- 1Key Laboratory of Oasis Eco-Agriculture, College of Agriculture, Shihezi University, Shihezi, China
- 2Shenzhen Branch, Guangdong Laboratory for Lingnan Modern Agriculture, Genome Analysis Laboratory of the Ministry of Agriculture, Agricultural Genomics Institute at Shenzhen, Chinese Academy of Agricultural Sciences, Shenzhen, China
- 3CSIRO Agriculture and Food, Canberra, ACT, Australia
Verticillium wilt, caused by Verticillium dahliae, is one of the most damaging and widespread soil-borne cotton diseases. The molecular mechanisms underlying the cotton defense against V. dahliae remain largely elusive. Here, we compared the transcriptional differences between Upland cotton cultivars: one highly resistant (HR; Shidalukang 1) and one highly susceptible (HS; Junmian 1). This was done at multiple time points after V. dahliae inoculation, which identified 2010 and 1275 differentially expressed genes (DEGs) in HR and HS, respectively. Plant hormone signal transduction-related genes were enriched in HR, whereas genes related to lignin biosynthesis were enriched in both HR and HS. Weighted gene co-expression network analysis (WGCNA) using the 2868 non-redundant genes differentially expressed between the V. dahliae infected and uninfected samples in HR or HS identified 10 different gene network modules and 22 hub genes with a potential role in regulating cotton defense against V. dahliae infection. GhGDH2, encoding glutamate dehydrogenase (GDH), was selected for functional characterization. Suppressing the expression level of GhGDH2 by virus-induced gene silencing (VIGS) in HS led to inhibition of the salicylic acid (SA) biosynthesis/signaling pathways and activation of the jasmonic acid (JA) biosynthesis/signaling pathways, which resulted in an increase of 42.1% JA content and a reduction of 78.9% SA content in cotton roots, and consequently enhanced V. dahliae resistance. Our finding provides new insights on the molecular mechanisms of cotton resistance to V. dahliae infection and candidate genes for breeding V. dahliae resistance cotton cultivars by genetic modification.
Introduction
Cotton (Gossypium hirsutum L.) is an economically important crop of the Malvaceae family, providing a source of renewable natural fiber and oilseeds. Cotton is continuously subjected to various diseases during its growth and development, which influence the yield and quality of cotton fiber. Verticillium dahliae is a soil-borne fungus and cause of vascular disease known as Verticillium wilt, which is one of the most common and serious diseases in cotton-producing regions worldwide (Cai et al., 2009). V. dahliae can infect cotton by penetrating roots during the whole growing season and result in wilting and defoliation of leaves, and even plant death (Shaban et al., 2018). Breeding resistant cultivars are the most effective, economical, safe, and environmentally friendly method for controlling Verticillium wilt. Due to the lack of disease-resistant germplasm resources in G. hirsutum and variation in V. dahliae strains, the breeding progress on Verticillium wilt resistance is slow (Wang et al., 2016). Understanding the molecular mechanism of Verticillium wilt resistance and identifying Verticillium wilt resistance genes are of great significance for improving Verticillium wilt resistance in cotton by genetic engineering.
Gossypium barbadense is more resistant to V. dahliae than G. hirsutum (Zhang et al., 2013). Previous studies have shown that a few major genes control the resistance of G. barbadense to V. dahliae, whereas V. dahliae resistance in G. hirsutum seems to be controlled by multiple genes (Zhang et al., 2013; Li et al., 2017). Transcriptomic analysis is an effective and widely used approach to identify genes differentially expressed in different biological processes or response to environmental stresses (Wang et al., 2009). One of the powerful approaches to investigate the mechanisms of cotton resistance to V. dahliae is to analyze the basal transcriptome and its response to V. dahliae infection in cotton cultivars with different levels of resistance. RNA-sequencing studies with an interest on the mechanisms of cotton – V. dahliae interaction have been carried out using various V. dahliae strains in different cotton species (Xu et al., 2011b; Sun et al., 2013; Zhang et al., 2013, 2016; Dong et al., 2019; Li C. et al., 2019), but only one study compared the transcriptomic differences between resistant and susceptible G. hirsutum varieties (Zhang et al., 2017), in which phenylpropanoid biosynthesis and various plant hormone signaling pathways were found to play an important role in G. hirsutum defense against V. dahliae.
Previous studies on the interaction between plants and V. dahliae found changes in phenylpropanoid biosynthesis in plants after V. dahliae infection, in particular, the lignin biosynthesis pathway was found to be activated (Xu et al., 2011b; Zhang et al., 2013; Dhar et al., 2020). Inhibition of the expression of the lignin biosynthesis related genes GhHCT1 and GhLac1 enhanced susceptibility of cotton to V. dahliae due to decreased activity of lignin biosynthesis (Guo et al., 2016; Hu et al., 2018). Several genes such as GhERF1, GhARPL18A-6, GhCOMT, and GhSNAT1 have been shown to positively mediate cotton resistance to V. dahliae by increasing lignin content (Guo et al., 2016; Li et al., 2019; Zhang et al., 2019). Jasmonic acid (JA) and salicylic acid (SA) are the two important defense signal molecules in plant immune response (Thaler et al., 2012). SA and JA play a key role in resistance against biotrophic and necrotrophic pathogens, respectively. Antagonistic interaction between SA and JA in defense response has been well documented in Arabidopsis (Thaler et al., 2012). Various studies have revealed that both JA and SA positively regulate cotton defense against V. dahliae infection (Shaban et al., 2018; Dhar et al., 2020). However, it is still unclear whether SA and JA have an antagonistic relationship in the process of cotton response to V. dahliae infection.
To explore the molecular mechanisms associated with V. dahliae resistance in G. hirsutum, we have sequenced the transcriptomes of Shidalukang 1 (HR; V. dahliae-resistant) and Junmian 1 (HS; V. dahliae-susceptible) at different time points after V. dahliae infection. In our previous investigation, we found that the basal transcriptome landscape between HR and HS has a substantial role in their responses to V. dahliae infection and that V. dahliae resistance is affected by the metabolites of the lignin biosynthesis pathway (Xiong et al., 2021). Here, we focused on the changes of transcriptomes in response to V. dahliae infection in HR and HS, and identified significantly enriched gene ontology (GO) terms and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways using the genes induced or repressed by V. dahliae infection in HR and HS. Moreover, 22 hub genes associated with cotton response to V. dahliae infection were identified by weighted gene co-expression network analysis (WGCNA), and the function of one (GhGDH2) of the hub genes was characterized through virus-induced gene silencing (VIGS). This work enriched our knowledge on the molecular-defense networks of G. hirsutum in response to V. dahliae infection and identified candidate genes for enhancing cotton resistance to V. dahliae.
Materials and Methods
Plant Materials and Verticillium dahliae Inoculation
Two G. hirsutum varieties, Shidalukang 1 (V. dahliae-resistant; HR) and Junmian 1 (V. dahliae-susceptible; HS), from our own seed stock were used in RNA-seq analysis. Root and stem samples were collected at 0 (control), 12, 24, and 48 h after inoculated with V. dahliae and a mixture of 200 mg roots and 200 mg stems was used in RNA extraction and RNA-seq. The VIGS experiments and disease assays were carried out using HS. Plants were cultivated in a greenhouse under 24/22°C and a photoperiod scheme of 16-h light/8-h dark. Disease assays were performed using the highly aggressive defoliating V. dahliae strain V991 kindly provided by Associate Professor Yanfei Sun of the Shihezi University. Two-leaf stage seedlings were inoculated with V. dahliae by the root irrigation method (Xiong et al., 2019).
Transcriptome Analysis
Construction and sequencing of RNA-seq libraries (24 in total, i.e., 4 time points × 2 cultivars × 3 biological replicates) from HR and HS have been previously described (Xiong et al., 2021). The raw short-reads containing adapter and/or ploy-N and low quality were removed using the Trimmomatic v0.30 program (Bolger et al., 2014). All the clean reads were aligned to the TM-1 genome (Zhang et al., 2015) using TopHat v2.0.12. Genes induced or repressed after V. dahliae inoculation in HR and HS were identified using the R package DESeq2 (Love et al., 2014). FPKM (fragments per kilobase of transcript sequence per million base pairs sequenced) was used to represent gene expression level (Trapnell et al., 2010). Genes with a | log2 (fold-change)| ≥ 1 and an adjusted P value < 0.01 were considered to be differentially expressed. Venn diagram and heatmap were generated using the Novemagic server1. The raw data of RNA-seq are available in the National Center for Biotechnology Information (NCBI) under the accession no. PRJNA593765.
A total of 2010 and 1,275 differentially expressed genes (DEGs) from HR and HS were used in analyses of GO terms and KEGG pathways by the GOseq and KOBAS 2.0 R package, respectively (Young et al., 2010; Xie et al., 2011). GO terms and KEGG pathways with a corrected P value < 0.05 were regarded as significant and selected for further analysis.
Co-expression Network Analysis
The FPKM of the 2868 non-redundant DEGs from HR and HS was used to construct the co-expression networks by the R package WGCNA (Langfelder and Horvath, 2008). First, the network was constructed in accordance with the scale-free topology criterion using a β of 14 as a weight function. The co-expression modules were identified with the default settings, except that the TOMType was assigned, and minModuleSize and merge CutHeight were set to 30 and 0.25, respectively. Each module was assigned by a unique color; genes not assigned to any module were grouped into the gray module.
Salicylic Acid and Methyl Jasmonate Treatments
For the SA and methyl jasmonate (MeJA) treatment experiments, two-leaf stage seedlings of Shidalukang 1 were sprayed by 1 mmol L–1 SA or 100 μmol L–1 MeJA. Root samples were collected from five individual cotton seedlings at 0, 12, 24, 48, and 72 h after treatments. Moreover, the roots, stems, and leaves from five individual seedlings of Shidalukang 1 or Junmian 1 at two-leaf stage were collected to measure the expression profiles of GhGDH2 and GhGDH3. Three independent biological replicates were sampled at each time points for both SA and MeJA treatments.
RNA Extraction and Real-Time Quantitative PCR Analysis
Total RNA was isolated using an RN09-EASYspin RNA Plant Mini Kit (Aidlab, Beijing, China) from roots, stems, and leaves collected from seedlings treated with SA, MeJA, or V. dahliae, respectively. The quality and concentration of the extracted RNA were determined by a NanoPhotometer spectrophotometer (IMPLEN, CA, United States). Total RNA (3 μg) was used in synthesis of cDNA template using the PrimeScript II 1st Strand cDNA Synthesis Kit (TaKaRa, Dalian, China). Then, 2 μl of a 10-times diluted cDNA was used in quantitative PCR (qPCR) using the SYBR Green kit (Roche, Rotkreuz, Switzerland) on a Light-Cycler 480II (Roche). The qPCR program comprised an initial denaturation step at 95°C for 10 min, followed by 40 cycles at 95°C for 15 s and 60°C for 15 s, and 72°C for 15 s. qPrimerDB2 was used to design the real-time quantitative PCR (RT-qPCR) primers (Supplementary Table 1). GhUBQ7 (DQ116441.1) was used as the internal control and the relative gene expression level from three biologically independent samples was evaluated using the 2–Δ Δ CT method (Livak and Schmittgen, 2001).
Virus-Induced Gene Silencing
Tobacco rattle virus (TRV)-based vectors were used in the VIGS experiments to explore the function of GhGDH2 in cotton defense against V. dahliae. TRV:GhCHLI was used as a positive control for the VIGS experiment as previously reported (Xiong et al., 2019). A 405-bp fragment specific to GhGDH2 (Gh_D03G0992) was amplified by PCR from HR with gene-specific primers (Supplementary Table 1). VIGS was performed with the same procedures as previously described (Gao et al., 2013a). Approximately 14 days after infiltration, the roots of five TRV:00 and TRV:GhGDH2 plants were collected to measure the expression level of GhGDH2 by RT-qPCR. The RT-qPCR primer sequences for GhGDH2 are listed in Supplementary Table 1. Thirty TRV:00 and TRV:GhGDH2 plants at the two-leaf-stage were subjected to V. dahliae inoculation to compare their disease response.
