- 1Department of Respiratory Medicine, The Second Hospital of Jilin University, Changchun, China
- 2Department of Oncology and Hematology, The Second Hospital of Jilin University, Changchun, China
- 3Department of Anesthesiology, The Second Hospital of Jilin University, Changchun, China
- 4Department of Operation Room, The Second Hospital of Jilin University, Changchun, China
- 5Department of Urinary Surgery, The Second Hospital of Jilin University, Changchun, China
A limited number of studies have indicated an association between isoleucyl-tRNA synthetase 2 (IARS2) and tumorigenesis. We evaluated IARS2 protein expression in lung tumor tissues and paired non-tumor tissues. We found higher IARS2 expression in the tumor tissues, which was associated with the late Tumor and Node stages of the Tumor, Node, Metastasis staging system. Silencing IARS2 inhibited the activity of A549 and H1299 cells, resulting in G0/G1 stasis of A549 cells and mitochondrial apoptosis. IARS2 silencing was also found to inhibit NSCLC tumor growth in nude mice. Complementary DNA microarray analysis revealed 742 differentially expressed genes (507 upregulated and 235 downregulated) in IARS2-silenced A549 cells compared to controls. Ingenuity Pathway Analysis of the differential expression data suggested that multiple pathways are associated with IARS2 silencing in NSCLC cells; upstream analysis predicted the activation or inhibition of transcriptional regulators. Correlation analysis revealed that AKT and MTOR activities were significantly inhibited in IARS2-silenced cells, but were partially restored by the AKT-stimulating agent SC79. IARS2 appears to regulate lung cancer cell proliferation via the AKT/MTOR pathway. Our results help clarify the complex roles of IARS2 in tumorigenesis and suggest that it may be a novel regulator of lung cancer development.
Introduction
Lung cancer remains the most malignant tumor with the highest morbidity and mortality worldwide; for 2018, 2.1 million new lung cancer cases and 1.8 million deaths have been predicted, representing nearly 1 in 5 (18.4%) cancer deaths (1). Lung cancer is commonly classified into small cell carcinoma and non-small cell lung carcinoma (NSCLC) (2). In most countries, patients with adenocarcinoma are more common than those with squamous cell carcinoma (3). Active exploration of the pathogenesis of lung cancer has indicated that the identification of new lung cancer-related biomarkers may be crucial to improve the treatment and prognosis of patients with lung cancer.
The aminoacyl-tRNA synthetase (ARSs) class of evolutionarily ancient enzymes is widely found in organisms. ARSs are responsible for catalysing the esterification of the hydroxyl group of tRNAs with the carboxyl group of the corresponding amino acids to form aminoacyl tRNAs (4). Mammalian ARSs have evolved many new non-catalytic domains to perform non-canonical functions (5). ARSs play important roles in tumor pathogenesis by regulating tumor cell growth, differentiation, cell cycle, cytokine activity, RNA splicing, cell adhesion, and angiogenesis (6–8).
Isoleucyl-tRNA synthetase 2, mitochondrial (IARS2) is a nuclear gene encoding a mitochondrial ARS. The expression of IARS2 mRNA in human colon cancer tissues is higher than that in surrounding tissues. Knocking down the IARS2 gene inhibits the proliferation of colon cancer RKO cells, increases the proportion of cells in G0/G1 phase, and decreases the proportion of cells in S phase (9). In gastric cancer AGS cells, IARS2 knockdown inhibits proliferation and colony formation and induces cell cycle arrest in the G2/M phase (10). The expression of IARS2 in short-term survivors of glioblastoma is higher than that in long-term survivors, suggesting that high IARS2 expression is a risk factor for glioblastoma (11). These results suggest that IARS2 may be involved in the development and progression of tumors. However, the role of IARS2 in the development of NSCLC and its related molecular mechanisms are not well defined.
Materials and Methods
Tissue Specimens
We enrolled 56 patients with primary NSCLC who underwent surgery at the Second Hospital of Jilin University from May 2017 to August 2018. The patients had not received chemotherapy or radiation before surgery. All patients were diagnosed according to the World Health Organization's lung cancer criteria and staged according to the Tumor (T), Node (N), Metastasis (M) staging system for lung cancer (12). This study was approved by the Ethics Committee of the Second Hospital of Jilin University (Changchun, China) and all participants provided written informed consent. Lung cancer and corresponding tumor-adjacent lung tissue samples were collected during surgery and stored at −80°C. All healthy and cancerous tissues were re-evaluated by pathologists.
Cell Culture and Reagents
Human embryonic kidney 293 (HEK-293) cells and the human lung cell lines A549 and H1299 were obtained from the Chinese Academy of Medical Sciences (Beijing, China) and cultured in Dulbecco's modified Eagle's medium or RPMI-1640 (Gibco, Carlsbad, CA, USA) supplemented with 10% fetal bovine serum at 37°C in a humidified atmosphere containing 5% CO2. SC79 (catalog number S1023) was obtained from MedChemExpress (Monmouth Junction, NJ, USA).
Plasmids, Lentiviral Production, and Transduction
The IARS2 shRNA (shIARS2) lentiviral gene transfer vector pGCSIL-GFP, which encodes the enhanced green fluorescent protein sequence, was constructed by GeneChem (Shanghai, China). The hairpin sequence of shIARS2-1 was CCGGGTACTTGCAGTCATCCATTAATTCAAGAGATTAATGGATGACTGCAAGTACTTTTTG and the sequence of shIARS2-2 was CCGGGCTTAGGAATACACTTCGCTTCTCGAGAAGCGAAGTGTATTCCTAAGCTTTTT (GenBank accession number: NM_018060). The resulting constructs were verified by sequencing. A corresponding random shRNA sequence was used as a control for shIARS2. The vectors were transfected into HEK-293 cells using Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA) according to the manufacturer's instructions. Cell culture medium containing lentiviral particles was harvested 48 h post-transfection and passed through a 0.45-μm filter (Merck Millipore, Burlington, MA, USA). The resulting lentiviral particles were stored at −80°C until use. After transfection with lentivirus for 48 h, cells were screened in puromycin culture medium to establish cell lines with a stable knockout of the IARS2 gene.
Colony Formation Assay
Lentiviral vector-transduced A549 and H1299 cells were seeded in 6-well plates (400 cells/well). After 14 days, colonies (>50 cells/colony) were counted and individually imaged after staining with Giemsa (Beijing Solarbio Science & Technology, Beijing, China).
Cell Counting Kit-8 Assay
We determined the viability of A549 and H1299 cells using a Cell Counting Kit-8 (CCK-8) assay (Dojindo, Tokyo, Japan). Cells (2 × 103 cells/well) were seeded in 96-well plates and incubated at 37°C for 24 h. We added CCK-8 reagent (10 μL) to each well and incubated the plates for a further 1 h at 37°C. The optical density of each well was measured using a microplate reader (Thermo Fisher Scientific, Waltham, MA, USA) at a test wavelength of 450 nm.
