
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
REVIEW article
Front. Microbiol. , 21 February 2025
Sec. Antimicrobials, Resistance and Chemotherapy
Volume 16 - 2025 | https://doi.org/10.3389/fmicb.2025.1496716
This article is part of the Research Topic Global Dissemination and Evolution of Epidemic Multidrug-Resistant Gram-Negative Bacterial Pathogens: Surveillance, Diagnosis and Treatment, Volume III View all 14 articles
 Shaqiu Zhang1,2,3,4†
Shaqiu Zhang1,2,3,4† Jing Yang1†
Jing Yang1† Muhammad Abbas5†
Muhammad Abbas5† Qian Yang1
Qian Yang1 Qianlong Li1
Qianlong Li1 Mafeng Liu1,2,3,4
Mafeng Liu1,2,3,4 Dekang Zhu1,2,3,4
Dekang Zhu1,2,3,4 Mingshu Wang1,2,3,4
Mingshu Wang1,2,3,4 Bin Tian1,2,3,4
Bin Tian1,2,3,4 Anchun Cheng1,2,3,4*
Anchun Cheng1,2,3,4*β-lactam antibiotics are essential medications for treating human diseases. The spread of extended-spectrum β-lactamase-producing Enterobacteriaceae (ESBL-PE) exists globally in multiple reservoirs and the natural environment and poses an immense threat to public health. Plasmid incompatibility groups, such as IncFIA, IncI1, IncY, IncFIB, IncN, IncFIC, IncX4, IncB/O/K/Z, IncHI1/2, and IncA/C, which exist in humans, animals, and the environment, carrying blaCTX-M, blaTEM, and blaSHV genes. The ISEcp1 upstream and orf477 downstream of blaCTX-M genes, as well as other mobile genetic elements (MGEs) such as IS903 and IS26, are involved in capturing and mobilizing antibiotic-resistant genes (ARGs). The blaCTX-M-15 gene is the most common among all discussed reservoirs. The environmental reservoir and propagation mode of ESBL-PE are increasing and difficult to control. The reasons include but are not limited to bacterial adaptability and horizontal gene transfer (HGT) mediated by MGEs and plasmids. Conjugation is a pathway of HGT that is almost uncontrollable. MGEs and plasmids such as Tn3, IS1380 families, IncI1, IncK, and IncN are facilitating HGT of blaCTX-M genes. This review highlights the need to monitor trends in antimicrobial resistance (AMR) in the natural environment. Therefore, policies such as antibiotic management plans, training for healthcare providers and/or patients, cautious use of antibiotics, the need for epidemiological networks, pre-travel consultations, World Health Organization (WHO) infection control and biosafety guidelines, and other intervention measures are considered desirable.
ESBL-PE, first reported in Germany in 1983 (Paterson and Bonomo, 2005), are present globally in humans, animals, and the environment, and have been spreading particularly rapidly in the last two decades (Pitout and Laupland, 2008; Lee et al., 2020). ESBL-PE is a community-level problem in many regions, such as Japan, China, France, Vietnam, and the Netherlands, as well as in low-or-middle-income countries (Castillo-Tokumori et al., 2017; Flateau et al., 2018; Arcilla et al., 2019; Nguyen V. T. et al., 2019; Kwok et al., 2020; Oka et al., 2022). In 2022, the European Antimicrobial Resistance Surveillance Network (EARS-Net) collected AMR data for the eight bacterial species from 30 European countries. Of the eight species, Escherichia coli (E. coli) was shown to be the most prolific, followed by Staphylococcus aureus (S. aureus) and Klebsiella pneumoniae (K. pneumoniae) (European Centre for Disease Prevention and Control, 2020). In the same year, the China Antimicrobial Resistance Surveillance System received surveillance data from 1997 hospitals in China, and E. coli was the most common Gram-negative bacterium, followed by K. pneumoniae and Pseudomonas aeruginosa (P. aeruginosa), and 46.8% of E. coli were resistant to third-generation cephalosporins (Moran et al., 2023). ESBL-PE often affects US inpatients, with approximately 290,220 people affected per year (Gupta et al., 2019).
AMR poses a significant public health threat and is a key issue within the “One Health” approach. The emergence of ESBL-PE, a bacteria found in animals, humans, and the environment, exemplifies this concept (Perestrelo et al., 2023). The term “One Health” emerged in 2003–2004 during the SARS and H5N1 outbreaks, emphasizing the need for interdisciplinary action against diseases (Atlas, 2013). The COVID-19 pandemic underscores the interdependence of human, animal, and environmental health, which is also crucial for addressing AMR. ARGs in these hosts contribute to AMR, posing a pandemic risk (Despotovic et al., 2023). It is vital to integrate human, animal, and environmental health into the “One Health” framework to effectively tackle the growing challenge of AMR (Muloi et al., 2023).
We flagged E. coli as one of the main threats to public health and environmental safety, and its contribution to the spread of ARGs is linked with integrons, transposons, and transmissible plasmids (Zhang et al., 2020). Plasmids, as the core of ARG transmission within the “One Health” framework, are the foundation for the spread of antibiotic resistance among bacteria and within and between habitats (Castañeda-Barba et al., 2024). Plasmid-mediated colistin, carbapenem, and tigecycline resistance genes, accompanied by ESBL genes, represent a potential public health concern (Clemente et al., 2019; Yang et al., 2023). Further, we have recently reviewed the co-existence of ESBL genes with resistance to colistin and carbapenem (Zhang et al., 2021). The spread of ESBL genes in natural environments and in humans is a burning issue, because plasmid-mediated HGT transforms non-resistant bacterium into resistant ones (Kawamura et al., 2017). Insertion sequences (IS) such as ISEcp1, IS903, IS3, IS1380 families, and orf477 have been linked with the genetic environment of ESBL genes (Lifshitz-Ziv et al., 2018; Valizadeh et al., 2021; Awosile and Agbaje, 2021; Sultan et al., 2022). Furthermore, MGEs and plasmids like Tn3, IncI1, IncK, IncFII, and IncN are enhancing the spread of these genes in ESBL-PE (Maamar et al., 2016; Lifshitz-Ziv et al., 2018; Sultan et al., 2020; Berbers et al., 2023). Routes of transmission of ESBL-PE include but are not limited to hospital infections (Artero et al., 2017), fecal carriage (Tamta et al., 2020), travelers (Nakayama et al., 2019), agriculture (Richter et al., 2020), wastewaters (Li et al., 2016), poultry production (Tansawai et al., 2019), and wild animals (Fuentes-Castillo et al., 2023). Researchers recognize the ESBL-PE spread pathways are associated with direct contact with animals, a poor living environment, and handling of contaminated meat/carcasses. In addition, flows out of hospitals, wastewater treatment plants (WWTPs) in agricultural fields, and the natural environment represent dangers.
As depicted in Figure 1, ESBL genes evolve and spread dramatically in environmental settings through transmissible plasmids, transposons, and MGEs, compromising safety and public health. This framework is concluded during this organized review, and the objective was to identify available evidence related to the capture, transmission, and expression mechanisms of ESBL genes and associated risk factors (RFs), plasmids, or MGE-mediated HGT among bacteria that affect animals, humans, and the natural environment. People from developed areas when traveling to underdeveloped infected communities can enhance the spread of ESBL-PE to healthy people and the environment. Lastly, the preventive control and restrictive measures for the ESBL-PE are summarized.
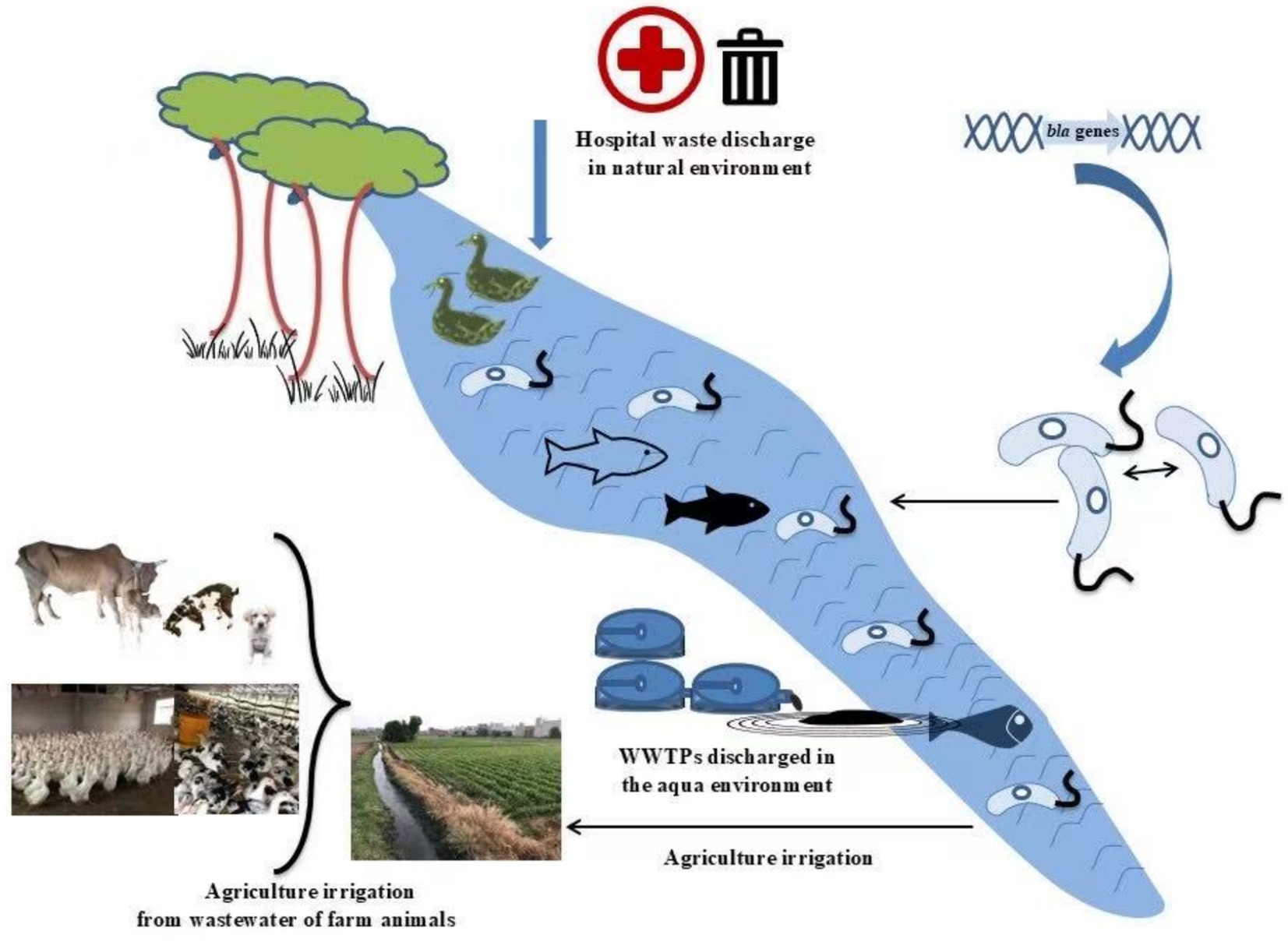
Figure 1. The illustration illustrates the release of ESBL-PE into the environment via hospital waste and WWTPs and their transmission between humans, animals, and the natural environment. These bacteria carry ARGs that may be passed between different hosts via plasmids and MGEs. The figure also shows how agricultural irrigation may facilitate the spread of these ARGs. This process highlights the complexity of transmission across species and environments and emphasizes the importance of implementing “One Health” strategies to monitor and control antibiotic resistance.
Published documents were accessed through databases such as PubMed, Science Direct, and Google Scholar. Keywords included, but were not limited to, ESBLs, Enterobacteriaceae, risk factors (RFs), risk, infection, travelers, healthcare workers (HCWs), illness, disease, farmworkers (FWs), environment, health care professionals, blaTEM, blaSHV, blaCTX-M, HGT, MGEs, Tn3, ISEcp1, antibiotic stewardship programs (ASPs), and occupational health. References cited were searched for additional related studies. The timeframe of the search was limited to English-language literature up to December 31, 2024, in order to ensure that the study incorporated the most recent research findings in the field.
The spread of ESBL-PE in medical settings has become a major public health issue in recent times (Table 1) (Karlowsky et al., 2022). High diversity in plasmidome in clinical isolates further complicates the situation (Neffe et al., 2022). For example, in Portugal, healthcare students were shown to carry blaCTX-M-1, as well as blaCTX-M-15 and blaCTX-M-8, and most of them are in IncFIA/FIB-type plasmids (Fournier et al., 2020). In Indonesia, blaCTX-M, blaTEM, and blaSHV, and primarily blaCTX-M-15, are circulating among medical students, located on different chromosome regions and plasmids (Rosantia et al., 2020). In Taiwan, blaSHV and blaCTX-M are predominantly prevalent and associated with IS elements as well as IncA/C-type plasmids (Chao et al., 2023). This may be due to the polluted environment in medical universities or communities.
Microbial aerosols contain a number of genes, namely blaTEM, followed by blaCTX-M and blaSHV, in the hospital environment (Wu B. et al., 2020). Interestingly, blaTEM is the most common and blaCTX-M gene is emerging in Enterobacteriaceae in hospitals (Jena et al., 2018; Perera et al., 2022). On environmental surfaces of hospital settings such as in urology, intensive care units, or orthopedic and surgical departments, ISEcp1 has been observed largely upstream of blaCTX-M-1, along with IS26 in two orientations and downstream of orf477 (Figure 2D) (Valizadeh et al., 2021). Isolates from outpatient department, wards, intensive care unit, cabins, and neonatal intensive care units carry the three ESBL genes (Jena et al., 2017). Upon discharge from hospital wards, patients colonize the blaCTX-M-15 gene at a higher level (Moremi et al., 2018; Díaz-Agero Pérez et al., 2019; Riccio et al., 2021). In addition, a statistically significant relationship between hospital antibiotic use and the prevalence of blaCTX-M-9, blaCTX-M-3, and blaSHV-5 was observed in hospital effluents (Korzeniewska and Harnisz, 2013a).
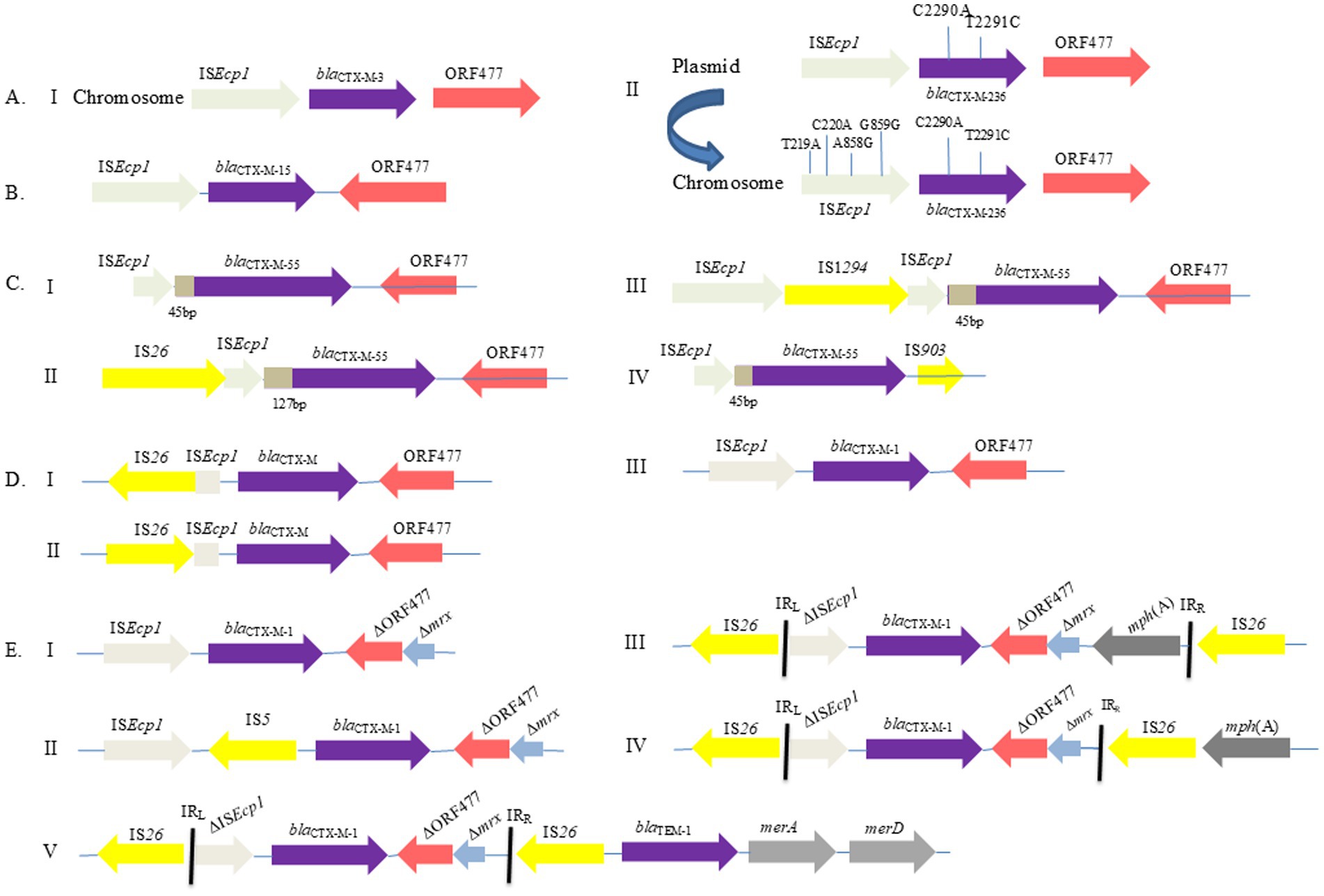
Figure 2. Illustration of the recently reported blaCTX-M genetic environment with transposition of ISEcp1 identified as a means of dissemination in different reservoirs. (A) Genetic environment of a newly identified variant of blaCTX-M-3 (blaCTX-M-236) gene; positioned on an IncM2-type plasmid and chromosome with identified allele and loci (Huang et al., 2021). (B) Genetic context of blaCTX-M-15 gene from the urban aquatic environment in India (Singh et al., 2018). (C) Four types of clinically oriented genetic environments noted for blaCTX-M-55 (Hu et al., 2018). (D) From the hospital environment, three patterns were noted, blaCTX-M genetic environment with IS elements (Valizadeh et al., 2021). (E) Schematic presentation of flanking regions of blaCTX-M-1 genes in plasmids prevalent in healthy humans and food animals (Wang et al., 2015).
Rehabilitation units in Israel, France, Italy, and Spain revealed that family members (FMs) were more at risk of ESBL-PE acquisition from their family patients than HCWs (Adler et al., 2014; March et al., 2017), which calls for ASPs and infection management strategies.
Approximately 150 million urinary tract infections (UTIs) are diagnosed worldwide annually (Flores-Mireles et al., 2015). A 2017 report showed that the WHO considered E. coli to be the main strain causing community- and hospital-acquired UTIs (Ala-Jaakkola et al., 2022). UTIs acquire nosocomial infections, which are associated with ESBL-PE. For instance, UTI-related major RFs include previous use of antibiotics, invasive devices, indwelling urinary catheters, prior hospitalization, and a history of recurrent UTI (Castillo-Tokumori et al., 2017; Alqasim et al., 2018; Al-Jamei et al., 2019; Goyal et al., 2019). From bloodstream infection of older patients, the blaTEM-1 gene was the most prevalent, followed by blaCTX-M-14, blaCTX-M-27, and blaCTX-M-65 genes located on the incompatible plasmid group, which includes FrepB, FIA, FIC, K types (Xiao et al., 2019). The extensive and uncontrolled usage of β-lactam antibiotics led to the emergence of drug resistance, because of this, the selection of antibiotics should consider antimicrobial activity, pharmacokinetics, and information on causal agents (Hashemizadeh et al., 2019).
