
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
MINI REVIEW article
Front. Pharmacol. , 27 February 2025
Sec. Pharmacogenetics and Pharmacogenomics
Volume 16 - 2025 | https://doi.org/10.3389/fphar.2025.1532971
 Adi Muradi Muhar1
Adi Muradi Muhar1 Adrian Joshua Velaro1
Adrian Joshua Velaro1 Arya Tjipta Prananda1
Arya Tjipta Prananda1 Sony Eka Nugraha2
Sony Eka Nugraha2 Princella Halim3
Princella Halim3 Rony Abdi Syahputra3*
Rony Abdi Syahputra3*Precision medicine has revolutionized the treatment of colorectal cancer by enabling a personalized approach tailored to each patient’s unique genetic characteristics. Genomic profiling allows for the identification of specific mutations in genes such as KRAS, BRAF, and PIK3CA, which play a crucial role in cell signaling pathways that regulate cell proliferation, apoptosis, and differentiation. This information enables doctors to select targeted therapies that inhibit specific molecular pathways, maximizing treatment effectiveness and minimizing side effects. Precision medicine also facilitates adaptive monitoring of tumor progression, allowing for adjustments in therapy to maintain treatment effectiveness. While challenges such as high costs, limited access to genomic technology, and the need for more representative genomic data for diverse populations remain, collaboration between researchers, medical practitioners, policymakers, and the pharmaceutical industry is crucial to ensure that precision medicine becomes a standard of care accessible to all. With continued advances and support, precision medicine has the potential to improve treatment outcomes, reduce morbidity and mortality rates, and enhance the quality of life for colorectal cancer patients worldwide.
The precision medicine approach has become a new milestone in the development of oncology, especially in the effort to conquer colorectal cancer. The development of genomic and bioinformatics technologies has opened up great opportunities to analyze the genetic characteristics of individuals in detail, allowing the identification of specific mutations present in cancer cells (Cortés-Ciriano et al., 2022). In colorectal cancer, common mutations that are often found include the genes KRAS, BRAF, PIK3CA, and TP53 (Jauhri et al., 2017; Sclafani et al., 2020; Afrăsânie et al., 2023; Marbun et al., 2022). These mutations play an important role in cell signaling pathways that regulate cell proliferation, apoptosis, and differentiation, which are critical points in cancer progression and aggressiveness (Feng et al., 2018). By identifying the unique genomic profile of each patient, precision medicine can be directed to target specific molecular pathways involved in the cancer process. This allows for the treatment of colorectal cancer with a more effective, efficient, and potentially reduced side effects approach compared to conventional therapies. In this approach, the genomic profile of each patient is not only a diagnostic guide but also serves as the basis for the most appropriate therapy selection (Malone et al., 2020; Naito et al., 2021). For example, patients with KRAS mutations are known not to respond well to EGFR (Epidermal Growth Factor Receptor) inhibitors, which are often used in colorectal cancer therapy (Martinelli et al., 2020; Sforza et al., 2016). With this genomic information, doctors can exclude EGFR inhibitor therapy and switch to other more appropriate therapies, such as BRAF inhibitors or MEK in cases where BRAF mutations are identified. This not only increases the likelihood of treatment success but also reduces the time it takes to determine the right therapy, so patients can benefit from faster and more personalized treatment (Prins et al., 2021).
In addition, precision medicine also allows for more adaptive monitoring of therapies. Using genomic profiling, doctors can monitor changes that occur in tumors during treatment. When cancer cells show resistance to a particular therapy, genomic information can be used to modify or change treatment strategies, thus maintaining the effectiveness of therapy as the disease progresses (Ramos and Bentires-Alj, 2015; Marine et al., 2020). This adaptive approach provides greater flexibility and opportunities to maintain control over cancer in the long term, as well as improve patients' quality of life as the treatment received is more targeted and minimizes side effects (Nikolaou et al., 2018; Salzman et al., 2018). However, the application of precision medicine in the treatment of colorectal cancer also faces various challenges. The cost of undergoing comprehensive genomic profiling is still relatively high, especially in developing countries, which can be a barrier for patients to access these genomic-based treatments (Ashenden et al., 2024). In addition, big data analysis and sophisticated bioinformatics infrastructure are needed to obtain accurate and relevant data (Tripathi et al., 2016). Access to these technologies is still limited and often concentrated in large health centers, which makes precision medicine difficult to access for a wider population, especially in rural areas.
Furthermore, genetic differences in different populations are also a challenge. For example, mutations commonly identified in Western populations do not necessarily have the same frequency in Asian or African populations (Mathieson and Reich, 2017; Gurdasani et al., 2019). For this reason, further research and data collection from various ethnic groups is urgently needed to accommodate this genetic diversity and ensure that precision medicine can be applied in an inclusive manner. To support the implementation of precision medicine, strong collaboration is needed between researchers, medical personnel, the government, and the pharmaceutical industry.
Researchers play a crucial role in developing and updating genomic knowledge as well as its analysis technologies. Meanwhile, medical personnel need to continue to update their insights in order to integrate the results of genomic analysis into daily clinical practice. The government, through supportive policies, also has a major role to play in facilitating access to this technology through subsidies, health infrastructure development, and regulations that support clinical research. On the other hand, the pharmaceutical industry needs to be involved in the development of drugs that are specific to the molecular targets found in colorectal cancers. This collaboration between stakeholders will be crucial in ensuring that precision medicine is widely accessible and able to have a significant impact in reducing morbidity and mortality due to colorectal cancer (Traversi et al., 2021; Khoury et al., 2022).
Overall, genomic profiling-based precision medicine offers a new paradigm in the treatment of colorectal cancer. By integrating genomic analysis, cancer therapies can be tailored to each patient’s unique characteristics, providing opportunities for more personalized and effective treatment. This approach not only offers hope to improve the effectiveness of treatment but also serves as the basis for the development of more advanced oncology therapies in the future. Despite still facing challenges in terms of cost, infrastructure, and accessibility, precision medicine is a promising step to reduce the incidence and mortality of colorectal cancer, as well as improve the quality of life of patients around the world.
Genomic profiling has become an important component in the treatment of colorectal cancer, allowing for a more precise approach to understanding and treating the disease (Dalal et al., 2020; Andrei et al., 2022). Through genomic technology, the identification of specific mutations in genes associated with the development of colorectal cancer, such as KRAS, BRAF, and PIK3CA, can be performed (Figure 1). Each of these genes has a significant role in cell signaling pathways that regulate the growth, proliferation, and survival of cancer cells. Mutations in these genes can have different impacts on cancer cells’ response to therapy, greatly influencing the selection of effective treatment strategies. For example, KRAS mutations are found in about 40% of colorectal cancer cases and directly affect the RAS-MAPK signaling pathway, which is responsible for controlling cell growth and division (Ahmad et al., 2021; Vitiello et al., 2019). These mutations cause the pathway to remain constitutively active, which in turn leads to uncontrolled proliferation of cancer cells. Patients with KRAS mutations generally do not respond to anti-EGFR (Epidermal Growth Factor Receptor) therapies, such as cetuximab and panitumumab, which work by inhibiting the EGFR pathway (Li et al., 2020; García-Foncillas et al., 2019; Braig et al., 2015). Therefore, information about KRAS mutations in the patient’s genomic profile allows doctors to avoid the use of anti-EGFR therapy, which is ineffective in these cases, and look for more appropriate therapeutic alternatives, such as chemotherapy combinations with other more specific target agents.