Flow Cytometry Analysis of Cell Cycle and Apoptosis
For cell cycle analysis, the lentiviral vector-transduced A549 and H1299 cells were labeled with propidium iodide (BD Biosciences, San Jose, CA, USA) and analyzed using flow cytometry. For apoptosis analysis, the cells were incubated with PE-conjugated annexin V and 7-AAD (BD Biosciences, San Jose, CA, USA), according to the manufacturer's guidelines, prior to flow cytometry.
cDNA Microarray Assay and Ingenuity Pathway Analysis
To profile the expression of IARS2-regulated genes, we first stably transduced the A549 cells with the lentiviral shIARS2-1 or control vector. The cells were subjected to RNA isolation. The RNA was quantified using a NanoDrop 2000 (Thermo Fisher Scientific, Waltham, MA, USA) and checked for quality with an Agilent Bioanalyzer 2100 (Agilent Technologies, Santa Clara, CA, USA). Then, the quality-checked RNA samples were labeled with the GeneChip 3′ IVT Express Kit (Affymetrix, Santa Clara, CA, USA) and hybridized to an Affymetrix GeneChip PrimeView Human Gene Expression Array, according to the manufacturer's protocols. The experiments were conducted by GeneChem (Shanghai, China). The data were analyzed using a GeneChip Scanner 3000 (Affymetrix). We set the thresholds to determine gene alterations after IARS2 silencing at a 2.0-fold change in expression and p < 0.05 (false discovery rate [FDR] < 0.05) vs. the control cells. Ingenuity Pathway Analysis (IPA; Ingenuity Systems; www.ingenuity.com; Redwood City, CA, USA) is an online software package used to identify canonical pathways and gene networks and to categorize specific physiological processes. The Ingenuity Pathway Knowledge Base was used for deep analysis of the global molecular network and discovery of interactions among the differentially expressed genes.
Western Blotting
We obtained cellular proteins by lysing lung tissue with radioimmunoprecipitation assay buffer supplemented with protease inhibitors and phosphatase inhibitors. We determined the protein concentrations using a bicinchoninic acid protein assay kit (Beyotime, Jiangsu, China) according to the instructions. Lysis proteins (30 μg) were separated by electrophoresis on 8–15% sodium dodecyl sulfate–polyacrylamide gels. We transferred the samples to polyvinylidene difluoride membranes(#PIVH00010, Merck Millipore, Burlington, MA, USA). The membranes were probed with primary antibodies and were then incubated with horseradish peroxidase-conjugated secondary antibodies. We purchased the antibodies against AKT (# 4691), phospho-AKT (S473; # 4060), phospho-AKT (T308; # 13038), BCL-2 (# 2872), BAX (# 5023), cleaved caspase 3 (# 9664), cleaved PARP (# 5625), caspase 9 (# 9508), mammalian target of rapamycin (MTOR; # 2983), and phospho-MTOR (Ser2448; # 5536) from Cell Signaling Technologies (Danvers, MA, USA). We purchased the antibodies against β-actin (# 60008-1-Ig) and IARS2 (# 17170-1-AP) from Proteintech (Rosemont, IL, USA). The signals were detected using an enhanced chemiluminescence detection kit (Merck Millipore, Burlington, MA, USA). Densitometric analysis was performed with ImageJ; relative values are displayed under their respective blots.
Animal Studies
Five-week-old BALB/c nude mice were maintained under specific-pathogen-free conditions. We subcutaneously injected A549 cells (5 × 106 cells; 2.5 × 107 cell/mL in 0.2 mL phosphate-buffered saline) that stably expressed shIARS2-1, shIARS2-2, or the negative control shRNA into the left flanks of the mice. We measured the tumors with electronic calipers and calculated the sizes with the formula: volume = length × width2 × 0.5. Animal experiments were approved by the Institutional Animal Care and Use Committee of Jilin University.
Statistical Analysis
Pearson's Chi-squared test was performed to determine the association of clinicopathological data with the expression of IARS2 proteins in NSCLC tissues. Statistical data are expressed as the mean ± standard deviation. Comparisons among groups were carried out with Student's t-test. A p < 0.05 was considered statistically significant. All data were analyzed using SPSS 19.0 software.
Results
IARS2 Expression Was Higher in NSCLC Tissues Than in Non-cancerous Controls
We measured IARS2 protein expression in 56 pairs of NSCLC and adjacent normal tissue samples. IARS2 expression was higher in NSCLC tissues, and the high expression rate of adenocarcinoma accounted for 72.73%, which was slightly higher than that of squamous cell carcinoma. Elevated IARS2 protein levels were associated with the T and N stages of advanced tumors, but not with gender, age, or smoking status (Figures 1A,B and Table 1).
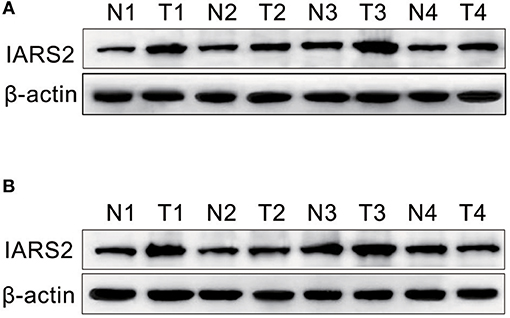
Figure 1. Western blotting of IARS2 protein expression in representative tissue samples from NSCLC (T) and non-tumor specimens (N). Total protein was extracted, subjected to western blotting analysis, and quantified using Image J software. (A) Squamous cell carcinoma tissue. (B) Adenocarcinoma tissue.
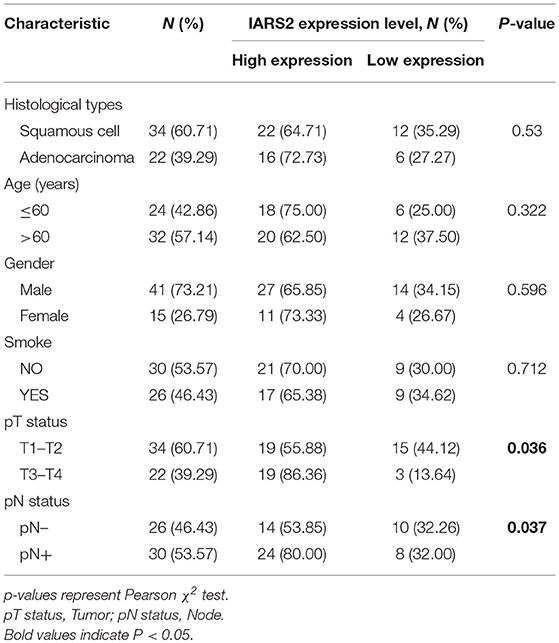
Table 1. Association of IARS2 with clinicopathological characteristics from 56 lung cancer patients.
IARS2 Knockdown Inhibited Lung Cancer Cell Viability and Colony Formation
We stably silenced IARS2 in A549 and H1299 cells. Compared with cells in the control group, cell proliferation activity of the shIARS2-1 and the shIARS2-2 groups decreased significantly (Figure 2A). Compared with control cells expressing the empty lentiviral expression vector, the shIARS2-1 and the shIARS2-2 groups had a lower clonogenic activity (Figure 2B).