Evaluation Verticillium dahliae Resistance of VIGS Cotton Plants
The ratio of diseased plants and disease index were calculated using thirty TRV:00 and TRV2:GhGDH2 plants at 14 and 21 days post-infection (dpi) according to the following formula: ratio of diseased plants = (the number of diseased plants/the total number of inoculated plants) × 100%. Disease index = [(Σ disease grade × the number of plants with the corresponding disease grade)/(4 × the total number of inoculated plants)] × 100. Cotton seedlings were classified into five grades (i.e., grades 0, 1, 2, 3, and 4) according to the phenotypes on the true leaves of TRV:00 and TRV2:GhGDH2 plants after V. dahliae infection (Gao et al., 2013a). The final results were the average of three independent experiments.
The stems above cotyledons of TRV:00 and TRV2:GhGDH2 plants were hand-sectioned at 14 dpi and observed using a stereomicroscope (SteREO Discovery.V20; ZEISS). For fungal recovery assay, ten 1-cm stem sections above the cotyledons of cotton plants were cut at 14 dpi and surface sterilized in 3% hypochlorite for 3 min. After rinsing three times with sterile water, the stem sections were plated on PDA medium for a week at 25°C in darkness. The relative fungal biomass was determined as previously described (Ellendorff et al., 2009). Briefly, the stems above the cotyledons of TRV:00 and TRV2:GhGDH2 plants were collected at 14 and 21 dpi for DNA extraction. V. dahliae specific primers ITS1-F and ST-Ve1-R and primers for the cotton UBQ7 gene were used to quantify fungal biomass by qPCR (Supplementary Table 1).
Measurement of JA and SA Content
The contents of endogenous SA and JA were measured as described previously with minor modifications (Sun et al., 2014). Briefly, at 72 h post-inoculation (hpi), root samples were collected from mock (inoculated with sterile water) and V. dahliae-inoculated TRV:00 and TRV:GhGDH2 plants. Approximately 200 mg of fresh root sample was ground to powder with liquid nitrogen and then extracted twice with 2 ml pre-cold 80% methanol overnight under dark at 4°C. The supernatant was dried by N2 at room temperature, then re-dissolved in 0.4 ml methanol. After filtering through a 0.22-μm membrane, the collected supernatant was used in quantification of the content of SA and JA by the high-performance liquid chromatography–tandem mass spectrometry (HPLC/MS) system (AB SCIEX Triple Quad 5500). Pure SA or JA (both ordered from Sigma) was used as a respective internal standard.
Data Analysis
SPSS (version 23.0; IBM, United States) was used for the statistical analysis. Data were checked for normality and homogeneity of variances with the Shapiro–Wilk test (α = 0.05) and the Levene test (α = 0.05), respectively. The differences between groups were compared using Student’s t-test (∗P < 0.05; ∗∗P < 0.01).
Results
Differentially Expressed Genes Upon Verticillium dahliae Infection
To characterize the genes responsive to V. dahliae infection in cotton cultivars with different levels of resistance, we compared the expression level of genes in the 12, 24, and 48 hpi samples with that in the uninfected sample (0 hpi). Based on the criteria of adjusted P value < 0.01 and | log2 (fold change)| ≥ 1, 775, 1004, and 1094 DEGs were observed at 12, 24, and 48 hpi in HR, respectively (Figure 1A), and 542, 655, and 683 DEGs were found at 12, 24, and 48 hpi in HS, respectively (Figure 1B). In total, 2010 and 1,275 non-redundant DEGs were identified in HR and HS, respectively (Supplementary Tables 2, 3). Among them, 182 and 119 genes were induced or repressed by V. dahliae infection at all three time points in HR and HS, respectively (Figures 1A,B). The number of DEGs was 48 hpi > 24 hpi > 12 hpi in both HR and HS, and with more DEGs in HR than in HS at each time point (Figure 1C). HR had more induced genes than repressed genes at all three time points, but HS showed the opposite trends (Figure 1C). Overall, 2868 non-redundant DEGs were detected in HR and HS with 417 DEGs common in both cultivars (Supplementary Figure 1 and Supplementary Table 4).
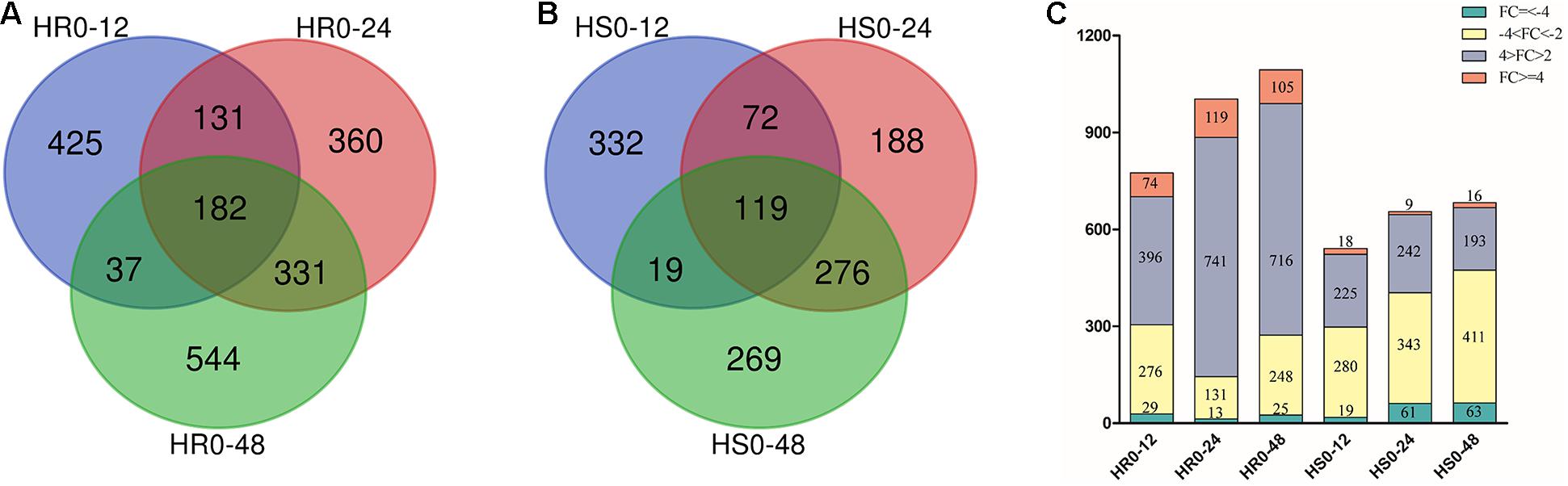
Figure 1. Comparison of the number of genes differentially expressed in Shidalukang 1 (HR) and Junmian 1 (HS) at 12, 24, and 48 h after Verticillium dahliae infection. (A,B) The number of differentially expressed genes (DEGs) unique to or overlapping between each time point in (A) HR and (B) HS following V. dahliae infection. (C) The number of DEGs with different fold-change in HR and HS after V. dahliae infection.
Functional Annotation of DEGs
To explore the potential functionality of the DEGs identified in HR and HS, we carried out GO enrichment analyses. The 2010 DEGs in HR were significantly enriched with 114 GO terms (Supplementary Table 5). Biological processes associated with single-organism metabolic process, oxidation–reduction process and oxidoreductase activity were the most significantly enriched GO terms in HR (Figure 2A and Supplementary Table 5). In HS, the 1,275 DEGs were significantly (corrected P value < 0.05) enriched with 41 GO terms, including single-organism metabolic process, catalytic activity, and oxidation–reduction process (Figure 2A and Supplementary Table 6). Notably, of the top 20 GO terms enriched in HR or HS, seven overlapped between HR and HS (Figure 2A), including heme binding, oxidation–reduction process, nucleic acid binding transcription factor activity, oxidoreductase activity, transcription factor activity sequence-specific DNA binding, and tetrapyrrole binding (Figure 2A). Biological processes involved in response to endogenous stimulus and hormone were only enriched in HR (Figure 2A).
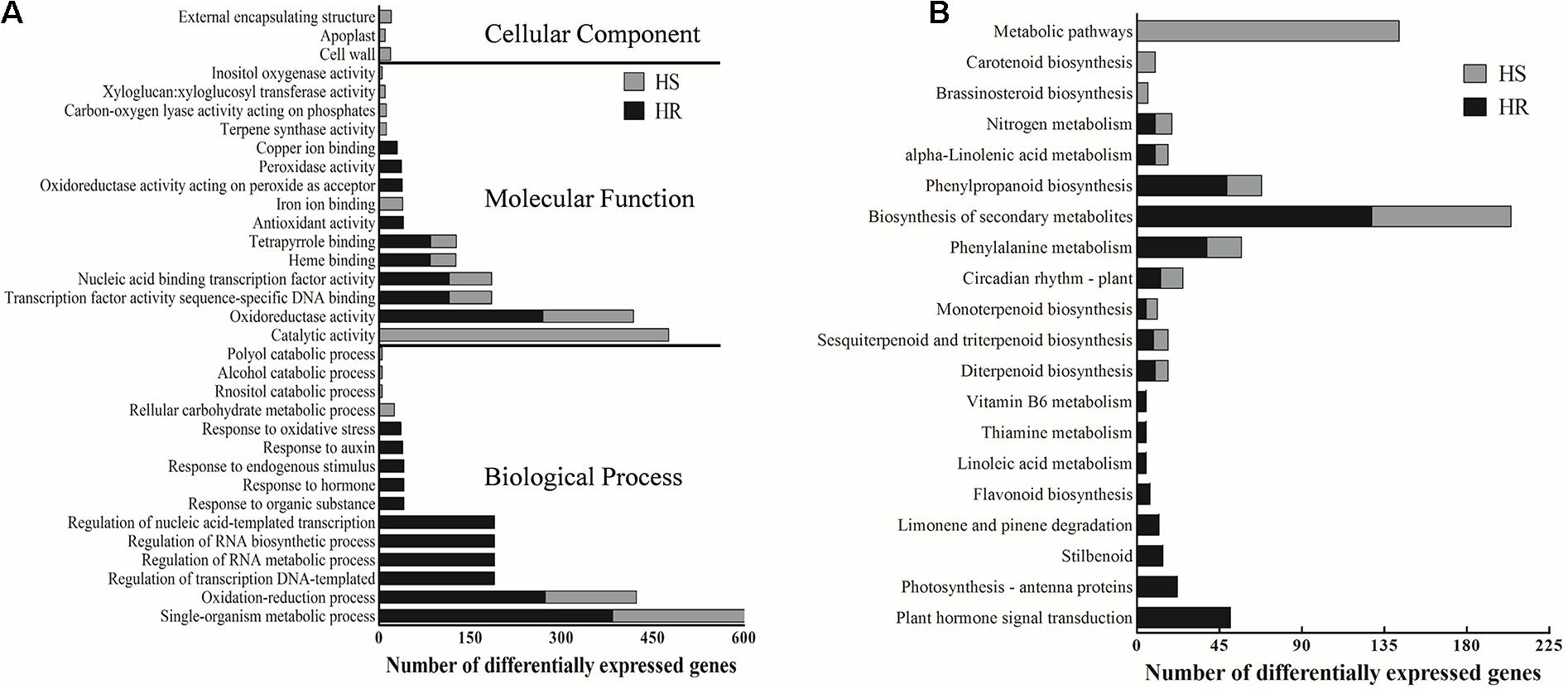
Figure 2. Gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis of the 2010 and 1,275 differentially expressed genes (DEGs) identified in Shidalukang 1 (HR) and Junmian 1 (HS) after Verticillium dahliae infection, respectively. (A) The top 20 significant (P < 0.05) GO terms enriched in the 2010 and 1,275 DEGs identified in HR and HS, respectively. (B) The significant (P < 0.05) KEGG pathways of the 2010 and 1275 DEGs identified in HR and HS, respectively.
Kyoto Encyclopedia of Genes and Genomes pathway analysis results showed that the 2010 DEGs identified in HR were significantly enriched with 17 pathways, while the 1275 DEGs identified in HS were enriched with 12 pathways (Figure 2B and Supplementary Tables 7, 8). Phenylpropanoid biosynthesis, phenylalanine metabolism, alpha-linolenic acid metabolism, diterpenoid biosynthesis, monoterpenoid biosynthesis, biosynthesis of secondary metabolites, circadian rhythm-plant, sesquiterpenoid and triterpenoid biosynthesis, and nitrogen metabolism were significantly enriched in both HR and HS (Figure 2B). Pathways related to plant hormone signal transduction and flavonoid biosynthesis were significantly enriched only in HR, and the brassinosteroid (BR) biosynthesis pathway was enriched only in HS (Figure 2B).