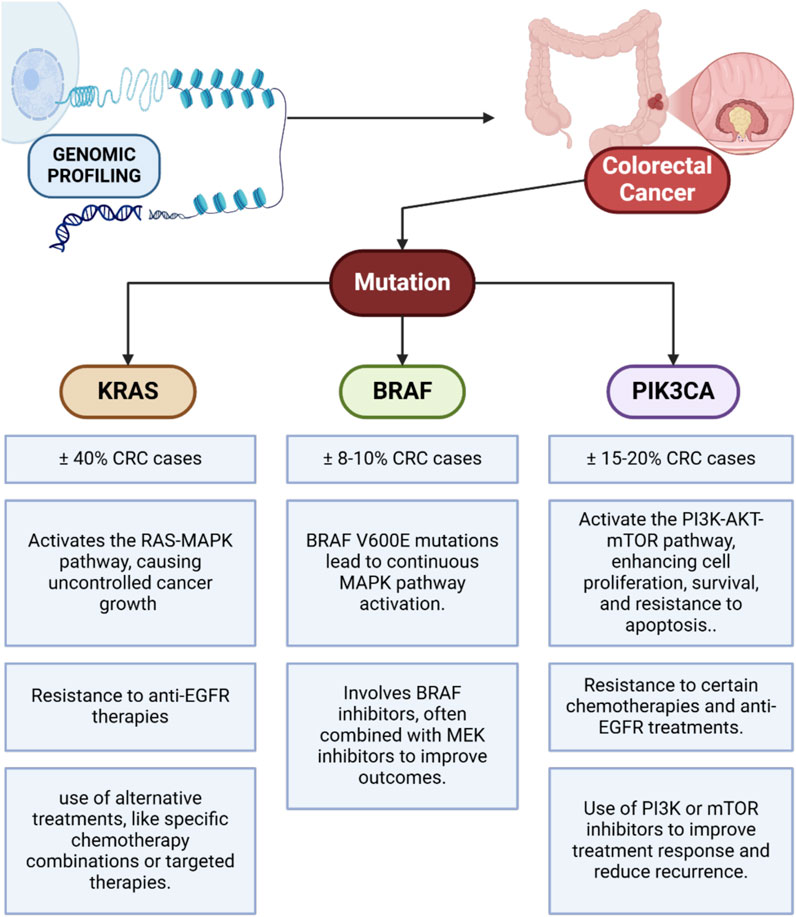
Figure 1. Identification of KRAS, BRAF, and PIK3CA mutations in colorectal cancer and their impact on cell signaling pathways and treatment strategies.
In addition to KRAS, mutations in BRAF also play an important role in colorectal cancer, although they are less common compared to KRAS mutations. BRAF V600E mutations, for example, occur in about 8%–10% of colorectal cancer patients and are usually associated with a poorer prognosis and higher levels of aggressiveness (Bond and Whitehall, 2018; Grothey et al., 2021). These mutations cause constitutive activation of the MAPK pathway, thus allowing for the growth of cancer cells that are more aggressive and resistant to some standard treatments. In patients with BRAF mutations, BRAF inhibitors, such as vemurafenib or dabrafenib, can be used to target these mutation pathways specifically (Dankner et al., 2018). The combination use of BRAF inhibitors with MEK inhibitors has also shown better results in inhibiting this pathway (Sanchez et al., 2018), providing a therapy option that is better tailored to the patient’s needs. Meanwhile, PIK3CA mutations contribute to the activation of the PI3K-AKT-mTOR pathway, which plays a role in proliferation, cell survival, and resistance to apoptosis (Hu et al., 2021; Rascio et al., 2021). PIK3CA mutations occur in about 15%–20% of colorectal cancer cases and are often associated with resistance to some types of chemotherapy and anti-EGFR therapy (Voutsadakis, 2021; Voutsadakis, 2023). By knowing the presence of these mutations, doctors may consider the use of PI3K inhibitors or mTOR as part of therapies directed specifically at these pathways, which may improve the patient’s response to treatment and reduce the risk of recurrence.
Overall, genomic profiling provides a deeper understanding of the specific molecular characteristics that drive the development of colorectal cancer. By knowing and analyzing relevant genetic mutations, doctors can tailor therapy individually to each patient, choose the most effective drugs and combination therapies, and avoid inappropriate treatments that may not provide benefits. This not only improves the effectiveness of treatment, but also reduces the potential for side effects, improves the patient’s quality of life, and opens up opportunities for adaptive monitoring of tumor progression (Fountzilas et al., 2022; Wang and Wang, 2023). Along with technological advances and research in the field of genomics, it is hoped that precision medicine can continue to develop and become the standard in the treatment of colorectal cancer, providing new hope for patients and encouraging better treatment outcomes.
Targeted treatment in colorectal cancer is a highly specific and molecular-based approach, designed to inhibit pathways or molecules that directly trigger cancer growth and development (Rastin et al., 2024; Frank et al., 2021; Yang et al., 2022). In the context of colorectal cancer treatment, this therapy relies heavily on identifying the patient’s genomic profile, allowing for more targeted and personalized treatment. Through this approach, the therapy is not only directed at cancer cells in general but also at specific mutations or pathways found in the patient’s genome, thus providing a higher potential for treatment success compared to conventional therapies (Dietel et al., 2015). One of the main targets in the treatment of colorectal cancer is the KRAS mutation, which is found in about 40% of patients. This mutation leads to the constitutive activation of the RAS-MAPK signaling pathway, an important pathway that regulates the process of cell proliferation and division (Ahmad et al., 2021; Vitiello et al., 2019). This overactivation allows cancer cells to grow and divide uncontrollably, ultimately worsening the development of cancer. Anti-EGFR therapy, such as cetuximab or panitumumab, is commonly used to target EGFR receptors on the surface of cancer cells but is not effective in patients with KRAS mutations (Janani et al., 2022; Kasi et al., 2023). Patients with KRAS mutations tend to show resistance to this therapy, as the RAS-MAPK signaling pathway remains active even when EGFR is inhibited (Adachi et al., 2021; Chen et al., 2021). Therefore, by knowing the presence of KRAS mutations through genomic profiling, doctors can immediately avoid the use of EGFR inhibitor therapy in these patients, thereby saving time and reducing the possibility of side effects of ineffective therapy.
Alternatively, patients with KRAS mutations may be given a combination of chemotherapy or therapies that target other more relevant pathways, such as MEK inhibitors that work in the RAS-MAPK pathway (Braicu et al., 2019; Dillon et al., 2021). In addition to KRAS, BRAF mutations also play a significant role in the pathogenesis of colorectal cancers, specifically the BRAF V600E mutation, which is found in about 8%–10% of cases (Bond and Whitehall, 2018; Grothey et al., 2021). These mutations cause constitutive activation of the MAPK pathway, which promotes aggressive proliferation of cancer cells. Patients with BRAF mutations generally have a poorer prognosis and higher levels of cancer aggressiveness and tend to be resistant to some standard therapies (Florent et al., 2023). However, the development of specific inhibitors, such as vemurafenib and dabrafenib, has opened up opportunities for patients with BRAF mutations. BRAF inhibitors work by over-targeting the overactive MAPK pathway, thereby inhibiting the growth of mutated cancer cells. Studies show that the combination of BRAF inhibitors with MEK inhibitors produces better results compared to BRAF inhibitors alone (Corcoran et al., 2018; Liu et al., 2017). This combination is able to suppress MAPK pathway activity at multiple points, reduce potential resistance, and provide longer clinical response times. This suggests that genomic profile-based targeted therapy, especially for patients with BRAF mutations, can provide more precise and effective therapy options. EGFR (Epidermal Growth Factor Receptor) is also an important target in the treatment of colorectal cancer, especially in patients without KRAS or BRAF (wild-type) mutations. EGFR is a receptor that triggers signaling pathways that play a role in the proliferation of cancer cells (Santos et al., 2021). EGFR inhibitors such as cetuximab or panitumumab work by inhibiting EGFR activity on the surface of cancer cells, thereby inhibiting signaling pathways that stimulate cell growth (Cai et al., 2020). In patients with KRAS and wild-type BRAF status, EGFR inhibitor therapy has shown significant results in suppressing the progression of colorectal cancer (Ros et al., 2021; Feng et al., 2023). This confirms that EGFR inhibitors can be effective in certain groups of patients, as evidenced by the presence of genomic profiling.
