
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Mol. Biosci. , 27 January 2022
Sec. Molecular Diagnostics and Therapeutics
Volume 8 - 2021 | https://doi.org/10.3389/fmolb.2021.792154
Immunotherapy can improve survival in a variety of cancers by modulating the interaction between tumors and the tumor immune microenvironment (TIME). V-set and transmembrane domain containing 2 like (VSTM2L) regulates interleukin (IL)-4 signaling pathway—which involves immune-related factors—and has been linked to some cancers. However, the expression profile and prognostic significance of VSTM2L in different cancers as well as its relationship to the TIME are not known. This study investigated the pan-cancer expression profile, prognostic value, and immunologic relevance of VSTM2L. VSTM2L expression in different cancers was analyzed using the Cancer Cell Line Encyclopedia (CCLE), Human Protein Atlas (HPA), Tumor Immune Estimation Resource (TIMER), The Cancer Genome Atlas (TCGA), and Genotype–Tissue Expression (GTEx) portal. We examined the association between VSTM2L expression and clinical outcomes by Kaplan–Meier and Cox regression analyses using TCGA and Kaplan–Meier Plotter, and the results were validated in a Gene Expression Omnibus cohort. The correlations between VSTM2L expression and immune cell infiltration, immunomodulators, tumor mutation burden (TMB), microsatellite instability (MSI), and immune and stromal scores across cancers were analyzed using TCGA, TIMER, and Tumor–Immune System Interactions and Drugbank databases (TISIDB). The results showed that VSTM2L expression varied across cancers and its aberrant expression was associated with clinical outcomes: upregulation of VSTM2L was positively associated with advanced stage and reduced overall survival (OS), disease-specific survival (DSS), progression-free interval (PFI), and disease-free interval (DFI) in stomach adenocarcinoma (STAD); and its upregulation was associated with early-stage disease and improved OS, DSS, PFI, and DFI in kidney renal papillary cell carcinoma (KIRP). VSTM2L expression level was correlated with immune cell infiltration, expression of immunomodulators, TMB, MSI, and immune and stromal scores in multiple cancers. In conclusion, VSTM2L has prognostic value in various cancers and can predict both poor (STAD) and good (KIRP) outcomes. The relationship between VSTM2L expression and immune markers suggests a role in modulating the TIME.
The interaction between tumors and the tumor immune microenvironment (TIME) influences the occurrence, progression, and treatment of cancers (Hinshaw and Shevde, 2019; Lei et al., 2020). Although cancer immunotherapy has improved the survival of cancer patients, treatment response rates are low (Binnewies et al., 2018). Clarifying tumor–TIME interactions can help to identify novel markers for predicting the response to immunotherapy or that can serve as pharmacologic targets (Binnewies et al., 2018; Taube et al., 2018; Petitprez et al., 2020).
V-set and transmembrane domain-containing two like (VSTM2L, also known as C20orf102) is expressed in multiple human tissues, with the highest expression observed in the cerebral cortex and pituitary. As a secreted protein that antagonizes the neuroprotective peptide humanin, VSTM2L has been implicated in neurodegenerative and metabolic diseases (Rossini et al., 2011). However, there have been few studies on the role of VSTM2L in cancer. VSTM2L was shown to be downregulated in Helicobacter pylori-positive gastric cancer compared to corresponding normal tissues (Hu et al., 2018); additionally, the CpG island methylation phenotype (CIMP)-related gene signature comprising VSTM2L and five other genes showed prognostic value in gastric cancer (Zeng et al., 2020). In locally advanced rectal cancer, elevated expression of VSTM2L conferred chemoradiotherapy resistance via regulation of interleukin (IL)-4 signaling pathway (Liu et al., 2021), which is involved in the immune response in cancer patients (Lee et al., 2009; Rajaraman et al., 2009; Siliņa et al., 2011; Li et al., 2019; Didonna et al., 2020; Wei et al., 2020).
The above observations suggest that VSTM2L plays an important role in cancer and likely modulates immunity, although this has not yet been reported. To address this point, in this study we used multiple databases, including TCGA, GEO, CCLE, HPA, TIMER, GTEx, and TISIDB in combination with Kaplan–Meier Plotter to perform a comprehensive and multidimensional pan-cancer analysis of the role of VSTM2L. We compared VSTM2L expression in different types of cancer and corresponding normal tissue. We also evaluated the prognostic value of VSTM2L in cancer and investigated the relationship between VSTM2L expression level and various aspects of the TIME including immune cell infiltration, expression of immunomodulators (such as immunostimulators, immunoinhibitors, and major histocompatibility complex (MHC) molecules), tumor mutation burden (TMB), microsatellite instability (MSI), and Immune/StromalScores.
VSTM2L expression level in human cancer cell lines was determined from RNA sequencing (RNA-seq) data in the Cancer Cell Line Encyclopedia (CCLE) database (https://portals.broadinstitute.org/ccle). The expression module in Tumor Immune Estimation Resource (TIMER; https://cistrome.shinyapps.io/timer/) was used to visualize differential expression of VSTM2L across 33 tumors in The Cancer Genome Atlas (TCGA) compared to adjacent normal tissue, including adrenocortical carcinoma (ACC), bladder urothelial carcinoma (BLCA), breast invasive carcinoma (BRCA), cervical squamous cell carcinoma (CESC), cholangiocarcinoma (CHOL), colon adenocarcinoma (COAD), lymphoid neoplasm diffuse large B cell lymphoma (DLBC), esophageal carcinoma (ESCA), glioblastoma multiforme (GBM), brain lower grade glioma (LGG), head and neck squamous cell carcinoma (HNSC), kidney chromophobe (KICH), kidney renal clear cell carcinoma (KIRC), kidney renal papillary cell carcinoma (KIRP), acute myeloid leukemia, liver hepatocellular carcinoma (LIHC), lung adenocarcinoma (LUAD), lung squamous cell carcinoma (LUSC), mesothelioma (MESO), ovarian serous cystadenocarcinoma (OV), pancreatic adenocarcinoma (PAAD), pheochromocytoma and paraganglioma (PCPG), prostate adenocarcinoma (PRAD), rectum adenocarcinoma (READ), sarcoma (SARC), skin cutaneous melanoma (SKCM), stomach adenocarcinoma (STAD), testicular germ cell tumors (TGCT), thyroid carcinoma (THCA), thymoma (THYM), uterine corpus endometrial carcinoma (UCEC), uterine carcinosarcoma (UCS), and uveal melanoma (UVM). Additionally, VSTM2L expression data of 31 normal tissues were obtained from the Genotype–Tissue Expression (GTEx) database (https://commonfund.nih.gov/GTex), and data for 33 cancer types, adjacent normal tissue, and corresponding clinical information of patients were extracted from TCGA datasets obtained from the University of California Santa Cruz Xena browser (https://xena.ucsc.edu/). Differential expression analyses of VSTM2L between cancer and normal tissues were performed by merging the mRNA expression data from GTEx with TCGA datasets, and data on protein expression detected by immunohistochemistry was obtained from Human Protein Atlas (HPA). VSTM2L expression was also evaluated with respect to clinical stage in 33 cancers. Whole RNA-seq data were normalized through log2 conversion.
