavior of WT and OT seedlings with respect to Cu, Zn tolerance, accumulation and translocation were compared. Both the OT and WT showed a prominent Cu, Zn accumulation in the roots under Cu, Zn stress conditions, and their accumulation differed significantly between the OT and WT seedlings. Compared with the OT seedlings, the WT showed a greater ability than OT to accumulate and translocate the metal to the aerial parts of the seedlings. In addition, the TI and AR in the OT in terms of Cu2+, Zn2+ were detected (Tables 7, 8). On the basis of the dry biomass of the total plants, the TI and AR revealed that the tolerance to Cu, Zn was much greater than that of the WT. Together, these results confirmed that VvOPR1 has a considerable potential to alleviate Cu2+, Zn2+ induced damage by reducing Cu2+, Zn2+ accumulation and translocation. The reduction translocation of Cu2+, Zn2+ from roots to the aerial organs could reduce the damaging effects of theses pollutants on leaf physiology and biochemistry.
A previous study has reported that ABA can reduce heavy metal stress by affecting heavy metal transport to the aerial parts (Perfus-Barbeoch et al., 2002). It is possible that ABA-induced stomatal closure suppressed of transpirational flow, resulting in a restriction of root-to-aerial translocation of metals (Bücker-Neto et al., 2017). Exogenous ABA application reduced the transport of Cd, Ni from the roots to the leaves, resulting in greater metal accumulation in the roots (Rubio et al., 1994), and in response to treatment with CdCl2, the ABA content rapidly increased in the leaves and roots of Cd-tolerant cultivar rice seedlings (Hsu and Kao, 2003). These findings are consistent with that the lower TF (Tables 7, 8) of OT seedlings, which accumulated higher ABA (Figure 7A) content, and were more tolerant of Cu, Zn stress than the WT. To date, studies are still in progress to characterize the OT seedlings with respect to the biochemical and molecular processes involved in the accumulation and translocation of Cu2+, Zn2+ to the aerial organs.
Recently, JA has been shown to be effective in improving plant tolerance to heavy metal stress (Yan et al., 2015; Piotrowska et al., 2009; Ahmad et al., 2017). In this study, the increase in JA levels in the leaves of the OT and WT seedlings in response to the Cu, Zn treatments was consistent with the increase in JA levels after Cu and Cd treatment (Maksymiec et al., 2005). The increase in JA caused by Cu, Zn stress could involve up-regulation of some JA biosynthesis genes, such as OsDAD1, OsPLA1, OsLOX2, OsAOS1, OsAOC1 and OsOPR7 (Figure 7C). These results suggested that JA is involved in Cu, Zn stress, however, no significant difference was defected in JA content or the expression level of JA synthesis genes was detected between OT and WT seedlings, so we concluded that neither the expression of JA synthesis genes nor the endogenous JA level was dependent on VvOPR1 overexpression (Figure 7C). In addition, similar results were detected for the expression levels of the JA signaling related genes OsJAR1 and OsCoI1b (Figure 7C). These findings indicate that VvOPR1 dose not regulate JA synthesis or signaling pathways.
In combination, we like to conclude that VvOPR1 may be an effective gene for improving Cu, Zn stress tolerance. Here, we first synthesized and characterized the characterization the OPRI gene VvOPR1, which regulates the expression of ABA biosynthesis and catabolism genes thus leading to increased endogenous ABA accumulation under Cu, Zn stress conditions, subsequently promoting the ABA signaling pathway, and reducing Cu, Zn accumulation and translocation. Moreover, VvOPR1 overexpression reduced the production of ROS (such as reduced H2O2 and MDA) and increased the ROS-scavenging system to confer tolerance to Cu, Zn stress. Therefore, we concluded that VvOPR1 may be an effective gene for improving Cu, Zn stress tolerance in plants. In addition, the function of VvOPR1 may lie in the metabolism of trans-(+)-OPDA, with consequent effects on the activities of the (ABA-dependent responsive and/or ROS) signaling pathway which this molecule mediates. Moreover, the presence of VvOPR1 did not induce any changing in JA synthesis or signaling pathways.
In summary, nine VvOPR genes were identified from grapevine genome and classified into two subfamilies. and the evolution was relatively conservative in the group; chromosome mapping confirmed that VvOPR family genes were only distributed on chromosome 11 and 18, gene structure analysis identified that the structure of VvOPRs was highly conservative, especially the conservative motif 6, 4, 2 and 3 were shared by all genes; promoter analysis revealed that the promoter region of OPR genes were rich in cis-element which response to growth and development, hormone signals and adversity. In addition, the results revealed that Cu, Zn had a relatively low toxicity, but could do damage to plants at some concentrations. Cu, Zn stress decreased the biomass, and photosynthetic activities due to increasing ROS. While VvOPR1 overexpression alleviated Cu, Zn stress and reduced the growth restriction. These biochemical mechanistic findings suggested in OT seedlings, VvOPR1 enhanced the photosynthetic capacity, promoted ABA synthesis and the ABA-dependent stress response pathway, improved the activities of ROS scavengers and the expression levels of their encoding genes, increased the accumulation of proline, AsA, GSH, while alleviated MDA and H2O2 accumulation. Moreover, VvOPR1 reduced Cu2+, Zn2+ accumulation, translocation. Together, ABA may play a crucial role in the response VvOPR1-overexpressing seedlings to Cu, Zn stress, VvOPR1 responds to Cu, Zn stress in an ABA-dependent manner to enhance tolerance to Cu, Zn stress, and this effect appears to be quite independent from JA synthesis or JA signaling. Despite the progress achieved, further work is needed to determine how ABA guides adaptation under Cu, Zn stress conditions.