Differentially Expressed Genes Involved in Plant Hormone Signal Transduction
Previous studies have reported that plant hormones, including SA, JA, ethylene (ET), and BR, positively regulate resistance to V. dahliae in cotton (Shaban et al., 2018). In this study, 12 DEGs involved in SA, JA, ET, and BR signaling pathways were identified (Figure 3A and Supplementary Table 9). RNA-seq and RT-qPCR results indicated that the expression level of GhJAZ1-2, GhERF1-1, and GhBAK1 was significantly increased in 12 to 48 hpi roots and stems of HR (except GhJAZ1-2 at 48 hpi), but decreased in HS (Figures 3A–D,F–H). In the roots of HR, GhPR1 showed upregulation at 24 hpi but downregulation at 48 hpi; in the stems of HR, GhPR1 showed constant but gradually reduced upregulation from 12 to 48 hpi (Figure 3E). In HS, a significant upregulation of GhPR1 was observed in the 48 hpi roots and 12–24 hpi stems (Figure 3I).
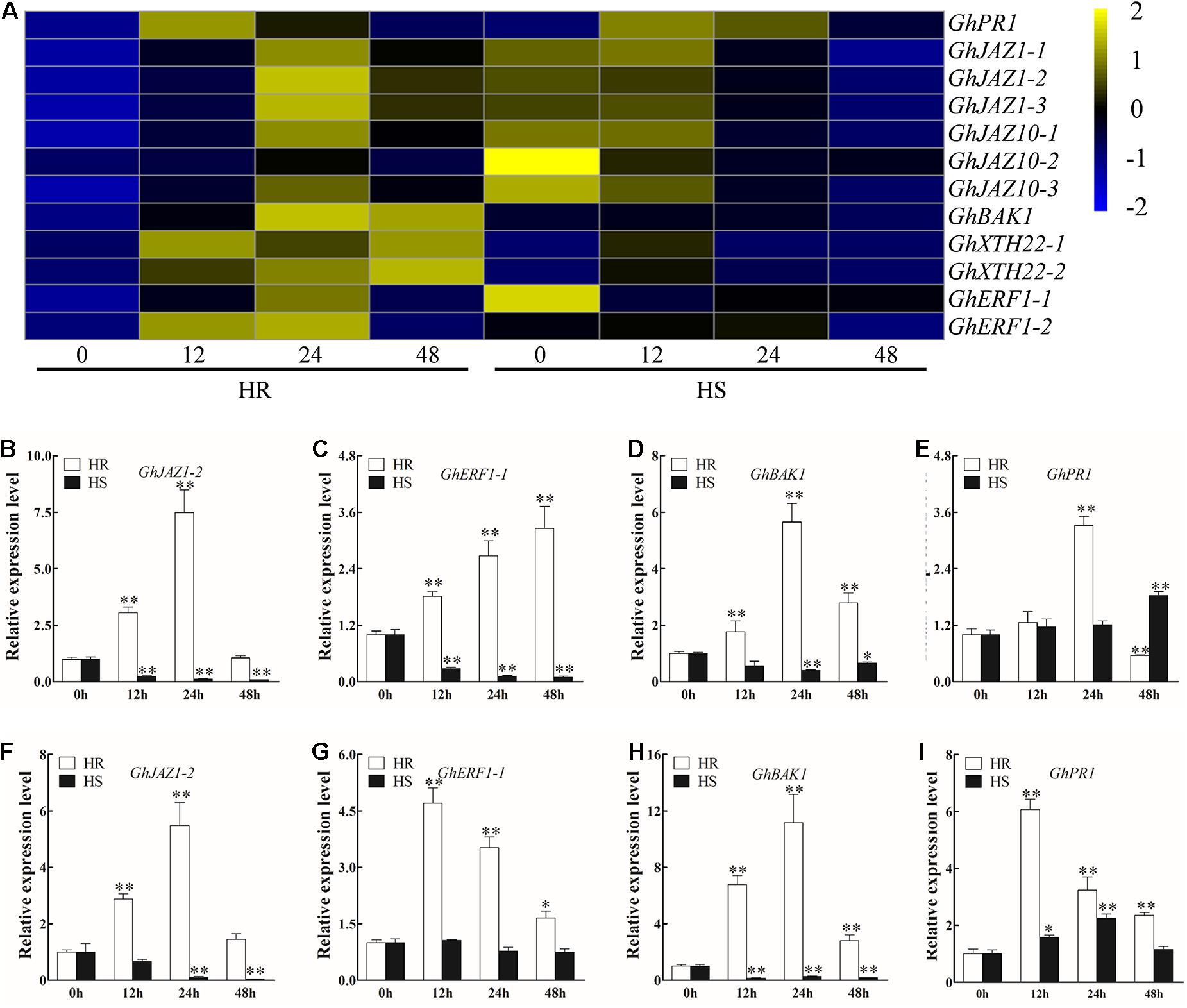
Figure 3. Transcriptional profile of plant hormone signal transduction-related genes after Verticillium dahliae infection. (A) Heatmap showing the expression pattern of plant hormone signal transduction-related genes in Shidalukang 1 (HR) and Junmian 1 (HS) at 0, 12, 24, and 48 h post-inoculation (hpi). The heatmap was generated using fragments per kilobase of transcript sequence per million base pairs sequenced values normalized by Z-score. (B–E) Relative expression levels of plant hormone signal transduction-related genes (B) GhJAZ1-2, (C) GhERF1-1, (D) GhBAK1, and (E) GhPR1 in the roots of HR and HS at 0, 12, 24, and 48 hpi. (F–I) Relative expression levels of plant hormone signal transduction–related genes (F) GhJAZ1-2, (G) GhERF1-1, (H) GhBAK1, and (I) GhPR1 in the stems of HR and HS at 0, 12, 24, and 48 hpi. Values represent mean ± SE of three independent biological replications (five individual plants per replicate). For each cultivar, the expression level at each time point was compared with that at 0 hpi that was set to 1. ∗ and ∗∗ represent significant differences from roots of the same cultivar at P < 0.05 and P < 0.01, respectively, based on Student’s t-test.
Lignin Biosynthesis-Related Genes
Lignin has been reported to play a critical role in plant immune response to pathogens, including to V. dahliae (Vanholme et al., 2010; Hu et al., 2019). Transcriptome data showed that most DEGs involved in lignin biosynthesis, including two Gh4CL, four GhCOMT, two GhF5H, and one GhCAD, were increased in both HR and HS following V. dahliae infection, although GhC4H1 and GhCOMT1-5 were repressed and activated in HS and HR, respectively (Figure 4A and Supplementary Table 9). Four (GhC4H1, GhCOMT1-1, GhF5H1-1, and GhCAD1) of those 11 genes were selected for RT-qPCR verification. In agreement with the RNA-seq results, the expression level of all four genes was increased in at least one time point in the roots and stems of HR after V. dahliae infection. In HS, the RT-qPCR results of GhF5H1-1, GhCAD1, and GhC4H1 were in line with the RNA-seq results (Figures 4B–I), although an inconsistent result was observed for GhCOMT1-1 (Figures 4F,G).
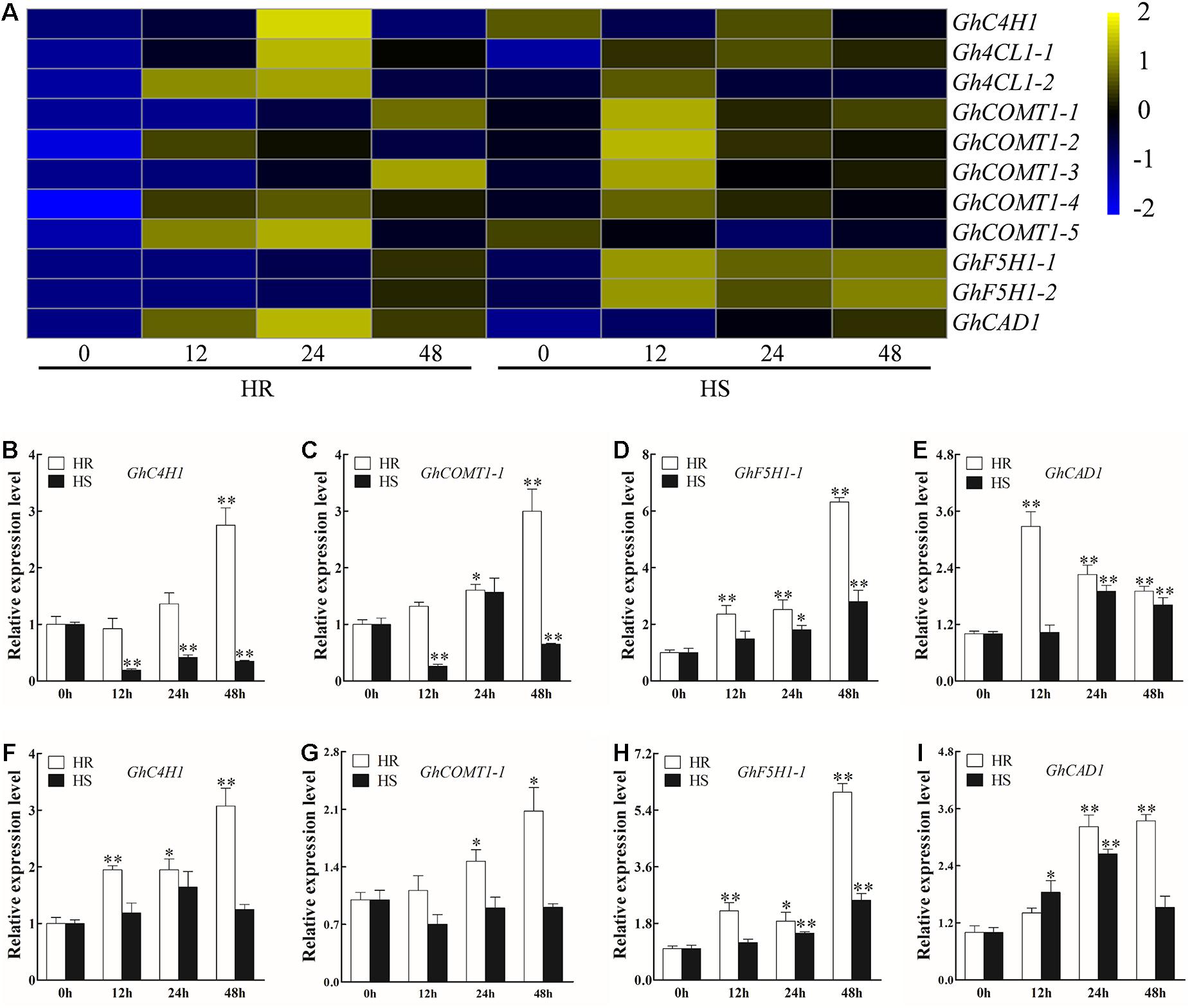
Figure 4. Transcriptional profile of genes involved in lignin biosynthesis after Verticillium dahliae infection. (A) Heatmap showing the expression pattern of lignin biosynthesis-related genes in HR and Junmian 1 (HS) at 0, 12, 24, and 48 h post-inoculation (hpi). The heatmap was generated using fragments per kilobase of transcript sequence per million base pairs sequenced values normalized by Z-score. (B–E) Real-time quantitative PCR (RT-qPCR) analysis of lignin biosynthesis-related genes (B) GhC4H1, (C) GhCOMT1-1, (D) GhF5H1-1, and (E) GhCAD1 in the roots of HR and HS at 0, 12, 24, and 48 hpi. (F–I) RT-qPCR analysis of lignin biosynthesis-related genes (F) GhC4H1, (G) GhCOMT1-1, (H) GhF5H1-1, and (I) GhCAD1 in the stems of HR and HS at 0, 12, 24, and 48 hpi. For each cultivar, the expression level at each time point was compared with that at 0 hpi that was set to 1. Values represent mean ± SE of three independent biological replications (five individual plants per replicate). ∗ and ∗∗ represent significant differences from roots of the same cultivar at P < 0.05 and P < 0.01, respectively, based on Student’s t-test.