However, it is important to note that although targeted therapy shows significant success, resistance can develop over time due to secondary mutations or adaptation of cancer cells to the given inhibitor (Haider et al., 2020; Labrie et al., 2022). Some patients may develop additional mutations or other resistance mechanisms that make targeted therapy less effective. For example, resistance to EGFR inhibitors may arise through the activation of alternative pathways or the development of new mutations that avoid inhibitor effects (Zhou et al., 2021; Dong et al., 2021; Cooper et al., 2022). To address this, regular monitoring of a patient’s genomic profile can help doctors identify changes in tumor characteristics and adjust therapy adaptively. Thus, this approach allows for more flexible and responsive treatment to cancer dynamics, ultimately increasing the chances of long-term treatment success.
Targeted treatment based on genomic profiles is becoming increasingly important along with technological advances and research in the field of oncology (Chakravarty and Solit, 2021; Chakraborty et al., 2024). By understanding the relevant molecular pathways and specific mutations in colorectal cancer, doctors can develop a more effective and tailored treatment plan for each patient. This approach not only increases the efficacy of the therapy but also minimizes unnecessary side effects, improves the patient’s quality of life, and reduces the risk of recurrence. Genomic profiling paves the way for a new era in colorectal cancer treatment, where therapies are no longer generic but based on the unique characteristics of each patient, bringing new hope to patients and enabling better and measurable treatment outcomes. In the future, with more and more clinical and genomic data available, targeted treatment is expected to become an integral part of colorectal cancer treatment protocols, making it the standard in oncology clinical practice. Collaboration between scientists, medical personnel, and healthcare providers is essential to ensure that this genomic profiling technology is accessible to more patients around the world.
Precision medicine’s approach to colorectal cancer treatment has brought significant breakthroughs in terms of treatment effectiveness and personalization of therapy (Figure 2). By recognizing specific mutations in each patient’s genome, precision medicine provides an opportunity to target specific molecular pathways or genes that play a role in cancer cell growth, thereby maximizing the effectiveness of the therapy and minimizing side effects (Dharani and Kamaraj, 2024). In the context of colorectal cancer, this approach is very important because this cancer often involves a variety of different genetic mutations in each patient, such as mutations in the KRAS, BRAF, and PIK3CA genes (Jauhri et al., 2017; Sclafani et al., 2020). This in-depth understanding of genomic profiles allows doctors to select therapies that suit the specific characteristics of cancer cells in each patient, resulting in more precise effects and a direct impact on the success of therapy.
One of the main benefits of precision medicine is the increased effectiveness of treatment. Targeted therapies allow doctors to target mutations that trigger specific signaling pathways, such as the RAS-MAPK or PI3K-AKT pathways, that directly affect the growth and development of cancer (Yip and Papa, 2021; Karagiannakos et al., 2022). By knowing the presence of certain mutations, such as the KRAS mutation that makes patients resistant to anti-EGFR therapy, doctors can avoid the use of these ineffective therapies and switch to other, more appropriate approaches. Similarly, in patients with BRAF mutations, therapies that target those mutations, such as BRAF inhibitors and MEK, can provide better outcomes compared to conventional chemotherapy (Molina-Cerrillo et al., 2020; Djanani et al., 2020). By targeting specific molecular pathways, precision medicine enables more targeted therapies, accelerates clinical responses, and provides a higher chance of achieving positive outcomes.
In addition, precision medicine reduces the side effects that are common to conventional therapies, such as chemotherapy and radiotherapy, which attack cancer cells as well as healthy cells at the same time. Because precision medicine is designed to target specific mutations or pathways, these therapies can be delivered in lower doses and directed at cancer cells without damaging the surrounding healthy tissue (Gu et al., 2021; Bashraheel et al., 2020). This not only improves the patient’s tolerance to treatment but also reduces the risk of complications and improves the patient’s overall quality of life. Patients undergoing genomic profile-based therapy tend to experience fewer side effects, which ultimately impact treatment adherence and sustainability (Rodriguez Castells et al., 2023). The quality of life of patients is also improved because they do not have to face the severe side effects that often arise with non-specific therapies, allowing them to live a more productive and comfortable life during treatment (Faulkner et al., 2020).
Precision medicine also opens up opportunities for more adaptive therapy monitoring. As technology advances, doctors can now monitor tumor responses to therapy more dynamically and make adjustments in case of mutations or resistance changes (Edsjö et al., 2023; Ciardiello et al., 2022). For example, in cases where cancer cells develop resistance to certain inhibitors, genomic information can be used to replace or modify therapies to remain effective (Bukowski et al., 2020). This flexibility is crucial in the management of colorectal cancer, which often develops aggressively and varies from individual to individual. This adaptive monitoring also allows for early detection of new changes or mutations that may emerge over time, so that doctors can respond quickly to those changes and prevent drug resistance, which is one of the leading factors in cancer therapy failure.
However, despite these significant benefits, precision medicine in colorectal cancer also faces a number of major challenges that limit its implementation in the clinic. One of the biggest challenges is the high cost of genomic testing. The genomic profiling process requires advanced sequencing technology and detailed bioinformatics analysis, which is still relatively expensive (Arora and Tollefsbol, 2021). These technologies include state-of-the-art sequencing tools and expensive data analysis software, which makes genomic testing very expensive for most patients. In developing countries, where health budgets and financial support are limited, this price can be a major obstacle, making precision medicine accessible to only a small portion of the population (Adeniji et al., 2021; Lu et al., 2023). This condition leads to an access gap, where patients who are unable to afford it or are in a region with limited resources cannot enjoy the same benefits of precision medicine as other patients in advanced health centers. In addition to the cost problem, patient accessibility to this technology is also still very limited (Mateo et al., 2022; Ho et al., 2020). Most hospitals and health centers in various countries do not yet have the infrastructure to conduct genomic testing. These facilities are generally only available in large health centers or in developed countries, so patients in remote areas or with limited access have to travel long distances to get this test. This has an impact on delays in diagnosis and treatment for patients in areas that do not have such facilities, which ultimately affects their treatment outcomes and quality of life (Keeling et al., 2020). This accessibility issue requires serious attention, including infrastructure development and equitable allocation of resources across the region.
Population diversity is also an important challenge in precision medicine. Most of the genomic data that exists today comes from Western populations, making it less representative of Asian, African, or other ethnic groups (Atutornu et al., 2022; Cerdeña et al., 2022). Genetic variability among different ethnic groups affects the distribution of mutations and responses to therapy. The lack of data from diverse populations can lead to bias in therapy and affect the effectiveness of precision medicine if applied to other populations. For example, the frequency of KRAS or BRAF mutations in colorectal cancer may differ between different populations, so effective therapies for one population may not necessarily have the same effectiveness in another (Habashy et al., 2023; Levin-Sparenberg et al., 2020; Booker et al., 2024). Therefore, additional research and data collection from various ethnic groups is needed to expand the scope and effectiveness of precision medicine in the treatment of colorectal cancer globally.