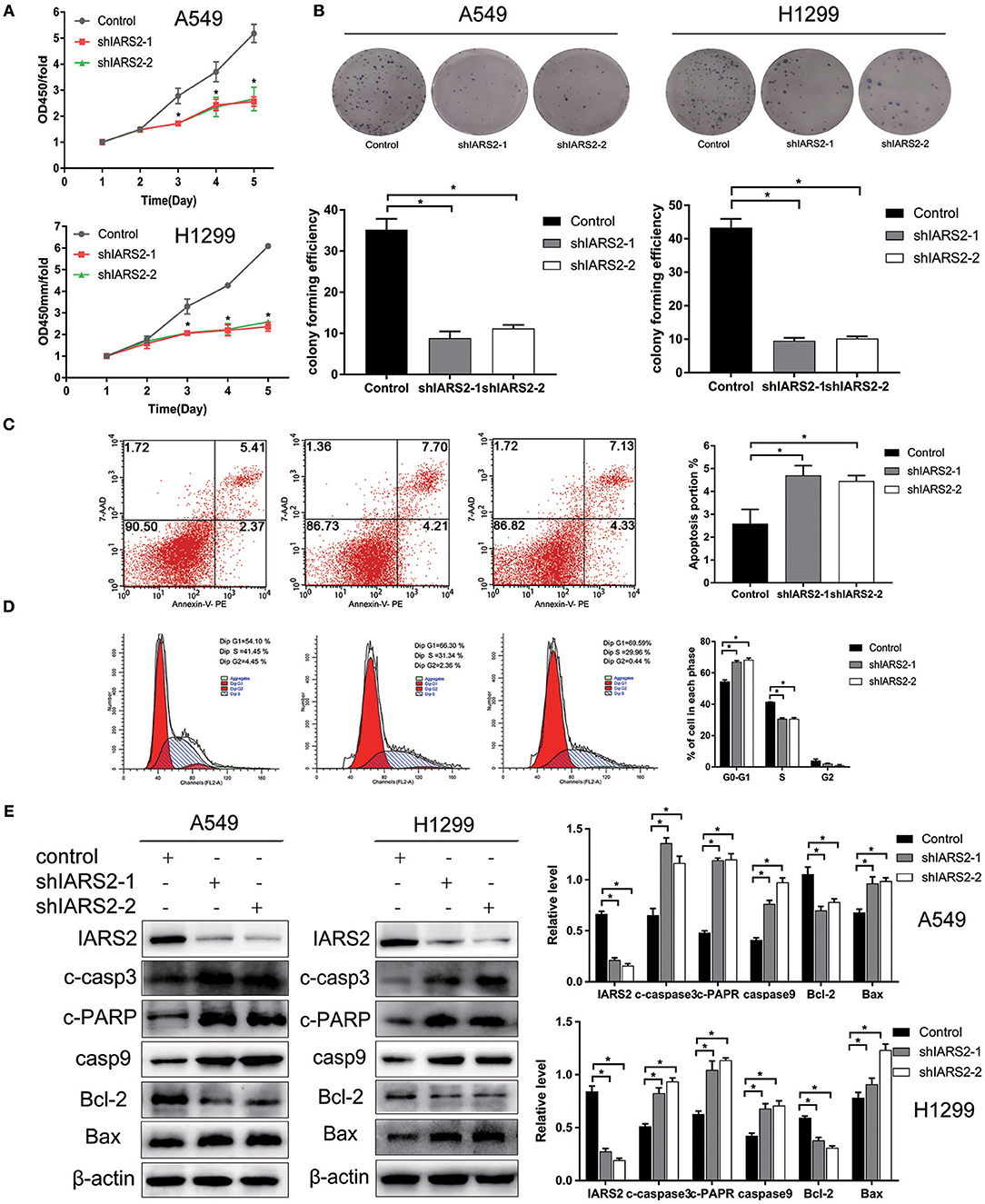
Figure 2. Effects of IARS2 knockdown on cell proliferation, cell cycle progression and apoptosis in NSCLC cells. (A) Cell Counting Kit-8 assay was used to evaluate A549 and H1299 cell growth after IARS2 knockdown. (B) Colony formation assay was used to evaluate A549 and H1299 cell growth after IARS2 knockdown. (C) Apoptosis was evaluated using flow cytometry in IARS2 knockdown and control A549 cells. Representative flow-cytograms are shown, and apoptotic rates were derived as percentages of Annexin V-PE positive cells. (D) Cell cycle was assessed in A549 cells using flow cytometry after transfection with the indicated shRNAs. Representative flow-cytograms are shown, as well as diagrams quantifying cell fractions in the G0/G1, S, and G2/M phases. (E) Differential expression of apoptosis regulatory proteins associated with IARS2 knockdown. Western blotting analysis was performed to compare expression levels of various apoptosis-related proteins between the IARS2 and control groups. *p < 0.05 compared with the control group.
Effect of IARS2 on Apoptosis and Cell Cycle
IARS2 silencing inhibited proliferation and promoted apoptosis of A549 cell (Figure 2C). In addition, compared to control cells, the proportion of G1 cells increased and that of S cells decreased (Figure 2D).
Analysis of Mitochondrial Apoptotic Pathway-Related Proteins in IARS2 Mediated-Apoptosis
To further illustrate the role of mitochondrial apoptosis as a downstream molecular mechanism of IARS2, we show that the expression of cleaved caspase 3, cleaved PARP, cleaved caspase 9, and BAX increased, whereas the expression of BCL-2 decreased, in IARS2-silenced cells. This suggests that IARS2-mediated growth suppression, at least in part, occurs via modulation of the mitochondrial apoptotic pathways (Figure 2E).
IARS2-Regulated Gene Expression Profiling
An Affymetrix GeneChip PrimeView Human cDNA microarray analysis was performed to profile the expression of IARS2-regulated genes in A549 cells (GEO accession number is GSE130007). IARS2 expression regulated the expression of 742 genes in A549 cells, of which 507 were upregulated and 235 were downregulated upon IARS2 silencing (Figure 3).