It is important to underline the role of ISEcp1 in the dissemination of ARGs. Recently, using upstream genetic structures of ISEcp1-blaCTX-M, plasmids have been shown to contain blaCTX-M transposition units. This information is useful to understand the location of ISEcp1 and classification of bacterial isolates harboring blaCTX-M (Yagi et al., 2021). A clinical bone biopsy revealed a chromosomal and IncM2-type plasmid harbored blaCTX-M-236 gene, and the chromosomally located blaCTX-M-3 gene was found with ISEcp1 upstream and orf477 downstream in all copies of Enterobacter hormaechei (Figure 2A) (Huang et al., 2021). In blood samples of inpatients, the blaCTX-M-15 gene exists downstream of ISEcp1 while blaKPC-3 was between ISKpn7 and ISKpn6; this was also the case with other transposons such as Tn4401a, tnpR, and tnpA in K. pneumoniae (Piccirilli et al., 2020). In infant blood, blaCTX-M-14, and blaCTX-M-64 were carried by ISEcp1-mediated transposons Tn6503a and Tn6502 (Yang et al., 2023). Likewise, in sputum samples, the blaCTX-M-24 gene occurred with an ISEcp1-type element and carried IncFII plasmid (Yang et al., 2020). Further, a patient’s wound and fecal isolates in Switzerland was shown to exhibit K. pneumoniae ST23 and E. coli ST1 with the IncFII plasmid harboring blaCTX-M-14, which was inserted between ISEcp1 upstream and truncated IS903B downstream (Xavier et al., 2019). Transposition of ISEcp1 and other MGEs such as Tn2, Tn3, and IS26 were observed with blaCTX-M-15 in the IncF- and IncI1-type plasmids (Zong et al., 2015), and the acquisition of these plasmids can make treatment options for resistant infections more difficult (Xavier et al., 2019). The plasmid- and ISEcp1-mediated transmission of blaCTX-M genes plays a significant role in the continued dissemination of ESBL resistance among E. coli populations that have the capacity to enter hospitals. Other RFs include hosts and sites of infection, such as infants infected with ESBL-PE (Marando et al., 2018) and the rectovaginal colonization of maternal ESBL-PE (Neemann et al., 2020). The ESBL-PE clones exist on newborns and on the hands of HCWs, which indicates a possible route of transmission among them (Dashti et al., 2010), as well as cross-contamination of environmental ESBL-PE (Boyer et al., 2015). The risk of ESBL-PE is higher in patients colonized with K. pneumoniae as compared to E. coli, and the presence of underlying diseases, such as malignant tumors, congestive heart failure, severe liver diseases, and peptic ulcers, can increase the risk further (Denkel et al., 2020). The risk of acquiring ESBL-producing E. coli (ESBL-EC) is higher in patients of an advanced age and with co-morbid medical conditions. Women are more at risk than men, and urban residents are at higher risk than rural residents (Laupland et al., 2008). Person-to-person communication (Ewers et al., 2012), mobile phones of HCWs (Loyola et al., 2016), ESBL-PE nasal colonization with hands (Founou et al., 2019), and feces in health care settings (NGuyen T. T. H. et al., 2019) are vital channels of AMR distribution.
The distribution of antibiotic-resistant bacteria (ARB) changes and increases outside clinical settings. The prevalence of ESBL genes and plasmids in livestock, FWs, and farm environments are summarized in Table 2. Food animals are an essential part of our diet because they cover our protein requirements. From food animals and farms, ESBL-PE have been well documented in several countries, such as the United Kingdom, Germany, the Netherlands, Lebanon, Malaysia, and Thailand (Randall et al., 2011; Huijbers et al., 2014; Fischer et al., 2014; Gonggrijp et al., 2016; Tansawai et al., 2019; Seenama et al., 2019; Mobasseri et al., 2019; Velasova et al., 2019). Studies in China have been conducted on poultry (Liao et al., 2013), waterfowl (Yang et al., 2019; Zhang et al., 2023a; Zhang et al., 2024), pigs (Tan et al., 2023), and free-range Tibetan yaks, where ESBL genes were significantly associated with other ARGs (Rehman et al., 2017; Lei et al., 2018). In the USA and Spain, environmental flies in dairy, beef, and poultry farms have been shown to harbor ESBL genes (Solà-Ginés et al., 2015; Poudel et al., 2019). Close contact with livestock and introduction of new animals to the farm can impact the transmission and prevalence of ESBL-PE with co-localization of ARGs (Velasova et al., 2019). In addition, animals in the low- and middle-income countries are reared in poor bio-containment circumstances and with limited or no control over the use of antimicrobials (Carrique-Mas et al., 2015). Therefore, the poor biosecurity and farm environmental contamination can be strong reasons for ESBL-PE colonization in poultry, cattle, and pig farms (Hiroi et al., 2012; Dandachi et al., 2018; Gay et al., 2018).
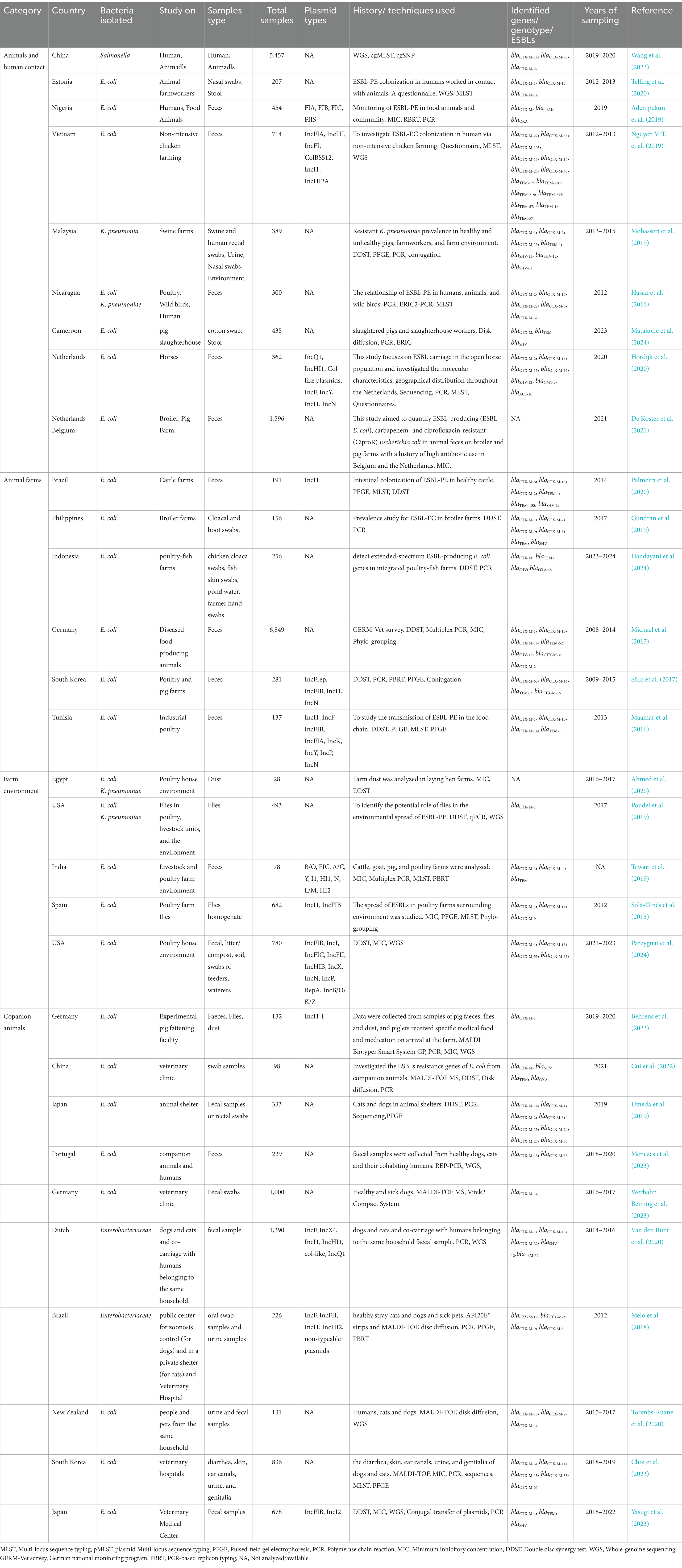
Table 2. Selected studies from animal farms and human and environmental settings reporting ESBL-PE in various countries.
ISEcp1 is often present upstream of the blaCTX-M genes while IS903 is in the downstream region in several replicon types, such as IncF, IncI1-Iγ, IncFIB, IncN, and IncP, which play a vital role in the capture, mobilization, and expression of blaCTX-M genes (Tamang et al., 2013; Ben Said et al., 2016). For instance, Salmonella enterica serovar Virchow isolated from carcasses and fecal samples of chickens (n = 256), pigs (n = 7), and cattle (n = 2) in South Korea exhibited the IncHI2 conjugative plasmid harboring the blaCTX-M-15 gene with transposition of upstream ISEcp1 and downstream orf477 (Na et al., 2020). Although the isolate rate is low, E. coli ST405 contains IncFII-type plasmid harboring blaCTX-M-14 gene with transposition of upstream ISEcp1 and downstream truncated IS903, which was isolated from pasture-based dairy farms in New Zealand (Burgess et al., 2021).
Specifically, from the perspective of poultry, air-borne dust affects the health status of laying hens (Gole et al., 2014). Meanwhile, ESBL-PE were recorded in the air-deposited dust on eggs in Egyptian hen coops (Ahmed et al., 2020). A low level of vertical transmission of ESBL genes has been reported from hatchery chicks, with a history of ESBL genes in parent poultry flocks (Projahn et al., 2017). Backyard poultry are involved in the dissemination of ESBL genes in poultry farms, FWs, and environmental settings by IncF-, IncY-, IncX-, IncHI1-, IncFIA-, and IncI1-type plasmids (Tansawai et al., 2019). Due to direct contact with broiler chickens, a higher level of ESBL-EC prevalence was recorded among the farm-based individuals compared to community living around organic Dutch broiler farms (Huijbers et al., 2015). Identical genes, such as blaCTX-M-65, and blaCTX-M-55, have been observed in non-intensive chicken farms and human beings. By whole-genome sequencing (WGS), 29 plasmid incompatibility groups of those 486 ESBL-EC isolates were noted, with IncFIA, IncFI, IncFII and ColBS512 predominant in human isolates, while Col156, IncQ1, and IncHI2A were predominant in chicken isolates (Nguyen V. T. et al., 2019). A recent study on a poultry farm showed the blaCTX-M-1 gene in IncI1/ST3 plasmids and/or chromosomes, which harbored an ISEcp1 element in the upstream region of this gene (Aldea et al., 2022). Thus, the plasmid-mediated transmission of ESBL genes plays an important role in AMR spread in the poultry production system and related environmental settings (Projahn et al., 2017; Oikarainen et al., 2019).
Plasmid-mediated ESBL genes transmission has been reported in pigs and pig FWs in several countries such as Denmark, Germany, Switzerland, and the Netherlands (Hammerum et al., 2014; Dohmen et al., 2015; Dahms et al., 2015; Kraemer et al., 2017). The fecal ARG load is higher in pig FWs compared to broiler FWs and livestock. For instance, blaCTX-M-1, blaCTX-M-14, and blaSHV-12 were observed in both humans and pigs on the same farms, while blaCTX-M-1 was detected on IncI1-, IncF-, or IncN-type plasmids (Hammerum et al., 2014). In a recent study, high clonal correlation of ESBL-EC strains was also observed in pig and human isolates from slaughterhouses (Matakone et al., 2024). The CTX-M-1 group is present in the piggery, air, and dust, which highlights the possibility of transmission between animals and humans after direct contact (Dohmen et al., 2017). Human and porcine isolates contained identical blaCTX-M-1 genes in E. coli ST453 with IncI1-type plasmid, showing the potential for clonal transmission (Dohmen et al., 2015). The rectal carriage of ESBL-PE occurred in pig-FWs, as in the general population of Germany. Isolation of clonally identical ST10/CTX-M-1 in FWs and the farm environment suggested the possibility of cross-transmission (Fischer et al., 2017). A study in Brandenburg, Germany, found that ESBL-EC isolated from flies showed genetic similarities to swine fecal isolates, indicating the fly-associated transport of diverse ESBL-EC from the pig farm into urban habitation areas (Behrens et al., 2023). Therefore, pigs and farmers can contribute to the spread of ESBL-PE, which can lead to the spillover of ARB to the community.
It is important to monitor the distribution of ESBL-PE through the food chain. Reports have shown the occurrences of ESBL-PE in poultry, swine, bovine, and fishes (Table 3). However, sequence data shows that ESBL-EC and K. pneumoniae isolates from human-polluted environments are genetically dissimilar from those polluting food items (Martak et al., 2022). The ESBL-PE with the highest prevalence of blaTEM, followed by blaCTX-M and blaSHV, was found in a variety of retail foods including frozen chicken, raw chicken meat, and pork. (Ye et al., 2018). For instance, 11 STs of E. coli carrying blaCTX-M-1, blaCTX-M-14, blaCTX-M-15, and blaTEM-1 genes were identified in fresh pork meat in Germany (Schill et al., 2017). The identification of E. coli isolates carrying blaCTX-M-8, blaCTX-M-14, blaCTX-M-15, and blaCTX-M-55 in retail meat products from supermarkets across the United Arab Emirates highlights a significant prevalence of these ARBs (Habib et al., 2023). Notably, one-day old chicks imported from Brazil to Uruguay were shown to act as the “Trojan horse,” allowing the latent spread of E. coli carrying mcr-9, blaCTX-M-2, blaCTX-M-8, blaCTX-M-15, blaCTX-M-55, and blaCMY-2 in poultry farms (Coppola et al., 2022). The E. coli ST48 isolated from camels was harboring blaCTX-M-15 in Kenya, which can threaten public health through direct contact or via the food chain (Nüesch-Inderbinen et al., 2020).
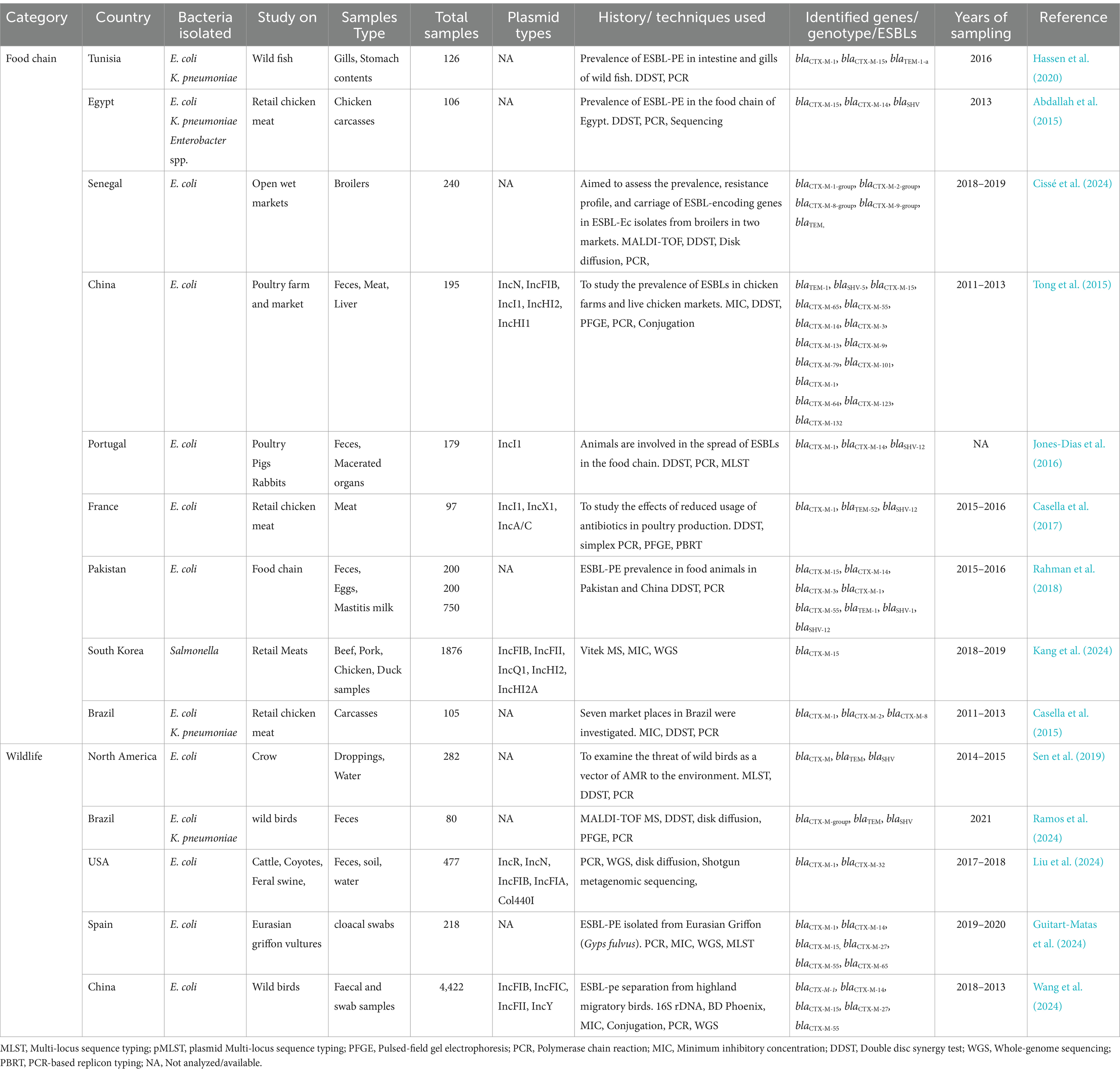
Table 3. Selected studies from food chain and wildlife environmental settings reporting ESBL-PE in various countries.
Since poultry is a common and affordable source of protein, the screening of industrial poultry production systems for ESBL-PE is essential (Maamar et al., 2016). For example, Finland stopped using antimicrobials in the broiler production system in 2009 (Päivärinta et al., 2016). Although at lower rates than other European countries, the various flocks of broiler chickens without antimicrobial usage also revealed seven ESBL-EC STs in Finland, harboring a range of plasmid incompatibility groups, such as IncB/O/K/Z, IncI1, IncFII, IncFIB, IncII, IncX1, and IncFIC (Päivärinta et al., 2020). Brazil is the third largest chicken meat producer and also has a huge local consumer market (Alves et al., 2012; Palmeira et al., 2020). In Brazil, the chicken meat acts as a reservoir of blaCTX-M-2, harboring a complex-class 1 integron, with ISCR1-linked sequence to dfr and/or aadA gene cassettes, which could be a community health hazard (Casella et al., 2015).
Beef and veal are another important food protein source. Worryingly, 68.2% of tested beef farms in Brazil were positive for ESBL-EC, with a high prevalence of blaCTX-M-8 carried by IncI1/pST113-type plasmids linked to ST648 and ST155 clones (Palmeira et al., 2020). Surprisingly, environmental contamination has been verified from sludge cultures on Dutch dairy farms, with seven different ESBL genes (Gonggrijp et al., 2016). High numbers of ESBL genes such as blaTEM-1, which poses a direct threat to the health of dairy cows and further affects the quality of dairy products, were observed in the bedding material (Yang et al., 2021), Finally, the presence of ISEcp1 with blaCTX-M in Indian dairy isolates highlights the potential role of the ISEcp1-like element in the spread of this particular gene into the food chain (Koovapra et al., 2016).
There are multifactorial reasons for the rapid spread of ESBL-PE, such as migration and human travel (Arcilla et al., 2017; Doi et al., 2017) and imported animal meat products (Kim et al., 2018). However, travel restrictions alone may not influence or limit the spread of ESBL genes (Howard-Jones et al., 2022). We have limited knowledge on the colonization of ESBL-PE through travel among different countries concerning humans and animals. For instance, imported meat in South Korea revealed a high prevalence of E. coli in poultry as well as pork and beef. blaCTX-M-94 has been reported in Brazilian and US poultry meats, blaCTX-M-58 was reported in pork meat in France, blaCTX-M-79, blaCTX-M-1 in Hungary, blaCTX-M-14/18 in Spain, blaCTX-M-1 and blaCTX-M-2 in Chile on multiple plasmid incompatibility groups, such as FIA, FIB, FIC, Frep, FIIA, HI1, and HI2 (Kim et al., 2018). In Ghana, E. coli ST10, ST38, and ST155 clones carrying blaCTX-M-15 and blaCTX-M-1 and K. pneumoniae ST2570, ST147, and ST15 clones carrying blaCTX-M-15 and blaCTX-M-14 have been identified in imported poultry meat from Belgium, Germany, Brazil, the USA, Poland, and the Netherlands (Eibach et al., 2018). Legal and illegal poultry/pig meat and matchmaking products imported into Germany from countries such as Brazil, Chile, and Egypt have shown evidence of E. coli ST101, ST117, ST7509, ST7845, and ST7602 clones primarily containing blaCTX-M-2 along with blaCTX-M-8, blaCTX-M-1, and blaCTX-M-9 (Müller et al., 2018). In sub-Saharan Africa, up to 54% of poultry meat from Europe and Brazil is regularly contaminated with ESBL-EC (Olaru et al., 2023). Further, from imported food products in the USA, S. enterica was isolated in frozen octopus carrying blaTEM-1-positve plasmid with IncN type, whereas blaTEM-1 and blaCTX-M-9 were harbored on IncI1 plasmid in frozen tilapia fish (Bae et al., 2015).