To address these challenges, cross-sector collaboration is needed between researchers, medical professionals, governments, and the pharmaceutical industry. Researchers can play a role in developing more affordable genomic technologies and expanding representative genomic databases. Meanwhile, medical professionals need to be trained in the interpretation and application of genomic results to support the implementation of precision medicine in the clinic. The government also has a big role to play in providing subsidies and supporting infrastructure development, as well as in designing policies that support equitable access to these technologies. The pharmaceutical industry, on the other hand, needs to be involved in the development of more affordable targeted medicines and promote global accessibility. Overall, although precision medicine offers promising prospects in the treatment of colorectal cancer, its implementation still faces challenges in terms of cost, accessibility, and diversity of genomic data. However, with continued advances and support from various sectors, precision medicine has the potential to become a new standard in cancer treatment, providing better treatment opportunities and improving the quality of life of colorectal cancer patients worldwide.
With the rapid development of technology and the increasing amount of genomic data available, the precision medicine approach has great potential to become the standard in the treatment of colorectal cancer in the future. Precision medicine allows doctors to tailor therapy based on each patient’s specific genomic profile, maximizing the effectiveness of treatment, and minimizing side effects. As one type of cancer that involves various genetic mutations such as in KRAS, BRAF, and PIK3CA, colorectal cancer greatly benefits from this genomics-based approach. In this context, the future of colorectal cancer treatment will focus on increasing the availability and accessibility of precision medicine for different populations and the development of more specific and effective therapies (Middleton et al., 2022; Bando et al., 2023).
One of the key factors driving precision medicine towards wider implementation is the increasing availability of genomic data. Today, large-scale genomics projects, such as the Human Genome Project and the Cancer Genome Atlas project, have produced rich data on genetic variation in different types of cancer, including colorectal cancer (Ganini et al., 2021; ICGC/TCGA Pan-Cancer Analysis of Whole Genomes Consortium, 2020; Guo et al., 2021). This data includes not only the most common mutations but also minor variations that may affect a patient’s response to therapy. With more and more data available, doctors can understand more about the specific mutation profiles in colorectal cancers and develop more accurate and effective therapeutic protocols. In addition, advances in sequencing technology, such as Next-Generation Sequencing (NGS), have enabled genomic profiling at higher speeds and lower costs (Satam et al., 2023; Gupta and Gupta, 2020). This technology makes the DNA sequencing process faster and more affordable so that genomic analysis can be applied more widely. In the future, it is hoped that this technology will become more affordable and accessible, allowing precision medicine to be integrated into clinical care standards for more colorectal cancer patients. With more affordable testing costs, precision medicine can be accessed by people from all walks of life, thereby increasing equity in cancer treatment.
For precision medicine to become the standard of care, close collaboration between researchers, medical practitioners, and policymakers is needed. Every stakeholder has a crucial role to play in realizing precision medicine that is more affordable, accessible, and has a real impact on colorectal cancer patients. Researchers in the fields of molecular biology, oncology, and bioinformatics have an important role to play in expanding the understanding of genetic mutations specific to colorectal cancer. They also need to develop more accurate and efficient algorithms for analyzing genomic data to assist doctors in making quick and precise clinical decisions (Hassan et al., 2022; Quazi, 2022). Broader and more representative research is needed to enrich genomic databases with data from various ethnic groups so that precision medicine can be applied in an inclusive manner (Tawfik et al., 2023). On the medical side, doctors and health workers need to be trained in the interpretation of genomic profile results and the application of targeted therapies in daily clinical practice. Many doctors may not have experience with precision medicine, so training and capacity building are essential. In addition, medical practitioners need to understand the importance of adaptive monitoring, where they must be prepared to modify or replace therapy if cancer cells show signs of resistance. Continuing education and expertise development in precision medicine can accelerate the integration of these approaches in clinical protocols. Policymakers have a strategic role to play in creating regulations and policies that support the development and access to precision medicine. Government subsidies for genomic testing, for example, can reduce the costs that patients have to bear, especially in developing countries. In addition, policies that encourage clinical research and the development of health infrastructure that supports precision medicine can accelerate the adoption of this approach across the region (Afzal et al., 2020). Policymakers also need to ensure that these technologies and healthcare services are available equally so that precision medicine is accessible to patients from different backgrounds and locations.
Although precision medicine offers a variety of benefits, there are still challenges that need to be overcome in order for this approach to become the standard in the treatment of colorectal cancer. In addition to the high cost, other challenges are the gap in access to technology and inequality in the availability of genomic data (Ory et al., 2023). Most of the current genomic data comes from Western populations, making it less representative of other populations, including Asian and African populations. Genetic variation between populations can affect the effectiveness of therapy, so additional research is needed to expand the scope of more inclusive genomic data (Sharif et al., 2020; Fernández-Rhodes et al., 2020). With more representative data, precision medicine can be applied more effectively and evenly around the world. In addition, to deal with potential drug resistance, precision medicine needs to develop a more responsive and adaptive monitoring system in the future (Beitler et al., 2022). Resistance to targeted therapy often develops as new mutations develop in cancer cells, which can reduce the effectiveness of treatment. Therefore, a more flexible monitoring system is needed to detect these changes early and adjust therapy according to the progression of the disease.
With strong collaboration between researchers, medical practitioners, and policymakers, precision medicine has great potential to become the standard in colorectal cancer treatment in the future. This process may take significant time and investment, but the results will provide great benefits to patients around the world (Denicolai and Previtali, 2020). With the widespread application of genomic technology, it is hoped that precision medicine can provide more effective, safe, and personalized therapy for colorectal cancer patients, improve their quality of life, and reduce morbidity and mortality rates. The success of precision medicine depends not only on technology but also on how this approach is integrated into the global health system. Policy support, infrastructure development, and education for healthcare professionals are key elements to make precision medicine a solution that is accessible to all patients, regardless of background or geographic location. Precision medicine is a promising future in colorectal cancer treatment, providing new hope for more effective and sustainable therapies, as well as pushing patients’ quality of life to a better level.
Genomic profiling-based precision medicine brings new promise in colorectal cancer treatment, enabling a more personalized and specific approach based on each patient’s genetic characteristics. With the ability to target specific genetic mutations or molecular pathways such as KRAS, BRAF, and PIK3CA, precision medicine is able to improve the effectiveness of treatment and reduce side effects, thereby significantly improving the patient’s quality of life. This approach not only focuses on more targeted treatments but also allows for adaptive monitoring of cancer cell progression, providing the flexibility to tailor therapy according to the patient’s changing condition. However, while precision medicine promises to have a major impact in treating colorectal cancer, its wider implementation still faces challenges, especially in terms of high costs, limited access to genomic technology, and the need for more representative genomic data for diverse populations. Therefore, continuous efforts are needed in terms of inclusive research, the development of more affordable technologies, and policy support from the government as well as cross-sector collaboration to expand access to precision medicine. With comprehensive support from various parties, precision medicine has the potential to become a standard of care that can be accessed by all levels of society, providing new hope for more effective and sustainable treatment of colorectal cancer.
