- 1Department of Microbiology and Immunology, The University of North Carolina at Chapel Hill, Chapel Hill, NC, United States
- 2Department of Genetics, The University of North Carolina at Chapel Hill, Chapel Hill, NC, United States
- 3Center for Gastrointestinal Biology and Disease, The University of North Carolina at Chapel Hill, Chapel Hill, NC, United States
- 4Lineberger Comprehensive Cancer Center, The University of North Carolina at Chapel Hill, Chapel Hill, NC, United States
Inflammatory bowel diseases (IBDs) and inflammation-associated colorectal cancer (CRC) are linked to blooms of adherent-invasive Escherichia coli (AIEC) in the intestinal microbiota. AIEC are functionally defined by their ability to adhere/invade epithelial cells and survive/replicate within macrophages. Changes in micronutrient availability can alter AIEC physiology and interactions with host cells. Thus, culturing AIEC for mechanistic investigations often involves precise nutrient formulation. We observed that the pro-inflammatory and pro-carcinogenic AIEC strain NC101 failed to grow in minimal media (MM). We hypothesized that NC101 was unable to synthesize a vital micronutrient normally found in the host gut. Through nutrient supplementation studies, we identified that NC101 is a nicotinic acid (NA) auxotroph. NA auxotrophy was not observed in the other non-toxigenic E. coli or AIEC strains we tested. Sequencing revealed NC101 has a missense mutation in nadA, a gene encoding quinolinate synthase A that is important for de novo nicotinamide adenine dinucleotide (NAD) biosynthesis. Correcting the identified nadA point mutation restored NC101 prototrophy without impacting AIEC function, including motility and AIEC-defining survival in macrophages. Our findings, along with the generation of a prototrophic NC101 strain, will greatly enhance the ability to perform in vitro functional studies that are needed for mechanistic investigations on the role of intestinal E. coli in digestive disease.
Introduction
Inflammatory bowel diseases (IBDs), including Crohn’s disease and ulcerative colitis, are a major global health concern that affects over 3 million adults in the United States alone (Kaplan, 2015; Dahlhamer et al., 2016). IBD is a chronic and multifactorial disease that is driven by aberrant immune responses to commensal microbes, genetic susceptibility, and environmental factors (Khor et al., 2011). IBD patients experience painful, chronic, and relapsing intestinal inflammation that can lead to life-threatening complications, including intestinal fibrosis and colorectal cancer (CRC) (Rieder et al., 2007; Nadeem et al., 2019). Experimental models have demonstrated that IBD and CRC can be driven by the intestinal microbiota and that specific microbes, such as Escherichia coli, are associated with human disease (Schirmer et al., 2019; Lopez et al., 2021). IBD and CRC have no single etiology and no cure (Sartor, 2006; Goethel et al., 2018). Therefore, understanding the function of disease-associated gut microbes may uncover novel therapeutic options for intestinal diseases, such as IBD and CRC.
Intestinal microbes influence the onset and progression of IBD and CRC via metabolite production and modulation of mucosal immunity (Sartor, 2008; Dogan et al., 2014; Baümler and Sperandio, 2016; Hughes et al., 2017; Caruso et al., 2020). E. coli are common inhabitants of the intestinal microbiota (Tenaillon et al., 2010; Mirsepasi-Lauridsen et al., 2019). Strain level differences can alter the pro-inflammatory or pro-carcinogenic potential of E. coli partly through changes in small-molecule production (Arthur et al., 2012; Dogan et al., 2014; Baümler and Sperandio, 2016; Hughes et al., 2017; Ellermann et al., 2019). A pathovar of E. coli, termed adherent-invasive E. coli (AIEC), is enriched in the gut microbiota of human IBD and CRC patients (Martinez-Medina and Garcia-Gil, 2014). AIEC exacerbate experimental colitis and promote CRC in a variety of murine models (Carvalho et al., 2008, 2012; Arthur et al., 2012; Small et al., 2013; Martinez-Medina et al., 2014; Dejea et al., 2018; Ellermann et al., 2019). There is no genetic definition for AIEC (Nash et al., 2010; Dogan et al., 2014; Martinez-Medina and Garcia-Gil, 2014). Instead, AIEC are classically defined by their ability to adhere/invade epithelial cells and survive/replicate within macrophages (Darfeuille-Michaud et al., 2004; Martinez-Medina and Garcia-Gil, 2014). Environmental conditions, including nutrient availability and intestinal inflammation, can alter AIEC behavior and impact intestinal colonization and disease (Dogan et al., 2014; Ellermann et al., 2015, 2019, 2020; Delmas et al., 2019; Ormsby et al., 2020). Therefore, the ability to precisely manipulate AIEC growth conditions is essential for in vitro studies investigating AIEC behavior and production of pro-inflammatory and pro-carcinogenic molecules.
Escherichia coli NC101 is a well-known AIEC strain utilized by numerous investigators to study how intestinal E. coli adapt to and influence the host during IBD and CRC (Arthur et al., 2012; Tchaptchet et al., 2013; Dogan et al., 2014; Dejea et al., 2018; Ellermann et al., 2019; Gogokhia et al., 2019; Schmitz et al., 2019). NC101 was originally isolated from a specific pathogen-free wild-type (WT) mouse at the North Carolina State University (Kim et al., 2005). Colonizing WT mice with NC101 does not induce intestinal pathology, even during monoassociation studies using gnotobiotic animals (Ellermann et al., 2019). However, despite a lack of traditional toxins and virulence factors, NC101 induces antigen-driven intestinal inflammation in genetically susceptible IBD mouse models (e.g., interleukin 10-deficient mice) (Kim et al., 2005). Thus, NC101 is considered a pathobiont and a highly relevant model organism for defining how susceptible individuals may mount inappropriate immune responses to seemingly innocuous intestinal E. coli.
NC101 adapts to the inflamed intestinal milieu by modulating the expression of its gene repertoire (Patwa et al., 2011; Arthur et al., 2014). Nutrient availability alters AIEC physiology, persistence in the microbiota, and production of pro-inflammatory and pro-carcinogenic mediators (Dogan et al., 2014; Ellermann et al., 2015, 2019, 2020; Delmas et al., 2019; Ormsby et al., 2020). Monoassociation studies with gnotobiotic mice have led to the discovery of several AIEC-derived host-influencing molecules (i.e., specialized metabolites) that drive inflammation and tumorigenesis, including yersiniabactin and colibactin (Arthur et al., 2012; Martinez-Medina and Garcia-Gil, 2014; Ellermann et al., 2019). Like many specialized metabolites, yersiniabactin and colibactin are produced via biosynthetic gene clusters that can be activated by changes in micronutrient availability, notably iron (Perry and Fetherston, 2011; Tronnet et al., 2016). The nature of AIEC-derived specialized metabolites makes them difficult to isolate and study in functional assays. Therefore, the repertoire of AIEC-derived metabolites and their impact on the host has been largely unexplored.
Variations in micronutrient availability can impact the virulence and physiology of AIEC (Sharon et al., 2014; Ellermann et al., 2015, 2020). Therefore, culturing AIEC for mechanistic studies necessitates using a simplified base medium that allows for precise nutrient manipulation. During our studies, we observed that modified M9 minimal media (MM) does not sustain NC101 growth in vitro. We hypothesized that NC101 was an auxotroph. Through nutrient supplementation studies, we discovered that NC101 requires nicotinic acid (NA, niacin, vitamin B3) for growth. NA auxotrophy was not observed in other non-toxigenic laboratory E. coli strains (K12 or 25922), AIEC, or non-AIEC human intestinal strains (Baumgart et al., 2007). Genetic evaluation revealed that NC101 has a missense mutation in the nicotinamide adenine dinucleotide (NAD) biosynthesis gene (nadA) that encodes for quinolinate synthase A. Importantly, we generated a prototrophic NC101 revertant strain that eliminated E. coli micronutrient restraints. Correcting NC101 auxotrophy had negligible impact on NC101 function, including motility and AIEC-defining survival in macrophages.
NC101 micronutrient constraints have limited our ability to perform in vitro functional studies, which often require careful nutrient manipulation. Overall, our findings will enable precise nutrient manipulation for mechanistic studies on auxotrophic microbiota members, like AIEC, Shigella spp., or uropathogenic E. coli (Prunier et al., 2007b; Li et al., 2012; Di Martino et al., 2013; Bouvet et al., 2017). Importantly, our work will facilitate in vitro functional assays and small-molecule purification efforts with the pro-inflammatory and pro-carcinogenic AIEC strain NC101. Furthermore, these studies will broadly improve our understanding of the microbiota in intestinal diseases like IBD and CRC.