Frequent trips between countries increases the spread of ESBL-PE (Table 4). International travel, the presence of gastrointestinal comorbidity, and prior UTIs are important in the acquisition of ESBL-PE (Strysko et al., 2016; Woerther et al., 2017; Wuerz et al., 2020). Travelers who visit relatives and friends are at lower risk of acquiring ESBL-PE (Schaumburg et al., 2019). Moreover, antibiotic therapy during travel (Angelin et al., 2015; Wuerz et al., 2020), travel diarrhea (López-Vélez et al., 2022), UTI during travel (Patjas and Kantele, 2022), and type of accommodation (Meurs et al., 2019) are important factors in ESBL-PE colonization. Reports confirmed colonization of blaCTX-M-15, blaCTX-M-3, blaCTX-M-55, blaCTX-M-14, and blaCTX-M-27 after traveling to Sub-Saharan Africa, North Africa, the Middle East, South Asia, South America, Central Asia, Central America, and Oceania from Japan (Mizuno et al., 2016; Nakayama et al., 2019). One study showed that 75% of AMR plasmids found in the feces of travelers with a history of travel to Laos contained ESBL genes (Snaith et al., 2023). Research showed German travelers to India received the highest acquisition of blaCTX-M-15, while travelers to South-East Asia were colonized with blaCTX-M-27, and Southern Africa and South America travelers acquired blaCTX-M-15 (Lübbert et al., 2015). Similarly, in another study from Germany, 21% of travelers to East Africa, 23% of travelers to Southeast Asia, and 42% to South Asia returned with ESBL-PE (Meurs et al., 2019). The UK’s findings supported the argument that travelers with a trip history to Asia or Africa carried blaCTX-M (Bevan et al., 2021; Otter et al., 2019). In Dutch travelers, higher colonization of blaCTX-M-15 and blaCTX-M-14/18 was recorded before and after travel to Asia, Africa, and Oceania (Arcilla et al., 2019). Finnish travelers with a travel history outside the Nordic countries were found to be predominantly positive with blaCTX-M-15 followed by blaCTX-M-1, blaCTX-M-9, blaTEM, and blaSHV (Kantele and Lääveri, 2021). The high rates of acquisition of the intestinal blaCTX-M-15, blaCTX-M-1, blaCTX-M-14, blaCTX-M-27, and blaCTX-M-65 carried by the IncF-type plasmids were observed in French soldiers who served in Afghanistan for 4–6 months without inter-individual transmission (Maataoui et al., 2019). Consequently, ongoing reporting and monitoring of ESBL-PE among travelers is required to mitigate the emerging challenge of AMR.
About 20 million hectares of arable land are irrigated with wastewater globally (Mateo-Sagasta et al., 2013); this wastewater comes from private use, slaughterhouses, animal farms, and hospital outflows that contain levels of ARGs (Dickin et al., 2016; Azuma et al., 2022). Thus, soils irrigated with untreated wastewater act as a reservoir for ARGs (Bougnom et al., 2020). For example, the occurrence of blaCTX-M-3, blaCTX-M-12, blaCTX-M-14, and blaCTX-M-15, as well as blaTEM and blaSHV, on spinach farms was alarming where river water was used to irrigate land (Richter et al., 2020). Wastewater from dairy farms in agricultural fields implies the spread of the bla genes in environmental settings. The blaTEM-1 gene was recorded in dairy waste even after the treatment of solid manure and dairy sewage (Yang et al., 2021).
Water from estuaries, wells, and springs used for human activities, watering animals, and crop irrigation without treatment is a source of CTX-M-15-producing E. coli ST10, ST38, and ST131 clones and Klebsiella isolates ST323, ST17, and ST405, which proposes a complex origin of contamination (Diab et al., 2018; Hooban et al., 2022). In addition, freshwater environments are vulnerable to antibiotic pollution from a variety of sources, including sewage, agricultural runoff, and farm leakage, creating an ideal environment for the colonization, multiplication, and spread of ARB and facilitating their accumulation, evolution, and spread (Cho et al., 2023). The level of ARB is highest in the river systems, followed by lakes, dams, ponds, and spring water (Nnadozie and Odume, 2019). The aquatic environment that receives wastewater in the Kashmir Valley, India, has shown coexistence of blaTEM and blaCTX-M-15 with MGEs such as ISEcp1, Tn21, and Tn3, associated with plasmid incompatibility groups such as B/O, HI1, HI2, N, I1, FIB, and FIA (Sultan et al., 2020; Sultan et al., 2022).
WWTP includes primary and secondary procedures to remove organic matter, pollutants, and microbes. However, these practices have some limitations in eradicating fecal coliforms and ARGs (Li et al., 2016; Smyth et al., 2020). Municipal-treated wastewater can be a potential source of MDR E. coli and ARGs in surface waters (Kutilova et al., 2021). ESBL genes, such as blaTEM, blaSHV-12, blaCTX-M-1, and blaCTX-M-15 carried by IncF-, IncI1-, IncHI1/2- and IncA/C-type plasmids, were identified in urban WWTP effluents (Smyth et al., 2020). The chloroform treatment process at WWTP reduces the number of E. coli, but there is still some ESBL-EC released into the environment through the effluent, the most prominent gene being blaCTX-M-9 (Xie et al., 2023). A wastewater treatment plant in Colombia receives domestic, hospital, and industrial wastewater and releases effluent into the natural water source. Samples of raw influent, recycled sludge, aeration tanks, and final effluents revealed diverse ESBL-EC clones with a high prevalence of blaTEM, followed by blaCTX-M-1, blaCTX-M-9, blaCTX-M-8/25, and blaCTX-M-2 (Aristizábal-Hoyos et al., 2019). UV disinfection is one of the practices used to eliminate ARB in WWTPs. UV treatment also reduced ARB in effluents (Silva I. et al., 2018). However, different genetic arrangements such as integrase gene, variants of blaCTX-M genes like blaCTX-M-15, blaCTX-M-32, blaCTX-M-1, blaCTX-M-27, and blaCTX-M-14 flanked by ISEcp1/IS26, IS903, and orf477, have been observed in UV-treated effluent (Silva I. et al., 2018). Weather can also affect the presence of ESBL-PE in WWTP effluents, as greater quantities were observed in colder months than in warmer months (Li et al., 2023). A range of bla genes were identified from wastewater, sewage sludge, and WWTP effluents including blaVIM, blaGES, blaCTX-M-9, and blaCTX-M-1 (Schages et al., 2020). Despite the treatment, WWTP effluents contained bacteria that could contaminate river water and air nearby grit tanks or bioreactors, primarily blaCTX-M-3, blaCTX-M-9, and blaCTX-M-1 followed by blaTEM-49, and blaSHV-2 genes (Korzeniewska and Harnisz, 2013b). Improved wastewater treatment is therefore needed to effectively monitor and control the spread of priority MDR pathogens and ESBL-PC in the population (Puljko et al., 2024).
Worrisomely, ARGs were abundant in untreated hospital wastewater, municipal WWTPs, and river sediments (Yuan et al., 2021). Untreated wastewater effluents from the sick bay contained blaCTXM-1, blaCTXM-2, blaCTXM-9, blaCTXM-8, blaCTXM-25, and blaTEM, which highlights the need for hospital-based wastewater treatment facilities (Adekanmbi et al., 2020). Moreover, a predominance of blaTEM, followed by blaSHV and blaCTX-M genes, have been noted from surface river water sites closer to human anthropogenic actions, such as hospital, industrial, and municipal wastes, from Ganga, India (Chaturvedi et al., 2020). Therefore, monitoring of AMR in WWTP effluents is necessary to understand the load of fecal coliforms, ARGs, MGEs, and ARB into the natural environment. This will strengthen our ability to decide on adding more treatment technologies to reduce the influx of ARGs to aquatic environments (Smyth et al., 2020).
Companion animals play a major role in the lives of many people and allow the transmission of ARB between people and animals (Dickson et al., 2019). Surprisingly, the blaCTX-M-15 gene was widespread in both humans and pets, with the possibility of gene exchange between them (Bogaerts et al., 2015). Occurrence of blaCTX-M-15 was highest in pets originating from shelters (Johansson et al., 2022), followed by livestock areas of county fairs, livestock auction markets, equine facilities, and dairy farms (Adams et al., 2018). Pets participate in the dissemination of the ESBL-EC. For instance, 13 distinct STs with the most common ST117, ST131, and ST38 clones containing eight variants of the blaCTX-M gene including blaCTX-M-14 and blaCTX-M-15 have been identified in animal shelters (Umeda et al., 2019). A German study indicated that 8.9% of dogs seen in veterinary clinics carried ESBL-EC, and that seven factors, including husbandry system, contact with puppies, and feeding raw meat, were significantly associated with ESBL-EP colonization (Werhahn Beining et al., 2023). The most common transmission and occurrence of blaCTX-M-1, followed by blaCTX-M-15 and blaCTX-M-32 harboring IncI1- and IncF-type plasmids, have been documented in dogs and their owners (Van den Bunt et al., 2020). A similar study on stray dogs and cats confirmed them as a significant source of MDR and ESBL-PE containing blaCTX-M-15, blaCTX-M-2, blaCTX-M-9, and blaCTX-M-8 with predominantly ST973, ST457, ST90, and ST2541 clones harboring different plasmid types (Melo et al., 2018). A pet like a dog can contribute to the spread and community transmission of ESBL-EC clonal types ST131, ST963, and ST69, as similar strains were isolated from family members and pets within the same household (Toombs-Ruane et al., 2020). In Korea, third-generation cephalosporin- and fluoroquinolone-resistant E. coli strains were widely distributed in cats and dogs, with E. coli ST131 harboring blaCTX-M-14 and blaCTX-M-15 genes and E. coli ST405 harboring the blaCMY-2 gene (Choi et al., 2023). Notably, ESBL-EC carrying silenced mcr-9 were isolated from companion cats in Japan, albeit not universally (Yasugi et al., 2023). Moreover, E. coli ST224 contains blaCTX-M-8 and identifies IncI1-, IncY-, IncFIA-, and IncFIB-type plasmids in domestic cats, which may create therapeutic limitations in veterinary medicine (Silva M. M. et al., 2018). Thus, pets are involved in the spread of ESBL-PE to other animals, the environment, and humans via saliva or direct contact (Melo et al., 2018). Therefore, strict monitoring of companion animals is required to limit the spread of ESBL-PE.
Rising populations lead to the destruction of natural wildlife habitats, which in turn unexpectedly increases the role of wildlife in ARG dissemination. This includes birds (Marcelino et al., 2019) and wild animals such as wild ungulates, wild boar (Sus scrofa), red deer (Cervus elaphus), fallow deer (Dama dama), and mouflon (Ovis aries subsp. musimon) (Torres et al., 2022). This is because animals need food reservoirs and water sources; thus, they travel from their native habitats to other places. This causes wildlife to come in to contact with anthropologically affected areas that may hold residual antibiotics and/or ARB (Carroll et al., 2015).
Wild birds are assumed as significant reservoirs and propagators of ARGs that can contaminate the environment (Dolejska and Literak, 2019). Migratory birds can further enhance ARG dissemination widely over long distances, which ultimately enhances worldwide occurrences of AMR (Hernando-Amado et al., 2019). For example, many reports have been documented for the existence of ARB in wild birds, included but not limited to geese and ducks (Li et al., 2024; Zhang et al., 2023b; Zhang et al., 2023c), gulls (Hernandez et al., 2013; Ruzauskas and Vaskeviciute, 2016), rooks (Oravcova et al., 2013), and passerines (Dolejská et al., 2008). Moreover, wild birds show higher ARGs in anthropologically stimulated environments than those existing in less inhabited zones (Miller et al., 2020; Marcelino et al., 2019). A recent study revealed the influence of microbial community structure, MGEs, and residual antibiotics on resistomes of bird feces and further found great interconnectivity of ARGs among the microbiomes of wild birds and their habitats (Luo et al., 2022).
Moreover, MGEs such as integrons, plasmids, and transposons are involved in the horizontal spread of ARGs in humans, animals, and the environment. ESBL-PE bacteria are increasingly found across various environments, with blaCTX-M-15 being widespread. MGEs, including plasmids and transposons, facilitate the spread of antibiotic resistance among bacteria (Zhang et al., 2020). MGEs play an important role in HGT and the association of ISCR1 elements recorded with blaCTX-M-15 (Ali et al., 2016), and the same gene has been recognized by both community and cattle linked by MGEs such as Tn3 or IS1380 families (Lifshitz-Ziv et al., 2018).
ISEcp1 has been identified to increase the expression of ESBL genes via a promoter in the upstream region of these genes (Vandecraen et al., 2017). The MGEs are widely prevalent and involve ARGs spread among various environments (Tamang et al., 2013; Jakobsen et al., 2015; Ben Said et al., 2016). The spread of blaCTX-M genes was enhanced by ISEcp1 element. It is linked to many blaCTX-M genes such as blaCTX-M-24, blaCTX-M-14, blaCTX-M-79, and blaCTX-M-22 (Tian et al., 2011). Recent studies have found that ISEcp1 and IS26 are the most common structures in the blaCTX-M-55 flanking environment, with ISEcp1-carrying blaCTX-M-55 being more prevalent but decreasing year by year in humans and IS26-carrying increasing year by year in animals and food (Yang et al., 2023). Previously, in food animals, ISEcp1 and IS903 were observed upstream and downstream to the blaCTX-M-14 gene while IS26 transposase was found upstream of the blaCTX-M-1 gene flanking a partially truncated ISEcp1 (Jouini et al., 2007). In the Indian urban aquatic environment, plasmid-mediated gene transmission between genera was observed through HGT; a genetic environment of blaCTX-M-15 is shown in Figure 2B (Singh et al., 2018).
Rapid HGT in clinical isolates containing blaCTX-M-55 with four different genetic contexts harboring upstream ISEcp1, truncation by IS26 and IS1294, and downstream orf477 and IS903 is shown in Figure 2C (Hu et al., 2018). Similar findings have been recorded from blood stream infections in elderly patients, where blaCTX-M is linked with ISEcp1, IS26, and IS903 elements with particular special sequences of region V (80 bp), Y(42 bp), and W(48 bp) for ISEcp1 (Xiao et al., 2019), as previously described (Eckert et al., 2006; Jouini et al., 2007). Plasmid-mediated dissemination of the blaCTX-M-1 gene from food-producing animals and healthy human links with the ISEcp1 IS5 and IS26 elements while the blaTEM gene associates with Tn2 and IS26 (Figure 2E) (Wang et al., 2015). Recently, high conjugational frequency showed ΔIS26-blaCTX-M-14-ΔIS903B transmission among E. coli clones and plasmids in pig farm anaerobic effluents (Tian et al., 2022).
Conjugational transfer is a well-known mechanism of HGT between organisms. The transferability of ESBL genes via conjugational experiments has been well documented (Tamang et al., 2013; Mobasseri et al., 2019; Palmeira et al., 2020). Interestingly, one human isolate was found to be a common ST3891 clone carrying blaCTX-M-1 and blaCTX-M-61 genes and was also shown in cattle isolated on the same farm. Furthermore, E. coli isolates derived from humans, cattle and pigs were found to carry identical blaCTX-M-1 and blaCTX-M-61 genes but exhibited distinct STs. This observation may indicate HGT of ARGs between these populations (Dahms et al., 2015). Moreover, ARGs such as blaCTX-M-1, blaCTX-M-3, blaCTX-M-9, blaCTX-M-14, blaCTX-M-15, blaTEM-49, and blaSHV-2 in WWTP effluents are transferrable in conjugational experiments (Korzeniewska and Harnisz, 2013b; Ben Said et al., 2016), while the blaCTX-M-1 conjugational transfer was related to the acquisition of IncI1- and IncN-type plasmids (Ben Said et al., 2016). It shows a high probability of HGT between bacteria in sewage and environmental settings. Likewise, different plasmid types such as IncI1 and IncK containing ESBL genes are pooled into farm animals and humans (De Been et al., 2014). Therefore, HGT needs urgent attention and can be reduced through intervention measures, legislative monitoring programs, and minimizing reservoirs or contaminants.
Control procedures can be based on the identification of resistant Enterobacteriaceae reservoirs (Figure 3). This includes screening and elucidation of ESBL-PE spread routes such as water, air, soil, farm manure, FWs, livestock, poultry, and wildlife (Guenther et al., 2011; Dahms et al., 2015; Dohmen et al., 2017; Wang et al., 2017). In addition to the appropriate use of antibiotics (Mobasseri et al., 2019), disinfection of hospital sewage (Korzeniewska and Harnisz, 2013b) is also recommended. Urinary catheters are a big source of ESBL-PE; direct contact with contaminated instruments can act as a vehicle for HGT. Therefore, it is important to disinfect, sterilize, and properly design instruments to monitor HGT (Al-Jamei et al., 2019). Minimum procedures required for infection control in hospitals should be mandatory, in accordance with WHO guidelines (World Health Organization, 2019). The installation of wastewater treatment facilities in hospitals could be carried out (Adekanmbi et al., 2020) to reduce the introduction of MDR bacteria and ESBL genes in the environment through hospital settings.
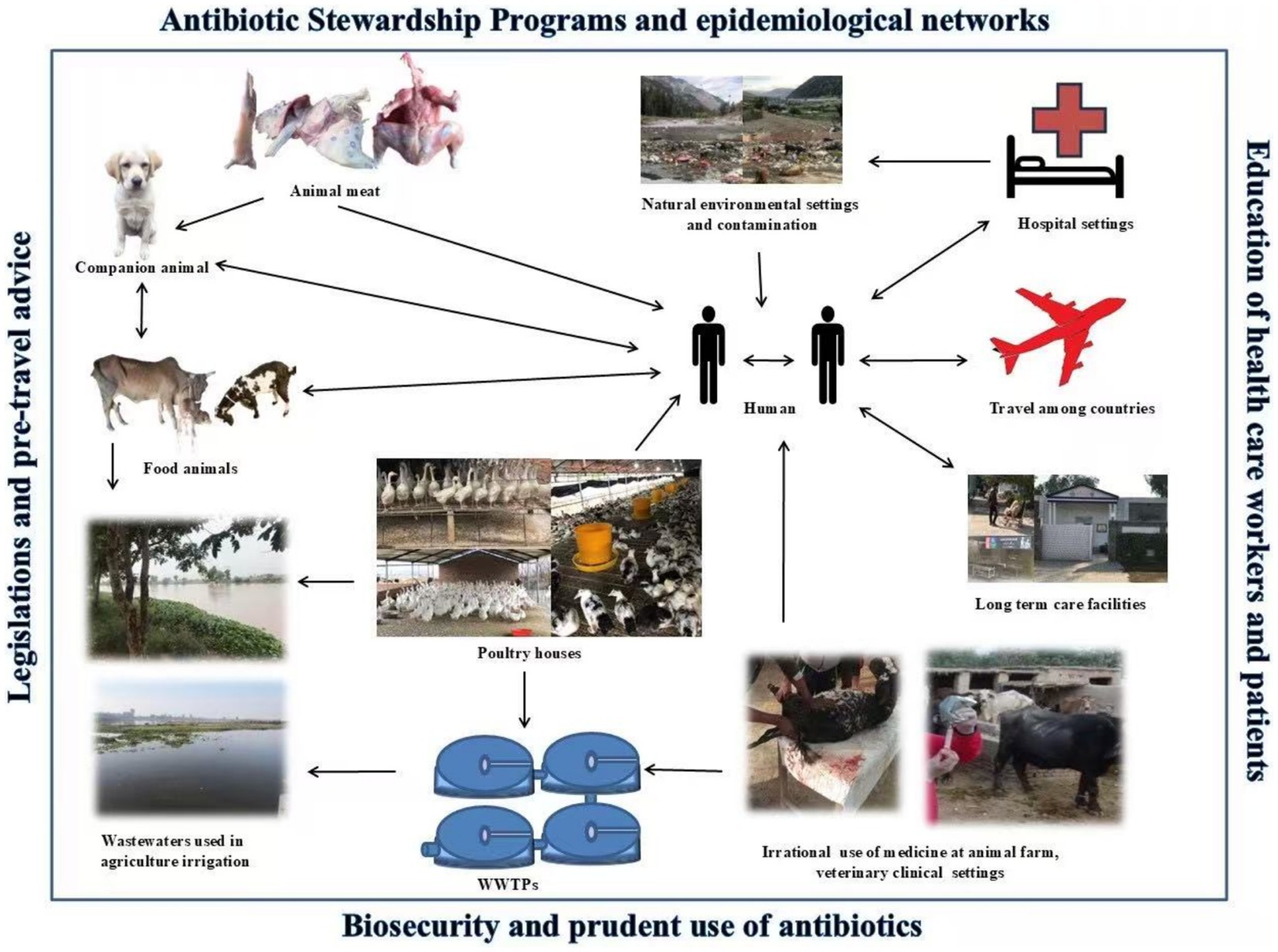
Figure 3. Illustrations of ESBL-PE spread routes in the natural environment. Potential transmission routes of ESBL-PE are indicated by arrows among reservoirs such as hospitals, long-term care facilities, livestock farms, meat, companion animals, travels, WWTPs, water resources, the agricultural field, and humans. The transmission of ESBL-PE in the natural environment can be a hazard to public health. Finally, interventions are given in the outer box which are required to control the AMR from a “One Health” perspective.