Weighted Gene Co-expression Network Analysis
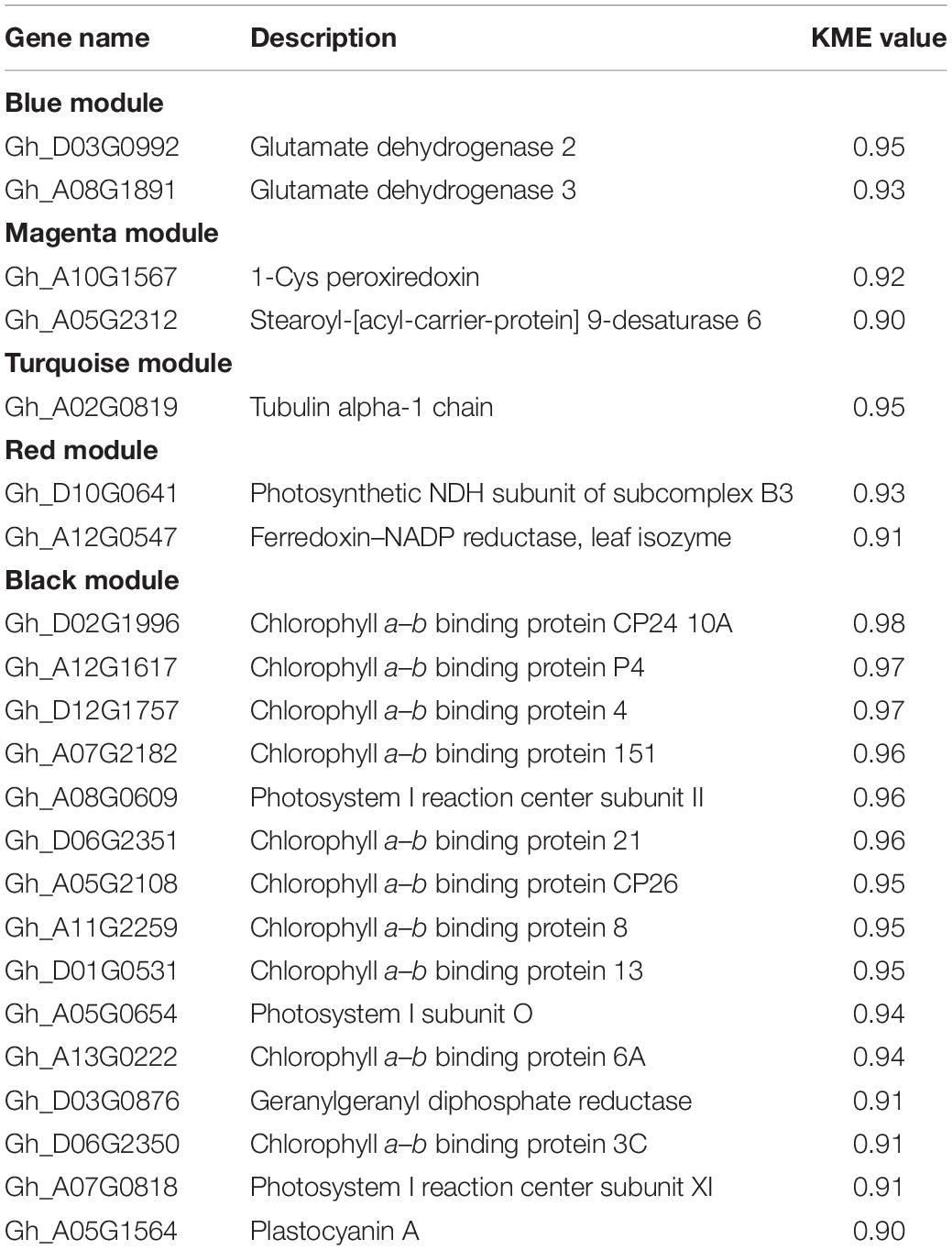
To determine the potential genes regulating cotton resistance to V. dahliae, 2868 non-redundant DEGs from HR and HS were subjected to WGCNA. A total of 10 gene network modules were obtained according to pairwise correlation analysis of gene expression across all samples (Figure 5A and Supplementary Table 10). As shown in Figure 4B, five module–trait relationships (indicated with red underlines) were highly significant (P value < 0.001) (Figure 5B). Genes in the blue module had a low expression level in HR0 while genes in the black and turquoise modules were expressed at lower levels in HS0. The transcript levels of the magenta module genes were quickly increased at 12 and 24 hpi in HR, but not significantly different after V. dahliae infection in HS. The transcript levels of the red module genes were rapidly increased at 12 hpi in both HR and HS (Figure 5B). In these five modules, a total of 22 hub genes related to V. dahliae resistance in cotton were identified using the criteria of KME > 0.9 and PPI > 5 (Table 1).
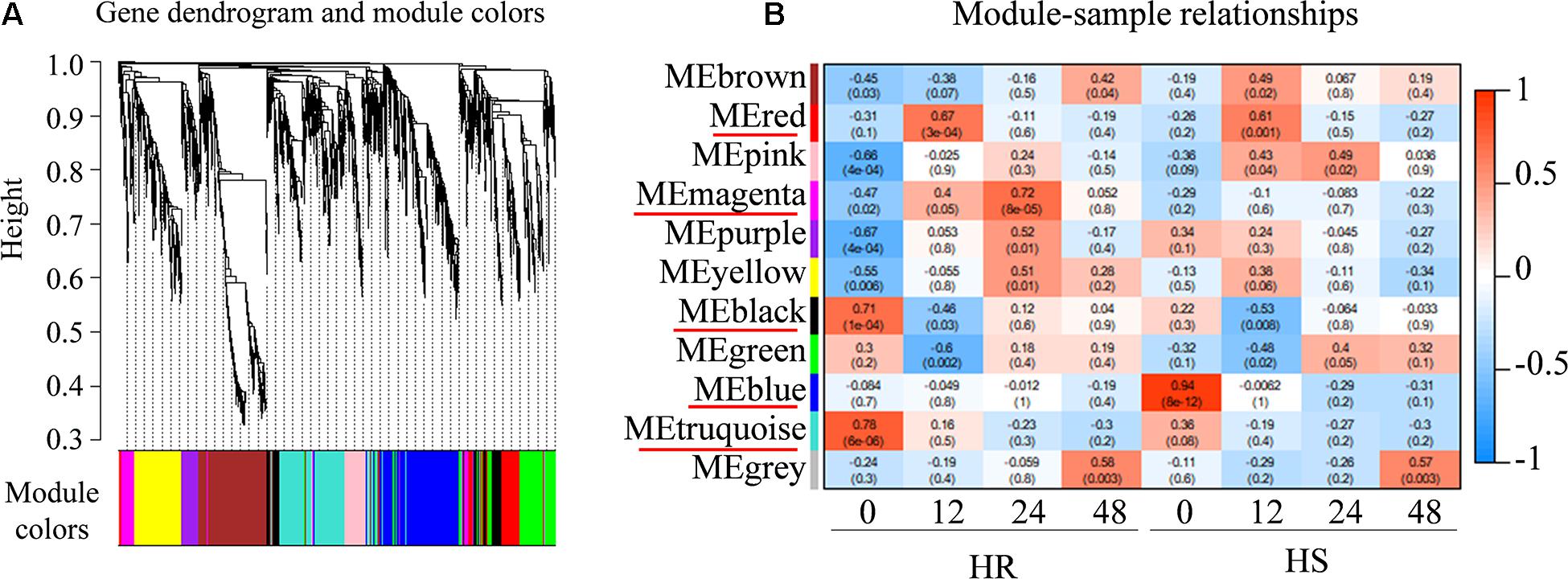
Figure 5. Weighted gene co-expression network analysis (WGCNA) of differentially expressed genes (DEGs) from Shidalukang 1 (HR) and Junmian 1 (HS) at 0, 12, 24, and 48 h post-inoculation. (A) Hierarchical cluster tree of WGCNA analysis. In total, 2,868 DEGs were grouped into 11 co-expression modules labeled with different colors based on the calculation of eigengenes. Each leaf in the tree and each major tree branch represent one gene and a distinct module, respectively. (B) Module–sample correlation. The color at the row–column intersection shows the level of correlation between each module and sample. The modules showing a high degree of correlation (P < 0.001) with samples are indicated by a red underline.
The Expression Profiles of GhGDH2 and GhGDH3
In our previous studies with a focus on the basal transcriptome difference between HR and HS, GhGDH3 (Gh_A08G1891) was characterized as a hub gene potentially associated with the defense response against V. dahliae infection in cotton (Xiong et al., 2021). Here, based on the genes induced or repressed by V. dahliae infection in HR and HS, GhGDH3 together with one of its homologs, GhGDH2 (Gh_D03G0992), were identified as hub genes of the gene networks related to V. dahliae response, suggesting that GhGDH2 and GhGDH3 may play an essential role in cotton defense against V. dahliae infection. To test this hypothesis, we analyzed their transcript levels in roots, stems, and leaves of uninfected HR and HS, and their response to treatment of SA or MeJA as well as to V. dahliae inoculation. The expression level of both GhGDH2 and GhGDH3 was significantly higher in HS than in HR (Figures 6A,B). In the three tissues analyzed, the transcript level of GhGDH2 in roots was higher than that in stems and leaves in both HR and HS, while the transcript level of GhGDH3 in the roots and leaves was significantly higher than that in stems (Figures 6A,B).
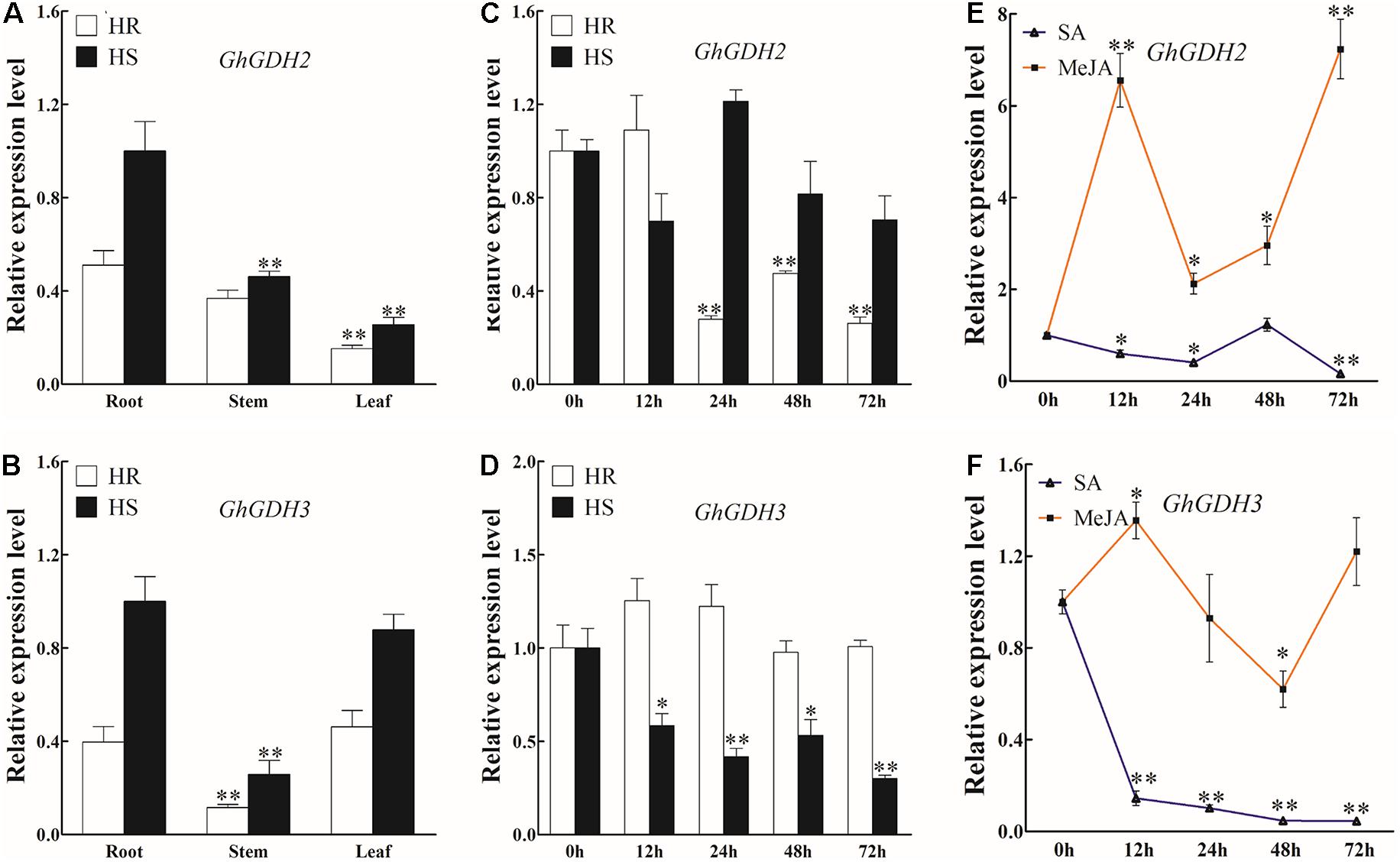
Figure 6. Expression profiles of GhGDH2 in different cotton tissues and under Verticillium dahliae, salicylic acid (SA), or methyl jasmonate (MeJA) treatment. (A,B) Expression profiling of (A) GhGDH2 and (B) GhGDH3 in roots, stems, and leaves in Shidalukang 1 (HR) and Junmian 1 (HS). Root, stem, and leaf samples from two-leaf stage cotton seedlings were collected. (C,D) Expression patterns of (C) GhGDH2 and (D) GhGDH3 in the roots of HS and HR at 0, 12, 24, 48, and 72 h post-inoculation (hpi). (E,F) Expression changes of (E) GhGDH2 and (F) GhGDH3 in the roots of HR after treatment with SA or MeJA. Samples collected at 0, 12, 24, 48, and 72 hpi after SA or MeJA treatment were used in reverse transcription quantitative PCR. GhUBQ7 was used as the control. Values represent mean ± SE of three independent biological replications (five individual plants per replicate). ∗ and ∗∗ represent significant differences from 0 hpi at P < 0.05 and P < 0.01, respectively, based on Student’s t-test.