AM: Conceptualization, Data curation, Formal Analysis, Funding acquisition, Investigation, Methodology, Project administration, Resources, Software, Supervision, Validation, Visualization, Writing–original draft, Writing–review and editing. AV: Conceptualization, Data curation, Formal Analysis, Funding acquisition, Investigation, Methodology, Project administration, Resources, Software, Supervision, Validation, Visualization, Writing–original draft, Writing–review and editing. AP: Conceptualization, Data curation, Formal Analysis, Funding acquisition, Investigation, Methodology, Project administration, Resources, Software, Supervision, Validation, Visualization, Writing–original draft, Writing–review and editing. SN: Conceptualization, Data curation, Formal Analysis, Funding acquisition, Investigation, Methodology, Project administration, Resources, Software, Supervision, Validation, Visualization, Writing–original draft, Writing–review and editing. PH: Conceptualization, Data curation, Formal Analysis, Funding acquisition, Investigation, Methodology, Project administration, Resources, Software, Supervision, Validation, Visualization, Writing–original draft, Writing–review and editing. RS: Project administration, Resources, Software, Supervision, Validation, Visualization, Writing–original draft, Writing–review and editing, Conceptualization, Data curation, Formal Analysis, Funding acquisition, Investigation, Methodology.
The author(s) declare that no financial support was received for the research, authorship, and/or publication of this article.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Adachi, Y., Kimura, R., Hirade, K., and Ebi, H. (2021). Escaping KRAS: Gaining autonomy and resistance to KRAS inhibition in KRAS mutant cancers. Cancers (Basel) 13, 5081. doi:10.3390/cancers13205081
Adeniji, A. A., Dulal, S., and Martin, M. G. (2021). Personalized medicine in oncology in the developing world: Barriers and concepts to improve status quo. World J. Oncol. 12, 50–60. doi:10.14740/wjon1345
Afrăsânie, V.-A., Marinca, M.-V., Gafton, B., Alexa-Stratulat, T., Rusu, A., Froicu, E.-M., et al. (2023). Clinical, pathological and molecular insights on KRAS, NRAS, BRAF, PIK3CA and TP53 mutations in metastatic colorectal cancer patients from Northeastern Romania. Int. J. Mol. Sci. 24, 12679. doi:10.3390/ijms241612679
Afzal, M., Riazul Islam, S. M., Hussain, M., and Lee, S. (2020). Precision medicine informatics: Principles, prospects, and challenges. IEEE Access 8, 13593–13612. doi:10.1109/access.2020.2965955
Ahmad, R., Singh, J. K., Wunnava, A., Al-Obeed, O., Abdulla, M., and Srivastava, S. K. (2021). Emerging trends in colorectal cancer: Dysregulated signaling pathways. Int. J. Mol. Med. 47, 1. (Review). doi:10.3892/ijmm.2021.4847
Andrei, P., Battuello, P., Grasso, G., Rovera, E., Tesio, N., and Bardelli, A. (2022). Integrated approaches for precision oncology in colorectal cancer: the more you know, the better. Semin. Cancer Biol. 84, 199–213. doi:10.1016/j.semcancer.2021.04.007
Arora, I., and Tollefsbol, T. O. (2021). Computational methods and next-generation sequencing approaches to analyze epigenetics data: profiling of methods and applications. Methods 187, 92–103. doi:10.1016/j.ymeth.2020.09.008
Ashenden, A. J., Chowdhury, A., Anastasi, L. T., Lam, K., Rozek, T., Ranieri, E., et al. (2024). The multi-omic approach to newborn screening: Opportunities and challenges. Int. J. Neonatal Screen 10, 42. doi:10.3390/ijns10030042
Atutornu, J., Milne, R., Costa, A., Patch, C., and Middleton, A. (2022). Towards equitable and trustworthy genomics research. EBioMedicine 76, 103879. doi:10.1016/j.ebiom.2022.103879
Bando, H., Ohtsu, A., and Yoshino, T. (2023). Therapeutic landscape and future direction of metastatic colorectal cancer. Nat. Rev. Gastroenterol. Hepatol. 20, 306–322. doi:10.1038/s41575-022-00736-1
Bashraheel, S. S., Domling, A., and Goda, S. K. (2020). Update on targeted cancer therapies, single or in combination, and their fine tuning for precision medicine. Biomed. Pharmacother. 125, 110009. doi:10.1016/j.biopha.2020.110009
Beitler, J. R., Thompson, B. T., Baron, R. M., Bastarache, J. A., Denlinger, L. C., Esserman, L., et al. (2022). Advancing precision medicine for acute respiratory distress syndrome. Lancet Respir. Med. 10, 107–120. doi:10.1016/S2213-2600(21)00157-0
Bond, C. E., and Whitehall, V. L. J. (2018). How the BRAF V600E mutation defines a distinct subgroup of colorectal cancer: Molecular and clinical implications. Gastroenterol. Res. Pract. 2018, 9250757. doi:10.1155/2018/9250757
Booker, B. D., Markt, S. C., Schumacher, F. R., Rose, J., Cooper, G., Selfridge, J. E., et al. (2024). Variation in KRAS/NRAS/BRAF-mutation status by age, sex, and race/ethnicity among a large cohort of patients with metastatic colorectal cancer (mCRC). J. Gastrointest. Cancer 55, 237–246. doi:10.1007/s12029-023-00954-z
Braicu, C., Buse, M., Busuioc, C., Drula, R., Gulei, D., Raduly, L., et al. (2019). A comprehensive review on MAPK: A promising therapeutic target in cancer. Cancers (Basel) 11, 1618. doi:10.3390/cancers11101618
Braig, F., März, M., Schieferdecker, A., Schulte, A., Voigt, M., Stein, A., et al. (2015). Epidermal growth factor receptor mutation mediates cross-resistance to panitumumab and cetuximab in gastrointestinal cancer. Oncotarget 6, 12035–12047. doi:10.18632/oncotarget.3574
Bukowski, K., Kciuk, M., and Kontek, R. (2020). Mechanisms of multidrug resistance in cancer chemotherapy. Int. J. Mol. Sci. 21, 3233. doi:10.3390/ijms21093233
Cai, W.-Q., Zeng, L.-S., Wang, L.-F., Wang, Y.-Y., Cheng, J.-T., Zhang, Y., et al. (2020). The latest battles between EGFR monoclonal antibodies and resistant tumor cells. Front. Oncol. 10, 1249. doi:10.3389/fonc.2020.01249
Cerdeña, J. P., Grubbs, V., and Non, A. L. (2022). Racialising genetic risk: assumptions, realities, and recommendations. Lancet 400, 2147–2154. doi:10.1016/S0140-6736(22)02040-2
Chakraborty, S., Sharma, G., Karmakar, S., and Banerjee, S. (2024). Multi-OMICS approaches in cancer biology: New era in cancer therapy. Biochim. Biophys. Acta Mol. Basis Dis. 1870, 167120. doi:10.1016/j.bbadis.2024.167120
Chakravarty, D., and Solit, D. B. (2021). Clinical cancer genomic profiling. Nat. Rev. Genet. 22, 483–501. doi:10.1038/s41576-021-00338-8