Other potential factors may include understanding the transfer of ARB at the livestock interface and other reservoirs (Lee et al., 2020). WWTPs emit ARB into the air, and bio-aerosol emissions can be reduced by covering aeration tanks and grit chambers to control environmental contamination (Korzeniewska and Harnisz, 2013b).
The prevalence of ESBL-PE is growing; therefore, management, specific hygiene measures, and optimal use of antibiotics should be implemented. There is a pressing need to develop approaches to determine the variability of AMR and trends in propagation mechanisms between multiple reservoirs (Figure 3). Training of doctors/staff in ASPs, communication procedures, hygiene, patient information, related management tools (Gillespie et al., 2013; Miclot et al., 2015; Aldrazi et al., 2020; Di Gennaro et al., 2020), excreta management, and strict implementation of rules (NGuyen T. T. H. et al., 2019) are necessary to prevent and control infections involving ESBLs. Studies on the dynamics of transmission, such as surgical site infections (Moremi et al., 2018), and epidemiological networks are also needed between hospital and community settings (Miclot et al., 2015). General practitioners can provide a toolkit that includes a list of infection prevention guidelines, infectious disease physician contacts, treatment procedures, general practitioners, and patient leaflets (Zucconi et al., 2018). Therefore, the spread of ESBL-PE can be measured and reduced to a minimum through active surveillance systems and infection control programs.
Global travel contributes to the spread of ESBL-PE among countries/continents. Active reviews and administrative segregation at the time of admission to medical services for patients who have travelled to Asia, especially with a 6-month history of travel in Southeast Asia and India, should be required (Lübbert et al., 2015; Raffelsberger et al., 2023). Pre-trip advice and awareness should be provided for those who have visited high-risk countries. That can include health safety (Tokinobu et al., 2020), personal hand hygiene procedures (Meurs et al., 2019; Maataoui et al., 2019), and guidelines on antibiotic use during travel (Angelin et al., 2015; Otter et al., 2019; Maataoui et al., 2019). Clinicians need to be careful about ESBL-PE acquired during patient handling. Risk-based monitoring should be carried out for both regular and long-term business visitors (Mizuno et al., 2016). Travel patients should be isolated after rapid recognition of ESBL-PE colonization to prevent further spread of the resistant organism (Lübbert et al., 2015). Thus, effective prevention can be achieved with a better understanding of the dissemination pathways of ESBL-PE in the natural environment (Figure 3).
Hygienic measures such as using specific work clothing, washing hands, taking baths at the end of a work shift, protecting any cuts or lesions on the skin, and wearing a respirator mask could be implemented (Kraemer et al., 2017). There is an urgent need for device, multi-faceted, multi-sectoral, and workable collaboration among stakeholders to contain the spread of ESBL-PE (Founou et al., 2019). Educating FWs, poultry producers, and sellers requires the promotion of hygiene and the prudent use of antibiotics in the poultry production system (Aworh et al., 2019). Finally, effects of municipal wastewater effluent on the spread of AMR in the aquatic environment can be reduced through the application of new treatment technologies, such as silver-based, mesoporous silica-based, titanium dioxide-based, chitosan-based, carbon-based, and clay-based nanomaterials (Ojemaye et al., 2020). The economical and biocompatible ionic liquid choline dihydrogen phosphate has shown to be a promising choice for the elimination of antibiotics from swine effluents (Álvarez et al., 2016). High temperatures in composted pig manure piles can reduce some MGEs such as Tn3, ISEcp1, IS613, IS256, tnpA-5, and tnpA-6 (Wu N. et al., 2020). In general, we are supposed to change our course of action towards a more prudent use of antibiotics. The alternative is to design new therapeutic drug molecules or to make improvements to existing antibiotics such as Halicin via modern machine learning approaches (Stokes et al., 2020). However, ensuring the availability of new drugs in the market is a harder task. One of the solutions is to reintroduce vulnerable flora as probiotics (Imperial and Ibana, 2016). Commensal bacteria can serve as beneficial partners, but strategies are needed to help them outcompete ARB.
The environment plays an important role in elevating selective pressure; for example, the overuse of antimicrobials in agricultural, human, and veterinary settings facilitates bacteria to gain novel mutations essential for survival (Udikovic-Kolic et al., 2014; Berendonk et al., 2015; Woolhouse et al., 2015). Studies have highlighted the link between antibiotic use and AMR within human beings or veterinary settings (Karkaba et al., 2017; Kwok et al., 2020). Thus, there is an urgent need to clarify the role of the clinical laboratory in success of the implementation of ASP to control community infections (Gambrah et al., 2021). For instance, a study in pastoralist out-patients recorded high-resistance ampicillin and 51.6% resistance to amoxicillin-clavulanic acid (Stanley et al., 2018), which highlights the importance of lab testing to improve the precision of antimicrobial therapy. Restricted use of antimicrobials such as cephalosporins has resulted in reduced ESBL usage from 29.5 to 9.5% in hospitals (Kurita et al., 2019). Patients show improved prognosis if they received the right therapy as per laboratory recommendations, as compared to patients who received inappropriate therapy (Fourie et al., 2018). The Ministry of Health, Malaysia, has approved 97 antimicrobials for humans and animals. Alarmingly, these antibiotics, including 3rd and 4th generation cephalosporins, are used in animals as well as in human prophylaxis and therapeutics (Mobasseri et al., 2019). Worryingly, large quantities of antimicrobials/medicines are discarded or unused each year, and most antibiotics used in humans and animals are excreted into the environment through urine and feces, which requires appropriate disposal to reduce the unwanted environmental as well as public health hazards attributed by pharmaceuticals (Zhang et al., 2015; Vatovec et al., 2021).
ASPs are appropriate for patient safety and efficiently reducing adverse drug effects and MDR (Bauer et al., 2019). Although an 100% solution to AMR is impossible, the devoted involvement of the healthcare system is required for the ultimate solution of the problem. The US Center for Disease Control and Prevention (CDC) analyzed effective ASPs in hospitals and recognized core elements for successful ASPs in 2014, which were further updated in 2019. These are hospital leadership commitment, accountability, drug expertise, action, tracking of antibiotic prescription, regular reporting of antibiotic use, resistance to HWs, educating doctors, nurses, and patients about AMR, prescription improvements, and interventions (CDC, 2020). Monitoring of ASPs is important, as results among institutions varied because of the difference in management and control systems. For instance, ASPs have been implemented in Spanish hospitals, and the Spanish Society of Clinical Microbiology and Infectious Diseases has defined the objectives of ASPs along with recommendations and execution (Rodríguez-Baño et al., 2012). As a result, a reduction in rates of UTIs was observed at the hospital level (Esteve-Palau et al., 2018). Further, imposing ASPs in primary health care centers is improving the use of antibiotics and subsequently causing a significant reduction in infections (Peñalva et al., 2020). In addition, education and monitoring of nurses improve the efficacy of ASPs (Gillespie et al., 2013). In Brazil, ASP-sparing β-lactam antibiotics was applied: results are promising, as overall antibiotic expenditure decreased by 53.6% after interventions (Zequinao et al., 2020). Overall, ASPs are effectively controlling and decreasing the use of antibiotics/antimicrobials by intervening on antibiotic prescriptions (Bolla et al., 2020; Velasco-Arnaiz et al., 2020). Therefore, the application of ASPs is encouraging and hereby recommended for its wider application from WHO.
This review updates the prevalence of plasmids in ESBL-PE across the three “One Health” compartments (human, animal, and environment). Key plasmid incompatibility groups include IncF (FIA, FIB, FIC, FI, FII), IncHI (HI1, HI2), IncI1, IncY, IncX (X, X1, X4), IncN, IncA/C, and others (IncB, IncO, IncK, IncZ). ESBL-PE is widely observed in reservoirs such as hospitals, wastewater, WWTPs, agriculture settings, international travel, and companion-, farm-, and food-animals, posing a threat to public health. ISEcp1 is present upstream and orf477 downstream to blaCTX-M genes with other MGEs, such as IS26 and IS903, playing important roles in the capturing, expression, and mobilization of blaCTX-M genes. The blaCTX-M-15 gene is most prevalent in multiple reservoirs. The HGT is a significant method of gene transfer that is difficult to control. MGEs such as Tn3 or IS1380 families and plasmids like IncI1, IncK, and IncN are involved in HGT. The spread of ESBL-PE can be reduced via interventions such as WWTPs facilities in hospitals, knowledge of spread routes, ASPs, and implementation of WHO infection control guidelines in hospitals. Laboratory recommendations of antibiotics should be used in human and veterinary clinics to avoid the overuse of antibiotics. The rise in ESBL-PE is largely due to the misuse of antibiotics in human and veterinary clinics. Policymakers should focus on the rising threat of ESBL-PE and timely implementation of policies as the choice of antibiotics becomes limited.
Competitive omission with probiotics, new drug development, and massive vaccination are future options to decrease ESBL-PE. Therefore, we require exploring the mechanisms involved in the colonization of ESBL-PE. Still, some diverse environments such as schools, offices, local buildings, livestock and poultry farms, and municipal settings need to be investigated for the prevalence of ESBL-PE.
SZ: Data curation, Methodology, Writing – original draft, Writing – review & editing. JY: Formal analysis, Investigation, Methodology, Supervision, Writing – original draft. MA: Conceptualization, Formal analysis, Methodology, Supervision, Writing – original draft. QY: Data curation, Investigation, Writing – original draft. QL: Investigation, Software, Writing – original draft. ML: Data curation, Methodology, Supervision, Writing – review & editing. DZ: Data curation, Methodology, Supervision, Writing – review & editing. MW: Formal analysis, Project administration, Validation, Writing – review & editing. BT: Formal analysis, Project administration, Validation, Writing – original draft. AC: Funding acquisition, Resources, Writing – original draft, Writing – review & editing.
The author(s) declare that financial support was received for the research, authorship, and/or publication of this article. This work was supported by the National Key Research and Development Program of China (2023YFD1800203), the earmarked fund for China Agriculture Research System (CARS-42-17), and the Program of Sichuan Veterinary Medicine and Drug Innovation Group of China Agricultural Research System (SCCXTD-2025-18).
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Abdallah, H. M., Reuland, E. A., Wintermans, B. B., Al Naiemi, N., Koek, A., Abdelwahab, A. M., et al. (2015). Extended-Spectrum β-lactamases and/or Carbapenemases-producing Enterobacteriaceae isolated from retail chicken meat in Zagazig, Egypt. PLoS One 10:e0136052. doi: 10.1371/journal.pone.0136052
Adams, R. J., Kim, S. S., Mollenkopf, D. F., Mathys, D. A., Schuenemann, G. M., Daniels, J. B., et al. (2018). Antimicrobial-resistant Enterobacteriaceae recovered from companion animal and livestock environments. Zoonoses Public Health 65, 519–527. doi: 10.1111/zph.12462
Adekanmbi, A. O., Akinpelu, M. O., Olaposi, A. V., and Oyelade, A. A. (2020). Diversity of extended Spectrum Beta-lactamase (ESBL) genes in Escherichia coli isolated from wastewater generated by a Sick Bay located in a university health care facility. Gene Rep. 20:100738. doi: 10.1016/j.genrep.2020.100738
Adenipekun, E. O., Jackson, C. R., Ramadan, H., Iwalokun, B. A., Frye, J. G., Barrett, J. B., et al. (2019). Plasmid replicons and β-lactamase-encoding genes of multidrug-resistant Escherichia coli isolated from humans and food animals in Lagos, Southwest Nigeria. Microb. Drug Resist. 25, 1410–1423. doi: 10.1089/mdr.2018.0305
Adler, A., Baraniak, A., Izdebski, R., Fiett, J., Salvia, A., Samso, J. V., et al. (2014). A multinational study of colonization with extended spectrum β-lactamase-producing Enterobacteriaceae in healthcare personnel and family members of carrier patients hospitalized in rehabilitation centres. Clin. Microbiol. Infect. 20, O516–O523. doi: 10.1111/1469-0691.12560
Ahmed, M. F. E., Ramadan, H., Seinige, D., Kehrenberg, C., Abd El-Wahab, A., Volkmann, N., et al. (2020). Occurrence of extended-spectrum beta-lactamase-producing Enterobacteriaceae, microbial loads, and endotoxin levels in dust from laying hen houses in Egypt. BMC Vet. Res. 16:301. doi: 10.1186/s12917-020-02510-4
Ala-Jaakkola, R., Laitila, A., Ouwehand, A. C., and Lehtoranta, L. (2022). Role of D-mannose in urinary tract infections - a narrative review. Nutr. J. 21:18. doi: 10.1186/s12937-022-00769-x
Aldea, I., Gibello, A., Hernández, M., Leekitcharoenphon, P., Bortolaia, V., and Moreno, M. A. (2022). Clonal and plasmid-mediated flow of ESBL/AmpC genes in Escherichia coli in a commercial laying hen farm. Vet. Microbiol. 270:109453. doi: 10.1016/j.vetmic.2022.109453
Aldrazi, F. A., Rabaan, A. A., Alsuliman, S. A., Aldrazi, H. A., Alabdalslam, M. J., Alsadiq, S. A., et al. (2020). ESBL expression and antibiotic resistance patterns in a hospital in Saudi Arabia: Do healthcare staff have the whole picture? J. Infect. Public Health 13, 759–766. doi: 10.1016/j.jiph.2019.12.001
Ali, T., Ur Rahman, S., Zhang, L., Shahid, M., Zhang, S., Liu, G., et al. (2016). ESBL-producing Escherichia coli from cows suffering mastitis in China contain clinical class 1 Integrons with CTX-M linked to ISCR1. Front. Microbiol. 7:1931. doi: 10.3389/FMICB.2016.01931
Al-Jamei, S. A., Albsoul, A. Y., Bakri, F. G., and Al-Bakri, A. G. (2019). Extended-spectrum β-lactamase producing E. coli in urinary tract infections: a two-center, cross-sectional study of prevalence, genotypes and risk factors in Amman, Jordan. J. Infect. Public Health 12, 21–25. doi: 10.1016/j.jiph.2018.07.011
Alqasim, A., Abu Jaffal, A., and Alyousef, A. A. (2018). Prevalence of multidrug resistance and extended-Spectrum β-lactamase carriage of clinical Uropathogenic Escherichia coli isolates in Riyadh, Saudi Arabia. Int. J. Microbiol 2018:851. doi: 10.1155/2018/3026851
Álvarez, M. S., Gómez, L., Ulloa, R. G., Deive, F. J., Sanromán, M. A., and Rodríguez, A. (2016). Antibiotics in swine husbandry effluents: laying the foundations for their efficient removal with a biocompatible ionic liquid. Chem. Eng. J. 298, 10–16. doi: 10.1016/j.cej.2016.04.014
Alves, J., Marques, V. V., Pereira, L. F. P., Hirooka, E. Y., and de Oliveira, T. C. R. M. (2012). Multiplex PCR for the detection of Campylobacter spp. and Salmonella spp. in chicken meat. J. Food Saf. 32, 345–350. doi: 10.1111/j.1745-4565.2012.00386.x
Angelin, M., Forsell, J., Granlund, M., Evengård, B., Palmgren, H., Johansson, A., et al. (2015). Risk factors for colonization with extended-spectrum beta-lactamase producing Enterobacteriaceae in healthcare students on clinical assignment abroad: a prospective study. Travel Med. Infect. Dis. 13, 223–229. doi: 10.1016/j.tmaid.2015.04.007
Arcilla, M. S., Van Hattem, J. M., Bootsma, M. C. J., van Genderen, P. J. J., Goorhuis, A., Grobusch, M. P., et al. (2017). Import and spread of extended-spectrum β-lactamase-producing Enterobacteriaceae by international travellers (COMBAT study): a prospective, multicentre cohort study. Lancet Infect. Dis. 17, 78–85. doi: 10.1016/S1473-3099(16)30319-X
Arcilla, M. S., Van Hattem, J. M., Bootsma, M. C. J., van Genderen, P. J. J., Goorhuis, A., Grobusch, M. P., et al. (2019). Prevalence and risk factors for carriage of ESBL-producing Enterobacteriaceae in a population of Dutch travellers: a cross-sectional study. Travel Med. Infect. Dis. 33:101547. doi: 10.1016/j.tmaid.2019.101547
Aristizábal-Hoyos, A. M., Rodríguez, E. A., Arias, L., and Jiménez, J. N. (2019). High clonal diversity of multidrug-resistant and extended spectrum beta-lactamase-producing Escherichia coli in a wastewater treatment plant. J. Environ. Manag. 245, 37–47. doi: 10.1016/j.jenvman.2019.05.073
Artero, A., Esparcia, A., Alberola, J., Madrazo, M., Nogueira, J. M., and Eiros, J. M. (2017). Prospective cohort study of risk factors for extended-spectrum ß-lactamase-producing Escherichia coli urinary tract infections in elderly patients admitted to hospital. Int. J. Clin. Pract. 71:e13001. doi: 10.1111/ijcp.13001
Atlas, R. M. (2013). One health: its origins and future. Curr. Top. Microbiol. Immunol. 365, 1–13. doi: 10.1007/82_2012_223
Aworh, M. K., Kwaga, J., Okolocha, E., Mba, N., and Thakur, S. (2019). Prevalence and risk factors for multi-drug resistant Escherichia coli among poultry workers in the Federal Capital Territory, Abuja Nigeria. PLoS One 14:e0225379. doi: 10.1371/journal.pone.0225379
Awosile, B. B., and Agbaje, M. (2021). Genetic environments of plasmid-mediated blaCTXM-15 Beta-lactamase gene in Enterobacteriaceae from Africa. Microbiol. Res. 12, 383–394. doi: 10.3390/MICROBIOLRES12020026
Azuma, T., Uchiyama, T., Zhang, D., Usui, M., and Hayashi, T. (2022). Distribution and characteristics of carbapenem-resistant and extended-spectrum β-lactamase (ESBL) producing Escherichia coli in hospital effluents, sewage treatment plants, and river water in an urban area of Japan. Sci. Total Environ. 839:156232. doi: 10.1016/j.scitotenv.2022.156232
Bae, D., Cheng, C. M., and Khan, A. A. (2015). Characterization of extended-spectrum β-lactamase (ESBL) producing non-typhoidal Salmonella (NTS) from imported food products. Int. J. Food Microbiol. 214, 12–17. doi: 10.1016/j.ijfoodmicro.2015.07.017
Bauer, K., Kullar, R., Gilchrist, M., and File, T. (2019). Antibiotics and adverse events: the role of antimicrobial stewardship programs in ‘doing no harm. Curr. Opin. Infect. Dis. 32, 553–558. doi: 10.1097/QCO.0000000000000604
Behrens, W., Kolte, B., Junker, V., Frentrup, M., Dolsdorf, C., Börger, M., et al. (2023). Bacterial genome sequencing tracks the housefly-associated dispersal of fluoroquinolone- and cephalosporin-resistant Escherichia coli from a pig farm. Environ. Microbiol. 25, 1174–1185. doi: 10.1111/1462-2920.16352
Ben Said, L., Jouini, A., Alonso, C. A., Klibi, N., Dziri, R., Boudabous, A., et al. (2016). Characteristics of extended-spectrum β-lactamase (ESBL)- and pAmpC beta-lactamase-producing Enterobacteriaceae of water samples in Tunisia. Sci. Total Environ. 550, 1103–1109. doi: 10.1016/j.scitotenv.2016.01.042
Berbers, B., Vanneste, K., Roosens, N. H. C. J., Marchal, K., Ceyssens, P. J., and De Keersmaecker, S. C. J. (2023). Using a combination of short- and long-read sequencing to investigate the diversity in plasmid- and chromosomally encoded extended-spectrum beta-lactamases (ESBLs) in clinical Shigella and Salmonella isolates in Belgium. Microb. Genomics 9:mgen000925. doi: 10.1099/mgen.0.000925
Berendonk, T. U., Manaia, C. M., Merlin, C., Fatta-Kassinos, D., Cytryn, E., Walsh, F., et al. (2015). Tackling antibiotic resistance: the environmental framework. Nat. Rev. Microbiol. 13, 310–317. doi: 10.1038/nrmicro3439
Bevan, E. R., Powell, M. J., Toleman, M. A., Thomas, C. M., Piddock, L. J. V., and Hawkey, P. M. (2021). Molecular characterization of plasmids encoding blaCTX-M from faecal Escherichia coli in travellers returning to the UK from South Asia. J. Hosp. Infect. 114, 134–143. doi: 10.1016/j.jhin.2021.03.030
Bogaerts, P., Huang, T.-D., Bouchahrouf, W., Bauraing, C., Berhin, C., El Garch, F., et al. (2015). Characterization of ESBL- and AmpC-producing Enterobacteriaceae from diseased companion animals in Europe. Microb. Drug Resist. 21, 643–650. doi: 10.1089/mdr.2014.0284
Bolla, C., Di Pietrantonj, C., Ferrando, E., Pernecco, A., Salerno, A., D’Orsi, M., et al. (2020). Example of antimicrobial stewardship program in a community hospital in Italy. Méd. Mal. Infect. 50, 342–345. doi: 10.1016/j.medmal.2019.11.008
Bonardi, S., Cabassi, C. S., Manfreda, G., Parisi, A., Fiaccadori, E., Sabatino, A., et al. (2022). Survey on Carbapenem-resistant Bacteria in pigs at slaughter and comparison with human clinical isolates in Italy. Antibiotics 11:777. doi: 10.3390/antibiotics11060777
Bougnom, B. P., Thiele-Bruhn, S., Ricci, V., Zongo, C., and Piddock, L. J. V. (2020). Raw wastewater irrigation for urban agriculture in three African cities increases the abundance of transferable antibiotic resistance genes in soil, including those encoding extended spectrum β-lactamases (ESBLs). Sci. Total Environ. 698:134201. doi: 10.1016/j.scitotenv.2019.134201
Boyer, A., Couallier, V., Clouzeau, B., Lasheras, A., Mzali, F., Kann, M., et al. (2015). Control of extended-spectrum β-lactamase-producing Enterobacteriaceae nosocomial acquisition in an intensive care unit: a time series regression analysis. Am. J. Infect. Control 43, 1296–1301. doi: 10.1016/j.ajic.2015.07.026
Burgess, S. A., Aplin, J., Biggs, P. J., Breckell, G., Benschop, J., Fayaz, A., et al. (2021). Characterisation of AmpC and extended-spectrum beta-lactamase producing E. coli from New Zealand dairy farms. Int. Dairy J. 117:104998. doi: 10.1016/j.idairyj.2021.104998
Carrique-Mas, J. J., Trung, N. V., Hoa, N. T., Mai, H. H., Thanh, T. H., Campbell, J. I., et al. (2015). Antimicrobial usage in chicken production in the Mekong Delta of Vietnam. Zoonoses Public Health 62, 70–78. doi: 10.1111/zph.12165
Carroll, D., Wang, J., Fanning, S., and McMahon, B. J. (2015). Antimicrobial resistance in wildlife: implications for public health. Zoonoses Public Health 62, 534–542. doi: 10.1111/zph.12182
Casella, T., Nogueira, M. C. L., Saras, E., Haenni, M., and Madec, J.-Y. (2017). High prevalence of ESBLs in retail chicken meat despite reduced use of antimicrobials in chicken production, France. Int. J. Food Microbiol. 257, 271–275. doi: 10.1016/j.ijfoodmicro.2017.07.005
Casella, T., Rodríguez, M. M., Takahashi, J. T., Ghiglione, B., Dropa, M., Assunção, E., et al. (2015). Detection of blaCTX-M-type genes in complex class 1 integrons carried by Enterobacteriaceae isolated from retail chicken meat in Brazil. Int. J. Food Microbiol. 197, 88–91. doi: 10.1016/j.ijfoodmicro.2014.12.001
Castañeda-Barba, S., Top, E. M., and Stalder, T. (2024). Plasmids, a molecular cornerstone of antimicrobial resistance in the one health era. Nat. Rev. Microbiol. 22, 18–32. doi: 10.1038/s41579-023-00926-x
Castillo-Tokumori, F., Irey-Salgado, C., and Málaga, G. (2017). Worrisome high frequency of extended-spectrum beta-lactamase-producing Escherichia coli in community-acquired urinary tract infections: a case–control study. Int. J. Infect. Dis. 55, 16–19. doi: 10.1016/j.ijid.2016.12.007
CDC Core Elements of Hospital Antibiotic Stewardship Programs | Antibiotic Use. (2020). Available at: https://www.cdc.gov/antibiotic-use/core-elements/hospital.html (Accessed December 23, 2020).