Upon V. dahliae infection, GhGDH2 was significantly downregulated by 52.4 to 73.9% from 24 to 72 hpi in HR and no significant change in HS (Figure 6C), and GhGDH3 showed an expression pattern opposite to GhGDH2 (Figure 6D). Both GhGDH2 and GhGDH3 exhibited a downregulation trend after SA treatment in HR, particularly GhGDH3, which decreased 6.9- and 22.3-fold at 12 and 72 hpi, respectively (Figures 6E,F). GhGDH2 and GhGDH3 responded similarly to treatment of MeJA from 12 to 72 hpi, i.e., an initial upregulation followed by a significant downregulation and then another wave of upregulation, but with different intensity (Figures 6E,F). As a result, the expression level of GhGDH2 at 12 to 72 hpi was significantly higher than that at 0 hpi, with an increase of 1.12- to 6.23-fold (Figure 6E), while the expression level of GhGDH3 was significantly upregulated and downregulated at 12 and 48 hpi, respectively (Figure 6F). These results indicated that GhGDH2 and GhGDH3 may play a potential role in cotton defense against V. dahliae, which could be achieved through SA- and JA-mediated signal transduction pathways.
Silencing of GhGDH2 Enhanced Resistance of Cotton to Verticillium dahliae
Given that GhGDH2 was significantly downregulated from 24 to 72 hpi and significantly induced by MeJA in HR, we examined whether it is possible to enhance V. dahliae resistance of susceptible cotton by knockdown of the expression level of GhGDH2. To verify this conjecture, we used VIGS to knock down the expression level of GhGDH2 in HS. The expression level of GhGDH2 was measured using roots from TRV:00 and TRV:GhGDH2 plants when TRV:GhCHLI plants (as positive control of the VIGS experiment) exhibited the expected yellowing leaf phenotype about 3 weeks after infiltration (Supplementary Figure 2). Compared with TRV:00 plants, TRV:GhGDH2 plants had an approximately 82% reduction of GhGHD2 expression (Figure 7A), and displayed less etiolated and wilted leaves at 21 dpi (Figure 7B). TRV:GhGDH2 plants had a significantly lower ratio of diseased plants (45.5%) and disease index (33.8%) than the TRV:00 plants (70.8 and 62.9%, respectively) at 21 dpi (Figures 7C,D). GhGDH2-silenced plants showed slight vascular browning in the stem at 14 dpi, whereas more severe diseased phenotype was evident in TRV:00 plants (Figure 7E). The fungal recovery assay showed that the number of fungal colonies in stem segments from TRV:00 plants was 20% more than that from TRV:GhGDH2 plants (Figure 7F). TRV:GhGDH2 plants accumulated less fungal biomass (0.3 and 1.1 at 14 and 21 dpi, respectively) than TRV:00 plants (1.0 and 3.1 at 14 and 21 dpi, respectively) (Figure 7G). These results indicated that GhGDH2 functions as a negative regulator of V. dahliae resistance.
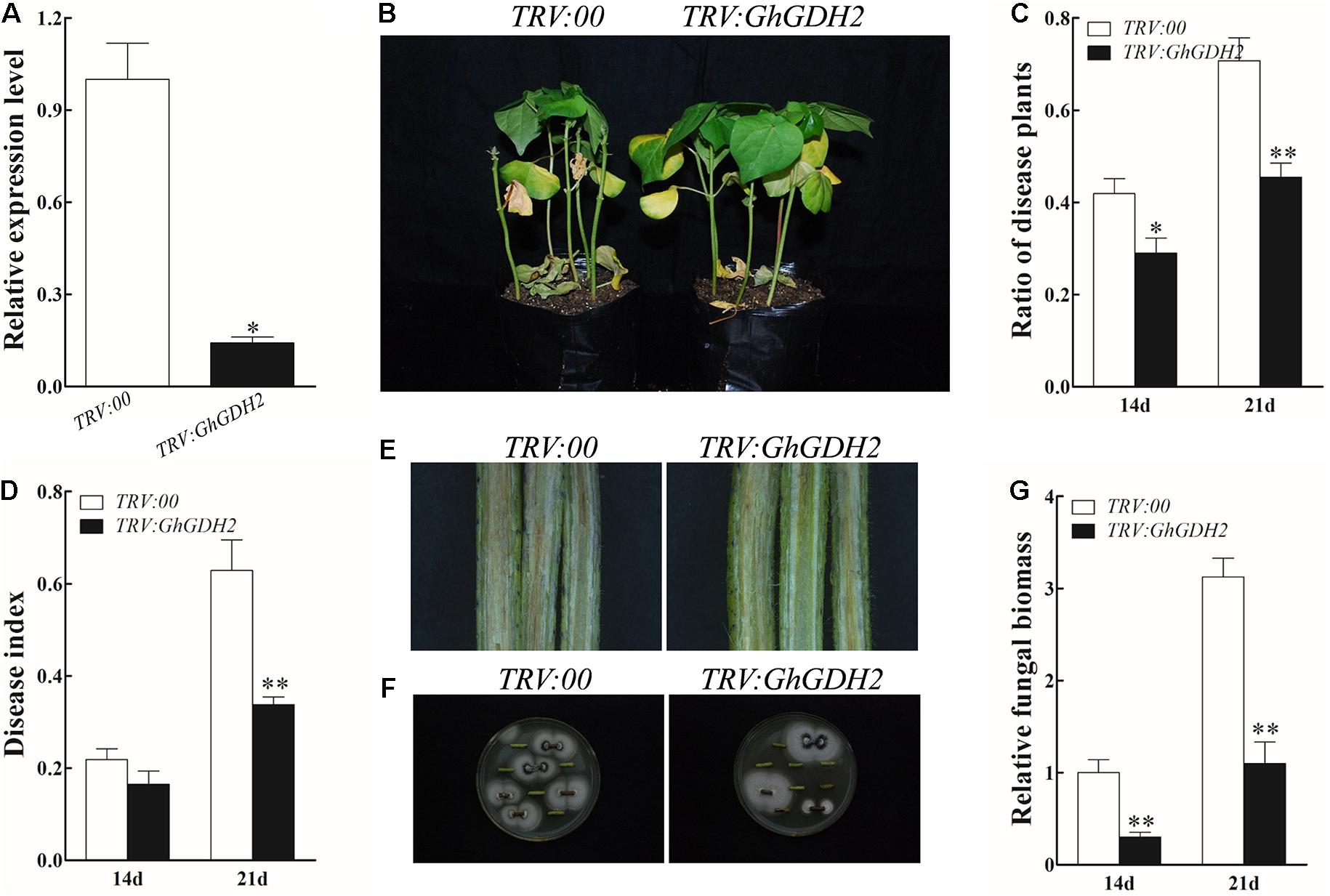
Figure 7. Knockdown of GhGDH2 in Verticillium dahliae-susceptible cultivar Junmian 1 enhanced disease resistance. (A) GhGDH2 expression in the TRV:00 and TRV:GhGDH2 plants. RNA was extracted from roots at 2 weeks after infiltration with the VIGS vector. (B) Disease phenotypes of the TRV:00 and TRV:GhGDH2 plants at 21 dpi. (C,D) Ratio of diseased plants and disease index of TRV:00 and TRV:GhGDH2 plants at 14 and 21 dpi. (E,F) Fungal recovery assay (E) and longitudinal sections of cotton stems (F) from TRV:00 and TRV:GhGDH2 plants at 14 dpi. (G) Comparison of fungal biomass. The roots from the TRV:00 and TRV: GhGDH2 plants at 14 and 21 dpi was used to extracted total DNA, which was used as a template for relative fungal biomass analysis by quantitative PCR analysis. Values represent mean ± SE of three independent biological replications (five individual plants per replicate). ∗ and ∗∗ represent significant differences from TRV:00 at P < 0.05 and P < 0.01, respectively, based on Student’s t-test.
Downregulation of GhGDH2 Positively Regulates JA Signaling Pathway, but Negatively Regulates SA Signaling Pathway
To explore whether the improvement of V. dahliae resistance in GhGDH2-silenced plants is mediated by JA and SA signaling pathways, we measured JA and SA concentrations in the roots of TRV:00 and TRV:GhGDH2 plants with or without V. dahliae inoculation. No significant difference in the content of JA and SA was detected between TRV:GhGDH2 and TRV:00 plants under the mock treatment (Figures 8A,B). At 72 hpi, the content of JA and SA in TRV:GhGDH2 plants increased and decreased 42.1 and 78.9%, respectively, compared with that in TRV:00 plants (Figures 8A,B). Consistent with this result, JA biosynthesis genes (GhAOS1 and GhLOX1) and JA response genes (GhPR4 and GhPDF1.2) were significantly induced in TRV:GhGDH2 plants at 72 hpi (Figures 8C–F), while the expression levels of SA biosynthesis (GhPAL1 and GhICS1) and response genes (GhNPR1 and GhPR1) were significantly downregulated in TRV:GhGDH2 plants at 72 hpi (Figures 8G–J). These results were consistent with those observed in HR (Figures 6C,E) and support the notion that GhGDH2 plays a negative role in V. dahliae resistance by activating JA signaling pathway and repressing SA signaling pathway.