Materials and Methods
Bacterial Strains
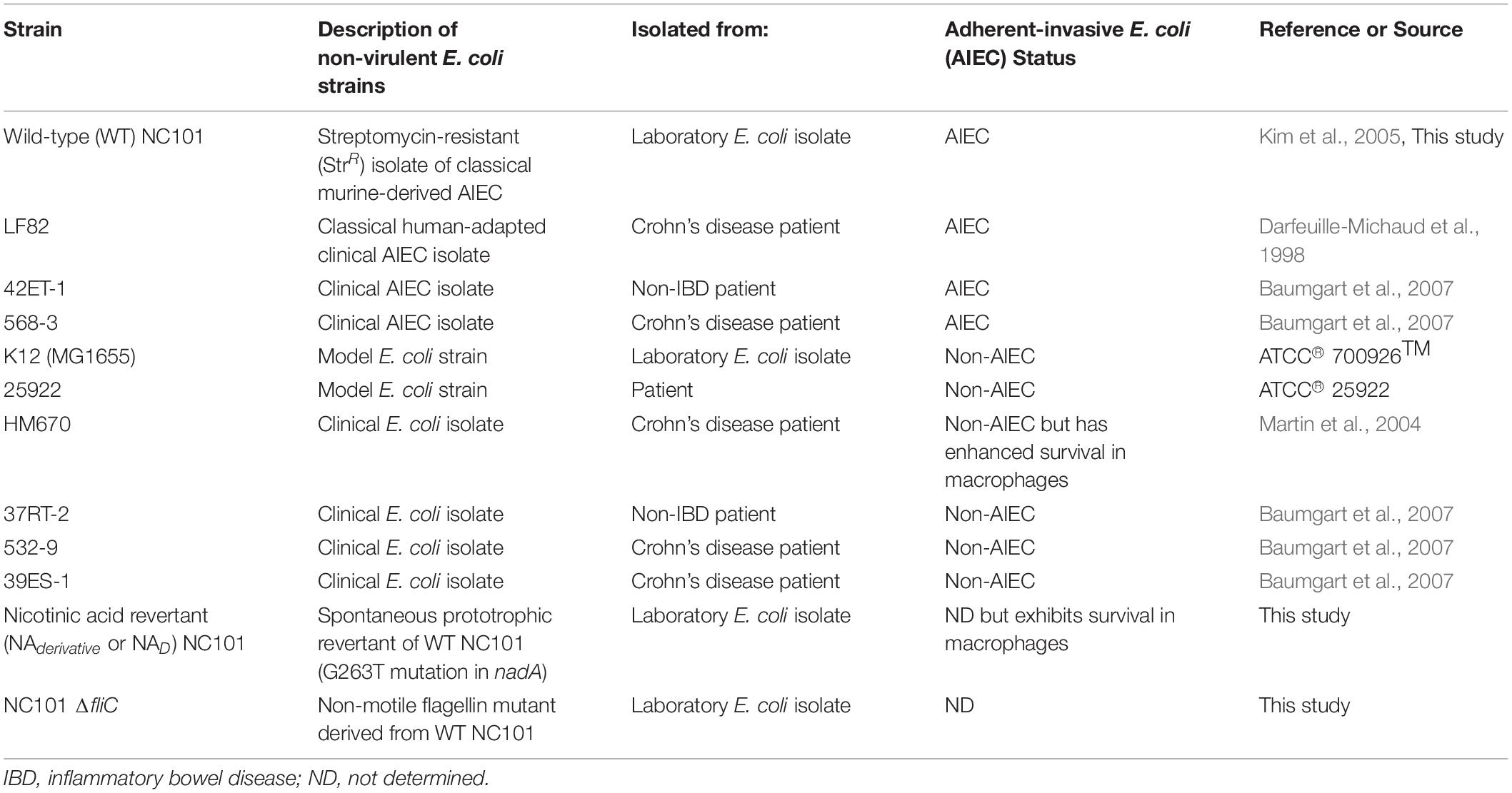
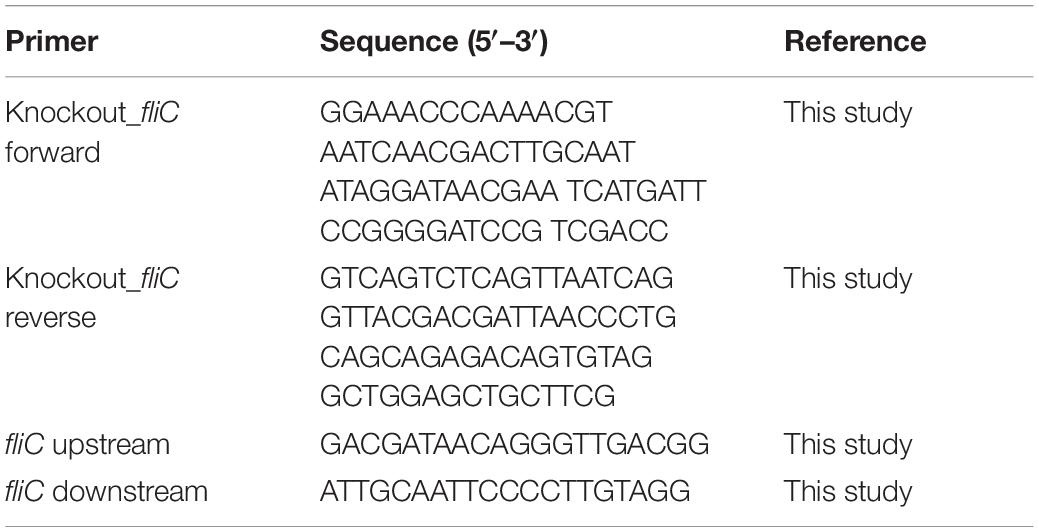
Descriptions of E. coli strains used in this study are listed in Table 1. NC101 ΔfliC was generated using the λ-red recombinase method, as previously described (Datsenko and Wanner, 2000; Ellermann et al., 2019). Primers used for ΔfliC generation are listed in Table 2.
Media Composition
M9 Minimal-Defined Media (MM)–5X M9 Salts
64 g Na2HPO4×7H2O, 15 g anhydrous KH2PO4, 2.5 g NaCl, and 5 g NH4Cl brought to 1 L in diH2O (Sambrook and Russell, 2006).
Complete MM
0.1 mM CaCl2, 1X M9 salts, 2 mM MgSO4, 0.4% glycerol, and 0.2% casamino acids (CAA, Sigma #2240) brought to 1 L in diH2O. Where indicated, the following were added at these final concentrations: NA (Sigma #N4126), 50 μg/L; L-aspartate (L-asp), 200 mg/L; nicotinamide (Nm), 50 μg/L; and NAD, 1 μg/L.
Vitamin Mix, 100X Stock
2 mg folic acid, 10 mg pyridoxine hydrochloride, 5 mg riboflavin, 2 mg biotin, 5 mg thiamine, 5 mg NA, 5 mg calcium pantothenate, 0.1 mg vitamin B12, 5 mg p-aminobenzoic acid, 5 mg thioctic acid, and 900 mg monopotassium phosphate brought to 1 L in diH2O and aliquoted into 10-ml stocks. One 10-ml stock was used per liter of media. Formulation is from ATCC and is based on Wolfe’s Vitamin solution (ATCC® MD-VSTM).
Overnight Cultures
Bacterial strains were preserved at -80°C and grown overnight at 37°C on Luria–Bertani (LB; Fisher Sc. #BP9722-2) agar plates. Isolated colonies were transferred to MM and grown overnight (>15 h) at 37°C with shaking at 220 rpm.
Growth Assays
Overnight cultures were centrifuged and washed three times with 1X phosphate buffered saline (PBS) to remove any trace compounds contained in the culture. Cells were resuspended and normalized by optical density (OD600) in test media. Cultures were grown at 37°C with shaking at 220 rpm. For passaging assays, OD600 was recorded at the indicated time points (2, 4, 8, or 24 h). For growth curves, OD600 was recorded every 45 min for 6 h, and a final time point was recorded at 24 h.
Spontaneous Prototrophic Revertant Generation
Wild-type NC101 was grown overnight in MM + NA, 5 ml of the culture was centrifuged, and the supernatant was discarded. The cell pellet was washed twice with 1X PBS and resuspended in 500 μl 1X PBS. A 100-μl spot was spread onto each of five MM agar plates without NA. Plates were incubated at 37°C and monitored for growth of revertant colonies (Li et al., 2012). Colonies were grown on MM without NA to confirm prototrophy, and isolates were preserved at −80°C. Whole-genome sequencing was performed to determine the location and nature of the mutation(s) leading to reversion. The revertant used in these studies was termed NC101 NADerivative (NAD).
NC101 Genome Assembly
A complete NC101 genome was assembled from nanopore sequence using Minimap2 and Miniasm (Li, 2016). The assembly was circularized and polished four times with Racon (Vaser et al., 2017) followed by once with Medaka (Oxford Nanopore Technologies)1. Matched Illumina sequence data were used to polish the resulting assembly using FMLRC (Wang et al., 2018) with parameters “-k 21 -K 30 -m 3 -f 0.05 -B 10.” The final polished genome was rotated and linearized such that it starts at the origin of replication.