Kaplan–Meier survival and Cox regression analyses were used to evaluate the prognostic value of VSTM2L expression for overall survival (OS), disease-specific survival (DSS), progression-free interval (PFI), and disease-free interval (DFI) in 33 cancer types using the “survival” and “survminer” packages of R software. According to the median VSTM2L expression in each cancer type, patients were divided into high and low expression groups and Kaplan–Meier survival analysis was carried out using prognostic data. Kaplan–Meier Plotter (http://kmplot.com/analysis/) was used to examine the influence of VSTM2L on OS in patients with STAD or KIRP based on clinicopathologic factors and cell content, respectively. Gene Expression Omnibus (GEO) data (GSE84437 and GSE2748) were used to assess the prognostic value of VSTM2L in STAD and KIRP. Hazard ratio (HR) with 95% confidence intervals (CIs) and the log-rank p value were determined by Cox regression analysis. A log p value < .05 was considered statistically significant.
We used TIMER (https://cistrome.shinyapps.io/timer/) to quantify the association between VSTM2L expression in diverse cancer types and the abundance of six tumor-infiltrating immune cells (TIIC) types including B cells, cluster of differentiation (CD)8 + T cells, CD4+ T cells, macrophages, neutrophils, and dendritic cells in the tumor mass as well as other immune cells, which was adjusted by purity. Gene lists of three types of immunomodulator including immunoinhibitors, immunostimulators, and MHC molecules were obtained from the Tumor–immune System Interactions and Drugbank (TISIDB) database (http://cis.hku.hk/TISIDB/index.php). TIMER was used to examine the correlation between VSTM2L expression level and immunomodulator genes in various cancers. Spearman correlations between VSTM2L expression and immune and stromal scores (ImmuneScore and StromalScore, respectively) were determined.
TMB was defined as the total incidences of mutation per million base pairs and MSI was calculated as the number of deletion or insertion events occurring in repeating sequences of genes; both were obtained from TCGA. The Spearman correlation test was used to determine the correlations between VSTM2L expression and TMB and MSI in 33 cancers types.
VSTM2L expression levels in tumor and normal tissues across cancers were compared by wilcoxon test. Kaplan-Meier analysis and univariate Cox regression analysis were used to evaluate the correlation between VSTM2L expression and patient prognosis. Correlations between VSTM2L expression and TIICs, immunomodulators, ImmuneScore, StromalScore, TMB, and MSI were determined by Spearman correlation analysis. The threshold for statistical significance was set as p < .05.
GTEx data showed that VSTM2L mRNA was widely expressed in normal tissues, with the highest expression in spleen, brain, and pituitary and the lowest expression in bone marrow and blood (Figure 1A). RNA-seq data in the CCLE database showed that VSTM2L was highly expressed in Ewing sarcoma cell line (Figure 1B). We analyzed VSTM2L expression in different cancers; the TIMER data across all TCGA tumors showed that compared to corresponding normal tissue, VSTM2L was significantly upregulated in BRCA, KIRP, LUAD, PAAD, PCPG, PRAD, and THCA and downregulated in COAD, GBM, KICH, KIRC, LUSC, STAD, and UCEC (Figure 1C). Analysis of combined TCGA and GTEx data revealed similar trends in VSTM2L expression as observed in the TIMER data, but it also showed that VSTM2L was significantly upregulated in ACC, OV, SKCM, and UCS and downregulated in ESCA, LGG, LIHC, and TGCT relative to matched normal tissue (Figure 1D). Advanced tumor stage was more closely associated with VSTM2L expression in BLCA, COAD, KIRC, STAD, and THCA, while the opposite was true for KIRP (Figure 1E). No association was found between VSTM2L expression and cancer stage in other cancer types (Supplementary Figure S1). VSTM2L protein expression in STAD and COAD and corresponding normal tissue detected by immunohistochemistry was determined from HPA datasets. Consistent with the mRNA levels, VSTM2L showed lower expression in both gastric cancer and colon cancer tissues than in normal tissues (Figure 1F).
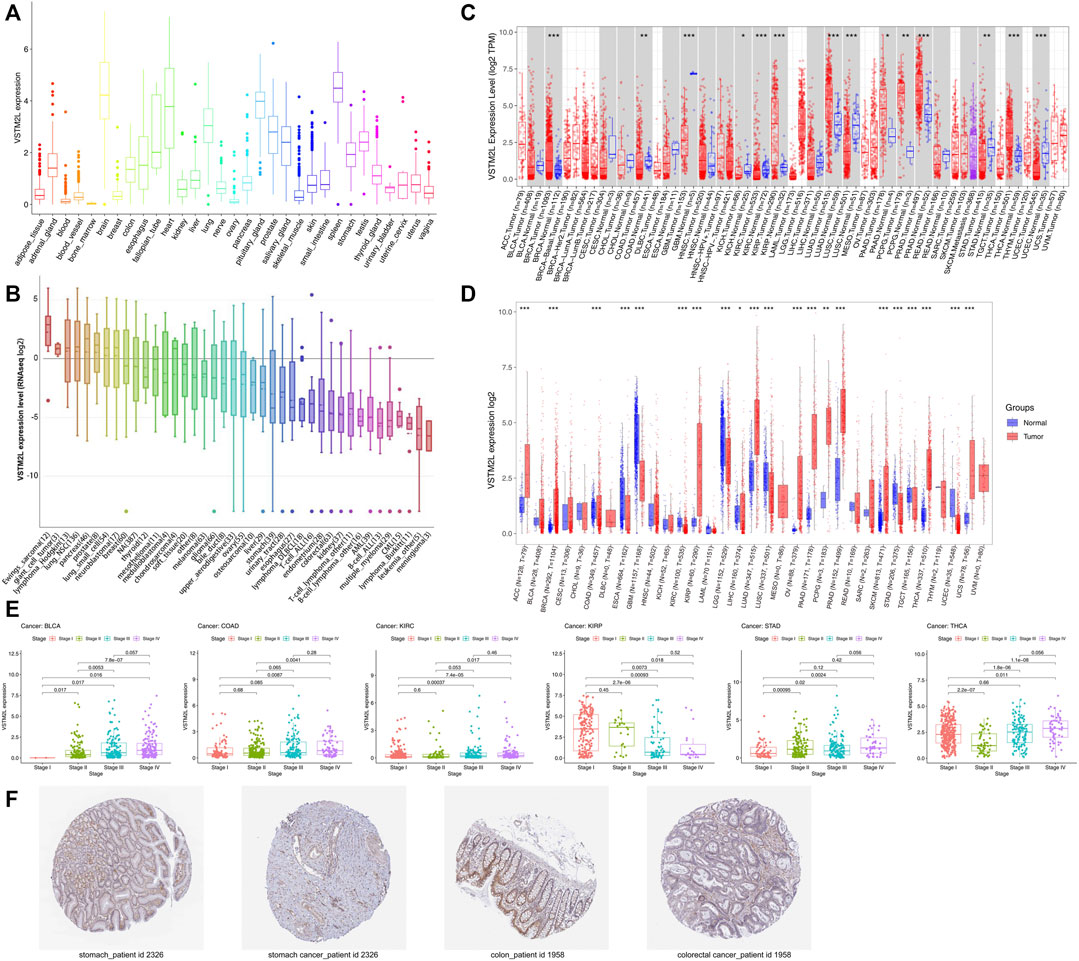
FIGURE 1. VSTM2L expression in different cancers. (A) RNA-seq data of VSTM2L expression level in human cancer cell lines in the CCLE database. (B) VSTM2L expression data of 31 normal tissues from GTEx datasets. (C) Differential expression of VSTM2L in various cancer types based on TCGA data in TIMER. (D) VSTM2L expression level in 33 cancer types from combined GTE and TCGA data. (E) Relationship between VSTM2L expression level and clinical stage in BLCA, COAD, KIRC, KIRP, STAD, and THCA. (F) Immunohistochemical detection of VSTM2L protein expression in the tumor and corresponding normal tissue of a STAD patient (ID: 2,326) and COAD patient (ID: 1958) in the HPA database. *p < .05, **p < .01, ***p < .001.