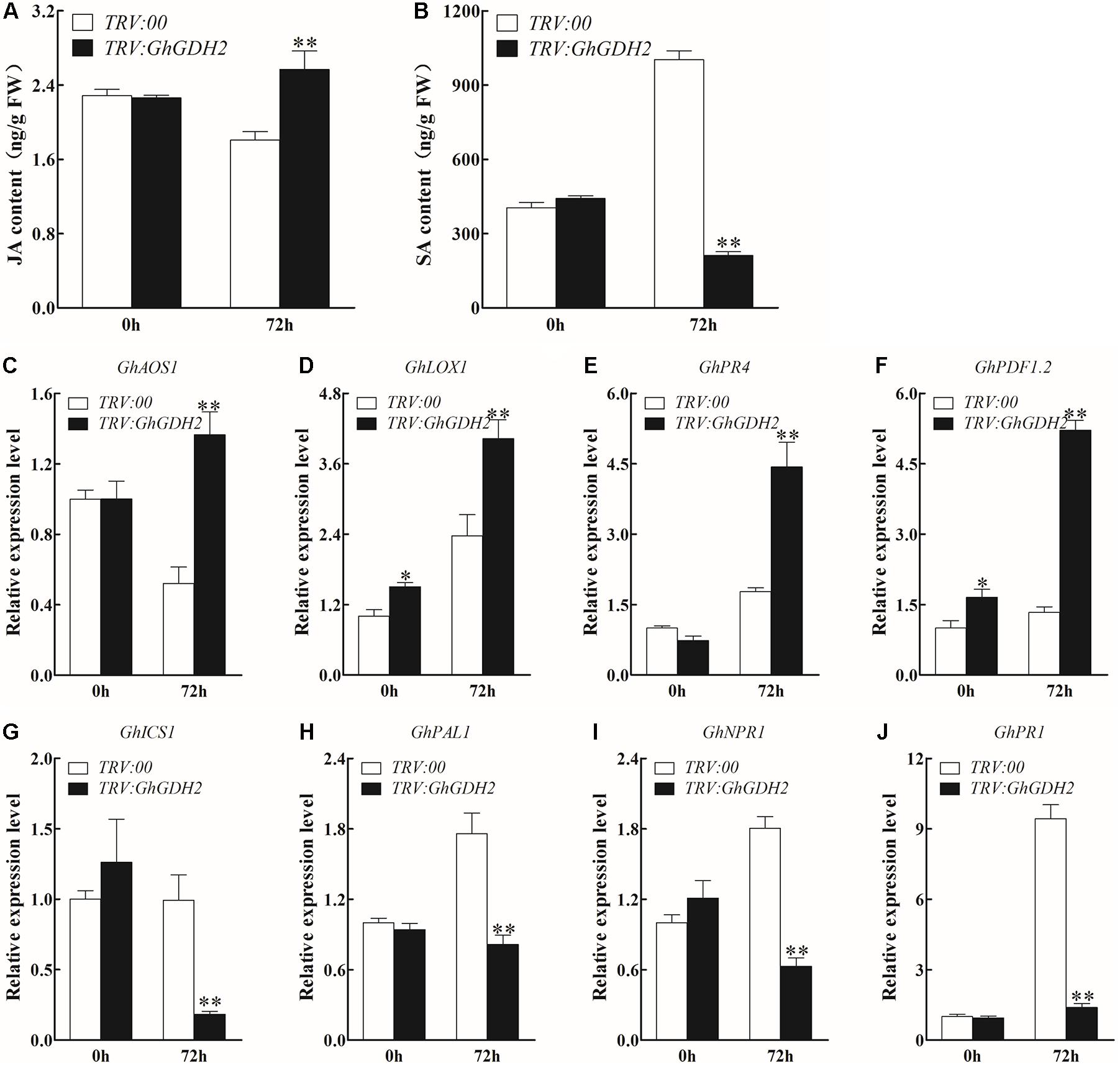
Figure 8. The change of jasmonic acid (JA) and salicylic (SA) signaling pathway in GhGDH2-silenced cotton plants. (A,B) Comparison of the (A) JA and (B) SA contents in TRV:00 and TRV:GhGDH2 plants at 72 h post-inoculation (hpi). (C–F) The relative expression levels of JA biosynthesis and response genes in TRV:00 and TRV:GhGDH2 plants at 72 hpi. (G–J) The relative expression level of SA biosynthesis and response genes in TRV: 00 and TRV:GhGDH2 plants at 72 hpi. Values represent mean ± SE of three independent biological replications (five individual plants per replicate). ∗ and ∗∗ represent significant differences from 0 h at P < 0.05 and P < 0.01, respectively, based on Student’s t-test.
Discussion
Hub Genes for Verticillium dahliae Resistance in Cotton
Mining V. dahliae-resistant genes is of great importance to breed disease-resistant cotton cultivars. We have used comparative transcriptome analysis to identify genes differentially expressed between Upland cotton cultivars resistant (HR) or susceptible (HS) to V. dahliae under uninfected or infected conditions. DEGs were subjected to investigate gene networks and hub genes associated with disease resistance. While our previous study was focused on the basal difference between HR and HS (Xiong et al., 2021), this study explored the transcriptomic responses of HR and HS upon V. dahliae infection. WGCNA is an effective method that have been previously used to identify hub genes involved in various biological processes and disease resistance in many plant species, such as soybeans (Du et al., 2017), rice (Tan et al., 2017), and cotton (Sun et al., 2019; Xiong et al., 2021). In this study, 22 hub genes related to V. dahliae resistance were identified based on WGCNA, including two genes encoding GDH (Figure 5B and Table 1).
Glutamate dehydrogenase is a key enzyme catalyzing the reversible reduction of glutamate into 2-oxoglutarate, which plays vital roles in plant amino acid metabolism (Fontaine et al., 2012; Seifi et al., 2013; Tsai et al., 2016). Studies have reported induction of GDHs by pathogen invasion in sunflower (Dulermo et al., 2009), tobacco (Pageau et al., 2006), and common bean (Tavernier et al., 2007). On the basis of identification of 22 hub genes (Figure 5B and Table 1), we analyzed the expression patterns of GhGDH2 and GhGDH3 after V. dahliae infection, or treatment of SA or MeJA (Figure 6), and found that GhGDH2 negatively regulates V. dahliae resistance in cotton, which was confirmed by the finding that knockdown of the expression level of GhGDH2 in cultivar susceptible to V. dahliae was able to enhance disease resistance (Figure 7). Therefore, GhGDH2 is a candidate gene for breeding cotton cultivars’ resistance to V. dahliae by downregulating its expression in susceptible elite cultivars through RNAi or gene editing. Gene editing would be the ideal strategy as the dysfunctional GhGDH2 created by the technology can be transferred to any other cultivars by crossing. GDH genes have been reported to be engaged in plant defense responses to pathogen infections (Pageau et al., 2006; Tavernier et al., 2007; Dulermo et al., 2009). Here, we found that GhGDH3 of cotton is also potentially involved in the response to V. dahliae infection but may have a function different from GhGDH2 as it showed downregulation in cultivar susceptible to V. dahliae infection (Figure 6D) and was repressed by MeJA at 48 hpi (Figure 6F). It is of our interest to further explore the potential role of GhGDH3 in regulating V. dahliae resistance.
In cotton, several studies have associated activating JA signaling pathways with V. dahliae resistance (Li et al., 2014; Sun et al., 2014; Xiong et al., 2019). Our results support those previous results that the expression of genes related to JA biosynthesis and response were significantly increased in TRV:GhGDH2 plants than in TRV:00 plants, as well as JA content at 72 hpi, and showed an enhanced V. dahliae resistance (Figures 8A,C–F). JA and SA often showed an antagonistic interaction in plant defense (Durrant and Dong, 2004; Thaler et al., 2012). In general, accumulation of JA content results in decreased level of SA and inhibition of SA signaling pathways after pathogen infection (Shim et al., 2013). We have previously observed this phenomenon in cotton for GhWRKY70A05 (Xiong et al., 2019); here, we demonstrated another such example in which downregulation of GhGDH2 promoted JA signaling pathways, and inhibited SA biosynthesis (Figure 8B) and signaling pathways (Figures 8G–J) upon V. dahliae infection, consequently improving V. dahliae resistance in cotton.
In addition to GDH genes, several other hub genes are of interest. Stearoyl-[acyl-carrier-protein]-desaturase (SAD) is involved in biosynthesis of fatty acids and produces monounsaturated fatty acids in plant cells (Kachroo et al., 2007). There are seven SAD genes in the Arabidopsis genome, some of which have been recognized to mediate plant defense response by regulating SA and JA signaling pathways (Kachroo et al., 2003, 2007). In our study, GhSAD6 (Gh_A05G2312), showing 73% amino acid sequence identity with AtSAD6, was one of the hub genes in the magenta module (Table 1). The expression level of GhSAD6 was significantly induced following V. dahliae infection (Figure 5B). Previously, a large number of photosynthesis-related gene have been reported to be downregulated by pathogens (de Torres Zabala et al., 2015). Our KEGG pathway analysis found that genes involved in photosynthesis were significantly enriched in HR, and their expression was significantly downregulated after V. dahliae infection in HR (Figure 2B and Supplementary Table 7). In the black module, several genes related to photosynthesis were identified as hub genes, including 10 chlorophyll a–b binding proteins, two photosystems I reaction center subunit and one photosystem I subunit O, and two photosystems I reaction center subunit (Table 1). These results indicate that the V. dahliae resistance observed in HR may be partly due to its better ability (compared with HS) to slow down growth and to divert more energy to defense, as downregulation of the metabolic pathways related to plant growth and development contributes to plant defense against pathogens (Attaran et al., 2014; Huot et al., 2014; Dhar et al., 2020).
Cotton Resistance to Verticillium dahliae Is Co-regulated by Various Plant Hormones
Jasmonic acid, SA, and ET are three important signaling molecules crucial for modulating plant defense response to pathogens (Koornneef and Pieterse, 2008), including V. dahliae (Dhar et al., 2020). Recent studies showed that JA signaling pathway positively regulates V. dahliae resistance in cotton (Gao et al., 2013a; Xiong et al., 2019). Two JAZ genes (GbJAZ1 and GbJAZ10) which act as repressors of JA signaling pathways are downregulated in G. barbadense cv Hai7124 after V. dahliae infection (Xu et al., 2011b). Different from this result, upregulated and downregulated expressions of three GhJAZ1 and three GhJAZ10 were found in HR and HS after V. dahliae infection, respectively (Figure 3A and Supplementary Table 9). One possible reason is that JA signaling pathway may have different regulatory functions in G. hirsutum and G. barbadense regarding their role in resistance to V. dahliae (Duan et al., 2016). RT-qPCR analyses revealed that the relative expression of a SA response gene GhPR1 was significantly induced in both resistant (HR) and susceptible (HS) varieties upon V. dahliae infection (Figures 3A,E,I), indicating that SA signaling pathway was activated by V. dahliae infection. A similar result has been reported in a previous study (Zhang et al., 2013). The upregulation of three GhJAZ1 and three GhJAZ10 in HR might be a result of the antagonistic interaction between the SA and JA signaling pathways (Durrant and Dong, 2004). Meanwhile, GhJAZ2 was also upregulated in G. hirsutum and negatively regulated cotton resistance to V. dahliae (He et al., 2018). These studies imply that GhJAZ1 and GhJAZ10 might be pivotal genes mediating cotton defense response against V. dahliae infection.
Ethylene is the only plant gas hormone that has been reported to be involved in various biological processes in plants. It often interacts synergistically with JA to advance resistance against necrotrophic pathogens (Zhu et al., 2011). ERF genes are important downstream response factors of the ET signaling pathway and participated in plant responses to pathogens (Amorim et al., 2016). Several reports have demonstrated upregulation of the ERF expression in cotton after V. dahliae infection (Wang et al., 2011; Xu et al., 2011a; Guo et al., 2016). Consistent with those results, upregulation and downregulation of three ERF genes were found in HR and HS after V. dahliae infection, respectively (Figure 3A and Supplementary Table 9). Moreover, knockdown of GhERF1-like and GhERF6 significantly decreases V. dahliae resistance in cotton by inhibiting the ET signaling pathway (Yang et al., 2015; Guo et al., 2016). Thus, the activation of ERF in the ET signaling pathway is beneficial to improve cotton resistance to V. dahliae.
Besides JA, SA, and ET, BR also is an important plant hormone, which has been reported to participate in the process of cotton defense against V. dahliae infection (Gao et al., 2013a; Shaban et al., 2018). BAK1, a leucine-rich-repeat receptor-like protein kinase, has been recognized to modulate BR signaling pathway (Li et al., 2002). In Arabidopsis, bak1-4 mutant plants are more susceptible to V. dahliae (Fradin et al., 2011). Silencing the expression of BAK1 through VIGS significantly decreases cotton resistance to V. dahliae (Gao et al., 2013b). Our results showed that several genes involved in the BR signaling pathway, for example, one BAK1 gene, were increased in HR but decreased in HS after V. dahliae infection (Figures 3A,D,H and Supplementary Table 9), suggesting that BR might have positive regulatory effects upon V. dahliae infection.