Whole-Genome Sequencing
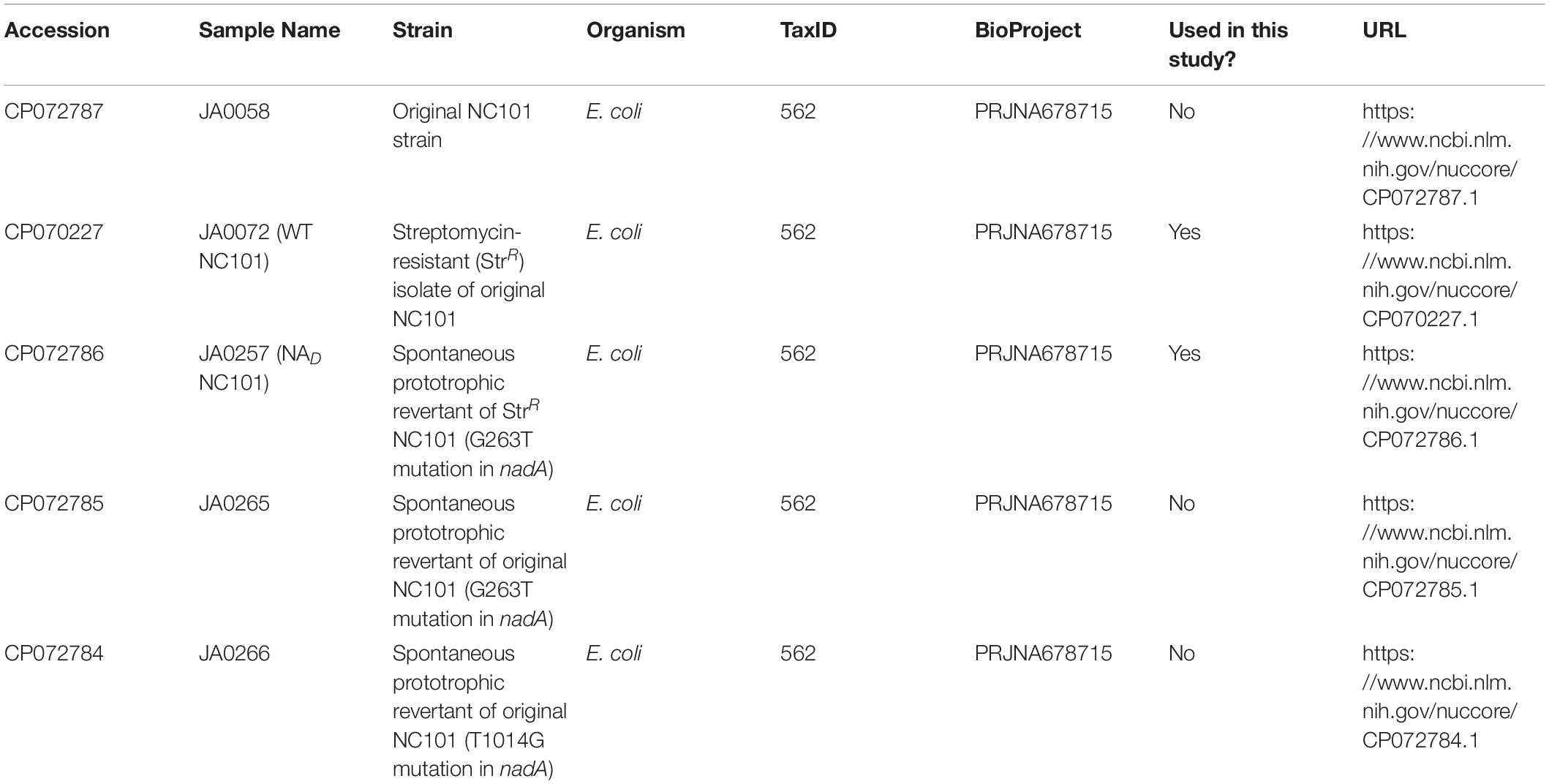
Three spontaneous prototrophic revertants, including NAD, were sent for whole-genome sequencing. Samples were sent to the Microbial Genome Sequencing Center (MiGS), formerly at the University of Pittsburgh, for genomic DNA extraction and Illumina 2×150 paired end sequencing on the NextSeq 550 platform. Sequencing reads were mapped to our closed NC101 genome using CLC Genomic Workbench7.5.1 with average coverage of 85× for JA0257, 62× for JA0265, and 75× for JA0266. Sequences for LF82 (NC_011993.1) and K12 (NC_000913.3) were obtained from the National Center for Biotechnology Information (NCBI), and all alignments were analyzed via Geneious Prime version 2020.1.2. Assembled sequences from this study were deposited in NCBI, and repository information is listed in Table 3.
Motility
Isolates were grown overnight, as described above. A 1-μl spot was used to inoculate the center of MM soft agar plates (MM + 0.25% agar) with NA. Plates were incubated at 37°C for 8 h, and the diameters of motility swarms were measured.
Macrophage Survival Assays
Bacterial intramacrophage survival was measured using the standard gentamicin protection assay for AIEC bacteria (Darfeuille-Michaud et al., 2004; Ellermann et al., 2015). The J774A.1 murine macrophage-like cell line was used as a model and maintained according to ATCC standards in Dulbecco’s modified Eagle’s medium (DMEM) + 10% heat-inactivated fetal bovine serum (FBS) (DMEM, Gibco #11995-065). J774A.1 cells were seeded at 2 × 105 cells/ml in 1 ml media into 24-well plates (Falcon #353047) and grown overnight. The next day, bacterial overnight cultures were subcultured in MM with and without NA for 3 h. Before infection, J774A.1 monolayers were washed twice with 1X PBS. Then, subcultured bacteria were added at a multiplicity of infection (MOI) = 10 in cell culture media with and without NA. Plates were spun at 180 × g for 5 min. Prepared bacterial cultures were serial diluted and plated on LB agar plates to validate infection dose.
After a 30-min incubation at 37°C with 5% CO2, infected cultures were washed twice with 1X PBS, and gentamicin-laden media was added (100 μg/ml gentamicin for 1 h time point and 20 μg/ml for 24 h in DMEM + 10% FBS with and without NA). At 1 and 24 h, cells were washed twice with 1X PBS, and 500 μl of 1% Triton X-100 in diH2O was added to each well for 5 min. Samples were mixed, serial diluted, and plated on LB agar plates to determine viable colony-forming units (CFUs). Percent intracellular bacteria = [(CFU/ml at 24 h)/(CFU/ml at 1 h] × 100.
Statistics
Statistical analysis was performed using Prism version 9.0.0 (GraphPad software San Diego, CA, United States). A Welch’s t-test was used when two experimental groups were compared, and a one-way ANOVA with Dunnett’s T3 multiple comparisons test was used when three or more experimental groups were compared. Differences with a p-value less than 0.05 were considered significant. All experiments included at least three biological replicates with 1–2 technical replicates each, per time point.
Data Availability
Assembled sequences from our whole-genome sequencing, above, were deposited in NCBI, and repository information is listed in Table 3.
Results
The Pro-carcinogenic Adherent-Invasive E. coli Strain NC101 Requires NA to Sustain Growth
During in vitro studies to evaluate AIEC function in long-term culture (24+ h), we attempted to passage NC101 in modified M9 MM that includes glycerol and casamino acids. NC101 can successfully be subcultured from LB agar or broth, a rich medium, to MM (Ellermann et al., 2015, 2019). However, NC101 failed to grow when subcultured from MM to MM (Figures 1A–C). We hypothesized that NC101 was an auxotroph, unable to synthesize a key nutrient found in the murine gut. Shigella spp., a transient gut pathogen and close relative of E. coli, are generally NA (niacin, vitamin B3) auxotrophs (Ahmed et al., 1988; Prunier et al., 2007b; Di Martino et al., 2013; Bouvet et al., 2017). Thus, we specifically tested whether vitamin supplementation could restore NC101 growth in MM. Supplementing MM with a complex vitamin mix (VM) restored NC101 growth at 8 and 24 h (Figures 1A,B).
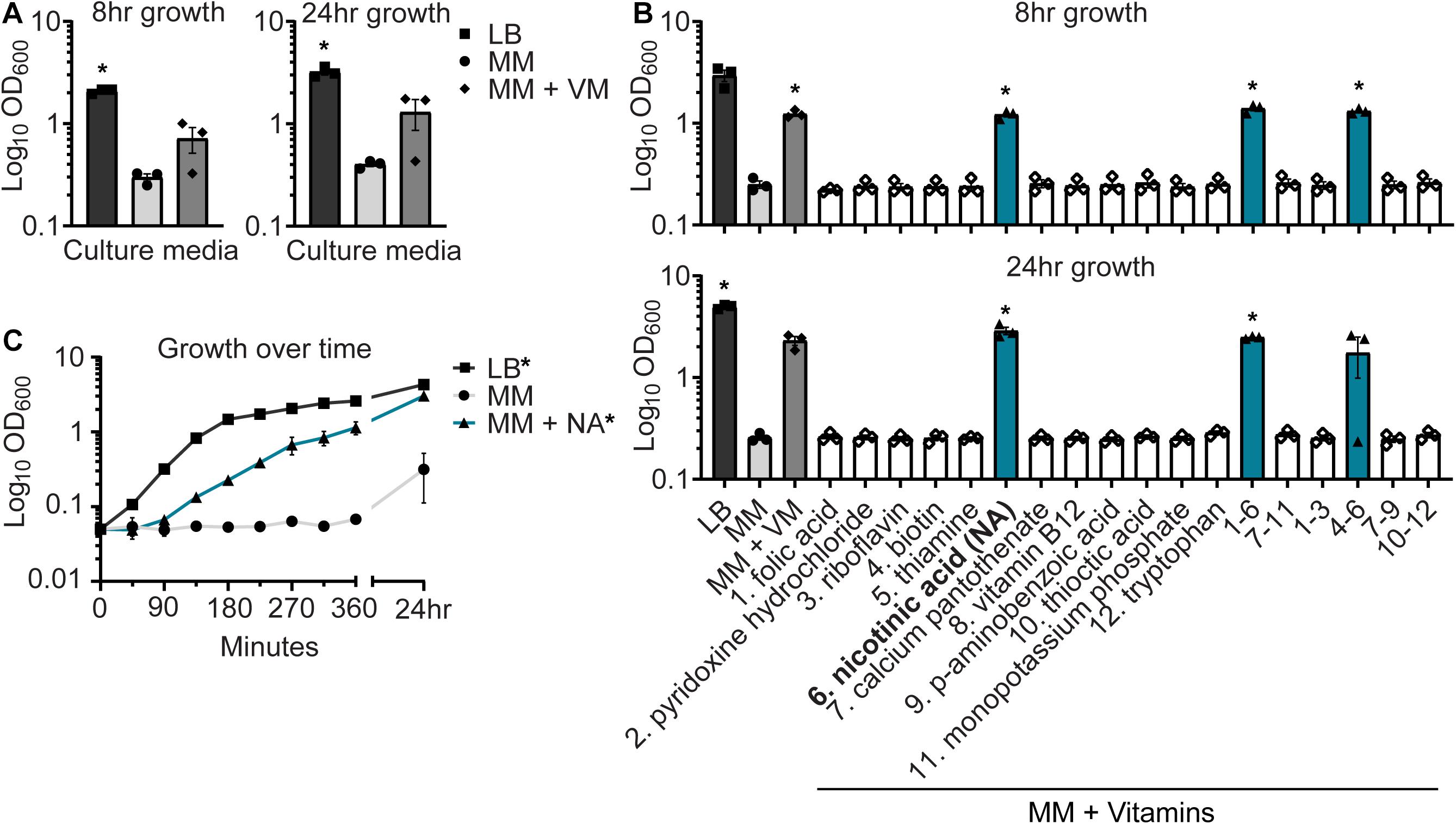
Figure 1. Nicotinic acid restores growth of E. coli NC101 in minimal media (MM). (A) Wild-type NC101 was grown in Luria–Bertani (LB) broth, MM, or MM + vitamin mix (VM). Growth was measured at 8 and 24 h by culture optical density, OD600. (B) NC101 was grown in LB, MM, MM + VM, or MM supplemented with individual or combinations of vitamins and tryptophan. OD600 was assessed at 8 and 24 h. (C) Growth curve of NC101 in LB, MM, or MM + NA. (A,B) Bars (n = 3) or (C) points (n = 4) depict mean ± SEM. Significance (*) is shown compared to NC101 growth in MM at (A,B) each time point or (C) 24 h and was determined at p < 0.05 using a one-way ANOVA with Dunnett’s T3 multiple comparisons test.