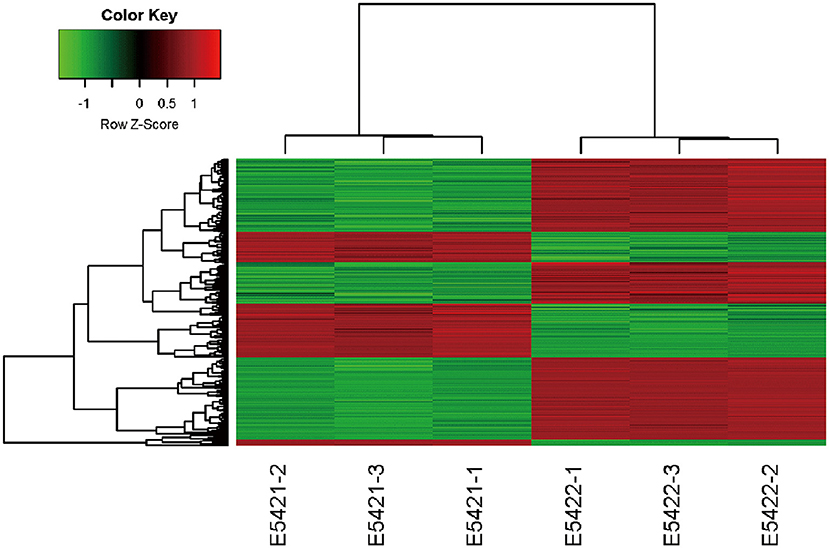
Figure 3. Heat map of the 2-way hierarchical clustering of genes and samples. Each row represents a gene and each column represents a sample. The gene clustering tree is shown on the left and the sample clustering tree appears at the top. Red, up-regulated in the IARS2-silenced vs. control cells; green down-regulated in the IARS2-silenced vs. control cells.
Functional Analysis of Differentially Expressed Genes Relative to Classical Pathways, Upstream Regulators, and Disease
Using IPA, we examined the relationships between the differentially expressed genes and canonical pathways. Our analysis revealed a highly significant overlap of the differentially regulated genes with 353 canonical pathways connected with apoptosis, cancer, cell cycle regulation, cellular immune responses, cellular growth, proliferation, and development. Figure 4A shows the 41 signaling pathways with the highest levels of significance.
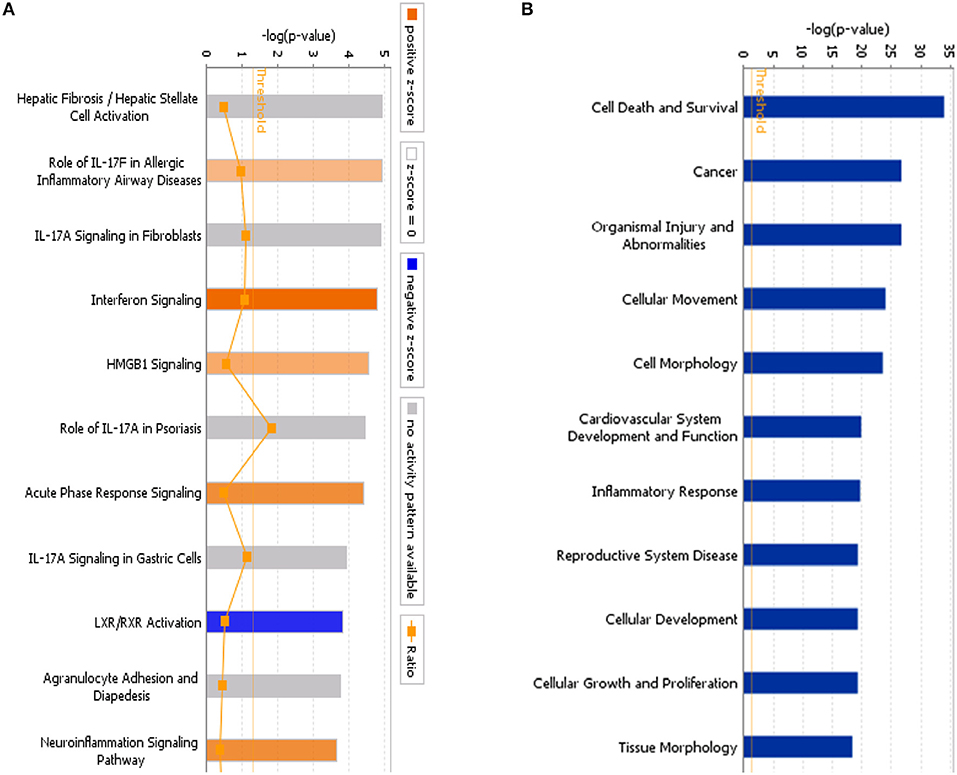
Figure 4. (A) Classical pathway enrichment analysis. The orange label indicates pathway activation (z-score >0), the blue label indicates pathway suppression (z-score <0), and the shades of orange and blue indicate the degree of activation or inhibition (the absolute value of the z-score). The ratio represents the number of differentially expressed genes in this signaling pathway and the number of all genes in the pathway. (B) Disease and functional enrichment analysis statistics. This figure shows the differentially expressed genes in the IARS2-silenced cells that are significantly enriched in disease and function. The abscissa is the name of the path and the ordinate is the level of significance of the enrichment (the negative logarithm of the base 10).
IPA identified 259 diseases or functions predicted to be activated upon IARS2 silencing, of which the top 5 were cell migration, cell movement, leukocyte migration, homing of cells, and chemotaxis. Of the 132 diseases or functions that were predicted to be inhibited, the top 5 were intestinal cancer, gastrointestinal tract cancer, infection of mammalia, morbidity or mortality, and organismal death (Figures 4B, 5).
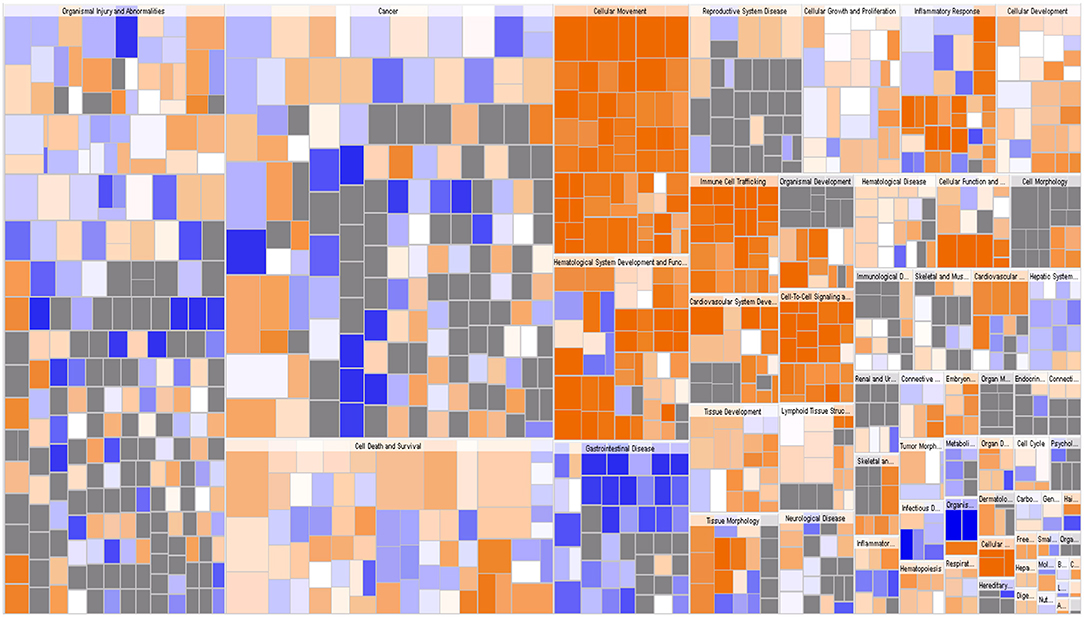
Figure 5. Disease and function heat map. This figure demonstrates the relationships between changes in differentially expressed gene levels and the activation and inhibition of diseases and functions. Orange indicates that the disease or functional status is activated (z-score >0), blue indicates that the disease or functional status is suppressed (z-score <0), and gray indicates that the disease or functional status was not determined (z-score cannot be calculated).