The Kaplan–Meier survival and Cox regression analyses based on TCGA data showed that the prognostic value of VSTM2L differed according to cancer types. High VSTM2L expression was associated with shorter OS in BLCA, KIRC, OV, STAD, THYM, UCEC, and UVM but predicted better outcome in KIRP (Figure 2A); VSTM2L expression had no prognostic value in the OS of other cancers (Supplementary Figure S2). Cox regression analysis showed that increased VSTM2L expression was correlated with shorter OS in KIRC (p = .047, HR = 1.166), LUSC (p = .049, HR = 1.099), MESO (p = .007, HR = 1.350), OV (p = .009 HR = 1.109), STAD (p = .023, HR = 1.129), UCEC (p = .010, HR = 1.255), and UVM (p = .010, HR = 1.883) (Figure 2B) and predicted a better outcome in KIRP (p = .003, HR = .790), LGG (p = .031, HR = .873), and PCPG (p < .001, HR = .461) (Figure 2B).
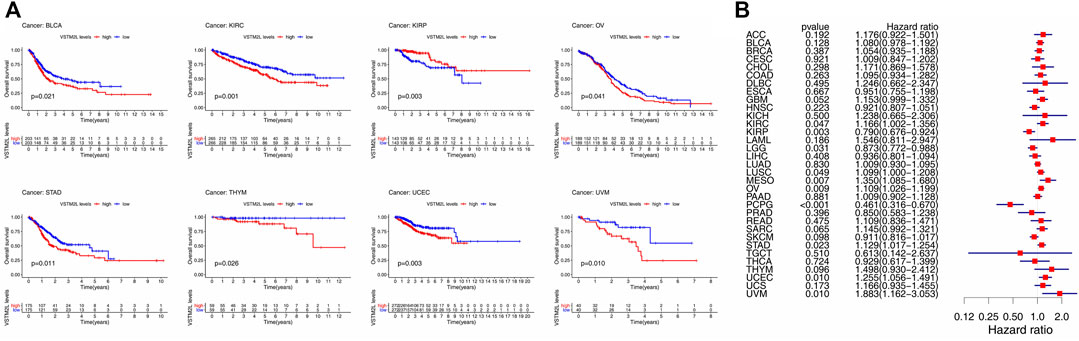
FIGURE 2. Kaplan–Meier survival and Cox regression analyses of the prognostic value of VSTM2L expression level for OS in different cancer types. (A) OS according to high and low VSTM2L expression in BLCA, KIRC, KIRP, OV, STAD, THYM, UCEC, and UVM from TCGA database. (B) Correlation between VSTM2L mRNA expression level and OS in various cancer types. *p < 0.05.
High VSTM2L expression was associated with shorter DSS in KIRC, OV, STAD, UCEC, and UVM and longer DSS in KIRP and PCPG in the Kaplan–Meier survival analysis (Figure 3A). The same results were obtained by Cox regression analysis, which also revealed a significant HR for GBM and LGG (Figure 3B).
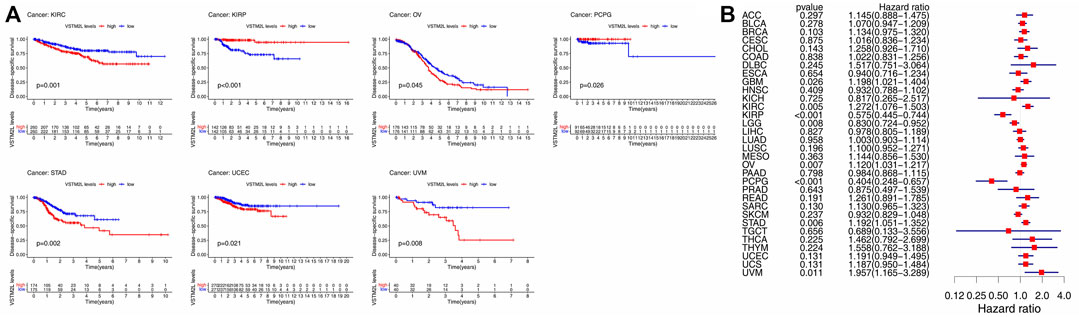
FIGURE 3. Kaplan–Meier survival and Cox regression analyses of the prognostic value of VSTM2L expression level for DSS in different cancer types. (A) DSS according to high and low VSTM2L expression in KIRC, KIRP, OV, PCPG, STAD, UCEC, and UVM from TCGA. (B) Correlation between VSTM2L mRNA expression level and DSS in various cancer types. *p < .05.
Elevated VSTM2L expression was linked to shorter PFI in DLBC, KIRC, STAD, and THCA, and shorter DFI in STAD; it was also associated with longer PFI in KIRP and longer DFI in KIRC, KIRP, and LIHC (Figure 4A and Figure 5A). Upregulation of VSTM2L was correlated with high HRs of PFI in DLBC, KIRC, LUSC, and STAD and high HRs of DFI in LUSC and STAD (Figure 4B and Figure 5B). Meanwhile, downregulation of VSTM2L expression was correlated with high HRs of PFI in KIRP, LGG, and PCPG and high HRs of DFI in KIRP.
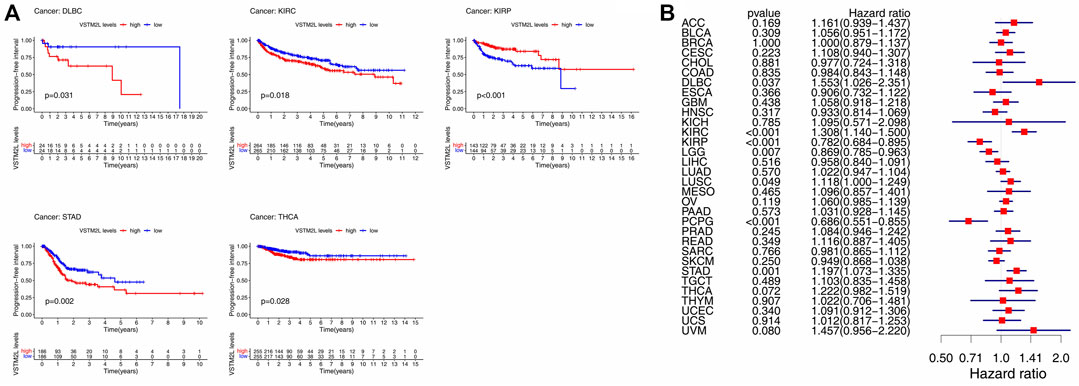
FIGURE 4. Kaplan–Meier survival and Cox regression analyses of the prognostic value of VSTM2L expression level for PFI in different cancer types. (A) PFI according to high and low VSTM2L expression in DLBC, KIRC, KIRP, STAD, and THCA from TCGA. (B) Correlation between VSTM2L mRNA expression level and PFI in various cancer types. *p < .05.