To identify which vitamins in the VM were essential for NC101 growth, we supplemented MM with individual or combinations of VM components and assessed NC101 growth. Tryptophan supplementation was also tested, as tryptophan metabolism can be influenced by host–microbe interactions in the gut (Agus et al., 2018). Only MM containing NA, alone or in combination, sustained NC101 growth in MM at 8 and 24 h (Figure 1B). Furthermore, NA alone restored normal NC101 growth kinetics in MM and significantly enhanced growth at 24 h (Figure 1C). NC101 grew when subcultured from MM to LB, indicating NC101 does not have a global growth defect (Figures 1A–C). Together, these data suggest that NC101 is an NA auxotroph.
Nicotinic Acid Auxotrophy Is Not a Defining Feature of Non-toxigenic E. coli
Resident non-toxigenic E. coli are common among the intestinal microbiota, and many are considered commensal strains (Tenaillon et al., 2010; Mirsepasi-Lauridsen et al., 2019). Yet, other E. coli (e.g., AIEC) are associated with chronic intestinal inflammation and may be referred to as pathobionts (Martinez-Medina and Garcia-Gil, 2014; Mirsepasi-Lauridsen et al., 2019). We questioned whether NA auxotrophy was shared across clinically derived non-toxigenic E. coli. In addition to evaluating model E. coli strains (K12 and 25922), we evaluated clinical specimens isolated from the intestinal mucosa of IBD or non-IBD patients (E. coli LF82, 42ET-1, 568-3, HM670, 37RT-2, 532-9, and 39ES-1) (Darfeuille-Michaud et al., 1998; Martin et al., 2004; Baumgart et al., 2007) (Table 1). These clinical isolates have been characterized in the lab from which they originated for AIEC status, and at least partial genome sequences are available for all strains (Baumgart et al., 2007). To determine the extent of NA dependency among these strains, we passaged isolates in MM with and without NA and assessed growth by measuring optical density (OD600) at 2, 4, 8, and 24 h (Figure 2A). We again observed that NC101 had a growth defect in MM, detectable at 2 h, and continuing through 24 h (Figure 2A). However, all examined laboratory strains and clinical isolates grew in MM with and without NA (Figure 2A). Therefore, NA auxotrophy does not appear to be a defining characteristic shared by non-toxigenic resident intestinal E. coli.
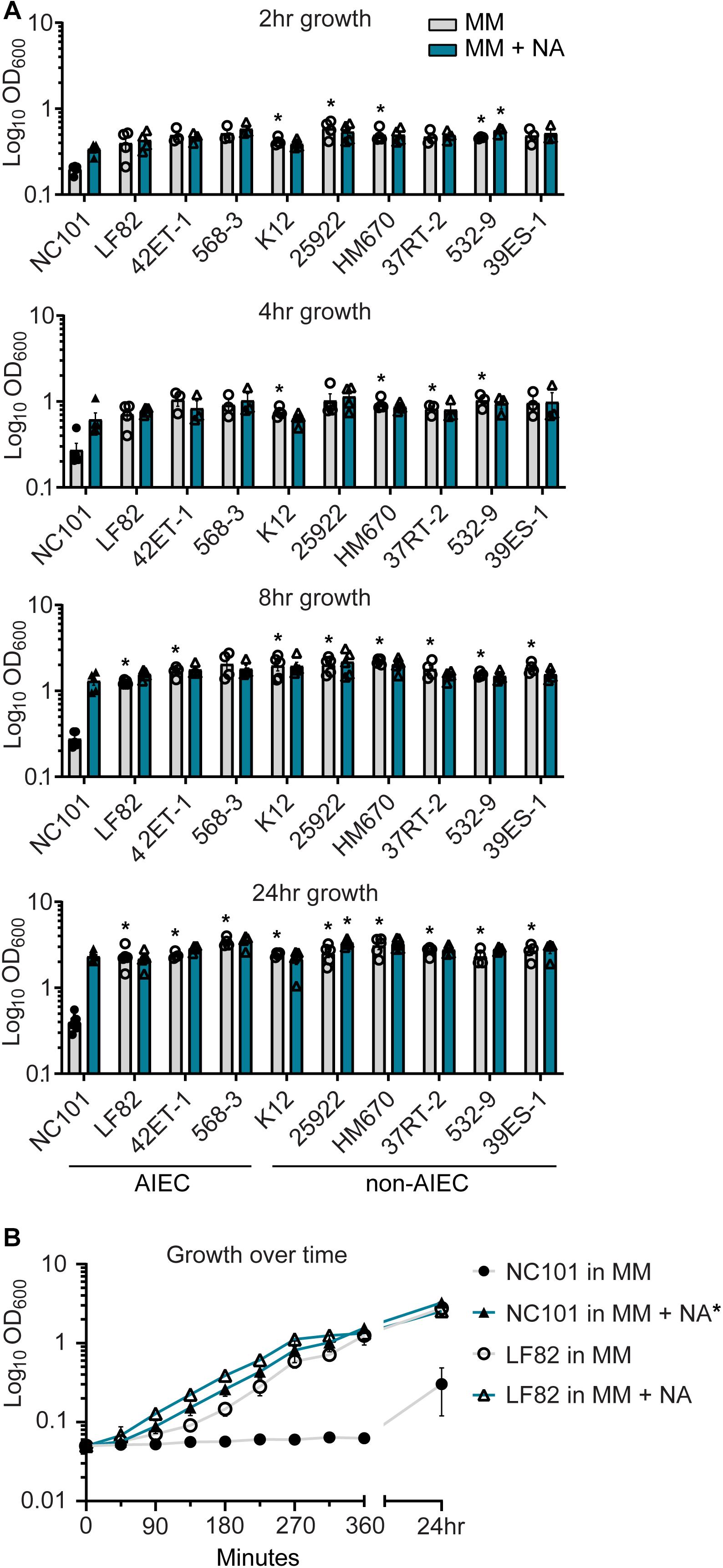
Figure 2. Nicotinic acid (NA) auxotrophy is not common among non-toxigenic Escherichia coli strains, including prototypic adherent-invasive E. coli (AIEC) LF82. (A) AIEC and non-AIEC E. coli isolates were grown in minimal media (MM) with and without NA. Growth was evaluated at 2, 4, 8, and 24 h by OD600. (B) Growth curve of wild-type NC101 and LF82 (human-derived AIEC) in MM with and without NA. (A) Bars (n = 3–6) or (B) points (n = 3) depict mean ± SEM. (A) Strains grown in MM were compared to NC101 grown in MM, and strains grown in MM + NA were compared to NC101 grown in MM + NA. Significance (*) is shown compared at (A) each time point or (B) 24 h and was determined at p < 0.05 using a one-way ANOVA with Dunnett’s T3 multiple comparisons test.
LF82 is a well-known human-derived AIEC strain that can grow in MM without NA (Darfeuille-Michaud et al., 1998) (Figures 2A,B). We directly compared the growth kinetics of NC101 and LF82 in MM with and without NA (Figure 2B). While early growth of LF82 in MM was minimally enhanced by NA, this difference was indistinguishable by 6 h (Figure 2B). Thus, the prototypic AIEC strain LF82 does not exhibit NA auxotrophy. Combined with our findings in Figure 2A, we conclude that NA auxotrophy is not an AIEC-defining feature.