The datasets presented in this study can be found in online repositories. The names of the repository/repositories and accession number(s) can be found in the article/Supplementary Material.
S-HY: Conceptualization, Data curation, Formal analysis, Methodology, Writing – original draft. Y-PC: Formal analysis, Methodology, Writing – review & editing. W-JS: Formal analysis, Resources, Writing – review & editing. XL: Formal analysis, Writing – review & editing. ZW: Conceptualization, Funding acquisition, Writing – review & editing. Q-HY: Conceptualization, Writing – review & editing.
The author(s) declare financial support was received for the research, authorship, and/or publication of this article. This study was supported by the Chongqing Science and Technology Commission Project (cstc2018jxjl80040), Chongqing Municipal Financial Commission Project (cqaas2023sjczsf011).
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fpls.2025.1509472/full#supplementary-material
Adrees, M., Ali, S., Rizwan, M., Ibrahim, M., Abbas, F., Farid, M., et al. (2015). The effect of excess copper on growth and physiology of important food crops: a review. Environ. Sci. pollut. Res. 22, 8148–8162. doi: 10.1007/s11356-015-4496-5
Agrawal, G. K., Jwa, N. S., Shibato, J., Han, O., Iwahashi, H., Rakwal, R. (2003). Diverse environmental cues transiently regulate OsOPR1 of the “octadecanoid pathway” revealing its importance in rice defense/stress and development. Biochem. Biophys. Res. Commun. 310, 1073–1082. doi: 10.1016/j.bbrc.2003.09.123
Ahmad, P., AlYemeni, M. N., Wijaya, L., Alam, P., Ahanger, M. A., Alamri, S. A. (2017). Jasmonic acid alleviates negative impacts of cadmium stress by modifying osmolytes and antioxidants in faba bean (Vicia faba L.). Arch. Agron. Soil Sci. 63, 1889–1899. doi: 10.1080/03650340.2017.1313406
Bailey, T. L., Boden, M., Buske, F. A., Frith, M., Grant, C. E., Clementi, L., et al. (2009). MEME Suite: tools for motif discovery and searching. Nucleic. Acids Res. 37, 202–208. doi: 10.1093/nar/gkp335
Baker, N. R., Rosenqvist, E. (2004). Applications of chlorophyll fluorescence can improve crop production strategies: an examination of future possibilities. J. Exp. Bot. 403, 1607–1621. doi: 10.1093/jxb/erh196
Bates, L. S., Waldren, R. P., Teare, I. D. (1973). Rapid determination of free proline for water-stress studies. Plant Soil 39, 205–207. doi: 10.1007/BF00018060
Beynon, E. R., Symons, Z. C., Jackson, R. G., Lorenz, A. L., Rylott, E. L., Bruce, N. C. (2009). The role of oxophytodienoate reductases in the Detoxification of the Explosive 2,4,6-trinitrotoluene by Arabidopsis. Plant Physiol. 151, 253–261. doi: 10.1104/pp.109.141598
Bibi, M., Hussain, M., Qureshi, M., Kousar, S., Ajaz, S. (2006). Heavy metal and nutrient ion accumulation by two black gram cultivars treated with copper and lead. J. Plant Nutt. 29, 913–920. doi: 10.1080/01904160600651837
Biesgen, C., Weiler, E. W. (1999). Structure and regulation of OPR1 and OPR2, two closely related genes encoding 12-oxophytodienoic acid-10,11-reductases from Arabidopsis thaliana. Planta 208, 155–165. doi: 10.1007/s004250050545
Bosch, M., Wright, L. P., Gershenzon, J., Wasternack, C., Hause, B., Schaller, A., et al. (2014). Jasmonic acid and its precursor 12-oxophytodienoic acid control different aspects of constitutive and induced herbivore defenses in tomato. Plant Physiol. 166, 396–410. doi: 10.1104/pp.114.237388
Breithaupt, C., Kurzbauer, R., Lilie, H., Schaller, A., Strassner, J., Huber, R., et al. (2006). Crystal structure of 12-oxophytodienoate reductase 3 from tomato: Self-inhibition by dimerization. Proc. Natl. Acad. Sci. 103, 14337–14342. doi: 10.1073/pnas.0606603103
Bücker-Neto, L., Paiva, A. L. S., MaChado, R. D., Arenhart, R. A., Margis-Pinheiro, M. (2017). Interactions between plant hormones and heavy metals responses. Genet. Mol. Biol. 40, 373–386. doi: 10.1590/1678-4685-gmb-2016-0087