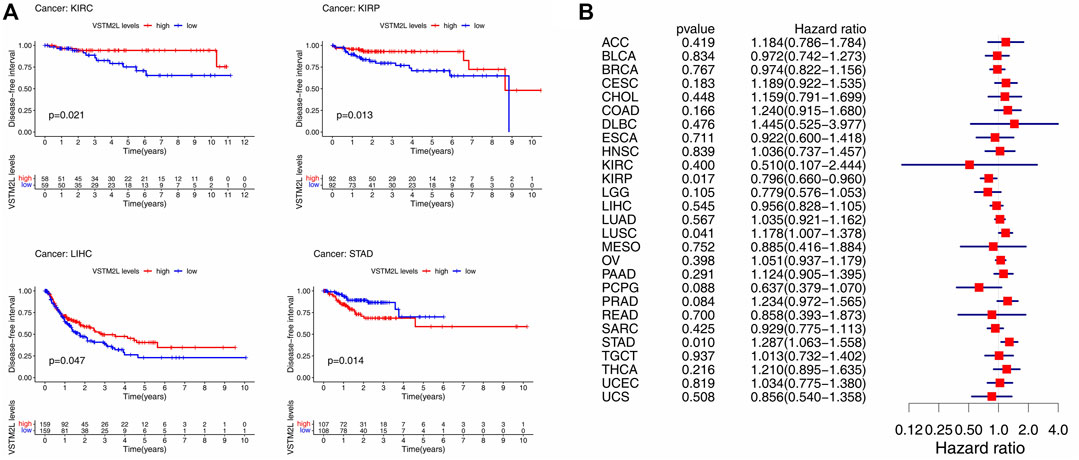
FIGURE 5. Kaplan–Meier survival and Cox regression analyses of the prognostic value of VSTM2L expression level for DFI in different cancer types. (A) DFI according to high and low VSTM2L expression in KIRC, KIRP, LIHC, and STAD from TCGA. (B) Correlation between VSTM2L mRNA expression level and DFI in various cancer types. *p < .05.
As the TIME is linked to prognosis and response to immunotherapy in cancer (Binnewies et al., 2018; Hinshaw and Shevde, 2019), we next examined the relationship between VSTM2L expression and immune cell infiltration (Figure 6, details are shown in Supplementary Figure S3) and the expression of immunomodulators in the TIME across 33 cancers extracted from TCGA datasets using TIMER. We found that VSTM2L was related to TIICs in most cancers with the exception of DLBC, GBM, and THYM. In CHOL and LUSC, VSTM2L expression was positively correlated with immune cell infiltration. Specifically, VSTM2L was positively correlated with B cells (r = .38, p = .0239), macrophages (r = .46, p = .0059), and neutrophils (r = .48, p = .0033) in CHOL and with CD4+ T cells (r = .33, p = 7.78E-14), neutrophils (r = .32, p = 4.54E-13), and dendritic cells (r = .34, p = 5.57E-14) in LUSC. On the contrary, VSTM2L was negatively correlated with CD4+ T cells (r = −.44, p = 3.88E-24), macrophages (r = −.42, p = 2.26E-21), and dendritic cells (r = −.33, p = 7.49E-14) in LGG and with B cells (r = −.36, p = 7.50E-06), CD8+ T cells (r = −.41, p = 1.97E-07), and dendritic cells (r = −.30, p = .0002) in TGCT.
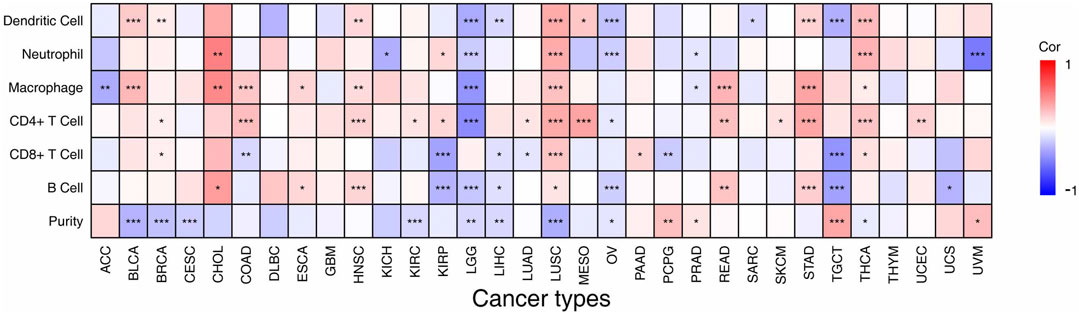
FIGURE 6. Association between VSTM2L expression level and abundance of TIICs in different cancer types in TIMER, shown as a purity-corrected partial Spearman’s rho value. *Padjusted < .05, **Padjusted < .01, ***Padjusted < .001.
We next examined the correlation between the expression of VSTM2L and immunomodulator genes including immunostimulators (Figure 7A, details are shown in Supplementary Figure S4), immunoinhibitors (Figure 7B, details are shown in Supplementary Figure S5), and MHC molecules (Figure 7C, details are shown in Supplementary Figure S6) in 33 tumors from the TISIDB database. VSTM2L expression was positively associated with that of immune checkpoint markers as well as immunostimulators in BLCA, BRCA, ESCA, HNSC, LUSC, STAD, and THCA, whereas a negative association was observed in KIRP and LGG. In particular, there was a complex relationship between VSTM2L and immunostimulators in TGCT (STING1: r = .59, p = 3.70E-14; NT5E: r = .62, p = 1.72E-16; IL6R: r = −.56, p = 5.58E-12), MESO (RAET1E: r = −.63, p = 7.74E-10; VSIR: r = −.60, p = 9.20E-09), and UVM (STING1: r = .73, p = 1.40E-13; ULBP1: r = .55, p = 6.49E-06); and between VSTM2L and immunoinhibitors in CHOL (PDCD1LG2: r = 0.52, p = .0054, ADORA2A: r = .49, p = .0071), TGCT (VTCN1: r = .58, p = 8.33E-14, PVRL2: r = .51, p = 4.03E-10, KDR: r = .60, p = 9.99E-15), THCA (VTCN1: r = .54, p = 9.62E-39), and MESO (TGFB1: r = .50, p = 6.50E-06). Immunoinhibitors that are targets of immunotherapies and were found here to be associated with VSTM2L in various cancers included CD274, also known as programmed death-ligand 1(PD-L1) (KIRP: r = −.38, p = 6.39E-10; TGCT: r = −.42, p = 2.05E-06); cytotoxic T lymphocyte-associated protein 4(CTLA-4) (TGCT: r = −.43, p = 2.97E-07); and PDCD1, also known as programmed death (PD)-1 (TGCT: r = −.42, p = 6.15E-07; UVM: r = .38, p = .0029). We also observed that VSTM2L expression was positively correlated with MHC molecules in BLCA, BRCA, CHOL, HNSC, LUSC, THCA, and UVM, while a negative association was observed in KIRC, LGG, SARC, and TGCT.