NC101 Has a Defect in the de novo NAD Biosynthesis Pathway
Nicotinic acid is a precursor for NAD biosynthesis (Bouvet et al., 2017). NAD is an electron carrier and an essential cofactor for bacterial metabolism (Bouvet et al., 2017). In E. coli and related bacteria, NAD can be synthesized de novo from L-aspartate (L-asp) through the generation of quinolinic acid (quinolinate, Qa). In this process, quinolinate synthases A and B (encoded by nadA and nadB, respectively) catalyze the oxidation of L-asp to iminoaspartate and condensation with dihydroxyacetone phosphate to generate quinolinate (Bouvet et al., 2017). Quinolinate is converted to nicotinic mononucleotide (NaMN) by a nadC encoded enzyme and ultimately NAD via enzymes encoded by nadD and nadE (Bouvet et al., 2017). NAD biosynthesis can also occur through salvage pathways that utilize vitamin precursors like NA or nicotinamide (Nm) (Liu et al., 1982; Di Martino et al., 2013; Bouvet et al., 2017) (Figure 3A).
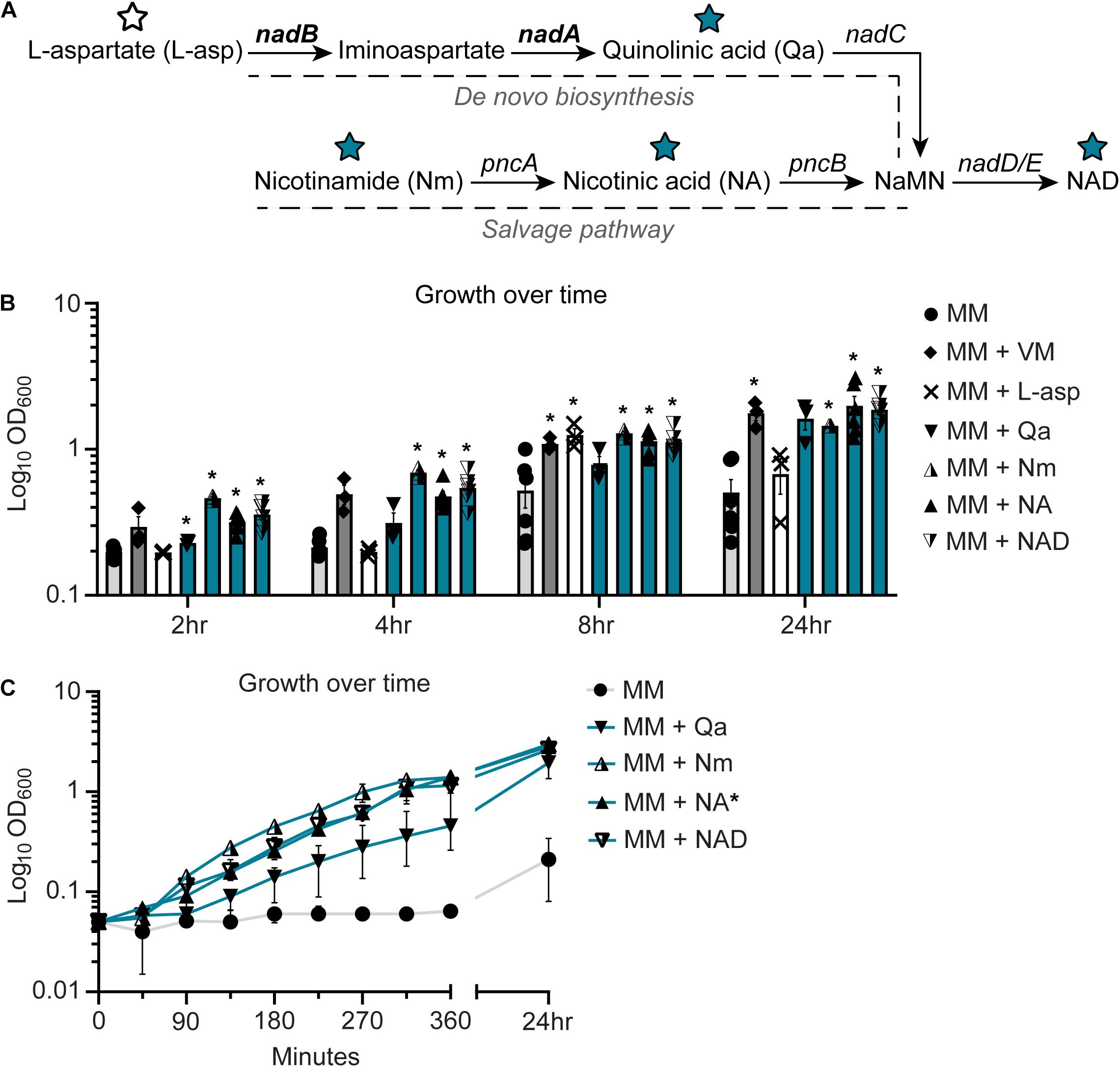
Figure 3. NC101 has a defect in the de novo nicotinamide adenine dinucleotide (NAD) biosynthesis pathway. (A) Illustration of NAD biosynthesis pathway in Escherichia coli, including pathway intermediates and genes involved. Intermediates tested in E. coli growth assays are indicated by stars (blue restored growth, white did not). Bolded genes, nadA and nadB, are predicted to be responsible for NC101 auxotrophy. (B) Wild-type NC101 was grown in minimal media (MM), MM + vitamin mix (VM), or MM + NAD biosynthesis pathway intermediates: L-aspartate (L-Asp), quinolinic acid (Qa), nicotinamide (Nm), nicotinic acid (NA), and NAD. Culture density was evaluated at 2, 4, 8, or 24 h by OD600. (C) Growth curve of NC101 in MM with or without Qa, Nm, NA, or NAD. (B) Bars (n = 3–6) or (C) points (n = 3) depict mean ± SEM. Significance (*) is shown compared to NC101 growth in MM at (B) each time point or (C) 24 h and was determined at p < 0.05 using a one-way ANOVA with Dunnett’s T3 multiple comparisons test.
Since, NA restored the growth of NC101, we predicted that NC101 had a defect within the NAD biosynthesis pathway. To determine whether this was the case, we assessed NC101 growth in MM supplemented with key NAD biosynthesis intermediates: L-asp, Qa, Nm, NA, and NAD. L-asp failed to consistently sustain NC101 growth in MM. Conversely, Qa sustained NC101 growth in MM, and Nm, NA, and NAD significantly restored growth across all time points (Figure 3B). Growth curves revealed the kinetics of enhanced NC101 growth in the presence of the restorative NAD biosynthesis intermediates: Qa, Nm, Na, and NAD (Figure 3C). When examining the NAD biosynthesis pathway, this indicated that NC101 likely had a defect in the NAD biosynthesis genes nadA or nadB (Figure 3A).
Nicotinic Acid Auxotrophy in NC101 Is Linked to a Mutation in NAD Biosynthesis Gene nadA
After our growth supplementation assays revealed a likely defect in nadA or nadB, we sought to identify the genetic factor(s) responsible for NA auxotrophy in NC101. We performed whole-genome sequencing on WT NC101 and compared the sequence to prototrophic E. coli, LF82 and K12. Sequencing revealed that WT NC101 has a missense mutation in nadA (T263G) that was associated with auxotrophy (Figure 4A).
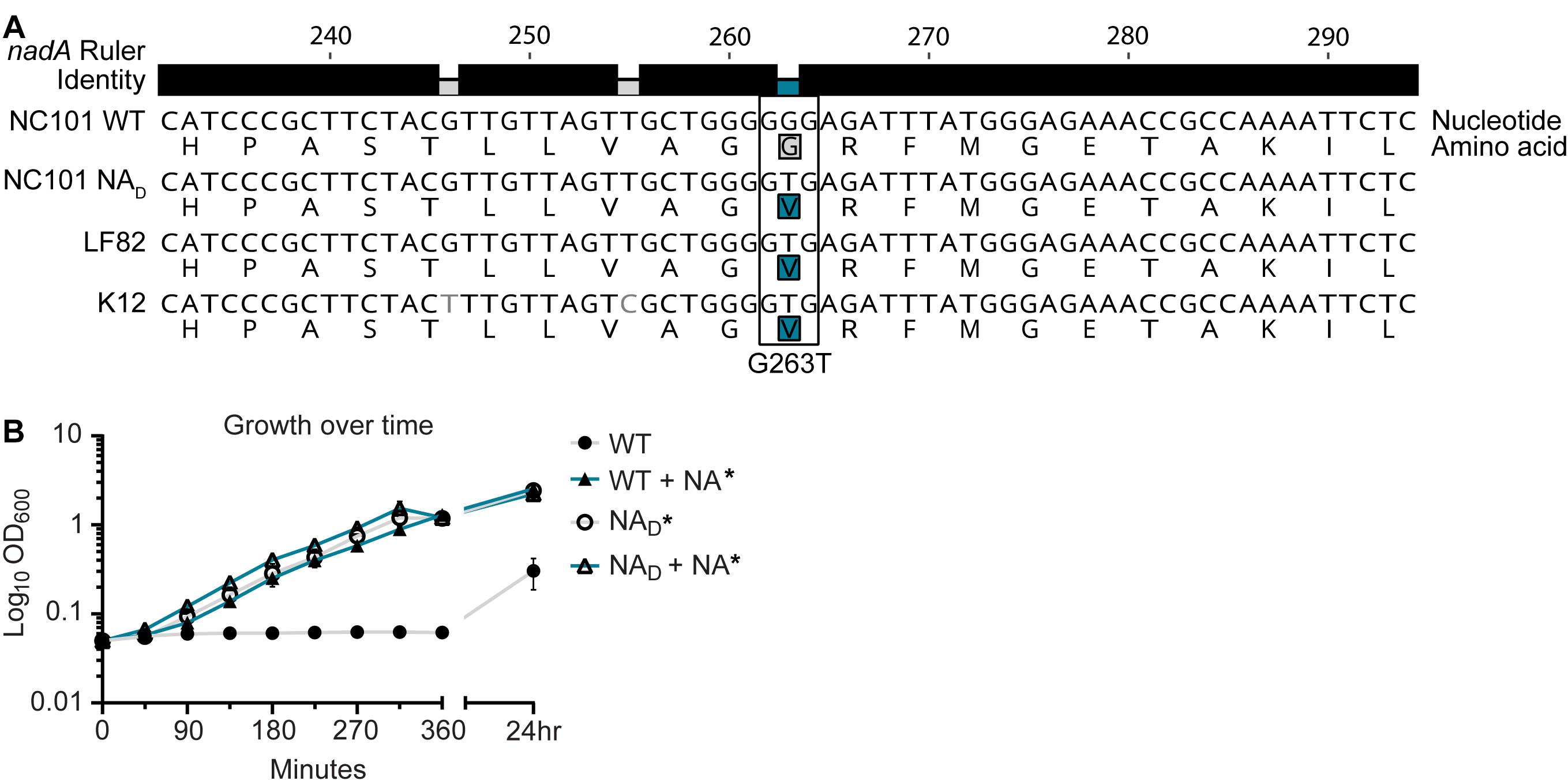
Figure 4. A missense mutation in nicotinamide adenine dinucleotide (NAD) biosynthesis gene nadA confers nicotinic acid (NA) auxotrophy in NC101. (A) Genetic alignment of partial nadA sequence from NA auxotrophic [Wild-type (WT) NC101] and prototrophic (NC101 NAD, LF82, and K12) E. coli. Nucleotide and amino acid sequences, noted by one-letter abbreviations, are shown. The ruler displays nucleotide position of coding sequence. The identity bar displays regions of similarity (black) or dissimilarity (gray or blue). The highlighted amino acids show the region of noted dissimilarity (nadA G263T) between NA auxotrophic (gray) and prototrophic (blue) E. coli. (B) A growth curve of WT NC101 and prototrophic revertant NC101 strain (NAD) in minimal media (MM) with and without NA. Growth was measured by culture optical density, OD600. Points depict mean ± SEM (n = 4). Significance (*) is shown compared to NC101 growth in MM at 24 h and was determined at p < 0.05 using a one-way ANOVA with Dunnett’s T3 multiple comparisons test.