Cannon, S. B., Mitra, A., Baumgarten, A., Young, N. D., May, G. (2004). The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. BMC Plant Biol. 4, 10. doi: 10.1186/1471-2229-4-10
Cho, S. C., Chao, Y. Y., Hong, C. Y., Kao, C. H. (2012). The role of hydrogen peroxide in cadmium-inhibited root growth of rice seedlings. Plant Growth Regul. 66, 27–35. doi: 10.1007/s10725-011-9625-7
Creelman, R. A., Mullet, J. E. (1995). Jasmonic acid distribution and action in plants: regulation during development and response to biotic and abiotic stress. Proc. Natl. Acad. Sci. U.S.A. 92, 4114–4119. doi: 10.1073/PNAS.92.10.4114
Dave, A., Hernandez, M. L., He, Z., Andriotis, V. M. E., Vaistij, F. E., Larson, T. R., et al. (2011). 12-oxo-phytodienoic acid accumulation during seed development represses seed germination in arabidopsis. Plant Cell 23, 583–599. doi: 10.1105/tpc.110.081489
Deng, X., Xia, Y., Hu, W., Zhang, H., Shen, Z. (2010). Cadmium-induced oxidative damage and protective effects of N-acetyl-l-cysteine against cadmium toxicity in Solanum nigrum L. J. Hazard. Mater. 180, 722–729. doi: 10.1016/j.jhazmat.2010.04.099
Dong, W., Wang, M., Xu, F., Quan, T., Peng, K., Xiao, L., et al. (2013). Wheat oxophytodienoate reductase gene TaOPR1 confers salinity tolerance via enhancement of abscisic acid signaling and reactive oxygen species scavenging. Plant Physiol. 161, 1217–1228. doi: 10.1104/pp.112.211854
Drazkiewicz, M., Skorzynska-Polit, E., Krupa, Z. (2003). Responses of the ascorbatee glutathione cycle to excess copper in Arabidopsis thaliana (L.). Plant Sci. 164, 195–202. doi: 10.1016/S0168-9452(02)00383-7
Druart, C., Scheifler, R., Millet, M., Vaufleury, A. (2012). Landsnail eggs bioassays: a new tool to assess embryotoxicity of contaminants in the solid, liquid or gaseous phase of soil. Appl. Soil Ecol. 53, 56–64. doi: 10.1016/j.apsoil.2011.11.006
El-Maarouf-Bouteau, H., Bailly, C. (2008). Oxidative signaling in seed germination and dormancy. Plant Signal. Behav. 3, 175–182. doi: 10.4161/psb.3.3.5539
Esterbauer, H. (1993). Cytotoxicity and genotoxicity of lipid-oxidation products. Am. J. Clin. Nutr. 57, 779S–786S. doi: 10.1093/ajcn/57.5.779S
Fattorini, L., Hause, B., Gutierrez, L., Veloccia, A., Della Rovere, F., Piacentini, D., et al. (2018). Jasmonate promotes auxin-induced adventitious rooting in dark-grown Arabidopsis thaliana seedlings and stem thin cell layers by a cross-talk with ethylene signaling and a modulation of xylogenesis. BMC Plant Biol. 18, 182. doi: 10.1186/s12870-018-1392-4
Fitzpatrick, T. B., Amrhein, N., Macheroux, P. (2003). Characterization of YqjM, an old yellow enzyme homolog from Bacillus subtilis involved in the oxidative stress response. J. Biol. Chem. 278, 19891–19897. doi: 10.1074/jbc.M211778200
Geng, A., Wang, X., Wu, L., Wang, F., Wu, Z., Yang, H., et al. (2018). Silicon improves growth and alleviates oxidative stress in rice seedlings (Oryza sativa L.) by strengthening antioxidant defense and enhancing protein metabolism under arsanilic acid exposure. Ecotoxicol. Environ. Saf. 158, 266–273. doi: 10.1016/j.ecoenv.2018.03.050
Ghavri, S. V., Singh, R. P. (2012). Growth, biomass production and remediation of copper contamination by Jatropha curcas plant in industrial wasteland soil. J. Enviro. Biol. 33, 207. doi: 10.1016/j.jclepro.2011.11.063
Guang, Y., Luo, S., Ahammed, G. J., Xiao, X., Li, J., Zhou, Y., et al. (2021). The OPR gene family in watermelon: Genome-wide identification and expression profiling under hormonal treatments and root-knot nematode infection. Plant Biol. 11, 80–88. doi: 10.1111/plb.13225
Guo, J., Wu, J., Ji, Q., Wang, C., Luo, L., Yuan, Y., et al. (2008). Genome-wide analysis of heat shock transcription factor families in rice and Arabidopsis. J. Genet. Genom. 35, 105–118. doi: 10.1016/S1673-8527(08)60016-8
Guo, A. Y., Zhu, Q. H., Chen, X., Luo, J. C. (2007). GSDS: a gene structure display server. Yi Chuan 29, 1023–1026. doi: 10.1360/yc-007-1023