Lignin Biosynthesis in Cotton Is Activated by Verticillium dahliae Infection
Kyoto Encyclopedia of Genes and Genomes pathway analysis indicated that phenylpropanoid biosynthesis is potentially implicated in defense against V. dahliae infection (Figure 2B). Lignin is biosynthesized by a main branch of the phenylpropanoid biosynthesis pathway and has a significant role in plant defense (Boerjan et al., 2003). Phenylalanine-ammonia lyase (PAL) and peroxidase are two important enzymes in the process of lignin biosynthesis, and their enzymatic activities were significantly increased in the Verticillium wilt-resistant tomato variety after V. dahliae infection (Yang et al., 2018; Hu et al., 2019). Studies have also reported the induction of lignin biosynthesis-related genes by pathogen invasion in Arabidopsis (Wang et al., 2018) and eggplant (Zhou et al., 2016). In the V. dahliae-resistant G. barbadense cv Hai7124, V. dahliae infection results in the upregulation of a large number of lignin biosynthesis-related genes including PAL, 4CL, F5H, CCoAOMT, and CAD (Xu et al., 2011b). In another V. dahliae-resistant G. barbadense cv Xinhai 15, the expression level of COMT is induced by V. dahliae (Zhang et al., 2013). We also found that 13 upregulated DEGs in HR were involved in lignin biosynthesis, such as C4H, 4CL, COMT, F5H, and CAD (Figure 4A and Supplementary Table 9). This finding is consistent with those of Zhang et al. (2013), indicating a role of genes related to lignin biosynthesis in V. dahliae resistance. Furthermore, numerous disease resistance genes positively regulate cotton resistance to V. dahliae by promoting lignin biosynthesis (Shaban et al., 2018). However, of the key genes involved in lignin biosynthesis, only Gh4CL30 and GhHCT1 have been functionally characterized in the context of V. dahliae infection (Guo et al., 2016; Xiong et al., 2021), the functions of other lignin biosynthesis-related genes in cotton resistance to V. dahliae are yet to be studied.
Conclusion
This study conducted a comparative transcriptomic analysis using V. dahliae-resistant (HR) and V. dahliae-susceptible (HS) cotton cultivar uninfected or infected with V. dahliae, and identified DEGs after V. dahliae infection in HR and HS. Our results support the notion of the importance of lignin and plant hormones in cotton defense against V. dahliae infection. Importantly, we demonstrated that GhGDH2 negatively modulate cotton resistance to V. dahliae and that it was possible to increase V. dahliae resistance of susceptible cotton cultivar by downregulating the expression level of GhGDH2. Our results also showed WGCNA being a particularly useful approach in identifying gene modules and candidate hub genes contributing to V. dahliae resistance.
Data Availability Statement
The datasets presented in this study can be found in online repositories. The names of the repository/repositories and accession number(s) can be found in the article/Supplementary Material.
Author Contributions
JS and FX conceived and designed the experiments. X-PX. performed the experiments and wrote the manuscript. S-CS performed the experiments and analysis of RNA-seq data. Q-HZ revised the manuscript. FL, X-YZ, and Y-JL performed the experiments. All authors contributed to the article and approved the submitted version.
Funding
This work was supported by the National Natural Science Foundation of China (Grant No. 31960438), the National Key Research and Development Program of China (Grant No. 2016YFD0100200), and the Specific Project for Crops Breeding of Shihezi University (Grant No. YZZX202001).
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fpls.2021.654676/full#supplementary-material
Supplementary Figure 1 | Venn diagram showing shared or unique differentially expressed genes in Shidalukang 1 (HR) and Junmian 1 (HS) after Verticillium dahliae infection.
Supplementary Figure 2 | Phenotypes of GhCHLI interference effect in Verticillium dahliae-susceptible cultivar Junmian 1 at 3 weeks after infiltration with Agrobacterium.
supplementary Table 1 | All primers used in this study.
supplementary Table 2 | Differentially expressed genes in Shidalukang 1 at 12, 24, and 48 h after Verticillium dahliae infection.
Supplementary Table 3 | Differentially expressed genes in Junmian 1 at 12, 24, and 48 h after Verticillium dahliae infection.
Supplementary Table 4 | Number of all differentially expressed genes induced by Verticillium dahliae in Shidalukang 1 and Junmian 1 at different time points.
Supplementary Table 5 | Gene ontology analysis of induced and repressed genes in Shidalukang 1 after infection with Verticillium dahliae.
Supplementary Table 6 | Gene ontology analysis of induced and repressed genes in Junmian 1 after infection with Verticillium dahliae.
Supplementary Table 7 | Kyoto Encyclopedia of Genes and Genomes pathways analysis of induced and repressed genes in Shidalukang 1 after infection with Verticillium dahliae.
Supplementary Table 8 | Kyoto Encyclopedia of Genes and Genomes pathways analysis of induced and repressed genes in Junmian 1 after Verticillium dahliae infection.
Supplementary Table 9 | The expression patterns of genes involved in the lignin biosynthesis and plant hormone signal transduction in Shidalukang 1 and Junmian 1 after Verticillium dahliae infection.
Supplementary Table 10 | List of all genes for each module.
Footnotes
References
Amorim, L. L. B., Romulo, D. F. D. S., Joao, P. B. N., Guida-Santos, M., Crovella, S., and Maria Benko-Iseppon, A. (2016). Transcription factors involved in plant resistance to pathogens. Curr. Protein Pept. Sci. 18, 335–351. doi: 10.2174/1389203717666160619185308
Attaran, E., Major, I. T., Cruz, J. A., Rosa, B. A., Koo, A. J., Chen, J., et al. (2014). Temporal dynamics of growth and photosynthesis suppression in response to jasmonate signaling. Plant Physiol. 165, 1302–1314. doi: 10.1104/pp.114.239004
Boerjan, W., Ralph, J., and Baucher, M. (2003). Lignin biosynthesis. Annu. Rev. Plant Biol. 54, 519–546. doi: 10.1146/annurev.arplant.54.031902.134938
Bolger, A. M., Lohse, M., and Usadel, B. (2014). Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120. doi: 10.1093/bioinformatics/btu170
Cai, F., He, H., Mo, C., Sun, Q., Yang, P., and Liu, G. (2009). Molecular research and genetic engineering of resistance to Verticillium wilt in cotton: a review. Afr. J. Biotechnol. 8, 7363–7372. doi: 10.5897/AJB2009.000-9571
de Torres Zabala, M., Littlejohn, G., Jayaraman, S., Studholme, D., Bailey, T., Lawson, T., et al. (2015). Chloroplasts play a central role in plant defence and are targeted by pathogen effectors. Nat. Plants 1:15074. doi: 10.1038/nplants.2015.74
Dhar, N., Chen, J. Y., Subbarao, K. V., and Klosterman, S. J. (2020). Hormone signaling and its interplay with development and defense responses in Verticillium-plant interactions. Front. Plant Sci. 11:584997. doi: 10.3389/fpls.2020.584997
Dong, Q., Magwanga, R. O., Cai, X., Lu, P., Nyangasi, Kirungu, J., et al. (2019). RNA-sequencing, physiological and RNAi analyses provide insights into the response mechanism of the ABC-mediated resistance to Verticillium dahliae infection in cotton. Genes 10:110. doi: 10.3390/genes10020110
Du, J., Wang, S., He, C., Zhou, B., Ruan, Y. L., and Shou, H. (2017). Identification of regulatory networks and hub genes controlling soybean seed set and size using RNA sequencing analysis. J. Exp. Bot. 68, 1955–1972. doi: 10.1093/jxb/erw460
Duan, X., Zhang, Z., Wang, J., and Zuo, K. (2016). Characterization of a novel cotton subtilase gene GbSBT1 in response to extracellular stimulations and its role in Verticillium resistance. PLoS One 11:e0153988. doi: 10.1371/journal.pone.0153988
Dulermo, T., Bligny, R., Gout, E., and Cotton, P. (2009). Amino acid changes during sunflower infection by the necrotrophic fungus B. cinerea. Plant Signal. Behav. 4, 859–861. doi: 10.4161/psb.4.9.9397
Durrant, W. E., and Dong, X. (2004). Systemic acquired resistance. Annu. Rev. Phytopathol. 42, 185–209. doi: 10.1146/annurev.phyto.42.040803.140421
Ellendorff, U., Fradin, E. F., De Jonge, R., and Thomma, B. (2009). RNA silencing is required for Arabidopsis defence against Verticillium wilt disease. J. Exp. Bot. 60, 591–602. doi: 10.1093/jxb/ern306
Fontaine, J. X., Tercé-Laforgue, T., Armengaud, P., Clément, G., Renou, J. P., Pelletier, S., et al. (2012). Characterization of a NADH-dependent glutamate dehydrogenase mutant of Arabidopsis demonstrates the key role of this enzyme in root carbon and nitrogen metabolism. Plant Cell 24, 4044–4065. doi: 10.1105/tpc.112.103689
Fradin, E. F., Abd-El-Haliem, A., Masini, L., van den Berg, G. C., Joosten, M. H., and Thomma, B. P. (2011). Interfamily transfer of tomato Ve1 mediates Verticillium resistance in Arabidopsis. Plant Physiol. 156, 2255–2265. doi: 10.1104/pp.111.180067
Gao, W., Long, L., Zhu, L. F., Xu, L., Gao, W. H., Sun, L. Q., et al. (2013a). Proteomic and virus-induced gene silencing (VIGS) Analyses reveal that gossypol, brassinosteroids, and jasmonic acid contribute to the resistance of cotton to Verticillium dahliae. Mol. Cell. Proteomics 12, 3690–3703. doi: 10.1074/mcp.M113.031013
Gao, X., Li, F., Li, M., Kianinejad, A. S., Dever, J. K., Wheeler, T. A., et al. (2013b). Cotton GhBAK1 mediates Verticillium wilt resistance and cell death. J. Integr. Plant Biol. 55, 586–596. doi: 10.1111/jipb.12064
Guo, W., Jin, L., Miao, Y., He, X., Hu, Q., Guo, K., et al. (2016). An ethylene response-related factor, GbERF1-like, from Gossypium barbadense improves resistance to Verticillium dahliae via activating lignin synthesis. Plant Mol. Biol. 91, 305–318. doi: 10.1007/s11103-016-0467-466
He, X., Zhu, L., Wassan, G. M., Wang, Y., Miao, Y., Shaban, M., et al. (2018). GhJAZ2 attenuates cotton resistance to biotic stresses via the inhibition of the transcriptional activity of GhbHLH171. Mol. Plant. Pathol. 19, 896–908. doi: 10.1111/mpp.12575
Hu, Q., Min, L., Yang, X., Jin, S., Zhang, L., Li, Y., et al. (2018). Laccase GhLac1 modulates broad-spectrum biotic stress tolerance via manipulating phenylpropanoid pathway and jasmonic acid synthesis. Plant Physiol. 176, 1808–1823. doi: 10.1104/pp.17.01628
Hu, X., Puri, K. D., Gurung, S., Klosterman, S. J., Wallis, C. M., Britton, M., et al. (2019). Proteome and metabolome analyses reveal differential responses in tomato -Verticillium dahliae-interactions. J. Proteomics 207:103449. doi: 10.1016/j.jprot.2019.103449
Huot, B., Yao, J., Montgomery, B. L., and He, S. Y. (2014). Growth-defense tradeoffs in plants: a balancing act to optimize fitness. Mol. Plant. 7, 1267–1287. doi: 10.1093/mp/ssu049
Kachroo, A., Lapchyk, L., Fukushige, H., Hildebrand, D., Klessig, D., and Kachroo, P. (2003). Plastidial fatty acid signaling modulates salicylic acid- and jasmonic acid-mediated defense pathways in the Arabidopsis ssi2 mutant. Plant Cell 15, 2952–2965. doi: 10.1105/tpc.017301