Chao, C. M., Lai, C. C., and Yu, W. L. (2023). Epidemiology of extended-spectrum β-lactamases in Enterobacterales in Taiwan for over two decades. Front. Microbiol. 13:1060050. doi: 10.3389/fmicb.2022.1060050
Chaturvedi, P., Chaurasia, D., Pandey, A., and Gupta, P. (2020). Co-occurrence of multidrug resistance, β-lactamase and plasmid mediated AmpC genes in bacteria isolated from river ganga, northern India. Environ. Pollut. 267:115502. doi: 10.1016/j.envpol.2020.115502
Cho, S., Jackson, C. R., and Frye, J. G. (2023). Freshwater environment as a reservoir of extended-spectrum β-lactamase-producing Enterobacteriaceae. J. Appl. Microbiol. 134:lxad034. doi: 10.1093/jambio/lxad034
Choi, J. H., Ali, M. S., Moon, B. Y., Kang, H. Y., Kim, S. J., Song, H. J., et al. (2023). Prevalence and characterization of extended-Spectrum β-lactamase-producing Escherichia coli isolated from dogs and cats in South Korea. Antibiotics 12:745. doi: 10.3390/antibiotics12040745
Cissé, A., Sambe Ba, B., Sow, O., Wane, A. A., Ndiaye, I., Fall, C., et al. (2024). Prevalence, antimicrobial susceptibility, and resistance genes of extended-Spectrum β-lactamase-producing Escherichia coli from broilers sold in open Markets of Dakar, Senegal. Microorganisms 12:2357. doi: 10.3390/microorganisms12112357
Clemente, L., Manageiro, V., Correia, I., Amaro, A., Albuquerque, T., and Themudo, P. (2019). Revealing mcr-1-positive ESBL-producing Escherichia coli strains among Enterobacteriaceae from food-producing animals (bovine, swine and poultry) and meat (bovine and swine), Portugal, 2010–2015. Int. J. Food Microbiol. 296, 37–42. doi: 10.1016/j.ijfoodmicro.2019.02.006
Coppola, N., Cordeiro, N. F., Trenchi, G., Esposito, F., Fuga, B., Fuentes-Castillo, D., et al. (2022). Imported one-day-old chicks as Trojan horses for multidrug-resistant priority pathogens harboring mcr-9, rmtG, and extended-Spectrum β-lactamase genes. Appl. Environ. Microbiol. 88:e0167521. doi: 10.1128/AEM.01675-21
Cui, L., Zhao, X., Li, R., Han, Y., Hao, G., Wang, G., et al. (2022). Companion animals as potential reservoirs of antibiotic resistant Diarrheagenic Escherichia coli in Shandong, China. Antibiotics 11:828. doi: 10.3390/antibiotics11060828
Dahms, C., Hübner, N.-O., Kossow, A., Mellmann, A., Dittmann, K., and Kramer, A. (2015). Occurrence of ESBL-producing Escherichia coli in livestock and farm Workers in Mecklenburg-Western Pomerania, Germany. PLoS One 10:e0143326. doi: 10.1371/journal.pone.0143326
Dandachi, I., Fayad, E., El-Bazzal, B., Sleiman, A., Rolain, J.-M., and Daoud, Z. (2018). Multi drug resistant organisms in chicken farms and their surrounding environment. J. Infect. Dev. Ctries. 12:7S. doi: 10.3855/jidc.10041
Dashti, A. A., Jadaon, M. M., Gomaa, H. H., Noronha, B., and Udo, E. E. (2010). Transmission of a Klebsiella pneumoniae clone harbouring genes for CTX-M-15-like and SHV-112 enzymes in a neonatal intensive care unit of a Kuwaiti hospital. J. Med. Microbiol. 59, 687–692. doi: 10.1099/jmm.0.019208-0
De Been, M., Lanza, V. F., De Toro, M., Scharringa, J., Dohmen, W., Du, Y., et al. (2014). Dissemination of cephalosporin resistance genes between Escherichia coli strains from farm animals and humans by specific plasmid lineages. PLoS Genet. 10:e1004776. doi: 10.1371/journal.pgen.1004776
De Koster, S., Ringenier, M., Lammens, C., Stegeman, A., Tobias, T., Velkers, F., et al. (2021). ESBL-producing, Carbapenem- and ciprofloxacin-resistant Escherichia coli in Belgian and Dutch broiler and pig farms: a cross-sectional and cross-border study. Antibiotics 10:945. doi: 10.3390/antibiotics10080945
Denkel, L. A., Maechler, F., Schwab, F., Kola, A., Weber, A., Gastmeier, P., et al. (2020). Infections caused by extended-spectrum β-lactamase-producing Enterobacterales after rectal colonization with ESBL-producing Escherichia coli or Klebsiella pneumoniae. Clin. Microbiol. Infect. 26, 1046–1051. doi: 10.1016/j.cmi.2019.11.025
Despotovic, M., de Nies, L., Busi, S. B., and Wilmes, P. (2023). Reservoirs of antimicrobial resistance in the context of one health. Curr. Opin. Microbiol. 73:102291. doi: 10.1016/j.mib.2023.102291
Di Gennaro, F., Marotta, C., Amicone, M., Bavaro, D. F., Bernaudo, F., Frisicale, E. M., et al. (2020). Italian young doctors’ knowledge, attitudes and practices on antibiotic use and resistance: a national cross-sectional survey. J. Glob. Antimicrob. Resist. 23, 167–173. doi: 10.1016/j.jgar.2020.08.022
Diab, M., Hamze, M., Bonnet, R., Saras, E., Madec, J.-Y., and Haenni, M. (2018). Extended-spectrum beta-lactamase (ESBL)- and carbapenemase-producing Enterobacteriaceae in water sources in Lebanon. Vet. Microbiol. 217, 97–103. doi: 10.1016/j.vetmic.2018.03.007
Díaz-Agero Pérez, C., López-Fresneña, N., Rincon Carlavilla, A. L., Hernandez Garcia, M., Ruiz-Garbajosa, P., and Aranaz-Andrés, J. M. (2019). Local prevalence of extended-spectrum beta-lactamase (ESBL) producing Enterobacteriaceae intestinal carriers at admission and co-expression of ESBL and OXA-48 carbapenemase in Klebsiella pneumoniae: a prevalence survey in a Spanish university hospital. BMJ Open 9:e024879. doi: 10.1136/bmjopen-2018-024879
Dickin, S. K., Schuster-Wallace, C. J., Qadir, M., and Pizzacalla, K. (2016). A review of health risks and pathways for exposure to wastewater use in agriculture. Environ. Health Perspect. 124, 900–909. doi: 10.1289/ehp.1509995
Dickson, A., Smith, M., Smith, F., Park, J., King, C., and Currie, K. (2019). Understanding the relationship between pet owners and their companion animals as a key context for antimicrobial resistance-related behaviours: an interpretative phenomenological analysis. Health Psychol. Behav. Med. 7, 45–61. doi: 10.1080/21642850.2019.1577738
Dohmen, W., Bonten, M. J. M., Bos, M. E. H., Van Marm, S., Scharringa, J., and Wagenaar, J. A. (2015). Carriage of extended-spectrum β-lactamases in pig farmers is associated with occurrence in pigs. Clin. Microbiol. Infect. 21, 917–923. doi: 10.1016/j.cmi.2015.05.032
Dohmen, W., Schmitt, H., Bonten, M., and Heederik, D. (2017). Air exposure as a possible route for ESBL in pig farmers. Environ. Res. 155, 359–364. doi: 10.1016/j.envres.2017.03.002
Doi, Y., Iovleva, A., and Bonomo, R. A. (2017). The ecology of extended-spectrum β-lactamases (ESBLs) in the developed world. J. Travel Med. 24, S44–S51. doi: 10.1093/jtm/taw102
Dolejska, M., and Literak, I. (2019). Wildlife is overlooked in the epidemiology of medically important antibiotic-resistant Bacteria. Antimicrob. Agents Chemother. 63, e01167–e01119. doi: 10.1128/AAC.01167-19
Dolejská, M., Šenk, D., Čížek, A., Rybaříková, J., Sychra, O., and Literák, I. (2008). Antimicrobial resistant Escherichia coli isolates in cattle and house sparrows on two Czech dairy farms. Res. Vet. Sci. 85, 491–494. doi: 10.1016/j.rvsc.2008.03.007
Eckert, C., Gautier, V., and Arlet, G. (2006). DNA sequence analysis of the genetic environment of various blaCTX-M genes. J. Antimicrob. Chemother. 57, 14–23. doi: 10.1093/JAC/DKI398
Eibach, D., Dekker, D., Gyau Boahen, K., Wiafe Akenten, C., Sarpong, N., and Belmar Campos, C. (2018). Extended-spectrum beta-lactamase-producing Escherichia coli and Klebsiella pneumoniae in local and imported poultry meat in Ghana. Vet. Microbiol. 217, 7–12. doi: 10.1016/J.VETMIC.2018.02.023
Esteve-Palau, E., Grau, S., Herrera, S., Sorlí, L., Montero, M., and Segura, C. (2018). Impact of an antimicrobial stewardship program on urinary tract infections caused by extended-spectrum β-lactamase-producing Escherichia coli. Rev. Esp. Quimioter. 31, 110–117
European Centre for Disease Prevention and Control Antimicrobial resistance in the EU/EEA (EARS-Net) Annual Epidemiological Report for 2019 (2020). Available at: https://www.ecdc.europa.eu/en/publications-data/surveillance-antimicrobial-resistance-europe-2019 (Accessed December 30, 2024).
Ewers, C., Bethe, A., Semmler, T., Guenther, S., and Wieler, L. H. (2012). Extended-spectrum β-lactamase-producing and AmpC-producing Escherichia coli from livestock and companion animals, and their putative impact on public health: a global perspective. Clin. Microbiol. Infect. 18, 646–655. doi: 10.1111/j.1469-0691.2012.03850.x
Fernández-Verdugo, A., Forcelledo, L., Rodríguez-Lozano, J., Rodríguez-Lucas, C., Barreiro-Hurlé, L., Canut, A., et al. (2020). Prospective multicentre study of rectal carriage of multidrug-resistant Enterobacteriaceae among health-care workers in Spain. Clin. Microbiol. Infect. 26, 649.e1–649.e4. doi: 10.1016/j.cmi.2020.01.015
Fischer, J., Hille, K., Ruddat, I., Mellmann, A., Köck, R., and Kreienbrock, L. (2017). Simultaneous occurrence of MRSA and ESBL-producing Enterobacteriaceae on pig farms and in nasal and stool samples from farmers. Vet. Microbiol. 200, 107–113. doi: 10.1016/j.vetmic.2016.05.021
Fischer, J., Rodríguez, I., Baumann, B., Guiral, E., Beutin, L., Schroeter, A., et al. (2014). blaCTX-M-15-carrying Escherichia coli and Salmonella isolates from livestock and food in Germany. J. Antimicrob. Chemother. 69, 2951–2958. doi: 10.1093/jac/dku270
Flateau, C., Duron-Martinaud, S., Haus-Cheymol, R., Bousquet, A., Delaune, D., Ficko, C., et al. (2018). Prevalence and risk factors for extended-Spectrum Beta-lactamase-producing Enterobacteriaceae in French military and civilian travelers: a cross-sectional analysis. Travel Med. Infect. Dis. 23, 44–48. doi: 10.1016/j.tmaid.2018.03.009
Flores-Mireles, A. L., Walker, J. N., Caparon, M., and Hultgren, S. J. (2015). Urinary tract infections: epidemiology, mechanisms of infection and treatment options. Nat. Rev. Microbiol. 13, 269–284. doi: 10.1038/nrmicro3432
Founou, L. L., Founou, R. C., Ntshobeni, N., Govinden, U., Bester, L. A., Chenia, H. Y., et al. (2019). Emergence and spread of extended Spectrum β-lactamase producing Enterobacteriaceae (ESBL-PE) in pigs and exposed workers: a multicentre comparative study between Cameroon and South Africa. Pathogens 8:10. doi: 10.3390/pathogens8010010
Fourie, T., Schellack, N., Bronkhorst, E., Coetzee, J., and Godman, B. (2018). Antibiotic prescribing practices in the presence of extended-spectrum β-lactamase (ESBL) positive organisms in an adult intensive care unit in South Africa – a pilot study. Alex. J. Med. 54, 541–547. doi: 10.1016/j.ajme.2018.09.001
Fournier, C., Aires de Sousa, M., Fuster Escriva, B., Sales, L., Nordmann, P., and Poirel, L. (2020). Epidemiology of extended-spectrum β-lactamase-producing Enterobacteriaceae among healthcare students, at the Portuguese red cross health School of Lisbon, Portugal. J. Glob. Antimicrob. Resist. 22, 733–737. doi: 10.1016/j.jgar.2020.07.004
Fuentes-Castillo, D., Castro-Tardón, D., Esposito, F., Neves, I., Rodrigues, L., Fontana, H., et al. (2023). Genomic evidences of gulls as reservoirs of critical priority CTX-M-producing Escherichia coli in Corcovado gulf, Patagonia. Sci. Total Environ. 874:162564. doi: 10.1016/j.scitotenv.2023.162564
Gambrah, E., Owusu-Ofori, A., Biney, E., Oppong, C., and Coffin, S. E. (2021). Diagnosis and treatment of urinary tract infections in hospitalized adults in Ghana: the role of the clinical microbiology laboratory in improving antimicrobial stewardship. Int. J. Infect. Dis. 102, 497–500. doi: 10.1016/j.ijid.2020.10.068
Gay, N., Leclaire, A., Laval, M., Miltgen, G., Jégo, M., Stéphane, R., et al. (2018). Risk factors of extended-Spectrum β-lactamase producing Enterobacteriaceae occurrence in farms in Reunion, Madagascar and Mayotte Islands, 2016–2017. Vet. Sci. 5:22. doi: 10.3390/vetsci5010022
Gillespie, E., Rodrigues, A., Wright, L., Williams, N., and Stuart, R. L. (2013). Improving antibiotic stewardship by involving nurses. Am. J. Infect. Control 41, 365–367. doi: 10.1016/j.ajic.2012.04.336
Gole, V. C., Chousalkar, K. K., Roberts, J. R., Sexton, M., May, D., Tan, J., et al. (2014). Effect of egg washing and correlation between eggshell characteristics and egg penetration by various Salmonella Typhimurium strains. PLoS One 9:e90987. doi: 10.1371/JOURNAL.PONE.0090987
Gonggrijp, M. A., Santman-Berends, I. M. G. A., Heuvelink, A. E., Buter, G. J., Van Schaik, G., Hage, J. J., et al. (2016). Prevalence and risk factors for extended-spectrum β-lactamase- and AmpC-producing Escherichia coli in dairy farms. J. Dairy Sci. 99, 9001–9013. doi: 10.3168/jds.2016-11134
Goyal, D., Dean, N., Neill, S., Jones, P., and Dascomb, K. (2019). Risk factors for community-acquired extended-Spectrum Beta-lactamase-producing Enterobacteriaceae infections-a retrospective study of symptomatic urinary tract Infections. Open Forum Infect. Dis. 6:357. doi: 10.1093/ofid/ofy357
Guenther, S., Ewers, C., and Wieler, L. H. (2011). Extended-Spectrum Beta-lactamases Producing E. coli in wildlife, yet another form of environmental pollution? Front. Microbiol. 2:246. doi: 10.3389/fmicb.2011.00246
Guitart-Matas, J., Espunyes, J., Illera, L., Gonzalez-Escalona, N., Ribas, M. P., Marco, I., et al. (2024). High-risk lineages of extended spectrum cephalosporinase producing Escherichia coli from Eurasian griffon vultures (Gyps fulvus) foraging in landfills in North-Eastern Spain. Sci. Total Environ. 909:168625. doi: 10.1016/j.scitotenv.2023.168625
Gundran, R. S., Cardenio, P. A., Villanueva, M. A., Sison, F. B., Benigno, C. C., and Kreausukon, K. (2019). Prevalence and distribution of blaCTX-M, blaSHV, blaTEM genes in extended-spectrum β-lactamase-producing E. coli isolates from broiler farms in the Philippines. BMC Vet. Res. 15, 15:227. doi: 10.1186/s12917-019-1975-9
Gupta, V., Ye, G., Olesky, M., Lawrence, K., Murray, J., and Yu, K. (2019). National prevalence estimates for resistant Enterobacteriaceae and Acinetobacter species in hospitalized patients in the United States. Int. J. Infect. Dis. 85, 203–211. doi: 10.1016/j.ijid.2019.06.017
Habib, I., Elbediwi, M., Mohamed, M. I., Ghazawi, A., Abdalla, A., and Khalifa, H. O. (2023). Enumeration, antimicrobial resistance and genomic characterization of extended-spectrum β-lactamases producing Escherichia coli from supermarket chicken meat in the United Arab Emirates. Int. J. Food Microbiol. 398:110224. doi: 10.1016/j.ijfoodmicro.2023.110224
Hammerum, A. M., Larsen, J., Andersen, V. D., Lester, C. H., Skytte, T. S. S., Hansen, F., et al. (2014). Characterization of extended-spectrum β-lactamase (ESBL)-producing Escherichia coli obtained from Danish pigs, pig farmers and their families from farms with high or no consumption of third- or fourth-generation cephalosporins. J. Antimicrob. Chemother. 69, 2650–2657. doi: 10.1093/jac/dku180
Handayani, K. S., Setiyono, A., Lukman, D. W., Pisestyani, H., and Rahayu, P. (2024). Distribution of extended-spectrum β-lactamase producing Escherichia coli genes in an integrated poultry-fish farming system in Bogor, Indonesia. Vet. World 17, 1596–1602. doi: 10.14202/vetworld.2024.1596-1602
Hasan, B., Laurell, K., Rakib, M. M., Ahlstedt, E., Hernandez, J., Caceres, M., et al. (2016). Fecal carriage of extended-Spectrum β-lactamases in healthy humans, poultry, and wild birds in León, Nicaragua-a shared Pool of blaCTX-M genes and possible interspecies clonal spread of extended-Spectrum β-lactamases-producing Escherichia coli. Microb. Drug Resist. 22, 682–687. doi: 10.1089/mdr.2015.0323