Hsu, Y. T., Kao, C. H. (2003). Role of abscisic acid in cadmium tolerance of rice (Oryza sativa L.) seedlings. Plant Cell Environ. 26, 867–874. doi: 10.1046/j.1365-3040.2003.01018.x
Hu, Q., Zhu, L., Zhang, X., Guan, Q., Xiao, S., Min, L., et al. (2018). GhCPK33 Negatively Regulates Defense against Verticillium dahliae by Phosphorylating GhOPR3. Plant Physiol. 178, 876–889. doi: 10.1104/pp.18.00737
Hung, K. T., Kao, C. H. (2003). Nitric oxide counteracts the senescence of rice leaves induced by abscisic acid. J. Plant Physiol. 160, 871–879. doi: 10.1078/0176-1617-01118
Hwang, S. G., Chen, H. C., Huang, W. Y., Chu, Y. C., Shii, C. T., Cheng, W. H. (2010). Ectopic expression of rice OsNCED3 in Arabidopsis increases ABA level and alters leaf morphology. Plant Sci. 178, 12–22. doi: 10.1016/j.plantsci.2009.09.014
Jakab, G., Ton, J., Flors, V., Zimmerli, L., Métraux, J. P., Mauch-Mani, B. (2005). Enhancing Aabidopsis salt and drought stress tolerance by chemical priming for its abscisic acid responses. Plant Physiol. 139, 267–274. doi: 10.1104/pp.105.065698
Juang, K. W., Lee, Y. I., Lai, H. Y., Wang, C. H., Chen, B. C. (2012). Copper accumulation, translocation, and toxic effects in grapevine cuttings. Environ. Sci. pollut. R. 19, 1315–1322. doi: 10.1007/s11356-011-0657-3
Karmous, I., Chaoui, A., Jaouani, K., Sheehan, D., Ferjani, E. E., Scoccianti, V., et al. (2014). Role of the ubiquitin-proteasome pathway and some peptidases during seed germination and copper stress in bean cotyledons. Plant Physiol. Biochem. 76, 77–85. doi: 10.1016/j.plaphy.2013.12.025
Khan, A. R., Uiiah, I., Khan, A. L., Park, G. S., Waqas, M., Hong, S. J., et al. (2015). Improvement in phytoremediation potential of Solanum nigrum under cadmium contamination through endophytic-assisted Serratia sp. RSC-14 inoculation. Environ. Sci. pollut. Res. Int. 22, 14032–14042. doi: 10.1007/s11356-015-4647-8
Khatun, S., Ali, M. B., Hahna, E. J., Paek, K. Y. (2008). Copper toxicity in Withania somnifera: Growth and antioxidant enzymes responses of in vitro grown plants. Environ. Exp. Bot. 64, 279–285. doi: 10.1016/j.envexpbot.2008.02.004
Larkin, M. A., Blackshields, G., Brown, N. P., Chenna, R., McGettigan, P. A., McWiliam, H., et al. (2007). Clustal W and clustal X version 2.0. Bioinformatics 23, 2947–2948. doi: 10.1093/bioinformatics/btm404
Li, W., Li., Y., Xu, Y., Kumar, S., Liu, Y., Zhu, G. (2024). Genome-wide identification, gene cloning, subcellular location and expression analysis of the OPR gene family under salt stress in sweetpotato. BMC Plant Biol. 24, 1171. doi: 10.1186/s12870-024-05887-8
Li, W., Liu, B., Yu, L., Feng, D., Wang, H., Wang, J. (2009). Phylogenetic analysis, structural evolution and functional divergence of the 12-oxo-phytodienoate acid reductase gene family in plants. BMC Evol. Biol. 9, 90. doi: 10.1186/1471-2148-9-90
Li, M., Xu, G., Xia, X., Wang, M., Yin, X., Zhang, B., et al. (2017). Deciphering the physiological and molecular mechanisms for copper tolerance in autotetraploid Arabidopsis. Plant Cell Rep. 36, 1585–1597. doi: 10.1007/s00299-017-2176-2
Li, X., Yang, Y., Jia, L., Chen, H., Wei, X. (2013). Zinc-induced oxidative damage, antioxidant enzyme response and proline metabolism in roots and leaves of wheat plants. Ecotoxicol. Environ. Saf. 89, 150–157. doi: 10.1016/j.ecoenv.2012.11.025
Li, W., Zhou, F., Liu, B., Feng, D., He, Y., Qi, K., et al. (2011). Comparative characterization, expression pattern and function analysis of the 12-oxo-phytodienoic acid reductase gene family in rice. Plant Cell Rep. 30, 981–995. doi: 10.1007/s00299-011-1002-5
Lichtenthaler, H. K. (1987). Chlorophylls and carotenoids: pigments of photosynthetic biomembranes. Meth. Enzymol. 148C, 350–382. doi: 10.1016/0076-6879(87)48036-1
Liechti, R., Farmer, E. E. (2006). Jasmonate biochemical pathway. Sci. STKE. 322, cm3. doi: 10.1126/stke.3222006cm3
Liu, S., Sun, R., Zhang, X., Feng, Z., Wei, F., Zhao, L., et al. (2020). Genome-wide analysis of OPR family genes in cotton identified a role for GhOPR9 in Verticillium dahliae resistance. Genes 11, 1134. doi: 10.3390/genes11101134