Kachroo, A., Shanklin, J., Whittle, E., Lapchyk, L., Hildebrand, D., and Kachroo, P. (2007). The Arabidopsis stearoyl-acyl carrier protein-desaturase family and the contribution of leaf isoforms to oleic acid synthesis. Plant Mol. Biol. 63, 257–271. doi: 10.1007/s11103-006-9086-y
Koornneef, A., and Pieterse, C. M. (2008). Cross talk in defense signaling. Plant Physiol. 146, 839–844. doi: 10.1104/pp.107.112029
Langfelder, P., and Horvath, S. (2008). WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics 9:559. doi: 10.1186/1471-2105-9-559
Li, C., He, Q., Zhang, F., Yu, J., Li, C., Zhao, T., et al. (2019). Melatonin enhances cotton immunity to Verticillium wilt via manipulating lignin and gossypol biosynthesis. Plant J. 100, 784–800. doi: 10.1111/tpj.14477
Li, C., He, X., Luo, X., Xu, L., Liu, L., Min, L., et al. (2014). Cotton WRKY1 mediates the plant defense-to-development transition during infection of cotton by Verticillium dahliae by activating JASMONATE ZIM-DOMAIN1 expression. Plant Physiol. 166, 2179–2194. doi: 10.1104/pp.114.246694
Li, J., Wen, J., Lease, K. A., Doke, J. T., Tax, F. E., and Walker, J. C. (2002). BAK1, an Arabidopsis LRR receptor-like protein kinase, interacts with BRI1 and modulates brassinosteroid signaling. Cell 110, 213–222. doi: 10.1016/s0092-8674(02)00812-817
Li, P. T., Rashid, M. H. O., Chen, T. T., Lu, Q. W., Ge, Q., Gong, W. K., et al. (2019). Transcriptomic and biochemical analysis of upland cotton (Gossypium hirsutum) and a chromosome segment substitution line from G. hirsutum x G. barbadense in response to Verticillium dahliae infection. BMC plant Biol. 19:19. doi: 10.1186/s12870-018-1619-1614
Li, T., Ma, X., Li, N., Zhou, L., Liu, Z., Han, H., et al. (2017). Genome-wide association study discovered candidate genes of Verticillium wilt resistance in upland cotton (Gossypium hirsutum L.). Plant Biotechnol. J. 15, 1520–1532. doi: 10.1111/pbi.12734
Livak, K. J., and Schmittgen, T. D. (2001). Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25, 402–408. doi: 10.1006/meth.2001.1262
Love, M. I., Huber, W., and Anders, S. (2014). Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15:550. doi: 10.1186/s13059-014-0550-558
Pageau, K., Reisdorf-Cren, M., Morot-Gaudry, J. F., and Masclaux-Daubresse, C. (2006). The two senescence-related markers, GS1 (cytosolic glutamine synthetase) and GDH (glutamate dehydrogenase), involved in nitrogen mobilization, are differentially regulated during pathogen attack and by stress hormones and reactive oxygen species in Nicotiana tabacum L. leaves. J. Exp. Bot. 57, 547–557. doi: 10.1093/jxb/erj035
Seifi, H. S., Van Bockhaven, J., Angenon, G., and Höfte, M. (2013). Glutamate metabolism in plant disease and defense: friend or foe? Mol. Plant Microbe Interact. 26, 475–485. doi: 10.1094/MPMI-07-12-0176-CR
Shaban, M., Miao, Y., Ullah, A., Khan, A. Q., Menghwar, H., Khan, A. H., et al. (2018). Physiological and molecular mechanism of defense in cotton against Verticillium dahliae. Plant Physiol. Biochem. 125, 193–204. doi: 10.1016/j.plaphy.2018.02.011
Shim, J. S., Jung, C., Lee, S., Min, K., Lee, Y. W., Choi, Y., et al. (2013). AtMYB44 regulates WRKY70 expression and modulates antagonistic interaction between salicylic acid and jasmonic acid signaling. Plant J. 73, 483–495. doi: 10.1111/tpj.12051
Sun, L., Zhu, L., Xu, L., Yuan, D., Min, L., and Zhang, X. (2014). Cotton cytochrome P450 CYP82D regulates systemic cell death by modulating the octadecanoid pathway. Nat. Commun. 5:5372. doi: 10.1038/ncomms6372
Sun, Q., Jiang, H., Zhu, X., Wang, W., He, X., Shi, Y., et al. (2013). Analysis of sea-island cotton and upland cotton in response to Verticillium dahliae infection by RNA sequencing. BMC Genomics 14:852. doi: 1186/1471-2164-14-852
Sun, S., Xiong, X., Zhu, Q., Li, Y., and Sun, J. (2019). Transcriptome sequencing and metabolome analysis reveal genes involved in pigmentation of green-colored cotton fibers. Int. J. Mol. Sci. 20:4838. doi: 10.3390/ijms20194838
Tan, M., Cheng, D., Yang, Y., Zhang, G., Qin, M., Chen, J., et al. (2017). Co-expression network analysis of the transcriptomes of rice roots exposed to various cadmium stresses reveals universal cadmium-responsive genes. BMC Plant Biol. 17:194. doi: 10.1186/s12870-017-1143-y
Tavernier, V., Cadiou, S., Pageau, K., Laugé, R., Reisdorf-Cren, M., Langin, T., et al. (2007). The plant nitrogen mobilization promoted by Colletotrichum lindemuthianum in Phaseolus leaves depends on fungus pathogenicity. J. Exp. Bot. 58, 3351–3360. doi: 10.1093/jxb/erm182
Thaler, J. S., Humphrey, P. T., and Whiteman, N. K. (2012). Evolution of jasmonate and salicylate signal crosstalk. Trends Plant. Sci. 17, 260–270. doi: 10.1016/j.tplants.2012.02.010
Trapnell, C., Williams, B. A., Pertea, G., Mortazavi, A., Kwan, G., van Baren, M. J., et al. (2010). Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515. doi: 10.1038/nbt.1621
Tsai, K. J., Lin, C. Y., Ting, C. Y., and Shih, M. C. (2016). Ethylene-regulated glutamate dehydrogenase fine-tunes metabolism during anoxia-reoxygenation. Plant Physiol. 172, 1548–1562. doi: 10.1104/pp.16.00985
Vanholme, R., Demedts, B., Morreel, K., Ralph, J., and Boerjan, W. (2010). Lignin biosynthesis and structure. Plant Physiol. 153, 895–905. doi: 10.1104/pp.110.155119
Wang, F. X., Luo, Y. M., Ye, Z. Q., Gao, X., Liang, J. N., Wang, Q., et al. (2018). iTRAQ-based proteomics analysis of autophagy-mediated immune responses against the vascular fungal pathogen Verticillium dahliae in Arabidopsis. Autophagy 14, 598–618. doi: 10.1080/15548627.2017.1423438
Wang, F. X., Ma, Y. P., Yang, C. L., Zhao, P. M., Yao, Y., Jian, G. L., et al. (2011). Proteomic analysis of the sea-island cotton roots infected by wilt pathogen Verticillium dahliae. Proteomics 11, 4296–4309. doi: 10.1002/pmic.201100062
Wang, Y., Liang, C., Wu, S., Zhang, X., Tang, J., Jian, G., et al. (2016). Significant improvement of cotton Verticillium wilt resistance by manipulating the expression of Gastrodia antifungal proteins. Mol. Plant. 9, 1436–1439. doi: 10.1016/j.molp.2016.06.013
Wang, Z., Gerstein, M., and Snyder, M. (2009). RNA-Seq: a revolutionary tool for transcriptomics. Nat. Rev. Genet. 10, 57–63. doi: 10.1038/nrg2484
Xie, C., Mao, X., Huang, J., Ding, Y., Wu, J., Dong, S., et al. (2011). KOBAS 2.0: a web server for annotation and identification of enriched pathways and diseases. Nucleic Acids Res. 39, W316–W322. doi: 10.1093/nar/gkr483
Xiong, X., Sun, S., Li, Y., Zhang, X., Sun, J., and Xue, F. (2019). The cotton WRKY transcription factor GhWRKY70 negatively regulates the defense response against Verticillium dahliae. Crop J. 7, 393–402. doi: 10.1016/j.cj.2018.10.005
Xiong, X., Sun, S., Zhu, Q., Zhang, X., Li, Y., Liu, F., et al. (2021). The cotton lignin biosynthetic gene Gh4CL30 regulates lignification, phenolic content and contributes to Verticillium wilt resistance. Mol. Plant Microbe Interact. 34, 240–254. doi: 10.1094/MPMI-03-20-0071-R
Xu, L., Zhu, L., Tu, L., Guo, X., Long, L., Sun, L., et al. (2011a). Differential gene expression in cotton defence response to Verticillium dahliae by SSH. J. Phytopathol. 159, 606–615. doi: 10.1111/j.1439-0434.2011.01813.x
Xu, L., Zhu, L., Tu, L., Liu, L., Yuan, D., and Jin, L. (2011b). Lignin metabolism has a central role in the resistance of cotton to the wilt fungus Verticillium dahliae as revealed by RNA-Seq-dependent transcriptional analysis and histochemistry. J. Exp. Bot. 62, 5607–5621. doi: 10.1093/jxb/err245
Yang, C., Liang, S., Wang, H., Han, L., Wang, F., Cheng, H., et al. (2015). Cotton major latex protein 28 functions as a positive regulator of the ethylene responsive factor 6 in defense against Verticillium dahliae. Mol. Plant. 8, 399–411. doi: 10.1016/j.molp.2014.11.023
Yang, C., Liang, Y., Qiu, D., Zeng, H., Yuan, J., and Yang, X. (2018). Lignin metabolism involves Botrytis cinerea BcGs1- induced defense response in tomato. BMC Plant Biol. 18:103. doi: 10.1186/s12870-018-1319-1310
Young, M., Wakefield, M., Smyth, G., and Oshlack, A. (2010). Gene ontology analysis for RNA-seq: accounting for selection bias. Genome Biol. 11:R14. doi: 10.1186/gb-2010-11-2-r14
Zhang, T., Hu, Y., Jiang, W., Fang, L., Guan, X., Chen, J., et al. (2015). Sequencing of allotetraploid cotton (Gossypium hirsutum L. acc. TM-1) provides a resource for fiber improvement. Nat. Biotechnol. 33, 531–537. doi: 10.1038/nbt.3207
Zhang, W., Zhang, H., Liu, K., Jian, G., Qi, F., and Si, N. (2017). Large-scale identification of Gossypium hirsutum genes associated with Verticillium dahliae by comparative transcriptomic and reverse genetics analysis. PLoS One 12:e0181609. doi: 10.1371/journal.pone.0181609
Zhang, W., Zhang, H., Qi, F., and Jian, G. (2016). Generation of transcriptome profiling and gene functional analysis in Gossypium hirsutum upon Verticillium dahliae infection. Biochem. Biophys. Res. Commun. 473, 879–885. doi: 10.1016/j.bbrc.2016.03.143
Zhang, Y., Jin, Y., Gong, Q., Li, Z., Zhao, L., Han, X., et al. (2019). Mechanismal analysis of resistance to Verticillium dahliae in upland cotton conferred by overexpression of RPL18A-6 (Ribosomal Protein L18A-6). Ind. Crop and Prod. 141:111742. doi: 10.1016/j.indcrop.2019.111742
Zhang, Y., Wang, X., Ding, Z., Ma, Q., Zhang, G., Zhang, S., et al. (2013). Transcriptome profiling of Gossypium barbadense inoculated with Verticillium dahliae provides a resource for cotton improvement. BMC Genomics 14:637. doi: 10.1186/1471-2164-14-637
Zhou, X., Bao, S., Liu, J., et al. (2016). De novo sequencing and analysis of the transcriptome of the wild eggplant species Solanum aculeatissimum in response to Verticillium dahliae. Plant Mol. Biol. Rep. 34, 1193–1203. doi: 10.1007/s11105-016-0998-997
Keywords: Gossypium hirsutum, Verticillium wilt, transcriptome, weighted gene co-expression network analysis, GhGDH2, jasmonic acid, salicylic acid
Citation: Xiong X-P, Sun S-C, Zhu Q-H, Zhang X-Y, Liu F, Li Y-J, Xue F and Sun J (2021) Transcriptome Analysis and RNA Interference Reveal GhGDH2 Regulating Cotton Resistance to Verticillium Wilt by JA and SA Signaling Pathways. Front. Plant Sci. 12:654676. doi: 10.3389/fpls.2021.654676
Received: 17 January 2021; Accepted: 18 March 2021;
Published: 11 June 2021.
Edited by:
Tina Kyndt, Ghent University, BelgiumReviewed by:
Steven J. Klosterman, United States Department of Agriculture, United StatesNataša Štajner, University of Ljubljana, Slovenia
Copyright © 2021 Xiong, Sun, Zhu, Zhang, Liu, Li, Xue and Sun. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Jie Sun, c3VuamllQHNoenUuZWR1LmNu; Fei Xue, eHVlZmVpQHNoenUuZWR1LmNu
†These authors have contributed equally to this work
 Xian-Peng Xiong1,2†
Xian-Peng Xiong1,2† Qian-Hao Zhu
Qian-Hao Zhu Jie Sun
Jie Sun