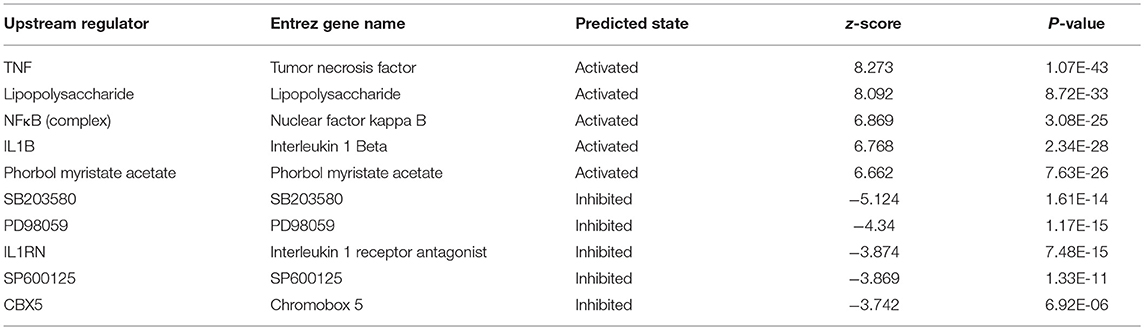
IPA uses the activation z-score algorithm to predict the activation or suppression of upstream regulators, reduce the instance of significant predictions resulting from random data, and analyse the relationships of genes to disease and function. In this study, 951 molecules (including transcription factors, cytokines, small RNAs, receptors, kinases, chemical molecules, and drugs) were predicted to be activators and 483 molecules were predicted to be inhibitors. Table 2 shows the IPA-predicted upstream activation or inhibitory molecules acting on the IARS2 gene (top 10).
IARS2-Activated AKT/MTOR Signaling
We found that the levels of AKT phosphorylation at Ser473 and Thr308 and MTOR phosphorylation in IARS2-silenced A549 and H1299 cells were significantly lower than those in the control group. This indicates that AKT/MTOR signaling pathway activation is significantly inhibited by IARS2 knockdown (Figure 6A, Figure S1).
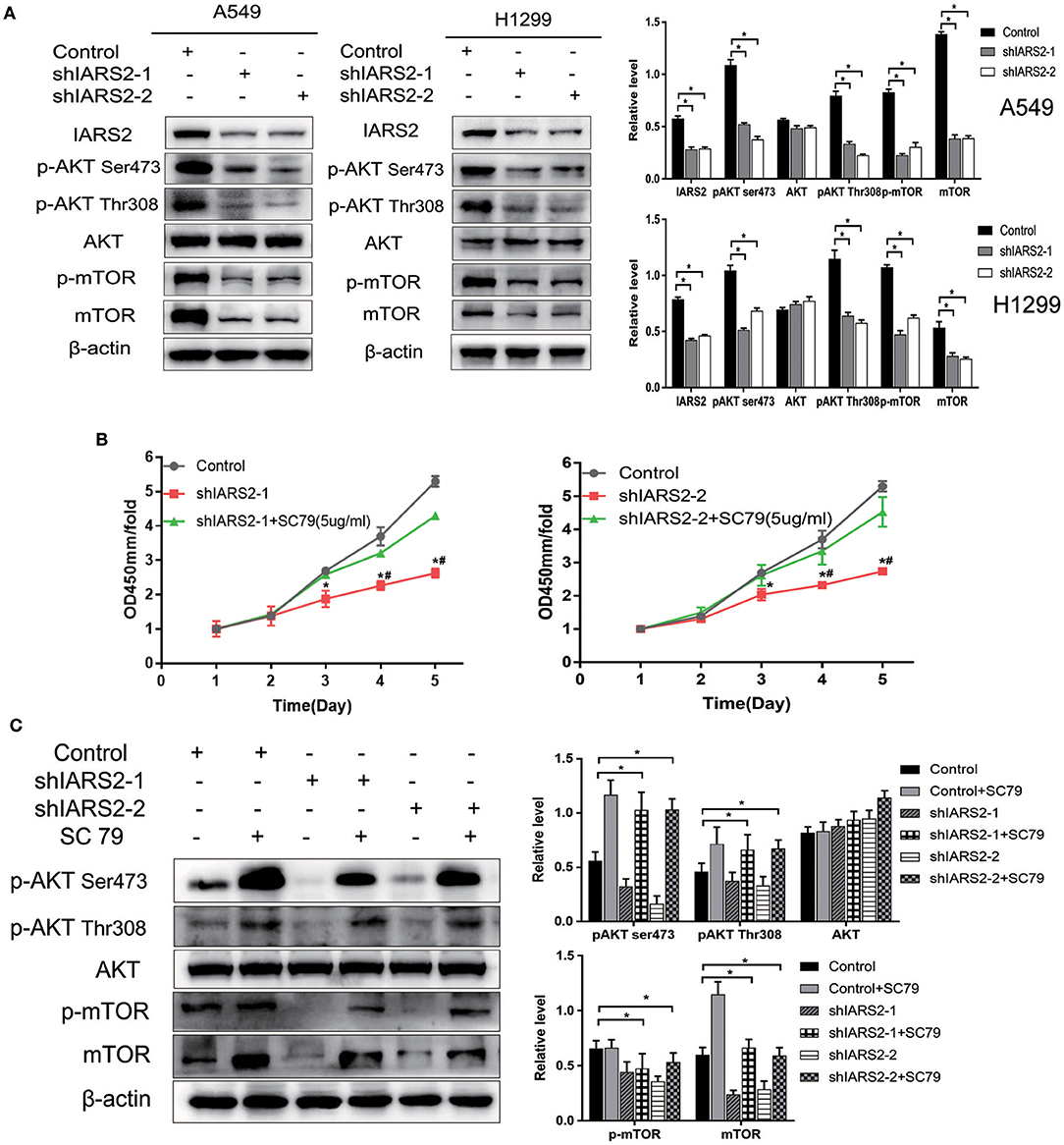
Figure 6. Correlation between IARS2 and AKT/MTOR signaling. (A) IARS2 activated AKT/MTOR signaling in A549 and H1299 cells. (B,C) The AKT activator SC79 partially restored AKT/MTOR signaling in A549 cells. CCK-8 assays and western blotting were performed to determine protein phosphorylation status after exposure to SC79 (5 μg/mL; 48 and 1 h, respectively). *, #p < 0.05. (*comparison with the control group; #comparison with the applied sc79 group).