Livak, K. J., Schmittgen, T. D. (2001). Analysis of relative gene expression data using real time quantitative PCR and the 2–ΔΔCt method. Methods 25, 402–408. doi: 10.1006/meth.2001.1262
Lou, Y. G., Baldwin, I. T. (2003). Manduca sexta recognition and resistance among allopolyploid Nicotiana host plants. Proc. Natl. Acad. Sci. U.S.A. 100, 14581–14586. doi: 10.2307/3147338
Lukatkin, A., Egorova, I., Michailova, I., Malec, P., Strzalka, K. (2014). Effect of copper on pro- and antioxidative reactions in radish (Raphanus sativus L.) in vitro and in vivo. J. Trace Elem. Med. Biol. 28, 80–86. doi: 10.1016/j.jtemb.2013.11.002
Maksymiec, W., Wianowska, D., Dawidowicz, A. L., Radkiewicz, S., Mardarowicz, M., Krupa, Z. (2005). The level of jasmonic acid in Arabidopsis thaliana and Phaseolus coccineus plants under heavy metal stress. J. Plant Physiol. 162, 1338–1346. doi: 10.1016/j.jplph.2005.01.013
Manousaki, E., Kadukova, J., Papadantonakis, N., Kalogerakis, N. (2008). Phytoextraction and phytoexcretion of Cd by the leaves of Tamarixs myrnensis growing on contaminated non-saline and saline soils. Environ. Res. 106, 326–332. doi: 10.1016/j.envres.2007.04.004
Matsui, H., Nakamura, G., Ishiga, Y., Toshima, H., Inagaki, Y., Toyoda, K., et al. (2004). Structure and expression of 12-oxophytodienoate reductase (OPR) subgroup I gene in pea, and oxidoreductase activity of their recombinant proteins. Mol. Genet. Genom. 271, 1–10. doi: 10.1007/s00438-003-0948-6
Miao, Y., Lv, D., Wang, P., Wang, X. C., Chen, J., Miao, C., et al. (2006). An Arabidopsis glutathione peroxidase functions as both a redox transducer and a scavenger in abscisic acid and drought stress responses. Plant Cell 18, 2749–2766. doi: 10.1105/tpc.106.044230
Michaud, A. M., Bravin, M. N., Galleguillos, M., Hinsinger, P. (2007). Copper uptake and phytotoxicity as assessed in situ for durum wheat (Tritcum turgidum durum L.) cultivated in cu-contaminated former vineyard soil. Plant Soil 229, 99–111. doi: 10.1007/s11104-007-9343-0
Mikkelsen, M. D., Pedas, P., Schiller, M., Vincze, E., Mills, R. F., Borg, S., et al. (2012). Barley HvHMA1 is a heavy metal pump involved in mobilizing organellar Zn and Cu and plays a role in metal loading into grains. PloS One 7, e49027. doi: 10.1371/journal.pone.0049027
Miotto, A., Ceretta, C. A., Brunetto, G., Nicoloso, F. T., Girotto, E., Farias, J. G., et al. (2014). Copper uptake, accumulation and physiological changes in adult grapevines in response to excess copper in soil. Plant Soil. 374, 593–610. doi: 10.1007/s11104-013-1886-7
Mirlean, N., Roisenberg, A., Chies, J. O. (2007). Metal contamination of vineyard soils in wet subtropics (southern Brazil). Environ. Poll. 149, 10–17. doi: 10.1016/j.envpol.2006.12.024
Mittler, R. (2002). Oxidative stress, antioxidants and stress tolerance. Trends Plant Sci. 7, 405–410. doi: 10.1016/s1360-1385(02)02312-9
Mittler, R., Vanderauwera, S., Suzuki, N., Miller, G., Tognetti, V. B., Vandepoele, K., et al. (2011). ROS signaling: the new wave? Trends Plant Sci. 16, 300–309. doi: 10.1016/j.tplants.2011.03.007
Montanini, B., Blaudez, D., Jeandroz, S., Sanders, D., Chalot, M. (2007). Phylogenetic and functional analysis of the Cation Difusion Facilitator (CDF) family: improved signature and prediction of substrate specificity. BMC Genom. 8, 107. doi: 10.1186/1471-2164-8-107
Moore, R., Purugganan, M. D. (2004). The early stages of duplicate gene evolution. Proc. Natl. Acad. Sci. U.S.A. 100, 15682–15687. doi: 10.1073/pnas.2535513100
Moravcová, Š., Tůma, J., Dučaiová, Z. K., Waligórski, P., Kula, M., Saja, D., et al. (2018). Influence of salicylic acid pretreatment on seeds germination and some defence mechanisms of Zea mays plants under copper stress. Plant Physiol. Biochem. 122, 19–30. doi: 10.1016/j.plaphy.2017.11.007
Mou, Y., Liu, Y., Tian, S., Guo, Q., Wang, C., Wen, S. (2019). Genome-Wide identification and characterization of the OPR Gene Family in Wheat (Triticum aestivum L.). Int. J. Mol. Sci. 20, 1914. doi: 10.3390/ijms20081914