Hashemizadeh, Z., Mohebi, S., Kalantar-Neyestanaki, D., Mansouri, S., Hosseini-Nave, H., and Bazargani, A. (2019). Prevalence of plasmid-mediated quinolone resistance and ESBLs genes in Escherichia coli isolated from urinary tract infections and fecal samples in Southeast Iran. Gene Rep. 17:100487. doi: 10.1016/j.genrep.2019.100487
Hassen, B., Jouini, A., Elbour, M., Hamrouni, S., and Maaroufi, A. (2020). Detection of extended-Spectrum β-lactamases (ESBL) producing Enterobacteriaceae from fish trapped in the lagoon area of Bizerte, Tunisia. Biomed. Res. Int. 2020:812. doi: 10.1155/2020/7132812
Hernandez, J., Johansson, A., Stedt, J., Bengtsson, S., Porczak, A., Granholm, S., et al. (2013). Characterization and comparison of extended-spectrum β-lactamase (ESBL) resistance genotypes and population structure of Escherichia coli isolated from Franklin’s gulls (Leucophaeus pipixcan) and humans in Chile. PloS one. 8:e76150. doi: 10.1371/journal.pone.0076150
Hernando-Amado, S., Coque, T. M., Baquero, F., and Martínez, J. L. (2019). Defining and combating antibiotic resistance from one health and Global Health perspectives. Nat. Microbiol. 4, 1432–1442. doi: 10.1038/s41564-019-0503-9
Hiroi, M., Matsui, S., Kubo, R., Iida, N., Noda, Y., Kanda, T., et al. (2012). Factors for occurrence of extended-spectrum β-lactamase-producing Escherichia coli in broilers. J. Vet. Med. Sci. 74, 1635–1637. doi: 10.1292/jvms.11-0479
Hooban, B., Fitzhenry, K., O’Connor, L., Miliotis, G., Joyce, A., Chueiri, A., et al. (2022). A longitudinal survey of antibiotic-resistant Enterobacterales in the Irish environment, 2019–2020. Sci. Total Environ. 828:154488. doi: 10.1016/j.scitotenv.2022.154488
Hordijk, J., Farmakioti, E., Smit, L. A. M., Duim, B., Graveland, H., Theelen, M. J. P., et al. (2020). Fecal carriage of extended-Spectrum-β-lactamase/AmpC-producing Escherichia coli in horses. Appl. Environ. Microbiol. 86, e02590–e02519. doi: 10.1128/AEM.02590-19
Howard-Jones, A. R., Anver, S. R., Tsang, K., and Branley, J. (2022). Invasive extended-spectrum beta-lactamase (ESBL) related infection rates are not influenced by international travel restrictions. Pathology (Phila.) 54, 792–795. doi: 10.1016/j.pathol.2021.11.005
Hu, X., Gou, J., Guo, X., Cao, Z., Li, Y., Jiao, H., et al. (2018). Genetic contexts related to the diffusion of plasmid-mediated CTX-M-55 extended-spectrum beta-lactamase isolated from Enterobacteriaceae in China. Ann. Clin. Microbiol. Antimicrob. 17, 12–17. doi: 10.1186/S12941-018-0265-X
Huang, Y. T., Yeh, T. K., Chen, W. H., Shih, P. W., Mao, Y. C., Lu, M. C., et al. (2021). Genome analysis of Enterobacter hormaechei identified ISEcp1 in association with blaCTX-M-236, a new blaCTX-M variant, located both in the chromosome and a plasmid. J. Glob. Antimicrob. Resist. 25, 37–39. doi: 10.1016/J.JGAR.2021.02.026
Huijbers, P. M. C., Graat, E. A. M., Haenen, A. P. J., van Santen, M. G., van Essen-Zandbergen, A., Mevius, D. J., et al. (2014). Extended-spectrum and AmpC β-lactamase-producing Escherichia coli in broilers and people living and/or working on broiler farms: prevalence, risk factors and molecular characteristics. J. Antimicrob. Chemother. 69, 2669–2675. doi: 10.1093/jac/dku178
Huijbers, P. M. C., van Hoek, A. H. A. M., Graat, E. A. M., Haenen, A. P. J., Florijn, A., Hengeveld, P. D., et al. (2015). Methicillin-resistant Staphylococcus aureus and extended-spectrum and AmpC β-lactamase-producing Escherichia coli in broilers and in people living and/or working on organic broiler farms. Vet. Microbiol. 176, 120–125. doi: 10.1016/j.vetmic.2014.12.010
Imperial, I. C. V. J., and Ibana, J. A. (2016). Addressing the antibiotic resistance problem with probiotics: reducing the risk of its double-edged sword effect. Front. Microbiol. 7:e01983. doi: 10.3389/fmicb.2016.01983
Jakobsen, L., Bortolaia, V., Bielak, E., Moodley, A., Olsen, S. S., and Hansen, D. S. (2015). Limited similarity between plasmids encoding CTX-M-1 β-lactamase in Escherichia coli from humans, pigs, cattle, organic poultry layers and horses in Denmark. J. Glob. Antimicrob. Resist. 3, 132–136. doi: 10.1016/J.JGAR.2015.03.009
Jena, J., Debata, N. K., Sahoo, R. K., Gaur, M., and Subudhi, E. (2018). Molecular characterization of extended spectrum β-lactamase-producing Enterobacteriaceae strains isolated from a tertiary care hospital. Microb. Pathog. 115, 112–116. doi: 10.1016/J.MICPATH.2017.12.056
Jena, J., Sahoo, R. K., Debata, N. K., and Subudhi, E. (2017). Prevalence of TEM, SHV, and CTX-M genes of extended-spectrum β-lactamase-producing Escherichia coli strains isolated from urinary tract infections in adults. 3 Biotech 7:244. doi: 10.1007/S13205-017-0879-2
Johansson, V., Nykäsenoja, S., Myllyniemi, A.-L., Rossow, H., and Heikinheimo, A. (2022). Genomic characterization of ESBL/AmpC-producing and high-risk clonal lineages of Escherichia coli and Klebsiella pneumoniae in imported dogs with shelter and stray background. J. Glob. Antimicrob. Resist. 30, 183–190. doi: 10.1016/j.jgar.2022.05.021
Jones-Dias, D., Manageiro, V., Martins, A. P., Ferreira, E., and Caniça, M. (2016). New class 2 Integron In2-4 among IncI1-positive Escherichia coli isolates carrying ESBL and PMAβ genes from food animals in Portugal. Foodborne Pathog. Dis. 13, 36–39. doi: 10.1089/fpd.2015.1972
Jouini, A., Vinué, L., Slama, K. B., Sáenz, Y., Klibi, N., and Hammami, S. (2007). Characterization of CTX-M and SHV extended-spectrum β-lactamases and associated resistance genes in Escherichia coli strains of food samples in Tunisia. J. Antimicrob. Chemother. 60, 1137–1141. doi: 10.1093/JAC/DKM316
Kang, H., Kim, H., Kim, H., Jeon, J. H., Kim, S., Park, Y., et al. (2024). Genetic characteristics of extended-Spectrum Beta-lactamase-producing Salmonella isolated from retail meats in South Korea. J. Microbiol. Biotechnol. 34, 1101–1108. doi: 10.4014/jmb.2312.12018
Kantele, A., and Lääveri, T. (2021). Extended-spectrum beta-lactamase-producing strains among diarrhoeagenic Escherichia coli-prospective traveller study with literature review. J. Travel Med. 29, 1–13. doi: 10.1093/JTM/TAAB042
Karkaba, A., Grinberg, A., Benschop, J., and Pleydell, E. (2017). Characterisation of extended-spectrum β-lactamase and AmpC β-lactamase-producing Enterobacteriaceae isolated from companion animals in New Zealand. N. Z. Vet. J. 65, 105–112. doi: 10.1080/00480169.2016.1271730
Karlowsky, J. A., Lob, S. H., DeRyke, C. A., Siddiqui, F., Young, K., and Motyl, M. R. (2022). Prevalence of ESBL non-CRE Escherichia coli and Klebsiella pneumoniae among clinical isolates collected by the SMART global surveillance programme from 2015 to 2019. Int. J. Antimicrob. Agents 59:106535. doi: 10.1016/j.ijantimicag.2022.106535
Kawamura, K., Nagano, N., Suzuki, M., Wachino, J., Kimura, K., and Arakawa, Y. (2017). ESBL-producing Escherichia coli and its rapid rise among healthy people. Food Saf. 5, 122–150. doi: 10.14252/foodsafetyfscj.2017011
Kim, Y. J., Moon, J. S., Oh, D. H., Chon, J. W., Song, B. R., and Lim, J. S. (2018). Genotypic characterization of ESBL-producing E. coli from imported meat in South Korea. Food Res. Int. 107, 158–164. doi: 10.1016/J.FOODRES.2017.12.023
Koovapra, S., Bandyopadhyay, S., Das, G., Bhattacharyya, D., Banerjee, J., and Mahanti, A. (2016). Molecular signature of extended spectrum β-lactamase producing Klebsiella pneumoniae isolated from bovine milk in eastern and North-Eastern India. Infect. Genet. Evol. 44, 395–402. doi: 10.1016/j.meegid.2016.07.032
Korzeniewska, E., and Harnisz, M. (2013a). Beta-lactamase-producing Enterobacteriaceae in hospital effluents. J. Environ. Manag. 123, 1–7. doi: 10.1016/j.jenvman.2013.03.024
Korzeniewska, E., and Harnisz, M. (2013b). Extended-spectrum beta-lactamase (ESBL)-positive Enterobacteriaceae in municipal sewage and their emission to the environment. J. Environ. Manag. 128, 904–911. doi: 10.1016/j.jenvman.2013.06.051
Kraemer, J. G., Pires, J., Kueffer, M., Semaani, E., Endimiani, A., and Hilty, M. (2017). Prevalence of extended-spectrum β-lactamase-producing Enterobacteriaceae and methicillin-resistant Staphylococcus aureus in pig farms in Switzerland. Sci. Total Environ. 603-604, 401–405. doi: 10.1016/j.scitotenv.2017.06.110
Kurita, G., Tsuyuki, Y., Murata, Y., and Takahashi, T. (2019). Reduced rates of antimicrobial resistance in Staphylococcus intermedius group and Escherichia coli isolated from diseased companion animals in an animal hospital after restriction of antimicrobial use. J. Infect. Chemother. 25, 531–536. doi: 10.1016/j.jiac.2019.02.017
Kutilova, I., Medvecky, M., Leekitcharoenphon, P., Munk, P., Masarikova, M., and Davidova-Gerzova, L. (2021). Extended-spectrum beta-lactamase-producing Escherichia coli and antimicrobial resistance in municipal and hospital wastewaters in Czech Republic: culture-based and metagenomic approaches. Environ. Res. 193:110487. doi: 10.1016/j.envres.2020.110487
Kwok, K.-O., Chan, E., Chung, P.-H., Tang, A., Wei, W.-I., and Zhu, C. (2020). Prevalence and associated factors for carriage of Enterobacteriaceae producing ESBLs or carbapenemase and methicillin-resistant Staphylococcus aureus in Hong Kong community. J. Infect. 81, 242–247. doi: 10.1016/j.jinf.2020.05.033
Laupland, K. B., Church, D. L., Vidakovich, J., Mucenski, M., and Pitout, J. D. D. (2008). Community-onset extended-spectrum β-lactamase (ESBL) producing Escherichia coli: importance of international travel. J. Infect. 57, 441–448. doi: 10.1016/j.jinf.2008.09.034
Lee, S., Mir, R. A., Park, S. H., Kim, D., Kim, H.-Y., Boughton, R. K., et al. (2020). Prevalence of extended-spectrum β-lactamases in the local farm environment and livestock: challenges to mitigate antimicrobial resistance. Crit. Rev. Microbiol. 46, 1–14. doi: 10.1080/1040841X.2020.1715339
Lei, L., Rehman, M. U., Huang, S., Zhang, L., Wang, L., Mehmood, K., et al. (2018). Antimicrobial resistance and prevalence of diarrheagenic Escherichia coli (DEC) in diarrheic yaks of Tibetan plateau, China. Acta Trop. 182, 111–114. doi: 10.1016/j.actatropica.2018.02.022
Li, J., Cheng, W., Xu, L., Jiao, Y., Baig, S. A., and Chen, H. (2016). Occurrence and removal of antibiotics and the corresponding resistance genes in wastewater treatment plants: effluents’ influence to downstream water environment. Environ. Sci. Pollut. Res. 23, 6826–6835. doi: 10.1007/s11356-015-5916-2
Li, Q., Yang, J., Wang, M., Jia, R., Chen, S., Liu, M., et al. (2024). Global distribution and genomic characteristics analysis of avian-derived mcr-1-positive Escherichia coli. Ecotoxicol. Environ. Saf. 285:117109. doi: 10.1016/j.ecoenv.2024.117109
Li, Q., Zou, H., Wang, D., Zhao, L., Meng, M., Wang, Z., et al. (2023). Tracking spatio-temporal distribution and transmission of antibiotic resistance in aquatic environments by using ESBL-producing Escherichia coli as an indicator. J. Environ. Manag. 344:118534. doi: 10.1016/j.jenvman.2023.118534
Liao, X.-P., Liu, B.-T., Yang, Q.-E., Sun, J., Li, L., Fang, L.-X., et al. (2013). Comparison of plasmids coharboring 16s rrna methylase and extended-spectrum β-lactamase genes among Escherichia coli isolates from pets and poultry. J. Food Prot. 76, 2018–2023. doi: 10.4315/0362-028X.JFP-13-200
Lifshitz-Ziv, S.-N., ParizadeMiriam, E. B., Gordon, M., Taran, D., and Adler, A. (2018). Distinctiveness and similarities between extended-Spectrum β-lactamase-producing Escherichia coli isolated from cattle and the Community in Israel. Microb. Drug Resist. 24, 868–875. doi: 10.1089/MDR.2017.0407
Liu, T., Lee, S., Kim, M., Fan, P., Boughton, R. K., Boucher, C., et al. (2024). A study at the wildlife-livestock interface unveils the potential of feral swine as a reservoir for extended-spectrum β-lactamase-producing Escherichia coli. J. Hazard. Mater. 473:134694. doi: 10.1016/j.jhazmat.2024.134694
López-Vélez, R., Lebens, M., Bundy, L., Barriga, J., and Steffen, R. (2022). Bacterial travellers’ diarrhoea: a narrative review of literature published over the past 10 years. Travel Med. Infect. Dis. 47:102293. doi: 10.1016/j.tmaid.2022.102293
Loyola, S., Gutierrez, L. R., Horna, G., Petersen, K., Agapito, J., Osada, J., et al. (2016). Extended-spectrum β-lactamase–producing Enterobacteriaceae in cell phones of health care workers from Peruvian pediatric and neonatal intensive care units. Am. J. Infect. Control 44, 910–916. doi: 10.1016/j.ajic.2016.02.020
Lübbert, C., Straube, L., Stein, C., Makarewicz, O., Schubert, S., Mössner, J., et al. (2015). Colonization with extended-spectrum beta-lactamase-producing and carbapenemase-producing Enterobacteriaceae in international travelers returning to Germany. Int. J. Med. Microbiol. 305, 148–156. doi: 10.1016/j.ijmm.2014.12.001
Luo, Y., Tan, L., Zhang, H., Bi, W., Zhao, L., Wang, X., et al. (2022). Characteristics of wild bird Resistomes and dissemination of antibiotic resistance genes in interconnected bird-habitat systems revealed by similarity of blaTEM polymorphic sequences. Environ. Sci. Technol. 56, 15084–15095. doi: 10.1021/acs.est.2c01633
Maamar, E., Hammami, S., Alonso, C. A., Dakhli, N., Abbassi, M. S., Ferjani, S., et al. (2016). High prevalence of extended-spectrum and plasmidic AmpC beta-lactamase-producing Escherichia coli from poultry in Tunisia. Int. J. Food Microbiol. 231, 69–75. doi: 10.1016/j.ijfoodmicro.2016.05.001
Maataoui, N., Mayet, A., Duron, S., Delacour, H., Mentré, F., Laouenan, C., et al. (2019). High acquisition rate of extended-spectrum β-lactamase-producing Enterobacteriaceae among French military personnel on mission abroad, without evidence of inter-individual transmission. Clin. Microbiol. Infect. 25, 631.e1–631.e9. doi: 10.1016/j.cmi.2018.07.030
Marando, R., Seni, J., Mirambo, M. M., Falgenhauer, L., Moremi, N., Mushi, M. F., et al. (2018). Predictors of the extended-spectrum-beta lactamases producing Enterobacteriaceae neonatal sepsis at a tertiary hospital, Tanzania. Int. J. Med. Microbiol. 308, 803–811. doi: 10.1016/j.ijmm.2018.06.012
Marcelino, V. R., Wille, M., Hurt, A. C., González-Acuña, D., Klaassen, M., Schlub, T. E., et al. (2019). Meta-transcriptomics reveals a diverse antibiotic resistance gene pool in avian microbiomes. BMC Biol. 17:31. doi: 10.1186/s12915-019-0649-1
March, A., Aschbacher, R., Sleghel, F., Soelva, G., Kaczor, M., Migliavacca, R., et al. (2017). Colonization of residents and staff of an Italian long-term care facility and an adjacent acute care hospital geriatric unit by multidrug-resistant bacteria. New Microbiol. 40, 258–263
Martak, D., Guther, J., Verschuuren, T. D., Valot, B., Conzelmann, N., Bunk, S., et al. (2022). Populations of extended-spectrum β-lactamase-producing Escherichia coli and Klebsiella pneumoniae are different in human-polluted environment and food items: a multicentre European study. Clin. Microbiol. Infect. 28, 447.e7–447.e14. doi: 10.1016/j.cmi.2021.07.022
Matakone, M., Founou, R. C., Founou, L. L., Dimani, B. D., Koudoum, P. L., Fonkoua, M. C., et al. (2024). Multi-drug resistant (MDR) and extended-spectrum β-lactamase (ESBL) producing Escherichia coli isolated from slaughtered pigs and slaughterhouse workers in Yaoundé, Cameroon. One Health 19:100885. doi: 10.1016/j.onehlt.2024.100885
Mateo-Sagasta, J., Medlicott, K., Qadir, M., Raschid-Sally, L., Drechsel, P., and Liebe, J. (2013). Safe use of wastewater in agriculture. Proc. Series 11:80.