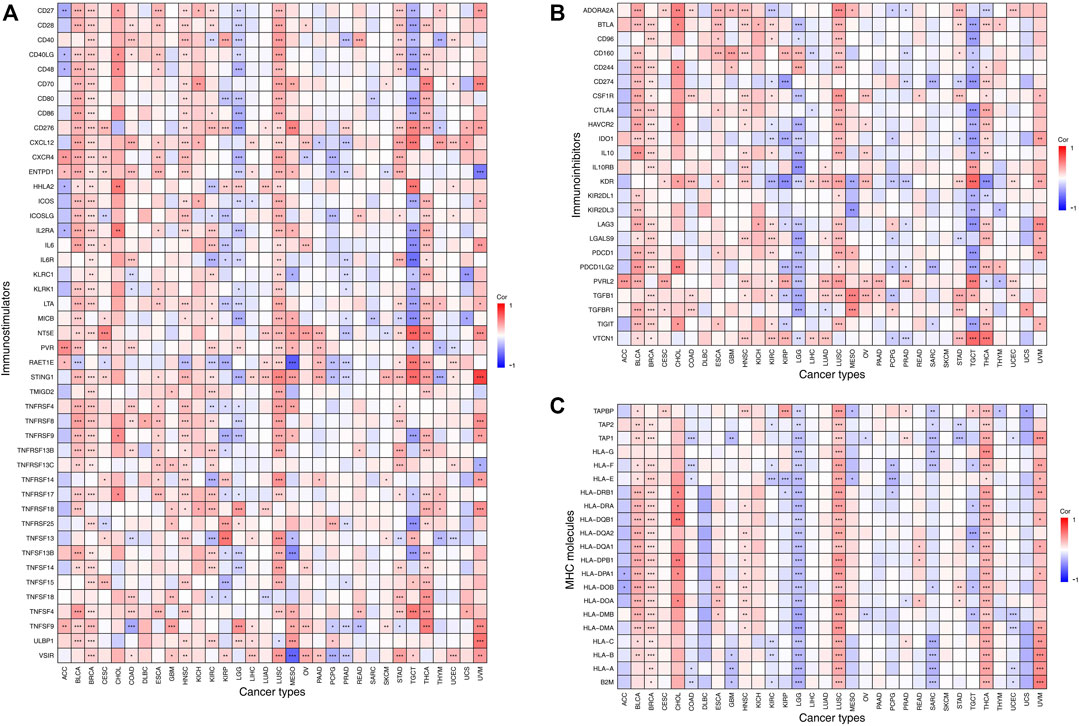
FIGURE 7. Association between expression levels of VSTM2L and immune checkpoint markers in TIMER, which includes immune checkpoint genes from the TISIDB database. Association between the expression of VSTM2L and immunostimulatory factors (A) and immunoinhibitory factors (B) and MHC molecules (C). *Padjusted < .05, **Padjusted < .01, ***Padjusted < .001.
VSTM2L expression in BLCA, BRCA, CHOL, ESCA, KIRC, LUSC, THCA, and UVM was positively correlated with ImmuneScore and StromalScore; and was negatively correlated with these scores in ACC, LGG, PAAD, and PCPG. Additionally, in TGCT, VSTM2L expression was negatively and positively correlated with ImmuneScore and StromalScore, respectively. The three cancer types showing the strongest correlation between VSTM2L expression and ImmuneScore were CHOL (r = .44, p = .0084), LGG (r = −.4, p = 2.20E-16), and LUSC (r = .39, p = 2.20E-16); and the top three cancers related to StromalScore were TGCT (r = .55, p = 2.20E-16), CHOL (r = .4, p = .0158), and BLCA (r = .36, p = 6.44E-14) (Figure 8A, details are shown in Supplementary Figure S7).

FIGURE 8. Correlation between VSTM2L expression level and markers of the TIME and tumor prognosis. (A) Correlation between VSTM2L expression and ImmuneScore and StromalScore. The color and area of each circle represent the degree and direction of correlation, with larger circles corresponding to a stronger correlation and red and blue corresponding to a positive and negative correlation, respectively. (B,C) Correlation between VSTM2L expression level and TMB (B) and MSI (C). *p < .05, **p < .01, ***p < .001.
TMB and MSI are useful prognostic markers and can predict the response to immunotherapy in various cancer types (Dudley et al., 2016; Chan et al., 2019; Franke et al., 2019). We examined the correlation between VSTM2L expression and TMB (Figure 8B, details are shown in Supplementary Figure S8) or MSI (Figure 8C, details are shown in Supplementary Figure S8) across 33 cancers and found that it was positively correlated with both markers in THCA and negatively correlated these markers in COAD, STAD, and UCEC. Significant correlations were also observed between VSTM2L upregulation and decreased TMB in other 10 cancer types (CESC, HNSC, LGG, LIHC, LUSC, READ, SARC, SKCM, THYM, and UCS) and increased TMB in four cancer types (ACC, KIRP, PRAD, and THCA). We also found that MESO and TGCT patients with high VSTM2L expression were more likely to have high MSI.
In order to evaluate the prognostic value of VSTM2L in STAD and KIRP, we examined survival data from the Kaplan–Meier Plotter database in relation to clinicopathologic factors (Table 1) and immune cell profile (Table 2). High VSTM2L expression level was correlated with worse OS in STAD patients who were female (HR = 1.86, p = .034), male (HR = 1.88, p = .0044), stage 3 (HR = 2.22, p = .0102), grade 2 (HR = 2.27, p = .0051), grade 3 (HR = 1.79, p = 0.0176), White (HR = 1.7, p = .0121), or Asian (HR = 3.87, p = .0029) or those with high TMB (HR = 1.89, p = .0083). On the other hand, high VSTM2L expression level was correlated with better OS in KIRP patients who were female (HR = .27, p = .0129), male (HR = .36, p = .0032), stage 3 (HR = .12, p = .0158), White (HR = .44, p = .0153), or Black/African American (HR = .13, p = .0079) or those with low TMB (HR = .21, p = .0002), with stage 1 patients (HR = 3.33, p = .0297) as an exception.
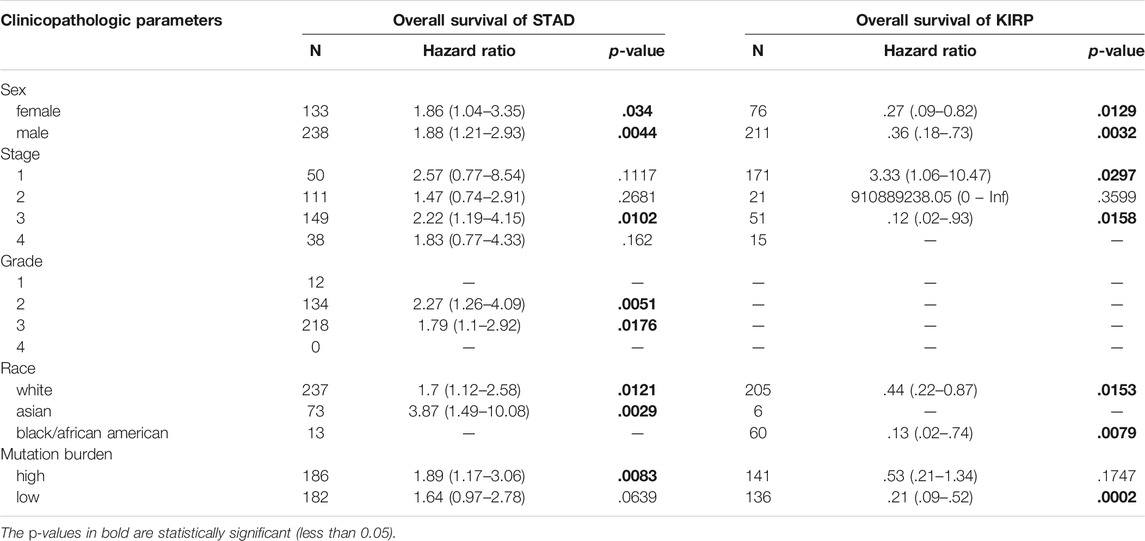
TABLE 1. Correlation of VSTM2L expression and overall survival in STAD and KIRP with different clinicopathologic parameters by Kaplan-Meier plotter.

TABLE 2. Correlation of VSTM2L expression and overall survival in STAD and KIRP according to the immune cell profile using Kaplan-Meier plotter.