Chen, K., Zhang, Y., Qian, L., and Wang, P. (2021). Emerging strategies to target RAS signaling in human cancer therapy. J. Hematol. Oncol. 14, 116. doi:10.1186/s13045-021-01127-w
Ciardiello, F., Ciardiello, D., Martini, G., Napolitano, S., Tabernero, J., and Cervantes, A. (2022). Clinical management of metastatic colorectal cancer in the era of precision medicine. CA Cancer J. Clin. 72, 372–401. doi:10.3322/caac.21728
Cooper, A. J., Sequist, L. V., and Lin, J. J. (2022). Third-generation EGFR and ALK inhibitors: mechanisms of resistance and management. Nat. Rev. Clin. Oncol. 19, 499–514. doi:10.1038/s41571-022-00639-9
Corcoran, R. B., André, T., Atreya, C. E., Schellens, J. H. M., Yoshino, T., Bendell, J. C., et al. (2018). Combined BRAF, EGFR, and MEK inhibition in patients with BRAFV600E-mutant colorectal cancer. Cancer Discov. 8, 428–443. doi:10.1158/2159-8290.CD-17-1226
Cortés-Ciriano, I., Gulhan, D. C., Lee, J. J.-K., Melloni, G. E. M., and Park, P. J. (2022). Computational analysis of cancer genome sequencing data. Nat. Rev. Genet. 23, 298–314. doi:10.1038/s41576-021-00431-y
Dalal, N., Jalandra, R., Sharma, M., Prakash, H., Makharia, G. K., Solanki, P. R., et al. (2020). Omics technologies for improved diagnosis and treatment of colorectal cancer: Technical advancement and major perspectives. Biomed. Pharmacother. 131, 110648. doi:10.1016/j.biopha.2020.110648
Dankner, M., Rose, A. A. N., Rajkumar, S., Siegel, P. M., and Watson, I. R. (2018). Classifying BRAF alterations in cancer: new rational therapeutic strategies for actionable mutations. Oncogene 37, 3183–3199. doi:10.1038/s41388-018-0171-x
Denicolai, S., and Previtali, P. (2020). Precision Medicine: Implications for value chains and business models in life sciences. Technol. Forecast Soc. Change 151, 119767. doi:10.1016/j.techfore.2019.119767
Dharani, S., and Kamaraj, R. (2024). A review of the regulatory challenges of personalized medicine. Cureus 16, e67891. doi:10.7759/cureus.67891
Dietel, M., Jöhrens, K., Laffert, M. V., Hummel, M., Bläker, H., Pfitzner, B. M., et al. (2015). A 2015 update on predictive molecular pathology and its role in targeted cancer therapy: a review focussing on clinical relevance. Cancer Gene Ther. 22, 417–430. doi:10.1038/cgt.2015.39
Dillon, M., Lopez, A., Lin, E., Sales, D., Perets, R., and Jain, P. (2021). Progress on Ras/MAPK signaling research and targeting in blood and solid cancers. Cancers (Basel) 13, 5059. doi:10.3390/cancers13205059
Djanani, A., Eller, S., Öfner, D., Troppmair, J., and Maglione, M. (2020). The role of BRAF in metastatic colorectal carcinoma-past, present, and future. Int. J. Mol. Sci. 21, 9001. doi:10.3390/ijms21239001
Dong, R.-F., Zhu, M.-L., Liu, M.-M., Xu, Y.-T., Yuan, L.-L., Bian, J., et al. (2021). EGFR mutation mediates resistance to EGFR tyrosine kinase inhibitors in NSCLC: From molecular mechanisms to clinical research. Pharmacol. Res. 167, 105583. doi:10.1016/j.phrs.2021.105583
Edsjö, A., Holmquist, L., Geoerger, B., Nowak, F., Gomon, G., Alix-Panabières, C., et al. (2023). Precision cancer medicine: Concepts, current practice, and future developments. J. Intern Med. 294, 455–481. doi:10.1111/joim.13709
Faulkner, E., Holtorf, A.-P., Walton, S., Liu, C. Y., Lin, H., Biltaj, E., et al. (2020). Being precise about precision medicine: What should value frameworks incorporate to address precision medicine? A report of the personalized precision medicine special interest group. Value Health 23, 529–539. doi:10.1016/j.jval.2019.11.010
Feng, J., Hu, Z., Xia, X., Liu, X., Lian, Z., Wang, H., et al. (2023). Feedback activation of EGFR/wild-type RAS signaling axis limits KRASG12D inhibitor efficacy in KRASG12D-mutated colorectal cancer. Oncogene 42, 1620–1633. doi:10.1038/s41388-023-02676-9
Feng, Y., Spezia, M., Huang, S., Yuan, C., Zeng, Z., Zhang, L., et al. (2018). Breast cancer development and progression: Risk factors, cancer stem cells, signaling pathways, genomics, and molecular pathogenesis. Genes Dis. 5, 77–106. doi:10.1016/j.gendis.2018.05.001
Fernández-Rhodes, L., Young, K. L., Lilly, A. G., Raffield, L. M., Highland, H. M., Wojcik, G. L., et al. (2020). Importance of genetic studies of cardiometabolic disease in diverse populations. Circ. Res. 126, 1816–1840. doi:10.1161/CIRCRESAHA.120.315893
Florent, L., Saby, C., Slimano, F., and Morjani, H. (2023). BRAF V600-mutated metastatic melanoma and targeted therapy resistance: An update of the current knowledge. Cancers (Basel) 15, 2607. doi:10.3390/cancers15092607
Fountzilas, E., Tsimberidou, A. M., Vo, H. H., and Kurzrock, R. (2022). Clinical trial design in the era of precision medicine. Genome Med. 14, 101. doi:10.1186/s13073-022-01102-1
Frank, M. H., Wilson, B. J., Gold, J. S., and Frank, N. Y. (2021). Clinical implications of colorectal cancer stem cells in the age of single-cell omics and targeted therapies. Gastroenterology 160, 1947–1960. doi:10.1053/j.gastro.2020.12.080
Ganini, C., Amelio, I., Bertolo, R., Bove, P., Buonomo, O. C., Candi, E., et al. (2021). Global mapping of cancers: The cancer genome Atlas and beyond. Mol. Oncol. 15, 2823–2840. doi:10.1002/1878-0261.13056
García-Foncillas, J., Sunakawa, Y., Aderka, D., Wainberg, Z., Ronga, P., Witzler, P., et al. (2019). Distinguishing features of cetuximab and panitumumab in colorectal cancer and other solid tumors. Front. Oncol. 9, 849. doi:10.3389/fonc.2019.00849
Grothey, A., Fakih, M., and Tabernero, J. (2021). Management of BRAF-mutant metastatic colorectal cancer: a review of treatment options and evidence-based guidelines. Ann. Oncol. 32, 959–967. doi:10.1016/j.annonc.2021.03.206
Gu, W., Meng, F., Haag, R., and Zhong, Z. (2021). Actively targeted nanomedicines for precision cancer therapy: Concept, construction, challenges and clinical translation. J. Control Release 329, 676–695. doi:10.1016/j.jconrel.2020.10.003
Guo, X., Lin, W., Wen, W., Huyghe, J., Bien, S., Cai, Q., et al. (2021). Identifying novel susceptibility genes for colorectal cancer risk from a transcriptome-wide association study of 125,478 subjects. Gastroenterology 160, 1164–1178.e6. doi:10.1053/j.gastro.2020.08.062
Gupta, A. K., and Gupta, U. D. (2020). “Next generation sequencing and its applications,” in Animal Biotechnology. Elsevier, 395–421. doi:10.1016/b978-0-12-811710-1.00018-5
Gurdasani, D., Barroso, I., Zeggini, E., and Sandhu, M. S. (2019). Genomics of disease risk in globally diverse populations. Nat. Rev. Genet. 20, 520–535. doi:10.1038/s41576-019-0144-0