Melo, L. C., Oresco, C., Leigue, L., Netto, H. M., Melville, P. A., Benites, N. R., et al. (2018). Prevalence and molecular features of ESBL/pAmpC-producing Enterobacteriaceae in healthy and diseased companion animals in Brazil. Vet. Microbiol. 221, 59–66. doi: 10.1016/j.vetmic.2018.05.017
Menezes, J., Frosini, S. M., Belas, A., Marques, C., da Silva, J. M., Amaral, A. J., et al. (2023). Longitudinal study of ESBL/AmpC-producing Enterobacterales strains sharing between cohabiting healthy companion animals and humans in Portugal and in the United Kingdom. Eur. J. Clin. Microbiol. Infect. Dis. 42, 1011–1024. doi: 10.1007/s10096-023-04629-2
Meurs, L., Lempp, F. S., Lippmann, N., Trawinski, H., Rodloff, A. C., Eckardt, M., et al. (2019). Intestinal colonization with extended-spectrum beta-lactamase producing Enterobacterales (ESBL-PE) during long distance travel: a cohort study in a German travel clinic (2016–2017). Travel Med. Infect. Dis. 33:101521. doi: 10.1016/j.tmaid.2019.101521
Michael, G. B., Kaspar, H., Siqueira, A. K., de Freitas Costa, E., Corbellini, L. G., Kadlec, K., et al. (2017). Extended-spectrum β-lactamase (ESBL)-producing Escherichia coli isolates collected from diseased food-producing animals in the GERM-vet monitoring program 2008-2014. Vet. Microbiol. 200, 142–150. doi: 10.1016/j.vetmic.2016.08.023
Miclot, C., Antoniotti, G., and Forestier, E. (2015). Study of family physicians’ practices in case of urinary tract infections caused by enterobacteriaceae secreting extended-spectrum-beta-lactamase. Méd. Mal. Infect. 45, 78–83. doi: 10.1016/j.medmal.2015.01.004
Miller, E. A., Ponder, J. B., Willette, M., Johnson, T. J., and VanderWaal, K. L. (2020). Merging metagenomics and spatial epidemiology to understand the distribution of antimicrobial resistance genes from Enterobacteriaceae in wild owls. Appl. Environ. Microbiol. 86, e00571–e00520. doi: 10.1128/AEM.00571-20
Mizuno, Y., Miura, Y., Yamaguchi, T., and Matsumoto, T. (2016). Extended-spectrum β-lactamase-producing Enterobacteriaceae colonisation in long-term overseas business travellers. Travel Med. Infect. Dis. 14, 561–567. doi: 10.1016/j.tmaid.2016.09.012
Mobasseri, G., Teh, C. S. J., Ooi, P. T., Tan, S. C., and Thong, K. L. (2019). Molecular characterization of multidrug-resistant and extended-Spectrum Beta-lactamase-producing Klebsiella pneumoniae isolated from swine farms in Malaysia. Microb. Drug Resist. Larchmt. 25, 1087–1098. doi: 10.1089/mdr.2018.0184
Moran, R. A., Baomo, L., Doughty, E. L., Guo, Y., Ba, X., van Schaik, W., et al. (2023). Extended-Spectrum β-lactamase genes traverse the Escherichia coli populations of intensive care unit patients, staff, and environment. Microbiol. Spectr. 11:e0507422. doi: 10.1128/spectrum.05074-22
Moremi, N., Claus, H., Rutta, L., Frosch, M., Vogel, U., and Mshana, S. E. (2018). High carriage rate of extended-spectrum beta-lactamase-producing Enterobacteriaceae among patients admitted for surgery in Tanzanian hospitals with a low rate of endogenous surgical site infections. J. Hosp. Infect. 100, 47–53. doi: 10.1016/j.jhin.2018.05.017
Müller, A., Jansen, W., Grabowski, N. T., Monecke, S., Ehricht, R., and Kehrenberg, C. (2018). ESBL- and AmpC-producing Escherichia coli from legally and illegally imported meat: characterization of isolates brought into the EU from third countries. Int. J. Food Microbiol. 283, 52–58. doi: 10.1016/j.ijfoodmicro.2018.06.009
Muloi, D. M., Jauneikaite, E., Anjum, M. F., Essack, S. Y., Singleton, D. A., Kasudi, M. R., et al. (2023). Exploiting genomics for antimicrobial resistance surveillance at one health interfaces. Lancet Microbe 4, e1056–e1062. doi: 10.1016/S2666-5247(23)00284-7
Na, S. H., Moon, D. C., Kang, H. Y., Song, H. J., Kim, S. J., Choi, J. H., et al. (2020). Molecular characteristics of extended-spectrum β-lactamase/AmpC-producing Salmonella enterica serovar Virchow isolated from food-producing animals during 2010–2017 in South Korea. Int. J. Food Microbiol. 322:108572. doi: 10.1016/j.ijfoodmicro.2020.108572
Nakayama, T., Kumeda, Y., Kawahara, R., and Yamamoto, Y. (2019). Quantification and long-term carriage study of human extended-spectrum/AmpC β-lactamase-producing Escherichia coli after international travel to Vietnam. J. Glob. Antimicrob. Resist. 21, 229–234. doi: 10.1016/j.jgar.2019.11.001
Neemann, K., Olateju, E. K., Izevbigie, N., Akaba, G., Olanipekun, G. M., Richard, J. C., et al. (2020). Neonatal outcomes associated with maternal recto-vaginal colonization with extended-spectrum β-lactamase producing Enterobacteriaceae in Nigeria: a prospective, cross-sectional study. Clin. Microbiol. Infect. 26, 463–469. doi: 10.1016/j.cmi.2019.07.013
Neffe, L., Abendroth, L., Bautsch, W., Häussler, S., and Tomasch, J. (2022). High plasmidome diversity of extended-spectrum beta-lactam-resistant Escherichia coli isolates collected during one year in one community hospital. Genomics 114:110368. doi: 10.1016/j.ygeno.2022.110368
Nguyen, T. T. H., Bourigault, C., Guillet, V., Buttes, A.-C. G., des Montassier, E., Batard, E., et al. (2019). Association between excreta management and incidence of extended-spectrum β-lactamase-producing Enterobacteriaceae: role of healthcare workers’ knowledge and practices. J. Hosp. Infect. 102, 31–36. doi: 10.1016/j.jhin.2018.12.006
Nguyen, V. T., Jamrozy, D., Matamoros, S., Carrique-Mas, J. J., Ho, H. M., Thai, Q. H., et al. (2019). Limited contribution of non-intensive chicken farming to ESBL-producing Escherichia coli colonization in humans in Vietnam: an epidemiological and genomic analysis. J. Antimicrob. Chemother. 74, 561–570. doi: 10.1093/jac/dky506
Nnadozie, C. F., and Odume, O. N. (2019). Freshwater environments as reservoirs of antibiotic resistant bacteria and their role in the dissemination of antibiotic resistance genes. Environ. Pollut. 254:113067. doi: 10.1016/j.envpol.2019.113067
Nüesch-Inderbinen, M., Kindle, P., Baschera, M., Liljander, A., Jores, J., Corman, V. M., et al. (2020). Antimicrobial resistant and extended-spectrum β-lactamase (ESBL) producing Escherichia coli isolated from fecal samples of African dromedary camels. Sci. Afr. 7:e00274. doi: 10.1016/j.sciaf.2020.e00274
Oikarainen, P. E., Pohjola, L. K., Pietola, E. S., and Heikinheimo, A. (2019). Direct vertical transmission of ESBL/pAmpC-producing Escherichia coli limited in poultry production pyramid. Vet. Microbiol. 231, 100–106. doi: 10.1016/j.vetmic.2019.03.001
Ojemaye, M. O., Adefisoye, M. A., and Okoh, A. I. (2020). Nanotechnology as a viable alternative for the removal of antimicrobial resistance determinants from discharged municipal effluents and associated watersheds: a review. J. Environ. Manag. 275:111234. doi: 10.1016/j.jenvman.2020.111234
Oka, K., Tetsuka, N., Morioka, H., Iguchi, M., Kawamura, K., Hayashi, K., et al. (2022). Genetic and epidemiological analysis of ESBL-producing Klebsiella pneumoniae in three Japanese university hospitals. J. Infect. Chemother. 28, 1286–1294. doi: 10.1016/j.jiac.2022.05.013
Olaru, I. D., Walther, B., and Schaumburg, F. (2023). Zoonotic sources and the spread of antimicrobial resistance from the perspective of low and middle-income countries. Infect. Dis. Poverty 12:59. doi: 10.1186/s40249-023-01113-z
Oravcova, V., Ghosh, A., Zurek, L., Bardon, J., Guenther, S., Cizek, A., et al. (2013). Vancomycin-resistant enterococci in rooks (Corvus frugilegus) wintering throughout Europe. Environ. Microbiol. 15, 548–556. doi: 10.1111/1462-2920.12002
Otter, J. A., Natale, A., Batra, R., Tosas Auguet, O., Dyakova, E., Goldenberg, S. D., et al. (2019). Individual- and community-level risk factors for ESBL Enterobacteriaceae colonization identified by universal admission screening in London. Clin. Microbiol. Infect. 25, 1259–1265. doi: 10.1016/j.cmi.2019.02.026
Päivärinta, M., Latvio, S., Fredriksson-Ahomaa, M., and Heikinheimo, A. (2020). Whole genome sequence analysis of antimicrobial resistance genes, multilocus sequence types and plasmid sequences in ESBL/AmpC Escherichia coli isolated from broiler caecum and meat. Int. J. Food Microbiol. 315:108361. doi: 10.1016/j.ijfoodmicro.2019.108361
Päivärinta, M., Pohjola, L., Fredriksson-Ahomaa, M., and Heikinheimo, A. (2016). Low occurrence of extended-Spectrum β-lactamase-producing Escherichia coli in Finnish food-producing animals. Zoonoses Public Health 63, 624–631. doi: 10.1111/zph.12277
Palmeira, J. D., Haenni, M., Metayer, V., Madec, J. Y., and Ferreira, H. M. N. (2020). Epidemic spread of IncI1/pST113 plasmid carrying the extended-Spectrum Beta-lactamase (ESBL) blaCTX-M-8 gene in Escherichia coli of Brazilian cattle. Vet. Microbiol. 243:108629. doi: 10.1016/j.vetmic.2020.108629
Parzygnat, J. L., Crespo, R., Koci, M. D., Dunn, R. R., Harden, L., Fosnaught, M., et al. (2024). Widespread prevalence of plasmid-mediated blaCTX-M type extended-spectrum beta-lactamase Escherichia coli in backyard broiler production systems in the United States. PLoS One 19:e0304599. doi: 10.1371/journal.pone.0304599
Paterson, D. L., and Bonomo, R. A. (2005). Extended-Spectrum β-lactamases: a clinical update. Clin. Microbiol. Rev. 18, 657–686. doi: 10.1128/CMR.18.4.657-686.2005
Patjas, A., and Kantele, A. (2022). International travel and travelers’ diarrhea – increased risk of urinary tract infection. Travel Med. Infect. Dis. 48:102331. doi: 10.1016/j.tmaid.2022.102331
Patjas, A., Martelius, A., Ollgren, J., and Kantele, A. (2024). International travel increases risk of urinary tract infection caused by extended-spectrum beta-lactamase-producing Enterobacterales-three-arm case-control study. J. Travel Med. 31:taad155. doi: 10.1093/jtm/taad155
Peñalva, G., Fernández-Urrusuno, R., Turmo, J. M., Hernández-Soto, R., Pajares, I., Carrión, L., et al. (2020). Long-term impact of an educational antimicrobial stewardship programme in primary care on infections caused by extended-spectrum β-lactamase-producing Escherichia coli in the community: an interrupted time-series analysis. Lancet Infect. Dis. 20, 199–207. doi: 10.1016/S1473-3099(19)30573-0
Perera, P. D. V. M., Gamage, S., de Silva, H. S. M., Jayatilleke, S. K., de Silva, N., Aydin, A., et al. (2022). Phenotypic and genotypic distribution of ESBL, AmpC β-lactamase and carbapenemase-producing Enterobacteriaceae in community acquired (CA-UTI) and hospital acquired urinary tract infections (HA-UTI) in Sri Lanka. J. Glob. Antimicrob. Resist. 30, 115–122. doi: 10.1016/j.jgar.2022.05.024
Perestrelo, S., Amaro, A., Brouwer, M. S. M., Clemente, L., Ribeiro Duarte, A. S., Kaesbohrer, A., et al. (2023). Building an international one health strain level database to characterise the epidemiology of AMR threats: ESBL-AmpC Producing E. coli as an example—challenges and perspectives. Antibiotics 12:552. doi: 10.3390/antibiotics12030552
Piccirilli, A., Perilli, M., Piccirilli, V., Segatore, B., Amicosante, G., Maccacaro, L., et al. (2020). Molecular characterization of carbapenem-resistant Klebsiella pneumoniae ST14 and ST512 causing bloodstream infections in ICU and surgery wards of a tertiary university hospital of Verona (northern Italy): co-production of KPC-3, OXA-48, and CTX-M-15 β-lactamases. Diagn. Microbiol. Infect. Dis. 96:114968. doi: 10.1016/J.DIAGMICROBIO.2019.114968
Pitout, J. D., and Laupland, K. B. (2008). Extended-spectrum β-lactamase-producing Enterobacteriaceae: an emerging public-health concern. Lancet Infect. Dis. 8, 159–166. doi: 10.1016/S1473-3099(08)70041-0
Poudel, A., Hathcock, T., Butaye, P., Kang, Y., Price, S., Macklin, K., et al. (2019). Multidrug-resistant Escherichia coli, Klebsiella pneumoniae and Staphylococcus spp. in houseflies and blowflies from farms and their environmental settings. Int. J. Environ. Res. Public Health 16:583. doi: 10.3390/ijerph16193583
Projahn, M., Daehre, K., Roesler, U., and Friese, A. (2017). Extended-Spectrum-Beta-lactamase- and plasmid-encoded Cephamycinase-producing Enterobacteria in the broiler hatchery as a potential mode of Pseudo-vertical transmission. Appl. Environ. Microbiol. 83, e02364–e02316. doi: 10.1128/AEM.02364-16
Puljko, A., Babić, I., Rozman, S. D., Barišić, I., Jelić, M., Maravić, A., et al. (2024). Treated municipal wastewater as a source of high-risk and emerging multidrug-resistant clones of E. Coli and other Enterobacterales producing extended-spectrum β-lactamases. Environ. Res. 243:117792. doi: 10.1016/j.envres.2023.117792
Raffelsberger, N., Buczek, D. J., Svendsen, K., Småbrekke, L., Pöntinen, A. K., Löhr, I. H., et al. (2023). Community carriage of ESBL-producing Escherichia coli and Klebsiella pneumoniae: a cross-sectional study of risk factors and comparative genomics of carriage and clinical isolates. mSphere 8:e0002523. doi: 10.1128/msphere.00025-23
Rahman, S., Ali, T., Muhammad, N., Umer, T., and Ayaz, S. (2018). Characterization and mechanism of dissemination of extended spectrum beta lactamase producers Escherichia coli in food producing animals in Pakistan and China, in 2018 15th International Bhurban Conference on Applied Sciences and Technology (IBCAST). Presented at the 2018 15th International Bhurban Conference on Applied Sciences and Technology (IBCAST), pp. 203–214.
Ramos, C. A., Ferreira, J. C., Ballaben, A. S., Filho, R. A. C. P., and Darini, A. L. D. C. (2024). Analysis of antibiotic resistance in gram-negative bacilli in wild and exotic healthy birds in Brazil: a warning sign. Vet. Microbiol. 296:110196. doi: 10.1016/j.vetmic.2024.110196
Randall, L. P., Clouting, C., Horton, R. A., Coldham, N. G., Wu, G., Clifton-Hadley, F. A., et al. (2011). Prevalence of Escherichia coli carrying extended-spectrum β-lactamases (CTX-M and TEM-52) from broiler chickens and turkeys in Great Britain between 2006 and 2009. J. Antimicrob. Chemother. 66, 86–95. doi: 10.1093/jac/dkq396
Rehman, M. U., Zhang, H., Iqbal, M. K., Nabi, F., Huang, S., Lan, Y., et al. (2017). Antibiotic resistance of Escherichia coli in free-ranging yaks (Bos grunniens) from Tibetan plateau, China. Pak. Vet. J. 37, 139–144.