AKT Activator SC79 Partially Restored AKT/MTOR Signaling
We treated shIARS2-1- and shIARS2-2-expressing A549 cells with the novel AKT activator SC79, which enhances phosphorylation of all AKT isoforms in a variety of cells. We performed CCK-8 assays and western blotting for protein phosphorylation status after exposure to SC79 (5 μg/mL; 48 and 1 h, respectively). SC79-induced AKT phosphorylation at Ser473 and Thr308, although the induction was slightly weaker than in control cells (Figures 6B,C). SC79 partially restored the IARS2 silencing-induced inhibition of lung cancer cell proliferation. Thus, the loss of IARS2 inhibits, at least in part, growth signaling cascades mediated by AKT.
IARS2 Regulated the Tumorigenic Capacity of Lung Cancer Cells
Finally, we assessed the tumorigenicity of IARS2-silenced cells in nude mice. All nude mice developed xenogenic tumors at the injection site (Figure 7A). In mice with IARS2-silenced cells, tumor growth was slower than in the control group (Figure 7B). Similarly, the weights of the excised gene-silenced xenograft tumors were lower than those of the control tumors (Figure 7C). Western blotting analyses of protein expression in tumor tissues (Figure 7D).
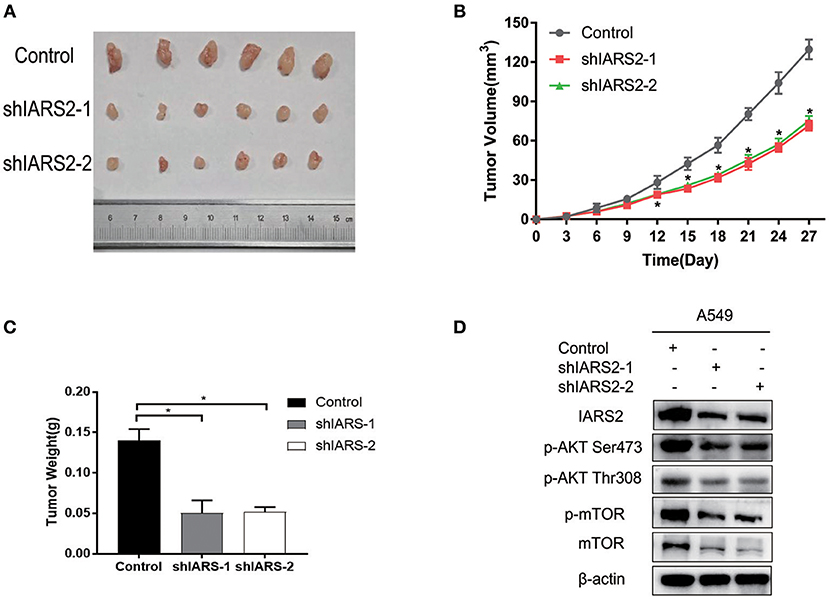
Figure 7. IARS2 silencing inhibits tumor growth in vivo. (A) Xenografts from mice in each group. (B) Tumor volumes were measured at the indicated times. (C) Mean weights of tumors obtained from mice. *p < 0.05 compared with the control group. (D)Western blotting analyses of protein expression in tumour tissues.
Discussion
Recently, a number of studies have shown that ARSs are associated with multiple tumors and play important roles in triggering or inhibiting tumors, including those associated with stomach, colon, lung, nasopharyngeal, oral, pancreatic, ovarian, prostate, colorectal, and breast cancers (13–22). The mitochondrial enzyme encoded by IARS2 (zone 4, band 1, chromosome 1) is synthesized in the cytoplasm and transported into the mitochondrion where it catalyses the binding of isoleucine to specific tRNAs for the completion of mitochondrial DNA translation. There are few reports on the relationship between IARS2 and disease; its role in tumor development remains unclear. In this study, we found that IARS2 is highly expressed in NSCLC tissues, particularly in adenocarcinoma patients and in tumor T and N stages. Detection of correlations between IARS2 expression and other clinical pathology data, such as gender or age, may require larger studies.
We found that knockdown of IARS2 inhibited the proliferation and clonal formation of A549 and H1299 cells, promoted apoptosis, and induced cell cycle arrest in A549 cells (G0/G1 phase) and H1299 cells (S phase), indicating that it affects DNA and protein synthesis, proliferation, and division in lung cancer cells. IARS2 knockdown significantly inhibits the proliferation and colony formation ability of gastric cancer AGS cells and induces cycle arrest at G2/M phase (10). Together, these results suggest that IARS2 may be a drug target for the treatment of NSCLC.
ARSs play important roles in mitochondrial protein synthesis, thereby contributing to mitochondrial oxidative phosphorylation (23). BCL2, BAX, and caspase 9 are important regulators of mitochondria-dependent apoptosis (24, 25). We found that IARS2 is involved in the mitochondrial apoptosis pathway. Apoptosis, cell cycle arrest, and cell proliferation inhibition are related. Our experimental results consistently indicate that IARS2 plays important roles in the growth, proliferation, and apoptosis of NSCLC cells.
Our cDNA microarray analysis revealed highly significant overlap of 353 IARS2-regulated canonical pathways connected with apoptosis, cancer, cell cycle regulation, and cellular immune responses, and cellular growth, proliferation, and development. Our results indicate that IARS2 is involved in the regulation of a variety of complex biological processes. These data may provide new clues for the study and treatment of lung cancer development, but the detailed mechanisms of IARS2 action require further research.
The AKT/MTOR pathway is a central regulator of cell proliferation, apoptosis, cell cycle, metabolism, and angiogenesis (26). Its activation is associated with tumorigenesis, tumor resistance, invasion, and metastasis. AKT/MTOR signaling pathway-related proteins are also abnormally expressed in liver, lung, breast, bladder, prostate, gastrointestinal, and ovarian cancers (27–33). Aberrant signaling pathway activation is associated with NSCLC and small cell lung cancer cells and cisplatin resistance (34, 35).
Mitochondrial stress leads to increased expression, activation, and nuclear localization of AKT (36). AKT-mediated signaling suppresses mitochondrial oxidation, inhibits apoptosis, and increases cancer cell proliferation (37, 38). Leucyl-tRNA synthetase reportedly initiates mTORC1 activation (39). Both leucyl-tRNA synthetase and IARS are class I ARSs that are presumed to have similar functions in cancer. In this study, IARS2 knockdown promoted mitochondria-dependent apoptosis, suggesting that IARS2 may promote lung cancer cell proliferation and inhibit apoptosis through abnormal activation of AKT/MTOR signaling.
IARS2 knockdown reduced AKT Ser473 and Thr308 and MTOR phosphorylation levels in lung cancer A549 and H1299 cells. Recent work suggested that mitochondrial stress leads to increased expression, activation, and nuclear localization of AKT. The Luo laboratory recently developed the novel AKT activator SC79, which enhances the phosphorylation of all AKT isoforms (40). Using SC79, we demonstrated that AKT activation moderately restored cell proliferation and decreased apoptosis in the absence of IARS.