To further validate the genetic determinants of NC101 NA auxotrophy, we generated a prototrophic strain by passaging WT NC101 on MM agar plates in the absence of NA (Li et al., 2012). Sequencing of a selected spontaneous prototrophic revertant, termed NADerivative or NAD NC101, revealed that NAD NC101 had a single nucleotide substitution in nadA (G263T, compared to WT) that matched the prototrophic E. coli strains LF82 and K12 (Figure 4A). It is important to note that NAD NC101 also had a silent mutation in an intergenic region that was absent from WT NC101, but we predict this mutation had no impact on NAD NC101 prototrophy (Accession #CP072786) (Table 3). To support that NA auxotrophy is due to the observed nadA mutation, our whole-genome sequencing revealed that two other NC101 spontaneous prototrophic revertants had missense mutations in nadA – one of which shared the same nadA (G263T, compared to WT) nucleotide substitution as NAD NC101 (accession #CP072785 and #CP072784) (Table 3).
To confirm that the NAD revertant restored prototrophy, we grew WT NC101 and NAD in MM with and without NA. NAD grew significantly better in MM vs. WT NC101 at 24 h (Figure 4B). Growth of NAD in the absence of NA was the equivalent to WT grown with added NA, as noted by overlapping growth curves (Figure 4B). The addition of NA did not significantly enhance NAD growth in MM, suggesting NA auxotrophy was successfully eliminated in this strain (Figure 4B). These findings are consistent with the literature, which indicates NadA is important for NAD biosynthesis, and mutations in nadA can drive NA auxotrophy in E. coli, Shigella spp., and Salmonella spp. (Bergthorsson and Roth, 2005; Prunier et al., 2007b; Di Martino et al., 2013; Bouvet et al., 2017). Therefore, our data support that NC101 NA auxotrophy is due to a mutation in nadA.
Correcting NA Auxotrophy in NC101 Has Negligible Impact on Bacterial Motility or AIEC-Associated Survival in Macrophages
To determine if correcting NA auxotrophy impacted AIEC physiology and interactions with mammalian cells, we assessed WT and NAD NC101 for motility and survival in macrophages. Motility is not an AIEC-defining feature, but hypermotility has recently been linked to changes in AIEC:host interactions (Martinez-Medina and Garcia-Gil, 2014; Elhenawy et al., 2019). Due to the WT NA auxotrophy, motility was only assessed on MM agar plates supplemented with NA. There was no significant difference in motility between WT and NAD NC101 in the presence of NA (Figure 5A). However, the motility of both WT and NAD NC101 differed significantly from the non-motile control mutant, NC101 ΔfliC (flagellar filament structural protein) (Elhenawy et al., 2019) (Figure 5A).
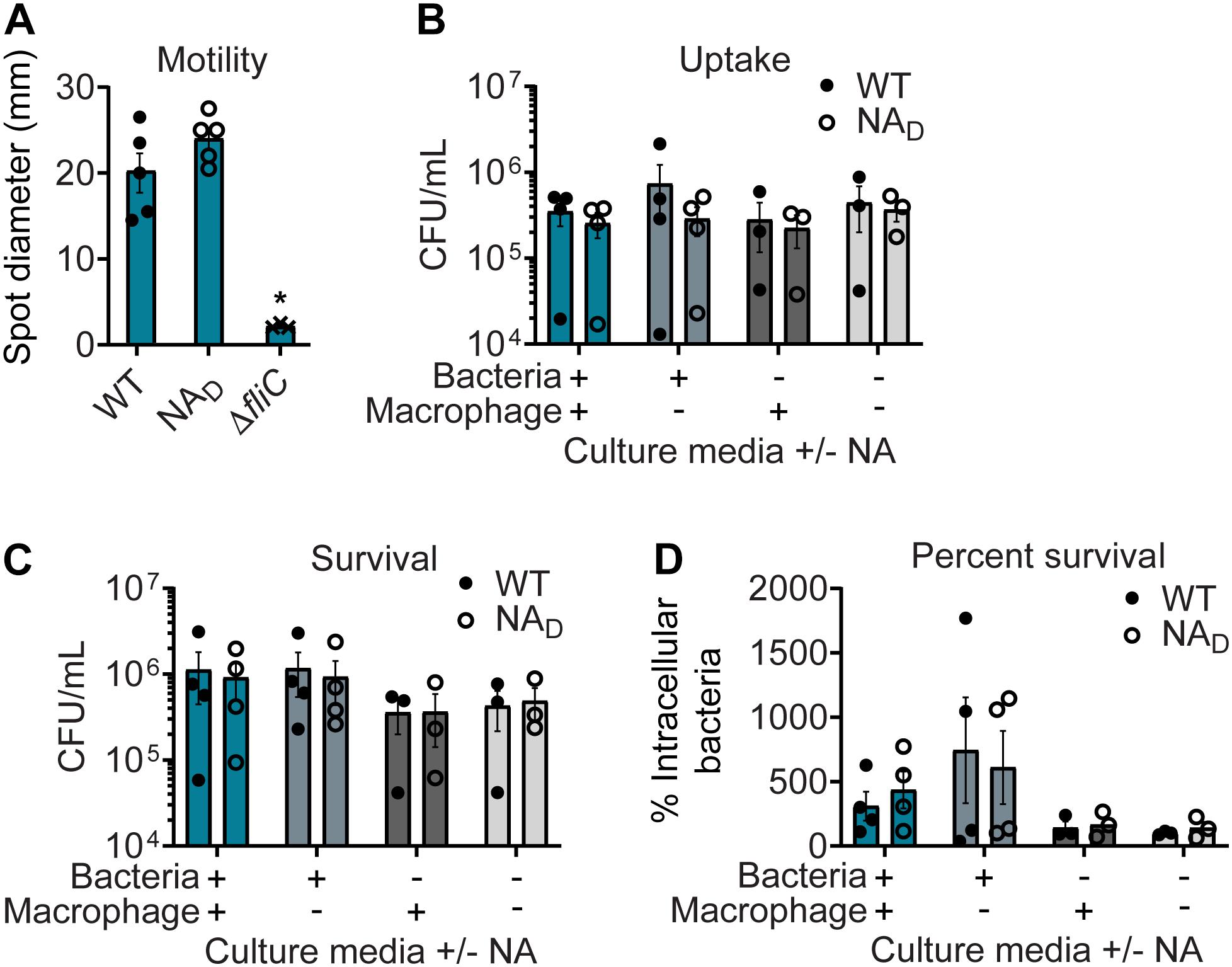
Figure 5. Correcting nicotinic acid (NA) auxotrophy in NC101 has minimal impact on in vitro adherent-invasive Escherichia coli (AIEC) function. (A) Wild-type (WT) NC101, prototrophic NC101 NAD, and non-motile control NC101 ΔfliC were grown on minimal media (MM) soft agar plates with NA. Diameters of motility swarm spots (mm) were measured at 8 h (n = 3–5). (B–D) J774A.1 murine macrophages were infected at a multiplicity of infection (MOI) = 10 with WT or NAD. Bacterial culture media before infection (“Bacteria”) or cell culture media during infection (“Macrophage”) were with or without NA supplementation. Numbers of bacteria are shown at (B) 1 h or (C) 24 h post-infection as Log10 colony-forming units (CFUs)/ml. (D) Percent survival = [(CFU/ml at 24 h)/(CFU/ml at 1 h] × 100 (n = 3–4). Bars depict mean ± SEM. Significance (*) is shown compared to WT NC101 and was determined at p < 0.05 using a (A) one-way ANOVA with Dunnett’s T3 multiple comparisons test or (B–D) Welch’s t-test.