VSTM2L expression level was negatively correlated with the OS of STAD patients with basophil enrichment (HR = 5.16, p = .0033) or reduction (HR = 1.51, p = .027); B cell enrichment (HR = 1.79, p = .017) or reduction (HR = 2.13, p = .0031); CD4+ memory T cell enrichment (HR = 1.86, p = .0069); CD8+ T cell reduction (HR = 2.74, p = 5.9E-05); eosinophil enrichment (HR = 1.69, p = .0101) or reduction (HR = 2.55, p = .0033); macrophage enrichment (HR = 1.73, p = .0235) or reduction (HR = 2.11, p = .0039); mesenchymal stem cell enrichment (HR = 1.73, p = .03) or reduction (HR = 2.12, p = .003); natural killer T cell enrichment (HR = 1.97, p = .0066) or reduction (HR = 2.38, p = .0064); regulatory T cell enrichment (HR = 1.99, p = .0006); type 1 T helper cell enrichment (HR = 2.28, p = .0015) or reduction (HR = 1.72, p = .0442); and type 2 T helper cell enrichment (HR = 1.68, p = .0027). VSTM2L expression was also positively correlated with the OS of KIRP patients with basophil enrichment (HR = .18, p = .0003) and reduction (HR = .37, p = .0095); B cell enrichment (HR = .31, p = .0012) and reduction (HR = .22, p = .0021); CD4+ memory T cell enrichment (HR = .11, p = .0088) and reduction (HR = .42, p = .0132); CD8+ T cell reduction (HR = .19, p = 6.7E-06); eosinophil reduction (HR = .09, p = .0034); macrophage enrichment (HR = .28, p = .00015); mesenchymal stem cell enrichment (HR = .25, p = .01) and reduction (HR = .28, p = .0005); natural killer T cell enrichment (HR = .26, p = .0003); regulatory T cell enrichment (HR = .21, p = .00075) and reduction (HR = .3, p = .0013); type 1 T helper cell enrichment (HR = .18, p = .001) and reduction (HR = .33, p = .0012); and type 2 T helper cell reduction (HR = .37, p = .0029).
GEO cohort data were used to identify potential markers of aberrant VSTM2L expression in STAD (GSE84437 dataset) and KIRP (GSE2748 dataset). The Kaplan–Meier survival analysis showed that increased VSTM2L expression was associated with poor OS in STAD and with improved OS in KIRP (Figure 9A). Univariate (Figure 9B) and multivariate (Figure 9C) Cox regression analyses showed that VSTM2L was an independent prognostic marker for survival when compared to clinical characteristics such as primary tumor and lymph nodes status for patients with STAD, and VSTM2L also showed great prognostic potential in KIRP. The time-dependent receiver operating characteristic (ROC) curve analysis of 1-year OS showed that VSTM2L had good predictive performance (area under roc curve, AUC = .692) in these patients (Figure 9D). High VSTM2L expression was also related to advanced tumor status (T2 vs T3, p = .0018; T2 vs. T4, p = .00053) and node status (N0 vs N2, p = .0074; N0 vs. N3, p = .017) in STAD patients irrespective of sex and age. In contrast, in KIRP, high VSTM2L expression was related to early tumor stage, better differentiation, and lower rate of metastasis (Figure 9E).
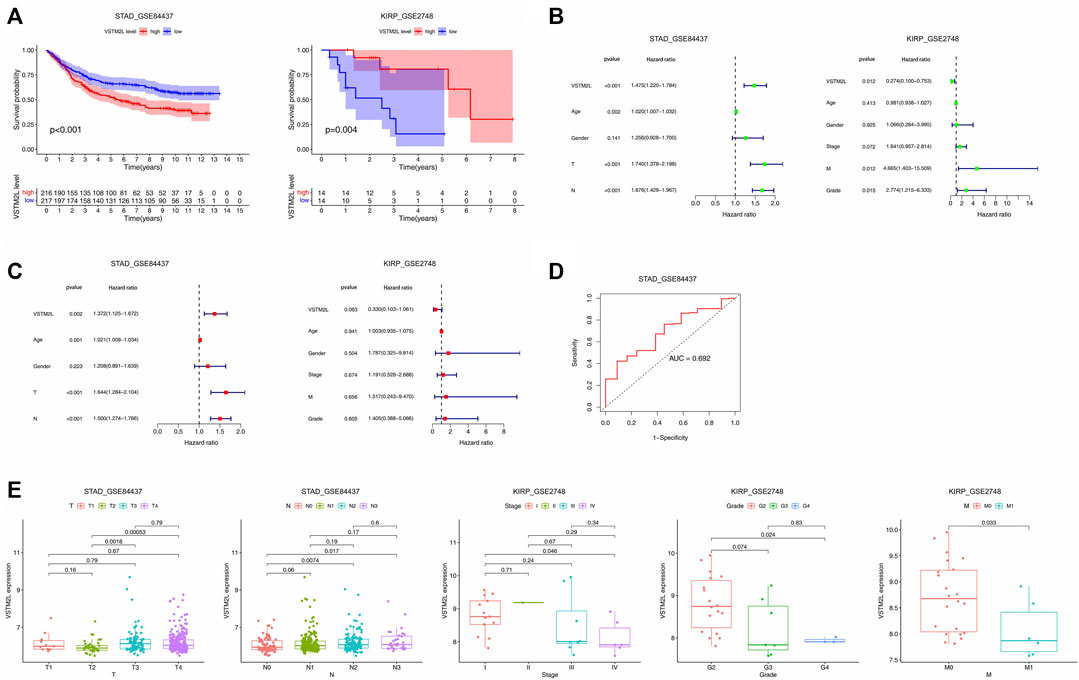
FIGURE 9. Prognostic value of VSTM2L expression in STAD and KIRP. (A) OS of STAD patients in the GSE84437 dataset and of KIRP patients in the GSE2748 dataset. (B,C) Univariate (B) and multivariate (C) Cox regression analyses of VSTM2L expression level and clinical characteristics of STAD and KIRP patients. (D) Time receiver operating characteristic curve (ROC) analysis with area under the ROC curve (AUC) representing the prognostic performance of VSTM2L expression for OS in STAD patients. (E) Relationship between VSTM2L and tumor–node–metastasis stage in STAD and KIRP patients.
The development of immunotherapy has greatly improved the prognosis of multiple cancers. However, its clinical benefits have not been fully confirmed due to a lack of useful markers (Steven et al., 2016; Sugie, 2018; Ganesh et al., 2019; Hegde and Chen, 2020; Walk et al., 2020). In this study, we found that VSTM2L expression differed between tumor and normal tissues and that this was linked to the clinical outcomes in various cancer types. VSTM2L was shown to be highly expressed in the cerebral cortex and pituitary and antagonized the neuroprotective peptide humanin (Rossini et al., 2011); this expression pattern was supported by our data. A higher mortality rate was observed in high-risk CIMP patients with elevated expression of prognostic genes including VSTM2L (Zeng et al., 2020); and in rectal cancer patients receiving preoperative chemoradiotherapy, high VSTM2L expression was correlated with poor therapeutic response and adverse clinical outcomes (Liu et al., 2021). It was previously reported that VSTM2L was expressed at a low level and predicted poor prognosis in gastric cancer and rectal cancer (Hu et al., 2018; Liu et al., 2021), which was confirmed by our observations in STAD. Our results demonstrated that VSTM2L was significantly upregulated in 11 cancers and downregulated in 11 cancers. VSTM2L had different prognostic values across cancers: high expression was related to worse prognosis in BLCA, DLBC, GBM, KIRC, LUSC, MESO, OV, STAD, THCA, THYM, UCEC, and UVM but was associated with a good prognosis in KIRP, LGG, and PCPG. Further study is needed to clarify the reasons for the dual role of VSTM2L in cancers.