Nagajyoti, P. C., Lee, K. D., Sreekanth, T. V. M. (2010). Heavy metals, occurrence and toxicity for plants: a review. Environ. Chem. Lett. 8, 199–216. doi: 10.1007/s10311-010-0297-8
Nakashima, K., Ito, Y., Yamaguchi-Shinozaki, K. (2009). Transcriptional regulatory networks in response to abiotic stresses in Arabidopsis and grasses. Plant Physiol. 149, 88–95. doi: 10.1104/pp.108.129791
Nanda, R., Agrawal, V. (2016). Elucidation of zinc and copper induced oxidative stress, DNA damage and activation of defence system during seed germination in Cassia angustifolia Vahl. Environ. Exp. Bot. 125, 31–41. doi: 10.1016/j.envexpbot.2016.02.001
Nie, W. F., Chen, Y., Tao, J., Li, Y., Zhou, Y., Yang, Y. (2022). Identification of the 12-oxophytoeienoic acid reductase (OPR) gene family in pepper (Capsicum annuum L.) and functional characterization of CaOPR6 in pepper fruit development and stress response. Genome 65, 537–545. doi: 10.1139/gen-2022-0037
Olmos, E., Martínez-Solano, J. R., Piqueras, A., Hellín, E. (2003). Early steps in the oxidative burst induced by cadmium in cultured tobacco cells (BY-2 line). J. Exp. Bot. 54, 291–301. doi: 10.1093/jxb/erg028
Panda, S. K., Chaudhury, I., Khan, M. H. (2003). Heavy metal induce lipid peroxidation and affect antioxidants in wheat leaves. Biol. Plant 46, 289–294. doi: 10.1023/A:1022871131698
Perfus-Barbeoch, L., Leonhardt, N., Vavasseur, A., Forestier, C. (2002). Heavy metal toxicity: Cadmium permeates through calcium channels and disturbs the plant water status. Plant J. 32, 539–548. doi: 10.1046/j.1365-313x.2002.01442.x
Piotrowska, A., Bajguz, A., Godlewska-Żyłkiewicz, B., Czerpak, R., Małgorzata, K. (2009). Jasmonic acid as modulator of lead toxicity in aquatic plant Wolffia arrhiza (Lemnaceae). Environ. Exp. Bot 66, 507–513. doi: 10.1016/j.envexpbot.2009.03.019
Radic, S., Babic, M., Skobic, D., Roje, V., Pevalek-Kozlina, B. (2010). Ecotoxicological effects of aluminum and zinc on growth and antioxidants in Lemna minor L. Ecotoxicol. Environ. Saf. 73, 336–342. doi: 10.1016/j.ecoenv.2009.10.014
Ramos, M., López-Acevedo, M. (2004). Zinc levels in vineyard soils from the Alt Pened`es-Anoia region (NE Spain) after compost application. Adv. Environ. Res. 8, 687–696. doi: 10.1016/s1093-0191(03)00041-8
Rehman, M., Liu, L., Wang, Q., Saleem, M. H., Bashir, S., Ullah, S., et al. (2019). Copper environmental toxicology, recent advances, and future outlook: A review. Environ. Sci. pollut. Res. Int. 26, 18003–18016. doi: 10.1007/s11356-019-05073-6
Rubio, M. I., Escrig, I., Martinez-Cortina, C., Lopez-Benet, F. J., Sanz, A. (1994). Cadmium and nickel accumulation in rice plants. Effects on mineral nutrition and possible interactions of abscisic and gibberellic acids. Plant Growth Regul. 14, 151–157. doi: 10.1007/BF00025217
Scalschi, L., Sanmartin, M., Camanes, G., Troncho, P., Sanchez-Serrano, J. J., Garcia-Agustin, P., et al. (2015). Silencing of OPR3 in tomato reveals the role of OPDA in callose deposition during the activation of defense responses against Botrytis cinerea. Plant J. 81, 304–315. doi: 10.1111/tpj.12728
Schaller, F. (2001). Enzymes of the biosynthesis of octadecanoid-derived signaling molecules. J. Exp. Bot. 52, 11–23. doi: 10.1093/jxb/52.354.11
Schaller, F., Hennig, P., Weiler, E. W. (1998). 12-oxophytodienoate-10,11-reductase: Occurrence of two isoenzymes of different specificity against stereoisomers of 12-oxophytodienoic acid. Plant Physiol. 118, 1345–1351. doi: 10.1104/pp.118.4.1345
Schaller, F., Weiler, E. W. (1997). Molecular cloning and characterization of 12-oxophytodienoate reductase, an enzyme of the octadecanoid signaling pathway from Arabidopsis thaliana. Structural and functional relationship to yeast old yellow enzyme. J. Biol. Chem. 272, 28066–28072. doi: 10.1074/jbc.272.44.28066