Conclusions
In summary, our results indicated that IARS2, an ancient protein synthesis enzyme, may play complex regulatory roles in lung cancer, which should be studied in depth. This study preliminarily revealed the important role of IARS2 in lung cancer pathogenesis and provided a solid hypothetical basis for considering IARS2 expression in the diagnosis and treatment of lung cancer.
Ethics Statement
This study was approved by the Ethics Committee of the Second Hospital of Jilin University (Changchun, China) and all participants provided written informed consent. The animal experiments were approved by the Institutional Animal Care and Use Committee of Jilin University.
Author Contributions
KW contributed conception and design of the study. HM, SC, and RW performed the samples collection. JL, CT, and MZ analyzed the data analysis. XD and XJ produced the main draft of the manuscript and made figures. KW and RL obtained funding for the study. All authors contributed to manuscript revision, read and approved the submitted version.
Funding
This study was supported by the Medical And Health Industry Development Guide Funds of Jilin Province (201603034YY to KW), the Special funds for industrial innovation in Jilin Province (2016C043-3 to KW), the Natural Science Foundation of Jilin Province (20180101103JC to RL), the Technology Research Funds of Jilin Province (20190303162SF to KW), and the Medical And Health Project Funds of Jilin Province (20191102012YY to RL).
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fonc.2019.00393/full#supplementary-material
Figure S1. Effect of silencing IARS2 on p-ERK/ERK.
References
1. Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. (2018) 68:394–424. doi: 10.3322/caac.21492
2. McIntyre A, Ganti AK. Lung cancer-A global perspective. J Surg Oncol. (2017) 115:550–4. doi: 10.1002/jso.24532
3. Cheng TY, Cramb SM, Baade PD, Youlden DR, Nwogu C, Reid ME. The international epidemiology of lung cancer: latest trends, disparities, and tumor characteristics. J Thorac Oncol. (2016) 11:1653–71. doi: 10.1016/j.jtho.2016.05.021
4. Brown MV, Reader JS, Tzima E. Mammalian aminoacyl-tRNA synthetases: cell signaling functions of the protein translation machinery. Vascul Pharmacol. (2010) 52:21–6. doi: 10.1016/j.vph.2009.11.009
5. Kim S, You S, Hwang D. Aminoacyl-tRNA synthetases and tumorigenesis: more than housekeeping. Nat Rev Cancer. (2011) 11:708–18. doi: 10.1038/nrc3124
6. Mirando AC, Francklyn CS, Lounsbury KM. Regulation of angiogenesis by aminoacyl-tRNA synthetases. Int J Mol Sci. (2014) 15:23725–48. doi: 10.3390/ijms151223725
7. Young HJ, Lee JW, Kim S. Function of membranous lysyl-tRNA synthetase and its implication for tumorigenesis. Biochim Biophys Acta. (2016) 1864:1707–13. doi: 10.1016/j.bbapap.2016.09.009
8. Kim D, Kwon NH, Kim S. Association of aminoacyl-tRNA synthetases with cancer. Top Curr Chem. (2014) 344:207–45. doi: 10.1007/128_2013_455
9. Zhong L, Zhang Y, Yang JY, Xiong LF, Shen T, Sa YL, et al. Expression of IARS2 gene in colon cancer and effect of its knockdown on biological behavior of RKO cells. Int J Clin Exp Pathol. (2015) 8:12151–9.
10. Fang Z, Wang X, Yan Q, Zhang S, Li Y. Knockdown of IARS2 suppressed growth of gastric cancer cells by regulating the phosphorylation of cell cycle-related proteins. Mol Cell Biochem. (2018) 443:93–100. doi: 10.1007/s11010-017-3213-8
11. Mazaris P, Hong X, Altshuler D, Schultz L, Poisson LM, Jain R, et al. Key determinants of short-term and long-term glioblastoma survival: a 14-year retrospective study of patients from the Hermelin Brain Tumor Center at Henry Ford Hospital. Clin Neurol Neurosurg. (2014) 120:103–12. doi: 10.1016/j.clineuro.2014.03.001
12. Goldstraw P, Crowley J, Chansky K, Giroux DJ, Groome PA, Rami-Porta R, et al. The IASLC Lung Cancer Staging Project: proposals for the revision of the TNM stage groupings in the forthcoming (seventh) edition of the TNM Classification of malignant tumours. J Thorac Oncol. (2007) 2:706–14. doi: 10.1097/JTO.0b013e31812f3c1a
13. Kim BH, Jung WY, Lee H, Kang Y, Jang YJ, Hong SW, et al. Lysyl-tRNA synthetase (KRS) expression in gastric carcinoma and tumor-associated inflammation. Ann Surg Oncol. (2014) 21:2020–7. doi: 10.1245/s10434-014-3522-z
14. Nam SH, Kim D, Lee MS, Lee D, Kwak TK, Kang M, et al. Noncanonical roles of membranous lysyl-tRNA synthetase in transducing cell-substrate signaling for invasive dissemination of colon cancer spheroids in 3D collagen I gels. Oncotarget. (2015) 6:21655–74. doi: 10.18632/oncotarget.4130
15. Shin SH, Kim HS, Jung SH, Xu HD, Jeong YB, Chung YJ. Implication of leucyl-tRNA synthetase 1 (LARS1) over-expression in growth and migration of lung cancer cells detected by siRNA targeted knock-down analysis. Exp Mol Med. (2008) 40:229–36. doi: 10.3858/emm.2008.40.2.229
16. Zhou W, Feng X, Li H, Wang L, Zhu B, Liu W, et al. Inactivation of LARS2, located at the commonly deleted region 3p21.3, by both epigenetic and genetic mechanisms in nasopharyngeal carcinoma. Acta Biochim Biophys Sin. (2009) 41:54–62. doi: 10.1093/abbs/gmn006
17. Lee CW, Chang KP, Chen YY, Liang Y, Hsueh C, Yu JS, et al. Overexpressed tryptophanyl-tRNA synthetase, an angiostatic protein, enhances oral cancer cell invasiveness. Oncotarget. (2015) 6:21979–92. doi: 10.18632/oncotarget.4273
18. Paley EL, Paley DE, Merkulova-Rainon T, Subbarayan PR. Hypoxia signature of splice forms of tryptophanyl-tRNA synthetase marks pancreatic cancer cells with distinct metastatic abilities. Pancreas. (2011) 40:1043–56. doi: 10.1097/MPA.0b013e318222e635