A key feature of AIEC is enhanced survival in macrophages, a characteristic linked to their pro-inflammatory activities (Darfeuille-Michaud et al., 2004; Kittana et al., 2019). To evaluate whether the identified nadA mutation impacted AIEC-defining survival in macrophages, WT and NAD NC101 were subcultured in MM with and without NA and used to infect macrophage cell cultures, which were also maintained in the presence or absence of NA. Despite the expected differences in culture densities between WT and NAD strains upon subculturing in MM without NA (Figure 1C), we could obtain a sufficient amount of WT NC101 to infect with an equivalent MOI for all experiments. Baseline macrophage cell culture media contains an excess of NA, so as expected, there were not WT NC101 survival defects in the infection assay. Importantly, we found there were no significant differences in AIEC intramacrophage uptake (1 h) or survival (24 h) between WT and NAD NC101 in the presence or absence of NA supplementation (Figures 5B–D). Therefore, eliminating NA auxotrophy in NAD NC101 had negligible impact on these AIEC-associated functions.
Discussion
Inflammatory bowel disease pathogenesis differs between patients in terms of the distribution of intestinal inflammation and clinical severity (Khor et al., 2011). The intestinal microbiota influences IBD development (Sartor, 2008; Dogan et al., 2014; Baümler and Sperandio, 2016; Hughes et al., 2017; Caruso et al., 2020). So, IBD heterogeneity could be driven by microbial strain level differences that alter the functional capacity of the intestinal microbiota (Khor et al., 2011). Common members of the mammalian microbiota, like E. coli, are associated with human intestinal disease (Tenaillon et al., 2010; Mirsepasi-Lauridsen et al., 2019). Analysis of metabolic features largely distinguishes commensal intestinal E. coli strains from those known to induce intestinal and extraintestinal disease (Monk et al., 2013). However, less is known about the distinguishing features of pathobionts, which only cause disease under specific circumstances (e.g., during chronic host inflammation) (Martinez-Medina and Garcia-Gil, 2014; Mirsepasi-Lauridsen et al., 2019).
AIEC are pathobionts that exacerbate intestinal disease (Martinez-Medina and Garcia-Gil, 2014; Mirsepasi-Lauridsen et al., 2019). The evolutionary history of AIEC differs from commensal E. coli strains, which is reflected by the pro-inflammatory nature of AIEC (Martinez-Medina and Garcia-Gil, 2014; Bouvet et al., 2017; Elhenawy et al., 2019; Mirsepasi-Lauridsen et al., 2019). There is some phylogenetic linkage between AIEC and extraintestinal pathogenic E. coli (Nash et al., 2010). However, despite their ability to induce disease, AIEC do not harbor traditional virulence factors commonly found in pathogenic E. coli (Baumgart et al., 2007). Furthermore, due to AIEC genetic diversity, the genetic determinants for their pro-inflammatory functions in IBD are mostly unknown (Nash et al., 2010; Dogan et al., 2014; Martinez-Medina and Garcia-Gil, 2014). So, strain-specific studies on AIEC function are imperative to understanding their role in disease development.
Herein, we illustrate that the AIEC strain NC101 (phylotype B2) is an NA auxotroph due to a missense mutation in NAD biosynthesis gene nadA. These findings are significant, as NC101 is an established AIEC often used for studies on IBD and CRC; yet, NA auxotrophy in NC101 has not been defined (Arthur et al., 2012; Tchaptchet et al., 2013; Dogan et al., 2014; Dejea et al., 2018; Ellermann et al., 2019; Gogokhia et al., 2019; Schmitz et al., 2019). The underlying advantage of the nadA mutation in NC101 is unclear, as many E. coli isolates are prototrophic (Figure 2; Bouvet et al., 2017). However, NA auxotrophy was found to be common among urinary tract E. coli isolates and the B2 phylotype of E. coli strains that are usual intestinal inhabitants (Domergue et al., 2005; Bouvet et al., 2017). These data suggest NA auxotrophy likely confers a strain-specific selective advantage in the host.
Due to their metabolic and phenotypic diversity, E. coli strains remarkably adapt to an array of environments (Bouvet et al., 2017). Microenvironmental pressures influence pathoadaptive mutations in a niche-specific manner (Di Martino et al., 2013; Elhenawy et al., 2019). Genome reduction or loss-of-function mutations may facilitate adaptation to the intestinal microenvironment, as the decrease in biosynthetic cost of compounds likely provides an advantage when key nutrients are consistently present within an environment (Moran, 2002; Nayfach et al., 2019). For example, in E. coli, the salvage pathway for NAD biosynthesis takes precedence over de novo NAD biosynthesis when precursors (i.e., NA and Nm) are readily available (Bouvet et al., 2017). So, it is possible that an abundance of NA in the murine diet permitted loss of NadA function and improved NC101 fitness. It is also possible that among the many stochastic mutations experienced by E. coli strains, this nadA mutation simply conferred no benefit or detriment, allowing NC101 to persist as a resident microbe of the murine gastrointestinal tract.
Besides reducing biosynthetic cost, NA/NAD play a key role in virulence and signaling across various microbial species, including E. coli, Shigella spp., Candida glabrata, Bordetella pertussis, and Legionella pneumophila (Domergue et al., 2005; Prunier et al., 2007a,b; Cannizzo et al., 2011; Li et al., 2012; Di Martino et al., 2013; Edwards et al., 2013; Bouvet et al., 2017). NA auxotrophy in these species is mostly due to mutations in nadA and nadB, which suggests that convergent evolution could have given rise to an NA requirement across microbes (Domergue et al., 2005; Prunier et al., 2007a,b; Cannizzo et al., 2011; Li et al., 2012; Di Martino et al., 2013; Edwards et al., 2013; Bouvet et al., 2017; Seif et al., 2020). However, the functional consequences of NA auxotrophy promoting mutations appear to be uniquely important to each microbial species and niche (Seif et al., 2020).
In Shigella spp., a pathogen but close relative of nonpathogenic E. coli, NAD biosynthesis gene products inhibit the secretion of type 3 secretion system (T3SS) effectors and decrease intracellular invasion (Prunier et al., 2007a,b; Di Martino et al., 2013). Thus, loss of functional NAD biosynthesis genes provides a virulence advantage in Shigella spp. (Prunier et al., 2007a,b). Conversely, our results demonstrate that NA auxotrophy does not impact NC101 survival in macrophages (a pro-inflammatory and key defining feature of AIEC) (Figure 5). Like many AIEC, NC101 lacks a T3SS (accession #CP072787). Thus, unlike Shigella spp., NC101 cellular invasion and intramacrophage survival are independent of T3SS effector delivery. However, NA auxotrophy may influence AIEC function in other ways.
As seen in some species, auxotrophy can improve cellular metabolic homeostasis, protect against intracellular accumulation of reactive oxygen species, regulate signal transduction, influence colonization, and impact microbial community stability (Utsumi et al., 1994; Edwards et al., 2013; Bouvet et al., 2017; Seif et al., 2020). In E. coli, NA can regulate the EvgA/EvgS two-component regulatory system that drives multidrug resistance and acid tolerance (Utsumi et al., 1994; Edwards et al., 2013). While it was shown in urinary tract infection models that NAD auxotrophy does not impact E. coli colonization, NA auxotrophic strains have been shown to outcompete prototrophic strains in coculture competition assays (Bouvet et al., 2017). Thus, the advantage of NC101 NA auxotrophy is likely context dependent and should be the subject of future studies. Interestingly, nadA/B were listed as potential virulence factors in an IBD patient-derived AIEC (Nash et al., 2010). So, while NA auxotrophy is not ubiquitous across all AIEC and E. coli isolates, NA auxotrophy may influence the functional features of certain AIEC. Therefore, correcting NA auxotrophy in other host-adapted E. coli could globally enhance functional studies and provide further understanding of pathobiont-specific features.
The intestinal microbiota comprises a large/diverse collection of host-associated microbes, microbial genes, and products (Lopez et al., 2021). Our lab and others have been interested in pro-inflammatory and pro-carcinogenic molecules derived from intestinal bacteria, namely, yersiniabactin and colibactin (Arthur et al., 2012; Ellermann et al., 2019). Many host-influencing microbia-derived molecules, often termed specialized metabolites, are produced by sophisticated multi-enzymatic machinery encoded by bacterial biosynthetic gene clusters. By nature, many of these specialized metabolites are difficult to isolate and purify in sufficient amounts for functional analysis. However, their production can often be activated by nutrient deficiency (Hood and Skaar, 2012; Sharon et al., 2014; Baümler and Sperandio, 2016). Therefore, studies on specialized metabolites and their interactions with host cells often require precise nutrient manipulation to study in vitro. To optimize the production of these unique bioactive molecules and reduce nonessential media components that complicate purification, we have identified the MM components necessary to grow the model AIEC NC101 and generated an NC101 strain no longer restricted by NA auxotrophy. This strain, NC101 NAD, can easily be cultured in MM for functional studies or used to purify AIEC specialized metabolites. Ultimately, this strain, NAD NC101, can now serve as a research tool to investigate how precise nutrient manipulation impacts AIEC behavior under MM conditions.
Auxotrophies have been used to identify strains and determine microbial lifestyles (e.g., commensal or pathogenic) (Seif et al., 2020). Thus, defining the nutritional needs of host-adapted microbes has been imperative for functional studies on the microbiota. However, the identification of in vitro nutrient requirements is often strain-specific and can be experimentally challenging, which complicates the identification of genetic determinants underlying auxotrophy (Seif et al., 2020).