Shu, W. S., Ye, Z. H., Lan, C. Y., Zhang, Z. Q., Wong, M. H. (2002). Lead, zinc and copper accumulation and tolerance in populations of Paspalum distichum and. Cynodon dactylon. Environ. pollut. 120, 445–453. doi: 10.1016/S0269-7491(02)00110-0
Song, S. Y., Chen, Y., Chen, J., Dai, X. Y., Zhang, W. H. (2011). Physiological mechanisms underlying OsNAC5-dependent tolerance of rice plants to abiotic stress. Planta 234, 331–345. doi: 10.1007/s00425-011-1403-2
Soudek, P., Petrova, S., Vankova, R., Song, J., Vanek, T. (2014). Accumulation of heavy metals using Sorghum sp. Chemosphere 104, 15–24. doi: 10.1016/j.chemosphere.2013.09.079
Stintzi, A., Browse, J. (2000). The Arabidopsis male-sterilemutant, opr3, lacks the 12-oxophytodienoic acid reductase required for jasmonate synthesis. Proc. Natl. Acad. Sci. U.S.A. 97, 10625–10630. doi: 10.1073/pnas.190264497
Strassner, J., Schaller, F., Frick, U. B., Howe, G. A., Weiler, E. W., Amrhein, N., et al. (2002). Characterization and cDNA-microarray expression analysis of 12-oxophytodienoate reductases reveals differential roles for octadecanoid biosynthesis in the local versus the systemic wound response. Plant J. 32, 585–601. doi: 10.1046/j.1365-313x.2002.01449.x
Tamura, K., Stecher, G., Pdterson, D., Filipski, A., Kuma, S. (2013). MEGA6: molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 30, 2725–2729. doi: 10.1093/molbev/mst197
Tang, X., Lowder, L. G., Zhang, T., Malzahn, A. A., Zheng, X., Voytas, D. F., et al. (2017). A CRISPR–Cpf1 system for efficient genome editing and transcriptional repression in plants. Nat. Plants 3, 17018. doi: 10.1038/nplants
Tanhan, P., Kruatrachue, M., Pokethitiyook, P., Chaiyarat, R. (2007). Uptake and accumulation of cadmium, lead and zinc by Siam weed [Chromol aenaodorata (L.) King & Robinson. Chemosphere 68, 323–329. doi: 10.1016/j.chemosphere.2006.12.064
Tewari, R. K., Kumar, P., Sharma, P. N. (2006). Antioxidant responses to enhanced generation of superoxide anion radical and hydrogen peroxide in the copper- stressed mulberry plants. Planta 223, 1145–1153. doi: 10.1007/s00425-005-0160-5
Thounaojam, T. C., Panda, P., Mazumdar, P., Kumar, D., Sharma, G. D., Sahoo, L., et al. (2012). Excess copper induced oxidative stress and response of antioxidants in rice. Plant Physiol. Biochem. 53, 33–39. doi: 10.1016/j.plaphy.2012.01.006
Toselli, M., Baldi, E., Marcolini, G., Malaguti, D., Quartieri, M., Sorrenti, G., et al. (2009). Response of potted grapevines to increasing soil copper concentration. Aust. J. Grape Wine R. 15, 85–92. doi: 10.1111/j.1755-0238.2008.00040.x
Trotter, E. W., Collinson, E. J., Dawes, I. W., Grant, C. M. (2006). Old yellow enzymes protect against acrolein toxicity in the Yeast Saccharomyces cerevisiae. Appl. Environ. Microbiol. 72, 4885–4892. doi: 10.1128/AEM.00526-06
Upadhyaya, R., Panda, S. K. (2010). Zinc reduces copper toxicity induced oxidative stress by promoting antioxidant defence in freshly grown aquatic duckweed Spirodela polyrhiza L. J. Hazard. Mater. 175, 1081–1084. doi: 10.1016/j.jhazmat.2009.10.016
Vishwakarma, K., Upadhyay, N., Kumar, N., Yadav, G., Singh, J., Mishra, R. K., et al. (2017). Abscisic acid signaling and abiotic stress tolerance in plants: a review on current knowledge and future prospects. Front. Plant Sci. 8. doi: 10.3389/fpls.2017.00161
Wang, Y., Wang, Y., Kai, W., Zhao, B., Chen, P., Sun, L., et al. (2014). Transcriptional regulation of abscisic acid signal core components during cucumber seed germination and under Cu2+, Zn2+, NaCl and simulated acid rain stresses. Plant Physiol. Biochem. 76, 67–76. doi: 10.1016/j.plaphy.2014.01.003
Wang, C., Wang, X., Kibet, N. K., Song, C., Zhang, C., Li, X., et al. (2011). Deep sequencing of grapevine flower and berry short RNA library for discovery of new microRNAs and validation of precise sequences of grapevine microRNAs deposited in miRBase. Physiol. Plant 143, 64–81. doi: 10.1111/j.1399-3054.2011.01481.x