As vital components of the TIME, TIICs are a marker for prognosis and response to immunotherapy in multiple malignancies (Hendry et al., 2017; Zhang and Zhang, 2020). Many other markers have been examined for their utility in revealing susceptibility to immunotherapy including MSI, TMB, and immune checkpoint markers (Vrána et al., 2018; Chan et al., 2019; Li et al., 2020). Components of the IL-4 signaling pathway such as ALOX5, EGR1, SPAG1, NCF2, and ATXN1 are known to affect the immune response (Lee et al., 2009; Rajaraman et al., 2009; Siliņa et al., 2011; Li et al., 2019; Didonna et al., 2020; Wei et al., 2020); their regulation by VSTM2L induced chemoradiotherapy resistance in rectal cancer through downstream IL-4 signaling which subsequently affects the progress of cell proliferation and apoptosis (Liu et al., 2021). ALOX5 enhances colorectal cancer cell growth while EGR1 promotes colorectal cancer (Li et al., 2019; Wei et al., 2020; Liu et al., 2021). NCF2 regulates the antiapoptotic role of p53 in cancer cells exhibited by increased apoptosis due to loss of NCF2 and the associated upregulation of ATXN1 during VSTM2L overexpression suggested a link with cancer regulation (Lee et al., 2009; Rajaraman et al., 2009; Siliņa et al., 2011; Didonna et al., 2020). We, therefore, investigated the relationship between aberrant VSTM2L expression and TIICs, immunomodulators, TMB, MSI, and immune and stromal scores in different cancers, which has not been previously reported. Immunostimulators, immunoinhibitors, and MHC molecules with the top three highest correlation scores were STING1, CXCL12, VSIR, KDR, PVRL2, ADORA2A, HLA-DMA, HLA-DOB, and HLA-DPB1. We also found that VSTM2L was positively correlated with TMB in ACC and KIRP and negatively correlated with both TMB and MSI in COAD, STAD, and UCEC. It is worth noting that VSTM2L had prognostic value in high TMB STAD patients and low TMB KIRP patients; the opposite effects may imply that VSTM2L has distinct immunomodulatory functions in these cancers. The correlation between VSTM2L and the immune scores in different cancer suggest that aberrant VSTM2L expression may alter the TIME (Hendry et al., 2017). Thus, the regulation of VSTM2L modulates cancer development and progression. Therefore, the dysregulation of VSTM2L and immune markers among cancer patients suggest a crucial role of VSTM2L in the modulation of TIME (Binnewies et al., 2018; Hinshaw and Shevde, 2019). Taken together, the significant expression of tumor-infiltrating immune cells, immunostimulators, immunoinhibitors, major histocompatibility complex molecules, among others in various cancers point to an association between changes in VSTM2L and disease prognosis and development.
Our results indicate that VSTM2L is a promising independent prognostic factor in STAD and KIRP. A previous study has shown that VSTM2L was downregulated in H. pylori-positive gastric cancer patients compared to patients who were H. pylori-negative in TCGA and was expressed at a lower level in gastric cancer tissue compared to adjacent normal tissue (Hu et al., 2018), which was supported by our work. However, in the same study, the Kaplan-Meier analysis of 72 gastric cancer patients showed no association between VSTM2L expression level and patient survival based on TCGA data, which differed from our findings, possibly because of the different number of available TCGA samples that were analyzed (Hu et al., 2018). A negative correlation was found between VSTM2L and CIMP, and a CIMP-related gene signature comprising six genes (VSTM2L, CST6, SLC7A2, RAB3B, IGFBP1, and EVX2) stratified gastric cancer patients into high‐ and low-risk groups with distinct prognoses (Zeng et al., 2020). We confirmed the different expression patterns of VSTM2L in STAD and KIRP compared to normal tissue using TIMER, TCGA, GTEx, and HPA. Surprisingly, using TCGA, Kaplan–Meier Plotter, and GEO data we found that VSTM2L predicted opposite clinical outcomes in STAD and KIRP. Elevated VSTM2L expression was related to worse prognosis including OS, DSS, DFI, or PFI in STAD; on the contrary, it was positively correlated with longer survival in KIRP. The prognostic role of VSTM2L in KIRP has not been previously reported. We used datasets from multiple databases to perform a pan-cancer analysis of the effects of VSTM2L in cancer, but the main limitation in our study is the lack of experimental data to support our findings.
In summary, we showed that VSTM2L has distinct expression patterns, prognostic value, and relationship with the TIME of different cancers. Kaplan–Meier survival and Cox regression analyses revealed that upregulation of VSTM2L was associated with poor prognosis in STAD and good OS in KIRP. Our results also demonstrate for the first time that aberrant expression of VSTM2L was associated with the TIME including TIICs, immunostimulators, immunoinhibitors, MHC molecules, TMB, MSI, and immune and stromal scores in various cancers. These findings provide a basis for more in-depth investigations of VSTM2L function and interaction with the TIME, and suggest that VSTM2L is a potential target for cancer immunotherapy.
Publicly available datasets were analyzed in this study. This data can be found here: https://xena.ucsc.edu/ https://commonfund.nih.gov/GTex https://portals.broadinstitute.org/ccle https://cistrome.shinyapps.io/timer/ http://cis.hku.hk/TISIDB/index.php http://kmplot.com/analysis/ https://www.ncbi.nlm.nih.gov/geo/.
XY designed and supervised the study. SZ, HX, and JY analyzed the data. SZ wrote the manuscript.
This work was supported by Science and Technology Project of Huizhou City (Grant No. 210809116941981).
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
We sincerely thank all researchers who participated in this study.