Habashy, P., Lea, V., Wilkinson, K., Wang, B., Wu, X.-J., Roberts, T. L., et al. (2023). KRAS and BRAF mutation rates and survival outcomes in colorectal cancer in an ethnically diverse patient cohort. Int. J. Mol. Sci. 24, 17509. doi:10.3390/ijms242417509
Haider, T., Pandey, V., Banjare, N., Gupta, P. N., and Soni, V. (2020). Drug resistance in cancer: mechanisms and tackling strategies. Pharmacol. Rep. 72, 1125–1151. doi:10.1007/s43440-020-00138-7
Hassan, M., Awan, F. M., Naz, A., deAndrés-Galiana, E. J., Alvarez, O., Cernea, A., et al. (2022). Innovations in genomics and big data analytics for personalized medicine and health care: A review. Int. J. Mol. Sci. 23, 4645. doi:10.3390/ijms23094645
Ho, D., Quake, S. R., McCabe, E. R. B., Chng, W. J., Chow, E. K., Ding, X., et al. (2020). Enabling technologies for personalized and precision medicine. Trends Biotechnol. 38, 497–518. doi:10.1016/j.tibtech.2019.12.021
Hu, H., Zhu, J., Zhong, Y., Geng, R., Ji, Y., Guan, Q., et al. (2021). PIK3CA mutation confers resistance to chemotherapy in triple-negative breast cancer by inhibiting apoptosis and activating the PI3K/AKT/mTOR signaling pathway. Ann. Transl. Med. 9, 410. doi:10.21037/atm-21-698
ICGC/TCGA Pan-Cancer Analysis of Whole Genomes Consortium (2020). Pan-cancer analysis of whole genomes. Nature 578, 82–93. doi:10.1038/s41586-020-1969-6
Janani, B., Vijayakumar, M., Priya, K., Kim, J. H., Prabakaran, D. S., Shahid, M., et al. (2022). EGFR-based targeted therapy for colorectal cancer-promises and challenges. Vaccines (Basel) 10, 499. doi:10.3390/vaccines10040499
Jauhri, M., Bhatnagar, A., Gupta, S., Bp, M., Minhas, S., Shokeen, Y., et al. (2017). Prevalence and coexistence of KRAS, BRAF, PIK3CA, NRAS, TP53, and APC mutations in Indian colorectal cancer patients: Next-generation sequencing-based cohort study. Tumour Biol. 39, 1010428317692265. doi:10.1177/1010428317692265
Karagiannakos, A., Adamaki, M., Tsintarakis, A., Vojtesek, B., Fåhraeus, R., Zoumpourlis, V., et al. (2022). Targeting oncogenic pathways in the era of personalized oncology: A systemic analysis reveals highly mutated signaling pathways in cancer patients and potential therapeutic targets. Cancers (Basel) 14, 664. doi:10.3390/cancers14030664
Kasi, P. M., Afable, M. G., Herting, C., Lukanowski, M., and Jin, Z. (2023). Anti-EGFR antibodies in the management of advanced colorectal cancer. Oncologist 28, 1034–1048. doi:10.1093/oncolo/oyad262
Keeling, P., Clark, J., and Finucane, S. (2020). Challenges in the clinical implementation of precision medicine companion diagnostics. Expert Rev. Mol. Diagn 20, 593–599. doi:10.1080/14737159.2020.1757436
Khoury, M. J., Bowen, S., Dotson, W. D., Drzymalla, E., Green, R. F., Goldstein, R., et al. (2022). Health equity in the implementation of genomics and precision medicine: A public health imperative. Genet. Med. 24, 1630–1639. doi:10.1016/j.gim.2022.04.009
Labrie, M., Brugge, J. S., Mills, G. B., and Zervantonakis, I. K. (2022). Therapy resistance: opportunities created by adaptive responses to targeted therapies in cancer. Nat. Rev. Cancer 22, 323–339. doi:10.1038/s41568-022-00454-5
Levin-Sparenberg, E., Bylsma, L. C., Lowe, K., Sangare, L., Fryzek, J. P., and Alexander, D. D. (2020). A systematic literature review and meta-analysis describing the prevalence of KRAS, NRAS, and BRAF gene mutations in metastatic colorectal cancer. Gastroenterol. Res. 13, 184–198. doi:10.14740/gr1167
Li, Q.-H., Wang, Y.-Z., Tu, J., Liu, C.-W., Yuan, Y.-J., Lin, R., et al. (2020). Anti-EGFR therapy in metastatic colorectal cancer: mechanisms and potential regimens of drug resistance. Gastroenterol. Rep. (Oxf) 8, 179–191. doi:10.1093/gastro/goaa026
Liu, M., Yang, X., Liu, J., Zhao, B., Cai, W., Li, Y., et al. (2017). Efficacy and safety of BRAF inhibition alone versus combined BRAF and MEK inhibition in melanoma: a meta-analysis of randomized controlled trials. Oncotarget 8, 32258–32269. doi:10.18632/oncotarget.15632
Lu, C. Y., Terry, V., and Thomas, D. M. (2023). Precision medicine: affording the successes of science. NPJ Precis. Oncol. 7, 3. doi:10.1038/s41698-022-00343-y
Malone, E. R., Oliva, M., Sabatini, P. J. B., Stockley, T. L., and Siu, L. L. (2020). Molecular profiling for precision cancer therapies. Genome Med. 12, 8. doi:10.1186/s13073-019-0703-1
Marbun, V. M. G., Erlina, L., and Lalisang, T. J. M. (2022). Genomic landscape of pathogenic mutation of APC, KRAS, TP53, PIK3CA, and MLH1 in Indonesian colorectal cancer. PLoS One 17, e0267090. doi:10.1371/journal.pone.0267090
Marine, J.-C., Dawson, S.-J., and Dawson, M. A. (2020). Non-genetic mechanisms of therapeutic resistance in cancer. Nat. Rev. Cancer 20, 743–756. doi:10.1038/s41568-020-00302-4
Martinelli, E., Ciardiello, D., Martini, G., Troiani, T., Cardone, C., Vitiello, P. P., et al. (2020). Implementing anti-epidermal growth factor receptor (EGFR) therapy in metastatic colorectal cancer: challenges and future perspectives. Ann. Oncol. 31, 30–40. doi:10.1016/j.annonc.2019.10.007
Mateo, J., Steuten, L., Aftimos, P., André, F., Davies, M., Garralda, E., et al. (2022). Delivering precision oncology to patients with cancer. Nat. Med. 28, 658–665. doi:10.1038/s41591-022-01717-2
Mathieson, I., and Reich, D. (2017). Differences in the rare variant spectrum among human populations. PLoS Genet. 13, e1006581. doi:10.1371/journal.pgen.1006581
Middleton, G., Robbins, H., Andre, F., and Swanton, C. (2022). A state-of-the-art review of stratified medicine in cancer: towards a future precision medicine strategy in cancer. Ann. Oncol. 33, 143–157. doi:10.1016/j.annonc.2021.11.004
Molina-Cerrillo, J., San Román, M., Pozas, J., Alonso-Gordoa, T., Pozas, M., Conde, E., et al. (2020). BRAF mutated colorectal cancer: New treatment approaches. Cancers (Basel) 12, 1571. doi:10.3390/cancers12061571
Naito, Y., Aburatani, H., Amano, T., Baba, E., Furukawa, T., Hayashida, T., et al. (2021). Clinical practice guidance for next-generation sequencing in cancer diagnosis and treatment (edition 2.1). Int. J. Clin. Oncol. 26, 233–283. doi:10.1007/s10147-020-01831-6
Nikolaou, M., Pavlopoulou, A., Georgakilas, A. G., and Kyrodimos, E. (2018). The challenge of drug resistance in cancer treatment: a current overview. Clin. Exp. Metastasis 35, 309–318. doi:10.1007/s10585-018-9903-0