Wang, Y., Yuan, G., Yuan, S., Duan, W., Wang, P., Bai, J., et al. (2016a). TaOPR2 encodes a 12-oxo-phytodienoic acid reductase involved in the biosynthesis of jasmonic acid in wheat (Triticum aestivum L.). Biochem. Biophys. Res. Commun. 470, 233–238. doi: 10.1016/j.bbrc.2016.01.043
Wang, F. B., Zhu, H., Chen, D. H., Li, Z. J., Peng, R. H., Quan, H. Y. (2016b). A grape bHLH transcription factor gene, VvbHLH1, increases the accumulation of flavonoids and enhances salt and drought tolerance in transgenic Arabidopsis thaliana. Plant Cell Tiss. 125, 387–398. doi: 10.1007/s11240-016-0953-1
Wellburn, A. R. (1994). The spectral determination of chlorophyll a and chlorophyll b, as well as total carotenoids, using various solvents with spectrophotometers of different resolution. J. Plant Physiol. 144, 307–313. doi: 10.1016/S0176-1617(11)81192-2
Wu, L., Wang, R., Li, M., Du, Z., Jin, Y., Jiang, W., et al. (2024). Functional analysis of a rice 12-oxo-phytodienoic acid reductase gene (OsOPR1) involved in Cd stress tolerance. Mol. Biol. Rep. 51, 198. doi: 10.1007/s11033-023-09159-w
Xin, Z., Zhang, J., Ge, L., Lei, S., Han, J., Zhang, X., et al. (2017). A putative 12-oxophytodienoate reductase gene CsOPR3 from Camellia sinensis, is involved in wound and herbivore infestation responses. Gene 615, 18–24. doi: 10.1016/j.gene.2017.03.013
Xiong, A. S., Yao, Q. H., Peng, R. H., Li, X., Fan, H. Q., Cheng, Z. M., et al. (2004). A simple, rapid, high-fidelity and cost-effective PCR-based two-step DNA synthesis method for long gene sequences. Nucleic. Acids Res. 32, e98. doi: 10.1093/nar/gnh094
Yan, Y., Christensen, S., Isakeit, T., Engelberth, J., Meeley, R., Hayward, A., et al. (2012). Disruption of OPR7 and OPR8 reveals the versatile functions of jasmonic acid in maize development and defense. Plant Cell 24, 1420–1436. doi: 10.1105/tpc.111.094151
Yan, Z., Zhang, W., Chen, J., Li, X. (2015). Methyl jasmonate alleviates cadmium toxicity in Solanum nigrum by regulating metal uptake and antioxidative capacity. Biol. Plant 59, 373–381. doi: 10.1007/s10535-015-0491-4
Yruela, I. (2005). Copper in plants. Braz. J. Plant Physiol. 17, 145–156. doi: 10.1590/S1677-04202005000100012
Yue, Y., Zhang, M., Zhang, J., Tian, X., Duan, L., Li, Z. (2012). Overexpression of the AtLOS5 gene increased abscisic acid level and drought tolerance in transgenic cotton. J. Exp. Bot. 63, 3741–3748. doi: 10.1093/jxb/ers069
Zalamena, J., Melo, G. W., Santos, H. P., Silva, L. S., Fialho, F. B., Brunetto, G. (2015). Physiological characterization of grapevine rootstocks grown in soil with increasing zinc doses. Rev. Bras. Eng. Agríc. Ambient. 19, 973–980. doi: 10.1590/1807-1929/agriambi.v19n10p973-980
Zhang, X., Henriques, R., Lin, S. S., Niu, Q. W., Chua, N. H. (2006). Agrobacterium-mediated transformation of Arabidopsis thaliana using the floral dip method. Nat. Protoc. 1, 641–646. doi: 10.1038/nprot.2006.97
Zhang, J. L., Simmons, C., Yalpani, N., Crane, V., Wilkinson, H., Kolomiets, M. (2005). Genomic analysis of the 12-oxo-phytodienoic acid reductase gene family of Zea mays. Plant Mol. Biol. 59, 323–343. doi: 10.1007/s11103-005-8883-z
Zheng, H., Pan, X., Deng, Y., Wu, H., Liu, P., Li, X. (2016). AtOPR3 specifically inhibits primary root growth in Arabidopsis under phosphate deficiency. Sci. Rep. 6, 24778. doi: 10.1038/srep24778
Keywords: reductase, grapevine, gene family, copper and zinc stress, mechanism
Citation: You S-H, Chen Y-P, Shi W-J, Li X, Wu Z and Yao Q-H (2025) Genome-wide analysis of OPR family genes in Vitis vinifera and the role of VvOPR1 in copper, zinc tolerance. Front. Plant Sci. 16:1509472. doi: 10.3389/fpls.2025.1509472
Received: 11 October 2024; Accepted: 05 February 2025;
Published: 26 February 2025.
Edited by:
Peng Wang, Jiangsu Province and Chinese Academy of Sciences, ChinaReviewed by:
Guo-Bnag Li, Sichuan Agricultural University, ChinaCopyright © 2025 You, Chen, Shi, Li, Wu and Yao. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Zheng Wu, d3poZW5nMDkyNUAxNjMuY29t; Quan-Hong Yao, eWFvcXVhbmhvbmc4OEAxNjMuY29t
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.