19. Wellman TL, Eckenstein M, Wong C, Rincon M, Ashikaga T, Mount SL, et al. Threonyl-tRNA synthetase overexpression correlates with angiogenic markers and progression of human ovarian cancer. BMC Cancer. (2014) 14:620. doi: 10.1186/1471-2407-14-620
20. Vellaichamy A, Sreekumar A, Strahler JR, Rajendiran T, Yu J, Varambally S, et al. Proteomic interrogation of androgen action in prostate cancer cells reveals roles of aminoacyl tRNA synthetases. PLoS ONE. (2009) 4:e7075. doi: 10.1371/journal.pone.0007075
21. Mallawaaratchy DM, Mactier S, Kaufman KL, Blomfield K, Christopherson RI. The phosphoinositide 3-kinase inhibitor LY294002, decreases aminoacyl-tRNA synthetases, chaperones and glycolytic enzymes in human HT-29 colorectal cancer cells. J Proteomics. (2012) 75:1590–9. doi: 10.1016/j.jprot.2011.11.032
22. He Y, Gong J, Wang Y, Qin Z, Jiang Y, Ma H, et al. Potentially functional polymorphisms in aminoacyl-tRNA synthetases genes are associated with breast cancer risk in a Chinese population. Mol Carcinog. (2015) 54:577–83. doi: 10.1002/mc.22128
23. Konovalova S, Tyynismaa H. Mitochondrial aminoacyl-tRNA synthetases in human disease. Mol Genet Metab. (2013) 108:206–11. doi: 10.1016/j.ymgme.2013.01.010
24. Lee JY, Jee SB, Park WY, Choi YJ, Kim B, Kim YH, et al. Tumor suppressor protein p53 promotes 2-methoxyestradiol-induced activation of Bak and Bax, leading to mitochondria-dependent apoptosis in human colon cancer HCT116 cells. J Microbiol Biotechnol. (2014) 24:1654–63. doi: 10.4014/jmb.1405.05062
25. Choi D, Hwang S, Lee E, Yoon S, Yoon BK, Bae D. Expression of mitochondria-dependent apoptosis genes (p53, Bax, and Bcl-2) in rat granulosa cells during follicular development. J Soc Gynecol Investig. (2004) 11:311–7. doi: 10.1016/j.jsgi.2004.01.015
26. Manfredi GI, Dicitore A, Gaudenzi G, Caraglia M, Persani L, Vitale G. PI3K/Akt/mTOR signaling in medullary thyroid cancer: a promising molecular target for cancer therapy. Endocrine. (2015) 48:363–70. doi: 10.1007/s12020-014-0380-1
27. Li H, Zeng J, Shen K. PI3K/AKT/mTOR signaling pathway as a therapeutic target for ovarian cancer. Arch Gynecol Obstet. (2014) 290:1067–78. doi: 10.1007/s00404-014-3377-3
28. Fumarola C, Bonelli MA, Petronini PG, Alfieri RR. Targeting PI3K/AKT/mTOR pathway in non-small cell lung cancer. Biochem Pharmacol. (2014) 90:197–207. doi: 10.1016/j.bcp.2014.05.011
29. Ciruelos Gil EM. Targeting the PI3K/AKT/mTOR pathway in estrogen receptor-positive breast cancer. Cancer Treat Rev. (2014) 40:862–71. doi: 10.1016/j.ctrv.2014.03.004
30. Costa C, Pereira S, Lima L, Peixoto A, Fernandes E, Neves D, et al. Abnormal protein glycosylation and activated PI3K/Akt/mTOR pathway: role in bladder cancer prognosis and targeted therapeutics. PLoS ONE. (2015) 10:e0141253. doi: 10.1371/journal.pone.0141253
31. Toren P, Zoubeidi A. Targeting the PI3K/Akt pathway in prostate cancer: challenges and opportunities (review). Int J Oncol. (2014) 45:1793–801. doi: 10.3892/ijo.2014.2601
32. Danielsen SA, Eide PW, Nesbakken A, Guren T, Leithe E, Lothe RA. Portrait of the PI3K/AKT pathway in colorectal cancer. Biochim Biophys Acta. (2015) 1855:104–21. doi: 10.1016/j.bbcan.2014.09.008
33. Mabuchi S, Kuroda H, Takahashi R, Sasano T. The PI3K/AKT/mTOR pathway as a therapeutic target in ovarian cancer. Gynecol Oncol. (2015) 137:173–9. doi: 10.1016/j.ygyno.2015.02.003
34. Umemura S, Mimaki S, Makinoshima H, Tada S, Ishii G, Ohmatsu H, et al. Therapeutic priority of the PI3K/AKT/mTOR pathway in small cell lung cancers as revealed by a comprehensive genomic analysis. J Thorac Oncol. (2014) 9:1324–31. doi: 10.1097/JTO.0000000000000250
35. Xia A, Li H, Li R, Lu L, Wu X. Co-treatment with BEZ235 enhances chemosensitivity of A549/DDP cells to cisplatin via inhibition of PI3K/Akt/mTOR signaling and downregulation of ERCC1 expression. Oncol Rep. (2018) 40:2353–62. doi: 10.3892/or.2018.6583
36. Guha M, Fang JK, Monks R, Birnbaum MJ, Avadhani NG. Activation of Akt is essential for the propagation of mitochondrial respiratory stress signaling and activation of the transcriptional coactivator heterogeneous ribonucleoprotein A2. Mol Biol Cell. (2010) 21:3578–89. doi: 10.1091/mbc.e10-03-0192
37. Chun SY, Johnson C, Washburn JG, Cruz-Correa MR, Dang DT, Dang LH. Oncogenic KRAS modulates mitochondrial metabolism in human colon cancer cells by inducing HIF-1alpha and HIF-2alpha target genes. Mol Cancer. (2010) 9:293. doi: 10.1186/1476-4598-9-293
38. Uchiumi T, Ohgaki K, Yagi M, Aoki Y, Sakai A, Matsumoto S, et al. ERAL1 is associated with mitochondrial ribosome and elimination of ERAL1 leads to mitochondrial dysfunction and growth retardation. Nucleic Acids Res. (2010) 38:5554–68. doi: 10.1093/nar/gkq305
39. Han JM, Jeong SJ, Park MC, Kim G, Kwon NH, Kim HK, et al. Leucyl-tRNA synthetase is an intracellular leucine sensor for the mTORC1-signaling pathway. Cell. (2012) 149:410–24. doi: 10.1016/j.cell.2012.02.044
Keywords: isoleucyl-tRNA synthetase 2, lung cancer, tumorigenesis, cDNA microarray, ingenuity pathway analysis, AKT, mammalian target of rapamycin
Citation: Di X, Jin X, Ma H, Wang R, Cong S, Tian C, Liu J, Zhao M, Li R and Wang K (2019) The Oncogene IARS2 Promotes Non-small Cell Lung Cancer Tumorigenesis by Activating the AKT/MTOR Pathway. Front. Oncol. 9:393. doi: 10.3389/fonc.2019.00393
Received: 13 December 2018; Accepted: 26 April 2019;
Published: 14 May 2019.
Edited by:
Emilio Hirsch, University of Turin, ItalyReviewed by:
Ana Clara Carrera, Spanish National Research Council (CSIC), SpainM. Christine Hollander, National Institutes of Health (NIH), United States
Copyright © 2019 Di, Jin, Ma, Wang, Cong, Tian, Liu, Zhao, Li and Wang. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Ke Wang, d2tlQGpsdS5lZHUuY24=
 Xin Di1
Xin Di1 Ke Wang
Ke Wang