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fmolb.2021.792154/full#supplementary-material
Binnewies, M., Roberts, E. W., Kersten, K., Chan, V., Fearon, D. F., Merad, M., et al. (2018). Understanding the Tumor Immune Microenvironment (TIME) for Effective Therapy. Nat. Med. 24 (5), 541–550. doi:10.1038/s41591-018-0014-x
Chan, T. A., Yarchoan, M., Jaffee, E., Swanton, C., Quezada, S. A., Stenzinger, A., et al. (2019). Development of Tumor Mutation burden as an Immunotherapy Biomarker: Utility for the Oncology Clinic. Ann. Oncol. 30 (1), 44–56. doi:10.1093/annonc/mdy495
Didonna, A., Canto Puig, E., Ma, Q., Matsunaga, A., Ho, B., Caillier, S. J., et al. (2020). Ataxin-1 Regulates B Cell Function and the Severity of Autoimmune Experimental Encephalomyelitis. Proc. Natl. Acad. Sci. USA 117 (38), 23742–23750. doi:10.1073/pnas.2003798117
Dudley, J. C., Lin, M.-T., Le, D. T., and Eshleman, J. R. (2016). Microsatellite Instability as a Biomarker for PD-1 Blockade. Clin. Cancer Res. 22 (4), 813–820. doi:10.1158/1078-0432.ccr-15-1678
Franke, A. J., Skelton, W. P., Starr, J. S., Parekh, H., Lee, J. J., Overman, M. J., et al. (2019). Immunotherapy for Colorectal Cancer: A Review of Current and Novel Therapeutic Approaches. J. Natl. Cancer Inst. 111 (11), 1131–1141. doi:10.1093/jnci/djz093
Ganesh, K., Stadler, Z. K., Cercek, A., Mendelsohn, R. B., Shia, J., Segal, N. H., et al. (2019). Immunotherapy in Colorectal Cancer: Rationale, Challenges and Potential. Nat. Rev. Gastroenterol. Hepatol. 16 (6), 361–375. doi:10.1038/s41575-019-0126-x
Hegde, P. S., and Chen, D. S. (2020). Top 10 Challenges in Cancer Immunotherapy. Immunity 52 (1), 17–35. doi:10.1016/j.immuni.2019.12.011
Hendry, S., Salgado, R., Gevaert, T., Russell, P. A., John, T., Thapa, B., et al. (2017). Assessing Tumor-Infiltrating Lymphocytes in Solid Tumors: A Practical Review for Pathologists and Proposal for a Standardized Method from the International Immunooncology Biomarkers Working Group: Part 1: Assessing the Host Immune Response, TILs in Invasive Breast Carcinoma and Ductal Carcinoma In Situ, Metastatic Tumor Deposits and Areas for Further Research. Adv. Anat. Pathol. 24 (5), 235–251. doi:10.1097/pap.0000000000000162
Hinshaw, D. C., and Shevde, L. A. (2019). The Tumor Microenvironment Innately Modulates Cancer Progression. Cancer Res. 79 (18), 4557–4566. doi:10.1158/0008-5472.can-18-3962
Hu, Y., He, C., Liu, J. P., Li, N. S., Peng, C., Yang‐Ou, Y. B., et al. (2018). Analysis of Key Genes and Signaling Pathways Involved inHelicobacter Pylori‐associated Gastric Cancer Based on the Cancer Genome Atlas Database andRNAsequencing Data. Helicobacter 23 (5), e12530. doi:10.1111/hel.12530
Lee, J.-Y., Park, A. K., Lee, K.-M., Park, S. K., Han, S., Han, W., et al. (2009). Candidate Gene Approach Evaluates Association between Innate Immunity Genes and Breast Cancer Risk in Korean Women. Carcinogenesis 30 (9), 1528–1531. doi:10.1093/carcin/bgp084
Lei, X., Lei, Y., Li, J.-K., Du, W.-X., Li, R.-G., Yang, J., et al. (2020). Immune Cells within the Tumor Microenvironment: Biological Functions and Roles in Cancer Immunotherapy. Cancer Lett. 470, 126–133. doi:10.1016/j.canlet.2019.11.009
Li, L., Ameri, A. H., Wang, S., Jansson, K. H., Casey, O. M., Yang, Q., et al. (2019). EGR1 Regulates Angiogenic and Osteoclastogenic Factors in Prostate Cancer and Promotes Metastasis. Oncogene 38 (35), 6241–6255. doi:10.1038/s41388-019-0873-8
Li, L., Feng, Q., and Wang, X. (2020). PreMSIm: An R Package for Predicting Microsatellite Instability from the Expression Profiling of a Gene Panel in Cancer. Comput. Struct. Biotechnol. J. 18, 668–675. doi:10.1016/j.csbj.2020.03.007
Liu, H., Zhang, Z., Zhen, P., and Zhou, M. (2021). High Expression of VSTM2L Induced Resistance to Chemoradiotherapy in Rectal Cancer through Downstream IL-4 Signaling. J. Immunol. Res. 2021, 1–17. doi:10.1155/2021/6657012
Petitprez, F., Meylan, M., de Reyniès, A., Sautès-Fridman, C., and Fridman, W. H. (2020). The Tumor Microenvironment in the Response to Immune Checkpoint Blockade Therapies. Front. Immunol. 11, 784. doi:10.3389/fimmu.2020.00784
Rajaraman, P., Brenner, A. V., Butler, M. A., Wang, S. S., Pfeiffer, R. M., Ruder, A. M., et al. (2009). Common Variation in Genes Related to Innate Immunity and Risk of Adult Glioma. Cancer Epidemiol. Biomarkers Prev. 18 (5), 1651–1658. doi:10.1158/1055-9965.epi-08-1041
Rossini, L., Hashimoto, Y., Suzuki, H., Kurita, M., Gianfriddo, M., Scali, C., et al. (2011). VSTM2L Is a Novel Secreted Antagonist of the Neuroprotective Peptide Humanin. FASEB j. 25 (6), 1983–2000. doi:10.1096/fj.10-163535
Siliņa, K., Zayakin, P., Kalniņa, Z., Ivanova, L., Meistere, I., Endzeliņš, E., et al. (2011). Sperm-associated Antigens as Targets for Cancer Immunotherapy: Expression Pattern And Humoral Immune Response In Cancer Patients. J. Immunother. 34 (1), 28–44. doi:10.1097/CJI.0b013e3181fb64fa
Steven, A., Fisher, S. A., and Robinson, B. W. (2016). Immunotherapy for Lung Cancer. Respirology 21 (5), 821–833. doi:10.1111/resp.12789
Sugie, T. (2018). Immunotherapy for Metastatic Breast Cancer. Chin. Clin. Oncol. 7 (3), 28. doi:10.21037/cco.2018.05.05
Taube, J. M., Galon, J., Sholl, L. M., Rodig, S. J., Cottrell, T. R., Giraldo, N. A., et al. (2018). Implications of the Tumor Immune Microenvironment for Staging and Therapeutics. Mod. Pathol. 31 (2), 214–234. doi:10.1038/modpathol.2017.156
Vrána, D., Matzenauer, M., Neoral, Č., Aujeský, R., Vrba, R., Melichar, B., et al. (2018). From Tumor Immunology to Immunotherapy in Gastric and Esophageal Cancer. Int. J. Mol. Sci. 20 (1), 13. doi:10.3390/ijms20010013
Walk, E. E., Yohe, S. L., Beckman, A., Schade, A., Zutter, M. M., Pfeifer, J., et al. (2020). The Cancer Immunotherapy Biomarker Testing Landscape. Arch. Pathol. Lab. Med. 144 (6), 706–724. doi:10.5858/arpa.2018-0584-CP
Wei, X., Wang, C., Feng, H., Li, B., Jiang, P., Yang, J., et al. (2020). Effects of ALOX5, IL6R and SFTPD Gene Polymorphisms on the Risk of Lung Cancer: A Case-Control Study in China. Int. Immunopharmacology 79, 106155. doi:10.1016/j.intimp.2019.106155
Zeng, Z., Xie, D., and Gong, J. (2020). Genome‐wide Identification of CpG Island Methylator Phenotype Related Gene Signature as a Novel Prognostic Biomarker of Gastric Cancer. PeerJ 8, e9624. doi:10.7717/peerj.9624
Keywords: VSTM2L, prognosis, immune cell infiltration, immunomodulators, pan-cancer
Citation: Zhang S, Xiong H, Yang J and Yuan X (2022) Pan-Cancer Analysis Reveals the Multidimensional Expression and Prognostic and Immunologic Roles of VSTM2L in Cancer. Front. Mol. Biosci. 8:792154. doi: 10.3389/fmolb.2021.792154
Received: 09 November 2021; Accepted: 29 December 2021;
Published: 27 January 2022.
Edited by:
Hem Chandra Jha, Indian Institute of Technology Indore, IndiaReviewed by:
Shajer Manzoor, University of Alabama at Birmingham, United StatesCopyright © 2022 Zhang, Xiong, Yang and Yuan. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Xia Yuan, WVgxMzcxOTY5NDAwNkAxNjMuY29t
†These authors share first authorship
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.