Riccio, M. E., Verschuuren, T., Conzelmann, N., Martak, D., Meunier, A., Salamanca, E., et al. (2021). Household acquisition and transmission of extended-spectrum β-lactamase (ESBL)-producing Enterobacteriaceae after hospital discharge of ESBL-positive index patients. Clin. Microbiol. Infect. 27, 1322–1329. doi: 10.1016/J.CMI.2020.12.024
Richter, L., du Plessis, E. M., Duvenage, S., and Korsten, L. (2020). Occurrence, phenotypic and molecular characterization of extended-Spectrum- and AmpC- β-lactamase producing Enterobacteriaceae isolated from selected commercial spinach supply chains in South Africa. Front. Microbiol. 11:638. doi: 10.3389/fmicb.2020.00638
Rodríguez-Baño, J., Paño-Pardo, J. R., Alvarez-Rocha, L., Asensio, Á., Calbo, E., Cercenado, E., et al. (2012). Programas de optimización de uso de antimicrobianos (PROA) en hospitales españoles: documento de consenso GEIH-SEIMC, SEFH y SEMPSPH. Microbiol. Clín. 30, 22.e1–22.e23. doi: 10.1016/j.eimc.2011.09.018
Rosantia, S., Higa, T., Yagi, N., Tokunaga, T., Higa, S., Yakabi, Y., et al. (2020). Characterization of CTX-M-type-extended-spectrum beta-lactamase (ESBL)-producing Enterobacteriaceae isolated from Indonesian undergraduate medical students of a university in Surabaya, Indonesia. J. Infect. Chemother. 26, 575–581. doi: 10.1016/j.jiac.2020.01.010
Ruzauskas, M., and Vaskeviciute, L. (2016). Detection of the mcr-1 gene in Escherichia coli prevalent in the migratory bird species Larus argentatus. J. Antimicrob. Chemother. 71, 2333–2334. doi: 10.1093/jac/dkw245
Schages, L., Wichern, F., Kalscheuer, R., and Bockmühl, D. (2020). Winter is coming – impact of temperature on the variation of beta-lactamase and mcr genes in a wastewater treatment plant. Sci. Total Environ. 712:136499. doi: 10.1016/j.scitotenv.2020.136499
Schaumburg, F., Sertic, S. M., Correa-Martinez, C., Mellmann, A., Köck, R., and Becker, K. (2019). Acquisition and colonization dynamics of antimicrobial-resistant bacteria during international travel: a prospective cohort study. Clin. Microbiol. Infect. 25, 1287.e1–1287.e7. doi: 10.1016/j.cmi.2019.03.002
Schill, F., Abdulmawjood, A., Klein, G., and Reich, F. (2017). Prevalence and characterization of extended-spectrum β-lactamase (ESBL) and AmpC β-lactamase producing Enterobacteriaceae in fresh pork meat at processing level in Germany. Int. J. Food Microbiol. 257, 58–66. doi: 10.1016/j.ijfoodmicro.2017.06.010
Seenama, C., Thamlikitkul, V., and Ratthawongjirakul, P. (2019). Multilocus sequence typing and blaESBL characterization of extended-spectrum beta-lactamase-producing Escherichia coli isolated from healthy humans and swine in northern Thailand. Infect. Drug Resist. 12, 2201–2214. doi: 10.2147/IDR.S209545
Sen, K., Berglund, T., Soares, M. A., Taheri, B., Ma, Y., Khalil, L., et al. (2019). Antibiotic resistance of E. coli isolated from a constructed wetland dominated by a crow roost, with emphasis on ESBL and AmpC Containing E. coli. Front. Microbiol. 10:1034. doi: 10.3389/fmicb.2019.01034
Shin, S. W., Jung, M., Won, H. G., Belaynehe, K. M., Yoon, I. J., and Yoo, H. S. (2017). Characteristics of transmissible CTX-M- and CMY-type β-lactamase-producing Escherichia coli isolates collected from pig and chicken farms in South Korea. J. Microbiol. Biotechnol. 27, 1716–1723. doi: 10.4014/jmb.1610.10006
Silva, M. M., Sellera, F. P., Fernandes, M. R., Moura, Q., Garino, F., Azevedo, S. S., et al. (2018). Genomic features of a highly virulent, ceftiofur-resistant, CTX-M-8-producing Escherichia coli ST224 causing fatal infection in a domestic cat. J. Glob. Antimicrob. Resist. 15, 252–253. doi: 10.1016/j.jgar.2018.10.023
Silva, I., Tacão, M., Tavares, R. D. S., Miranda, R., Araújo, S., Manaia, C. M., et al. (2018). Fate of cefotaxime-resistant Enterobacteriaceae and ESBL-producers over a full-scale wastewater treatment process with UV disinfection. Sci. Total Environ. 639, 1028–1037. doi: 10.1016/j.scitotenv.2018.05.229
Singh, N. S., Singhal, N., and Virdi, J. S. (2018). Genetic environment of blaTEM-1, blaCTX-M-15, blaCMY-42 and characterization of Integrons of Escherichia coli isolated from an Indian urban aquatic environment. Front. Microbiol. 9:382. doi: 10.3389/FMICB.2018.00382
Smyth, C., O’Flaherty, A., Walsh, F., and Do, T. T. (2020). Antibiotic resistant and extended-spectrum β-lactamase producing faecal coliforms in wastewater treatment plant effluent. Environ. Pollut. 262:114244. doi: 10.1016/j.envpol.2020.114244
Snaith, A. E., Dunn, S. J., Moran, R. A., Newton, P. N., Dance, D. A. B., Davong, V., et al. (2023). The highly diverse plasmid population found in Escherichia coli colonizing travellers to Laos and its role in antimicrobial resistance gene carriage. Microb. Genom. 9, –mgen001000. doi: 10.1099/mgen.0.001000
Solà-Ginés, M., González-López, J. J., Cameron-Veas, K., Piedra-Carrasco, N., Cerdà-Cuéllar, M., and Migura-Garcia, L. (2015). Houseflies (Musca domestica) as vectors for extended-Spectrum β-lactamase-producing Escherichia coli on Spanish broiler farms. Appl. Environ. Microbiol. 81, 3604–3611. doi: 10.1128/AEM.04252-14
Stanley, I. J., Kajumbula, H., Bazira, J., Kansiime, C., Rwego, I. B., Asiimwe, B. B., et al. (2018). Multidrug resistance among Escherichia coli and Klebsiella pneumoniae carried in the gut of out-patients from pastoralist communities of Kasese district, Uganda. PLoS One 13:93. doi: 10.1371/journal.pone.0200093
Stokes, J. M., Yang, K., Swanson, K., Jin, W., Cubillos-Ruiz, A., Donghia, N. M., et al. (2020). A deep learning approach to antibiotic discovery. Cell 180, 688–702.e13. doi: 10.1016/j.cell.2020.01.021
Strysko, J. P., Mony, V., Cleveland, J., Siddiqui, H., Homel, P., and Gagliardo, C. (2016). International travel is a risk factor for extended-spectrum β-lactamase-producing Enterobacteriaceae acquisition in children: a case-case-control study in an urban U.S. hospital. Travel Med. Infect. Dis. 14, 568–571. doi: 10.1016/j.tmaid.2016.11.012
Sultan, I., Ali, A., Gogry, F. A., Rather, I. A., Sabir, J. S. M., and Haq, Q. M. R. (2020). Bacterial isolates harboring antibiotics and heavy-metal resistance genes co-existing with mobile genetic elements in natural aquatic water bodies. Saudi J. Biol. Sci. 27, 2660–2668. doi: 10.1016/J.SJBS.2020.06.002
Sultan, I., Siddiqui, M. T., Gogry, F. A., Haq, Q., and Mohd, R. (2022). Molecular characterization of resistance determinants and mobile genetic elements of ESBL producing multidrug-resistant bacteria from freshwater lakes in Kashmir, India. Sci. Total Environ. 827:154221. doi: 10.1016/j.scitotenv.2022.154221
Tamang, M. D., Nam, H.-M., Gurung, M., Jang, G.-C., Kim, S.-R., Jung, S.-C., et al. (2013). Molecular characterization of CTX-M β-lactamase and associated addiction systems in Escherichia coli circulating among cattle, farm workers, and the farm environment. Appl. Environ. Microbiol. 79, 3898–3905. doi: 10.1128/AEM.00522-13
Tamta, S., Vinodh Kumar, O. R., Pruthvishree, B. S., Karthikeyan, R., Rupner, R. N., Chethan, G. E., et al. (2020). Faecal carriage of extended spectrum beta-lactamase (ESBL) and New Delhi metallo beta-lactamase (NDM) producing Escherichia coli between piglets and pig farmworkers. Comp. Immunol. Microbiol. Infect. Dis. 73:101564. doi: 10.1016/j.cimid.2020.101564
Tan, W., Lu, Y., Zhu, Z., Xu, Z., Zhang, Y., Huang, Q., et al. (2023). Cotransfer of resistance to cephalosporins, colistin, and fosfomycin mediated by an IncHI2/pSH16G4928-like plasmid in ESBL-producing monophasic Salmonella Typhimurium strains of pig origin. J. Appl. Microbiol. 134:lxac060. doi: 10.1093/jambio/lxac060
Tansawai, U., Walsh, T. R., and Niumsup, P. R. (2019). Extended spectrum ß-lactamase-producing Escherichia coli among backyard poultry farms, farmers, and environments in Thailand. Poult. Sci. 98, 2622–2631. doi: 10.3382/ps/pez009
Telling, K., Brauer, A., Laht, M., Kalmus, P., Toompere, K., Kisand, V., et al. (2020). Characteristics of extended-Spectrum Beta-lactamase-producing Enterobacteriaceae and contact to animals in Estonia. Microorganisms 8:1130. doi: 10.3390/microorganisms8081130
Tewari, R., Mitra, S., Ganaie, F., Das, S., Chakraborty, A., Venugopal, N., et al. (2019). Dissemination and characterisation of Escherichia coli producing extended-spectrum β-lactamases, AmpC β-lactamases and metallo-β-lactamases from livestock and poultry in Northeast India: a molecular surveillance approach. J. Glob. Antimicrob. Resist. 17, 209–215. doi: 10.1016/j.jgar.2018.12.025
Tian, S. F., Chu, Y. Z., Chen, B. Y., Nian, H., and Shang, H. (2011). ISEcp1 element in association with blaCTX-M genes of E. coli that produce extended-spectrum β-lactamase among the elderly in community settings. Enferm. Infecc. Microbiol. Clin. 29, 731–734. doi: 10.1016/J.EIMC.2011.07.011
Tian, T., Dai, S., Liu, D., Wang, Y., Qiao, W., Yang, M., et al. (2022). Occurrence and transfer characteristics of blaCTX-M genes among Escherichia coli in anaerobic digestion systems treating swine waste. Sci. Total Environ. 834:155321. doi: 10.1016/j.scitotenv.2022.155321
Tokinobu, A., Yamakawa, M., Tsuda, T., Matsushita, N., and Hashizume, M. (2020). Japanese tourists travelling in India have poor pre-travel preparedness. Travel Med. Infect. Dis. 33:101417. doi: 10.1016/j.tmaid.2019.05.003
Tong, P., Sun, Y., Ji, X., Du, X., Guo, X., Liu, J., et al. (2015). Characterization of antimicrobial resistance and extended-Spectrum β-lactamase genes in Escherichia coli isolated from chickens. Foodborne Pathog. Dis. 12, 345–352. doi: 10.1089/fpd.2014.1857
Toombs-Ruane, L. J., Benschop, J., French, N. P., Biggs, P. J., Midwinter, A. C., Marshall, J. C., et al. (2020). Carriage of extended-Spectrum-Beta-lactamase- and AmpC Beta-lactamase-producing Escherichia coli strains from humans and pets in the same households. Appl. Environ. Microbiol. 86, e01613–e01620. doi: 10.1128/AEM.01613-20
Toriro, R., Pallett, S. J. C., Nevin, W., Ross, T. M., Hale, I., Routledge, M., et al. (2024). Prevalence of extended-spectrum β-lactamase-producing Enterobacterales and carbapenemase-resistant Enterobacterales in British military cohorts. BMJ Military Health. military-2024-002837. doi: 10.1136/military-2024-002837
Torres, R. T., Cunha, M. V., Araujo, D., Ferreira, H., Fonseca, C., and Palmeira, J. D. (2022). A walk on the wild side: wild ungulates as potential reservoirs of multi-drug resistant bacteria and genes, including Escherichia coli harbouring CTX-M beta-lactamases. Environ. Pollut. 306:119367. doi: 10.1016/j.envpol.2022.119367
Udikovic-Kolic, N., Wichmann, F., Broderick, N. A., and Handelsman, J. (2014). Bloom of resident antibiotic-resistant bacteria in soil following manure fertilization. Proc. Natl. Acad. Sci. 111, 15202–15207. doi: 10.1073/pnas.1409836111
Umeda, K., Hase, A., Matsuo, M., Horimoto, T., and Ogasawara, J. (2019). Prevalence and genetic characterization of cephalosporin-resistant Enterobacteriaceae among dogs and cats in an animal shelter. J. Med. Microbiol. 68, 339–345. doi: 10.1099/jmm.0.000933
Valizadeh, S., Yousefi, B., Abdolshahi, A., Emadi, A., and Eslami, M. (2021). Determination of genetic relationship between environmental Escherichia coli with PFGE and investigation of IS element in blaCTX-M gene of these isolates. Microb. Pathog. 159:105154. doi: 10.1016/J.MICPATH.2021.105154
Van den Bunt, G., Fluit, A. C., Spaninks, M. P., Timmerman, A. J., Geurts, Y., Kant, A., et al. (2020). Faecal carriage, risk factors, acquisition and persistence of ESBL-producing Enterobacteriaceae in dogs and cats and co-carriage with humans belonging to the same household. J. Antimicrob. Chemother. 75, 342–350. doi: 10.1093/jac/dkz462
Vandecraen, J., Chandler, M., Aertsen, A., and Houdt, R.Van (2017). The impact of insertion sequences on bacterial genome plasticity and adaptability. 43, 709–730. doi: 10.1080/1040841X.2017.1303661
Vatovec, C., Kolodinsky, J., Callas, P., Hart, C., and Gallagher, K. (2021). Pharmaceutical pollution sources and solutions: survey of human and veterinary medication purchasing, use, and disposal. J. Environ. Manag. 285:112106. doi: 10.1016/j.jenvman.2021.112106
Velasco-Arnaiz, E., Simó-Nebot, S., Ríos-Barnés, M., López Ramos, M. G., Monsonís, M., Urrea-Ayala, M., et al. (2020). Benefits of a pediatric antimicrobial stewardship program in antimicrobial use and quality of prescriptions in a referral Children’s hospital. J. Pediatr. 225, 222–230.e1. doi: 10.1016/j.jpeds.2020.06.008
Velasova, M., Smith, R. P., Lemma, F., Horton, R. A., Duggett, N. A., Evans, J., et al. (2019). Detection of extended-spectrum β-lactam, AmpC and carbapenem resistance in Enterobacteriaceae in beef cattle in Great Britain in 2015. J. Appl. Microbiol. 126, 1081–1095. doi: 10.1111/jam.14211
Wang, Z., Jiang, Y., Xu, H., Jiao, X., Wang, J., Li, Q., et al. (2023). Poultry production as the main reservoir of ciprofloxacin- and tigecycline-resistant extended-spectrum β-lactamase (ESBL)-producing Salmonella enterica serovar Kentucky ST198.2-2 causing human infections in China. Appl. Environ. Microbiol. 89:e0094423. doi: 10.1128/aem.00944-23
Wang, D., Ji, X., Jiang, B., Yuan, Y., Liang, B., Sun, S., et al. (2024). Prevalence of antibiotic resistance and virulence genes in Escherichia coli carried by migratory birds on the Inner Mongolia plateau of northern China from 2018 to 2023. Microorganisms 12:1076. doi: 10.3390/microorganisms12061076
Wang, J., Ma, Z.-B., Zeng, Z.-L., Yang, X.-W., Huang, Y., and Liu, J.-H. (2017). The role of wildlife (wild birds) in the global transmission of antimicrobial resistance genes. Zool. Res. 38, 55–80. doi: 10.24272/j.issn.2095-8137.2017.003
Wang, J., Stephan, R., Zurfluh, K., Hächler, H., and Fanning, S. (2015). Characterization of the genetic environment of blaESBL genes, integrons and toxin-antitoxin systems identified on large transferrable plasmids in multi-drug resistant Escherichia coli. Front. Microbiol. 5:716. doi: 10.3389/FMICB.2014.00716
Werhahn Beining, M., Hartmann, M., Luebke-Becker, A., Guenther, S., Schaufler, K., Hille, K., et al. (2023). Carriage of extended Spectrum Beta lactamase-producing Escherichia coli: prevalence and factors associated with fecal colonization of dogs from a pet Clinic in Lower Saxony, Germany. Animals 13:584. doi: 10.3390/ani13040584
Woerther, P.-L., Andremont, A., and Kantele, A. (2017). Travel-acquired ESBL-producing Enterobacteriaceae: impact of colonization at individual and community level. J. Travel Med. 24, S29–S34. doi: 10.1093/jtm/taw101
Woolhouse, M., Ward, M., van Bunnik, B., and Farrar, J. (2015). Antimicrobial resistance in humans, livestock and the wider environment. Philos. Trans. R. Soc. B Biol. Sci. 370:20140083. doi: 10.1098/rstb.2014.0083
World Health Organization (2019). Minimum requirements for infection prevention and control programmes. Geneva: World Health Organization.
Wu, B., Qi, C., Wang, L., Yang, W., Zhou, D., Wang, M., et al. (2020). Detection of microbial aerosols in hospital wards and molecular identification and dissemination of drug resistance of Escherichia coli. Environ. Int. 137:105479. doi: 10.1016/j.envint.2020.105479
Wu, N., Xie, S., Zeng, M., Xu, X., Li, Y., Liu, X., et al. (2020). Impacts of pile temperature on antibiotic resistance, metal resistance and microbial community during swine manure composting. Sci. Total Environ. 744:140920. doi: 10.1016/J.SCITOTENV.2020.140920
Wuerz, T. C., Kassim, S. S., and Atkins, K. E. (2020). Acquisition of extended-spectrum beta-lactamase-producing Enterobacteriaceae (ESBL-PE) carriage after exposure to systemic antimicrobials during travel: systematic review and meta-analysis. Travel Med. Infect. Dis. 37:101823. doi: 10.1016/j.tmaid.2020.101823
Xavier, B. B., Renzi, G., Lammens, C., Cherkaoui, A., Goossens, H., Schrenzel, J., et al. (2019). Potential in vivo transfer of a blaCTX-M-14-harbouring plasmid established by combining long- and short-read sequencing. J. Microbiol. Methods 159, 1–4. doi: 10.1016/J.MIMET.2019.02.004
Xiao, L., Wang, X., Kong, N., Zhang, L., Cao, M., Sun, M., et al. (2019). Characterization of Beta-lactamases in bloodstream-infection Escherichia coli: dissemination of blaADC–162 and blaCMY–2 among Bacteria via an IncF plasmid. Front. Microbiol. 10:2175. doi: 10.3389/FMICB.2019.02175
Xie, H., Yamada, K., Tamai, S., Shimamoto, H., Nukazawa, K., and Suzuki, Y. (2023). Disappearance and prevalence of extended-spectrum β-lactamase-producing Escherichia coli and other coliforms in the wastewater treatment process. Environ. Sci. Pollut. Res. Int. 30, 83950–83960. doi: 10.1007/s11356-023-28382-3
Yagi, N., Hamamoto, K., Thi Bui, K. N., Ueda, S., Tawata, S., Le, D. T., et al. (2021). A high-throughput sequencing determination method for upstream genetic structure (UGS) of ISEcp1-blaCTX-M transposition unit and application of the UGS to classification of bacterial isolates possessing blaCTX-M. J. Infect. Chemother. 27, 1288–1294. doi: 10.1016/J.JIAC.2021.04.001
Yang, Y., Li, X., Zhang, Y., Liu, J., Hu, X., Nie, T., et al. (2020). Characterization of a hypervirulent multidrug-resistant ST23 Klebsiella pneumoniae carrying a blaCTX-M-24 IncFII plasmid and a pK2044-like plasmid. J. Glob. Antimicrob. Resist. 22, 674–679. doi: 10.1016/J.JGAR.2020.05.004
Yang, M., Liu, D., Li, X., Xiao, C., Mao, Y., He, J., et al. (2023). Characterizations of blaCTX-M-14 and blaCTX-M-64 in a clinical isolate of Escherichia coli from China. Front. Microbiol. 14:1158659. doi: 10.3389/fmicb.2023.1158659
Yang, H., Rehman, M. U., Zhang, S., Yang, J., Li, Y., Gao, J., et al. (2019). High prevalence of CTX-M belonging to ST410 and ST889 among ESBL producing E. coli isolates from waterfowl birds in China’s tropical island, Hainan. Acta Trop. 194, 30–35. doi: 10.1016/j.actatropica.2019.03.008
Yang, F., Tian, X., Han, B., Zhao, R., Li, J., and Zhang, K. (2021). Tracking high-risk β-lactamase gene (Bla gene) transfers in two Chinese intensive dairy farms. Environ. Pollut. 274:116593. doi: 10.1016/j.envpol.2021.116593
Yasugi, M., Hatoya, S., Motooka, D., Kondo, D., Akiyoshi, H., Horie, M., et al. (2023). Genetic and phenotypic analyses of mcr-harboring extended-spectrum β-lactamase-producing Escherichia coli isolates from companion dogs and cats in Japan. Vet. Microbiol. 280:109695. doi: 10.1016/j.vetmic.2023.109695
Ye, Q., Wu, Q., Zhang, S., Zhang, J., Yang, G., Wang, J., et al. (2018). Characterization of Extended-Spectrum β-Lactamase-Producing Enterobacteriaceae from Retail Food in China. Front. Microbiol. 9:1709 doi: 10.3389/fmicb.2018.01709
Yuan, W., Zhang, Y., Riaz, L., Yang, Q., Du, B., and Wang, R. (2021). Multiple antibiotic resistance and DNA methylation in Enterobacteriaceae isolates from different environments. J. Hazard. Mater. 402:123822. doi: 10.1016/j.jhazmat.2020.123822
Zequinao, T., Gasparetto, J., Oliveira, D., Dos, S., Silva, G. T., Telles, J. P., et al. (2020). A broad-spectrum beta-lactam-sparing stewardship program in a middle-income country public hospital: antibiotic use and expenditure outcomes and antimicrobial susceptibility profiles. Braz. J. Infect. Dis. 24, 221–230. doi: 10.1016/j.bjid.2020.05.005
Zhang, S., Abbas, M., Rehman, M. U., Huang, Y., Zhou, R., Gong, S., et al. (2020). Dissemination of antibiotic resistance genes (ARGs) via integrons in Escherichia coli: a risk to human health. Environ. Pollut. 266:115260. doi: 10.1016/j.envpol.2020.115260
Zhang, S., Abbas, M., Rehman, M. U., Wang, M., Jia, R., Chen, S., et al. (2021). Updates on the global dissemination of colistin-resistant Escherichia coli: an emerging threat to public health. Sci. Total Environ. 799:149280. doi: 10.1016/j.scitotenv.2021.149280
Zhang, S., Guo, X., Wang, Y., Zhong, Z., Wang, M., Jia, R., et al. (2023a). Implications of different waterfowl farming on cephalosporin resistance: investigating the role of blaCTX-M-55. Poult. Sci. 102:102929. doi: 10.1016/j.psj.2023.102929
Zhang, S., Shu, Y., Wang, Y., Zhong, Z., Wang, M., Jia, R., et al. (2023c). High rate of multidrug resistance and integrons in Escherichia coli isolates from diseased ducks in select regions of China. Poult. Sci. 102:102956. doi: 10.1016/j.psj.2023.102956
Zhang, S., Shu, Y., Yang, Z., Zhong, Z., Wang, M., Jia, R., et al. (2024). Decoding the enigma: unveiling the transmission characteristics of waterfowl-associated blaNDM-5-positive Escherichia coli in select regions of China. Front. Microbiol. 15:1501594. doi: 10.3389/fmicb.2024.1501594
Zhang, S., Wen, J., Wang, Y., Zhong, Z., Wang, M., Jia, R., et al. (2023b). Decoding the enigma: unveiling the molecular transmission of avian-associated tet(X4)-positive E. coli in Sichuan Province, China. Poult. Sci. 102:103142. doi: 10.1016/j.psj.2023.103142
Zhang, Q. Q., Ying, G. G., Pan, C. G., Liu, Y. S., and Zhao, J. L. (2015). Comprehensive evaluation of antibiotics emission and fate in the river basins of China: source analysis, multimedia modeling, and linkage to bacterial resistance. Environ. Sci. Technol. 49, 6772–6782. doi: 10.1021/acs.est.5b00729
Zong, Z., Ginn, A. N., Dobiasova, H., Iredell, J. R., and Partridge, S. R. (2015). Different IncI1 plasmids from Escherichia coli carry ISEcp1-blaCTX-M-15 associated with different Tn2-derived elements. Plasmid 80, 118–126. doi: 10.1016/J.PLASMID.2015.04.007
Zucconi, A., Courjon, J., Maruéjouls, C., Saintpère, F., Degand, N., Pandiani, L., et al. (2018). Managing ESBL-producing Enterobacteriaceae-related urinary tract infection in primary care: a tool kit for general practitioners. Eur. J. Clin. Microbiol. Infect. Dis. 37, 983–986. doi: 10.1007/s10096-018-3229-3
Keywords: antimicrobial resistance, Enterobacteriaceae, extended-spectrum β-lactamases, mobile genetic elements, horizontal gene transfer, plasmid incompatibility groups
Citation: Zhang S, Yang J, Abbas M, Yang Q, Li Q, Liu M, Zhu D, Wang M, Tian B and Cheng A (2025) Threats across boundaries: the spread of ESBL-positive Enterobacteriaceae bacteria and its challenge to the “one health” concept. Front. Microbiol. 16:1496716. doi: 10.3389/fmicb.2025.1496716
Received: 15 September 2024; Accepted: 05 February 2025;
Published: 21 February 2025.
Edited by:
João Pedro Rueda Furlan, Federal University of São Paulo, BrazilReviewed by:
Katiany Rizzieri Caleffi Ferracioli, State University of Maringá, BrazilCopyright © 2025 Zhang, Yang, Abbas, Yang, Li, Liu, Zhu, Wang, Tian and Cheng. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Anchun Cheng, Y2hlbmdhbmNodW5AdmlwLjE2My5jb20=
†These authors have contributed equally to this work
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.