Ory, M. G., Adepoju, O. E., Ramos, K. S., Silva, P. S., and Vollmer Dahlke, D. (2023). Health equity innovation in precision medicine: Current challenges and future directions. Front. Public Health 11, 1119736. doi:10.3389/fpubh.2023.1119736
Prins, B. P., Leitsalu, L., Pärna, K., Fischer, K., Metspalu, A., Haller, T., et al. (2021). Advances in genomic discovery and implications for personalized prevention and medicine: Estonia as example. J. Pers. Med. 11, 358. doi:10.3390/jpm11050358
Quazi, S. (2022). Artificial intelligence and machine learning in precision and genomic medicine. Med. Oncol. 39, 120. doi:10.1007/s12032-022-01711-1
Ramos, P., and Bentires-Alj, M. (2015). Mechanism-based cancer therapy: resistance to therapy, therapy for resistance. Oncogene 34, 3617–3626. doi:10.1038/onc.2014.314
Rascio, F., Spadaccino, F., Rocchetti, M. T., Castellano, G., Stallone, G., Netti, G. S., et al. (2021). The pathogenic role of PI3K/AKT pathway in cancer onset and drug resistance: An updated review. Cancers (Basel) 13, 3949. doi:10.3390/cancers13163949
Rastin, F., Javid, H., Oryani, M. A., Rezagholinejad, N., Afshari, A.-R., and Karimi-Shahri, M. (2024). Immunotherapy for colorectal cancer: Rational strategies and novel therapeutic progress. Int. Immunopharmacol. 126, 111055. doi:10.1016/j.intimp.2023.111055
Rodriguez Castells, M., Baraibar, I., Ros, J., Saoudi, N., Salvà, F., García, A., et al. (2023). The impact of clinical and translational research on the quality of life during the metastatic colorectal cancer patient journey. Front. Oncol. 13, 1272561. doi:10.3389/fonc.2023.1272561
Ros, J., Baraibar, I., Sardo, E., Mulet, N., Salvà, F., Argilés, G., et al. (2021). BRAF, MEK and EGFR inhibition as treatment strategies in BRAF V600E metastatic colorectal cancer. Ther. Adv. Med. Oncol. 13, 1758835921992974. doi:10.1177/1758835921992974
Salzman, R., Cook, F., Hunt, T., Malech, H. L., Reilly, P., Foss-Campbell, B., et al. (2018). Addressing the value of gene therapy and enhancing patient access to transformative treatments. Mol. Ther. 26, 2717–2726. doi:10.1016/j.ymthe.2018.10.017
Sanchez, J. N., Wang, T., and Cohen, M. S. (2018). BRAF and MEK inhibitors: Use and resistance in BRAF-mutated cancers. Drugs 78, 549–566. doi:10.1007/s40265-018-0884-8
Santos, E. da S., Nogueira, K. A. B., Fernandes, L. C. C., Martins, J. R. P., Reis, A. V. F., Neto, J. de B. V., et al. (2021). EGFR targeting for cancer therapy: Pharmacology and immunoconjugates with drugs and nanoparticles. Int. J. Pharm. 592, 120082. doi:10.1016/j.ijpharm.2020.120082
Satam, H., Joshi, K., Mangrolia, U., Waghoo, S., Zaidi, G., Rawool, S., et al. (2023). Next-generation sequencing technology: Current trends and advancements. Biol. (Basel) 12, 997. doi:10.3390/biology12070997
Sclafani, F., Wilson, S. H., Cunningham, D., Gonzalez De Castro, D., Kalaitzaki, E., Begum, R., et al. (2020). Analysis of KRAS, NRAS, BRAF, PIK3CA and TP53 mutations in a large prospective series of locally advanced rectal cancer patients. Int. J. Cancer 146, 94–102. doi:10.1002/ijc.32507
Sforza, V., Martinelli, E., Ciardiello, F., Gambardella, V., Napolitano, S., Martini, G., et al. (2016). Mechanisms of resistance to anti-epidermal growth factor receptor inhibitors in metastatic colorectal cancer. World J. Gastroenterol. 22, 6345–6361. doi:10.3748/wjg.v22.i28.6345
Sharif, S. M., Blyth, M., Ahmed, M., Sheridan, E., Saltus, R., Yu, J., et al. (2020). Enhancing inclusion of diverse populations in genomics: A competence framework. J. Genet. Couns. 29, 282–292. doi:10.1002/jgc4.1263
Tawfik, S. M., Elhosseiny, A. A., Galal, A. A., William, M. B., Qansuwa, E., Elbaz, R. M., et al. (2023). Health inequity in genomic personalized medicine in underrepresented populations: a look at the current evidence. Funct. Integr. Genomics 23, 54. doi:10.1007/s10142-023-00979-4
Traversi, D., Pulliero, A., Izzotti, A., Franchitti, E., Iacoviello, L., Gianfagna, F., et al. (2021). Precision medicine and public health: New challenges for effective and sustainable health. J. Pers. Med. 11, 135. doi:10.3390/jpm11020135
Tripathi, R., Sharma, P., Chakraborty, P., and Varadwaj, P. K. (2016). Next-generation sequencing revolution through big data analytics. Front. Life Sci. 9, 119–149. doi:10.1080/21553769.2016.1178180
Vitiello, P. P., Cardone, C., Martini, G., Ciardiello, D., Belli, V., Matrone, N., et al. (2019). Receptor tyrosine kinase-dependent PI3K activation is an escape mechanism to vertical suppression of the EGFR/RAS/MAPK pathway in KRAS-mutated human colorectal cancer cell lines. J. Exp. Clin. Cancer Res. 38, 41. doi:10.1186/s13046-019-1035-0
Voutsadakis, I. A. (2021). The landscape of PIK3CA mutations in colorectal cancer. Clin. Colorectal Cancer 20, 201–215. doi:10.1016/j.clcc.2021.02.003
Voutsadakis, I. A. (2023). KRAS mutated colorectal cancers with or without PIK3CA mutations: Clinical and molecular profiles inform current and future therapeutics. Crit. Rev. Oncol. Hematol. 186, 103987. doi:10.1016/j.critrevonc.2023.103987
Wang, R. C., and Wang, Z. (2023). Precision medicine: Disease subtyping and tailored treatment. Cancers (Basel) 15, 3837. doi:10.3390/cancers15153837
Yang, J., Griffin, A., Qiang, Z., and Ren, J. (2022). Organelle-targeted therapies: a comprehensive review on system design for enabling precision oncology. Signal Transduct. Target Ther. 7, 379. doi:10.1038/s41392-022-01243-0
Yip, H. Y. K., and Papa, A. (2021). Signaling pathways in cancer: Therapeutic targets, combinatorial treatments, and new developments. Cells 10, 659. doi:10.3390/cells10030659
Keywords: precision medicine, colorectal cancer, inflammation, BRAF mutations, genomics profiling
Citation: Muradi Muhar A, Velaro AJ, Prananda AT, Nugraha SE, Halim P and Syahputra RA (2025) Precision medicine in colorectal cancer: genomics profiling and targeted treatment. Front. Pharmacol. 16:1532971. doi: 10.3389/fphar.2025.1532971
Received: 27 November 2024; Accepted: 11 February 2025;
Published: 27 February 2025.
Edited by:
Sujit Nair, Phytoveda Pvt. Ltd., IndiaReviewed by:
Tamer A. Addissouky, University of Menoufia, EgyptCopyright © 2025 Muradi Muhar, Velaro, Prananda, Nugraha, Halim and Syahputra. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Rony Abdi Syahputra, cm9ueUB1c3UuYWMuaWQ=
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.