Our work highlights the genetic basis for NA auxotrophy in AIEC NC101. Correcting NC101 NA auxotrophy will (1) enable precise nutrient manipulation for in vitro studies on AIEC as they relate to IBD and CRC, (2) inform culture-based methods to evaluate the function of other auxotrophic gut microbiota members and their metabolites, and (3) facilitate small-molecule isolation and purification from the pro-inflammatory and pro-carcinogenic strain NC101. Intriguingly, our findings pose outstanding questions key to understanding the selective advantage of auxotrophy in microbiota members. Long-term, we expect our findings will contribute to the identification of microbiota-derived prognostic and therapeutic targets for human digestive diseases.
Data Availability Statement
The datasets presented in this study can be found in online repositories. The names of the repository/repositories and accession number(s) can be found in the article/Supplementary Material.
Author Contributions
LL, CJB, CAB, and JA contributed to the conceptualization and design of the study. LL was responsible for the data curation, formal analysis, visualization, and generation of the first draft. CJB, CAB, and JW wrote the sub-sections of the methods and materials. LL and JA were involved in the supervision and validation of the study. All authors involved in the investigation, reviewed, revised, and approved the submitted manuscript.
Funding
NC101 genome assembly was supported by Jeremy Wang NIH K01DK119582 (JW). The remaining body of this work was supported by funding from NIH R01DK124617 (JA), pilot funding from NIH P30DK034987 (subcontract JA), and a UNC Lineberger Comprehensive Cancer Center Innovation Award (JA).
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank the following colleagues for kindly gifting E. coli strains: R. Balfour Sartor, University of North Carolina (UNC) at Chapel Hill (E. coli NC101, LF82, and K12 MG1655); Kenneth W. Simpson, Cornell University (E. coli 42ET-1, 568-3, 37RT-2, 532-9, and 39ES-1); Leslie M. Hicks, UNC at Chapel Hill (E. coli 25922); Barry J. Campbell, University of Liverpool (E. coli HM670); Melissa Ellermann, University of South Carolina at Columbia (NC101 ΔfliC). We thank Kimberly Walker and Melissa Ellermann for critical reading of the manuscript.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fmicb.2021.670005/full#supplementary-material
Footnotes
References
Agus, A., Planchais, J., and Sokol, H. (2018). Gut microbiota regulation of tryptophan metabolism in health and disease. Cell Host Microbe 23, 716–724. doi: 10.1016/j.chom.2018.05.003
Ahmed, Z. U., Sarker, M. R., and Sack, D. A. (1988). Nutritional requirements of Shigellae for growth in a minimal medium. Infect. Immun. 56, 1007–1009. doi: 10.1128/iai.56.4.1007-1009.1988
Arthur, J. C., Gharaibeh, R. Z., Mühlbauer, M., Perez-Chanona, E., Uronis, J. M., McCafferty, J., et al. (2014). Microbial genomic analysis reveals the essential role of inflammation in bacteria-induced colorectal cancer. Nat. Commun. 5:4724. doi: 10.1038/ncomms5724
Arthur, J. C., Perez-Chanona, E., Mühlbauer, M., Tomkovich, S., Uronis, J. M., Fan, T.-J., et al. (2012). Intestinal inflammation targets cancer-inducing activity of the microbiota. Science 338, 120–123. doi: 10.1126/science.1224820
Baumgart, M., Dogan, B., Rishniw, M., Weitzman, G., Bosworth, B., Yantiss, R., et al. (2007). Culture independent analysis of ileal mucosa reveals a selective increase in invasive Escherichia coli of novel phylogeny relative to depletion of Clostridiales in Crohn’s disease involving the ileum. ISME J. 1, 403–418. doi: 10.1038/ismej.2007.52
Baümler, A. J., and Sperandio, V. (2016). Interactions between the microbiota and pathogenic bacteria in the gut. Nature 535, 85–93. doi: 10.1038/nature18849
Bergthorsson, U., and Roth, J. R. (2005). Natural isolates of Salmonella enterica serovar Dublin carry a single nadA missense mutation. J. Bacteriol. 187, 400–403. doi: 10.1128/JB.187.1.400-403.2005
Bouvet, O., Bourdelier, E., Glodt, J., Clermont, O., and Denamur, E. (2017). Diversity of the auxotrophic requirements in natural isolates of Escherichia coli. Microbiology 163, 891–899. doi: 10.1099/mic.0.000482
Cannizzo, E. S., Clement, C. C., Sahu, R., and Follo, C. (2011). Oxidative stress, inflam-aging and immunosenescence. J. Proteomics 74, 2313–2323. doi: 10.1016/j.jprot.2011.06.005
Caruso, R., Lo, B. C., and Núñez, G. (2020). Host–microbiota interactions in inflammatory bowel disease. Nat. Rev. Immunol. 20, 411–426. doi: 10.1038/s41577-019-0268-7
Carvalho, F. A., Barnich, N., Sauvanet, P., Darcha, C., Gelot, A., and Darfeuille-Michaud, A. (2008). Crohn’s disease-associated Escherichia coli LF82 aggravates colitis in injured mouse colon via signaling by flagellin. Inflamm. Bowel Dis. 14, 1051–1060. doi: 10.1002/ibd.20423
Carvalho, F. A., Koren, O., Goodrich, J. K., Johansson, M. E. V., Nalbantoglu, I., Aitken, J. D., et al. (2012). Transient inability to manage proteobacteria promotes chronic gut inflammation in TLR5-deficient mice. Cell Host Microbe 12, 139–152. doi: 10.1016/j.chom.2012.07.004
Dahlhamer, J. M., Zammitti, E. P., Ward, B. W., Wheaton, A. G., and Croft, J. B. (2016). Prevalence of inflammatory bowel disease among adults aged =18 years — United States, 2015. MMWR Morb. Mortal. Wkly. Rep. 65, 1166–1169. doi: 10.15585/mmwr.mm6542a3
Darfeuille-Michaud, A., Boudeau, J., Bulois, P., Neut, C., Glasser, A. L., Barnich, N., et al. (2004). High prevalence of adherent-invasive Escherichia coli associated with ileal mucosa in Crohn’s disease. Gastroenterology 127, 412–421. doi: 10.1053/j.gastro.2004.04.061
Darfeuille-Michaud, A., Neut, C., Barnich, N., Lederman, E., Di Martino, P., Desreumaux, P., et al. (1998). Presence of adherent Escherichia coli strains in ileal mucosa of patients with Crohn’s disease. Gastroenterology 115, 1405–1413. doi: 10.1016/S0016-5085(98)70019-8
Datsenko, K. A., and Wanner, B. L. (2000). One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl. Acad. Sci. U.S.A. 97, 6640–6645. doi: 10.1073/pnas.120163297
Dejea, C. M., Fathi, P., Craig, J. M., Boleij, A., Taddese, R., Geis, A. L., et al. (2018). Patients with familial adenomatous polyposis harbor colonic biofilms containing tumorigenic bacteria. Science 359, 592–597. doi: 10.1126/science.aah3648
Delmas, J., Gibold, L., Faïs, T., Batista, S., Leremboure, M., Sinel, C., et al. (2019). Metabolic adaptation of adherent-invasive Escherichia coli to exposure to bile salts. Sci. Rep. 9:2175. doi: 10.1038/s41598-019-38628-1
Di Martino, M. L., Fioravanti, R., Barbabella, G., Prosseda, G., Colonna, B., and Casalino, M. (2013). Molecular evolution of the nicotinic acid requirement within the Shigella/EIEC pathotype. Int. J. Med. Microbiol. 303, 651–661. doi: 10.1016/j.ijmm.2013.09.007
Dogan, B., Suzuki, H., Herlekar, D., Sartor, B. R. B., Campbell, B. J., Roberts, C. L., et al. (2014). Inflammation-associated adherent-invasive Escherichia coli are enriched in pathways for use of propanediol and iron and M-cell translocation. Inflamm. Bowel Dis. 20, 1919–1932. doi: 10.1097/MIB.0000000000000183
Domergue, R., Castano, I., De Las Penas, A., Zupancic, M., Lockatell, V., Hebel, J. R., et al. (2005). Nicotinic acid limitation regulates silencing of Candida adhesins during UTI. Science 308, 866–870.
Edwards, R. L., Bryan, A., Jules, M., Harada, K., Buchrieser, C., and Swansona, M. S. (2013). Nicotinic acid modulates Legionella pneumophila gene expression and induces virulence traits. Infect. Immun. 81, 945–955. doi: 10.1128/IAI.00999-12
Elhenawy, W., Tsai, C. N., and Coombes, B. K. (2019). Host-apecific adaptive diversification of Crohn’s disease-associated adherent-invasive Escherichia coli. Cell Host Microbe 25, 301–312.e5. doi: 10.1016/j.chom.2018.12.010
Ellermann, M., Gharaibeh, R. Z., Fulbright, L., Dogan, B., Moore, L. N., Broberg, C. A., et al. (2019). Yersiniabactin-producing adherent/invasive Escherichia coli promotes inflammation-associated fibrosis in gnotobiotic IL-10-/- mice. Infect. Immun. 87:e00587-19.
Ellermann, M., Gharaibeh, R. Z., Maharshak, N., Peréz-Chanona, E., Jobin, C., Carroll, I. M., et al. (2020). Dietary iron variably modulates assembly of the intestinal microbiota in colitis-resistant and colitis-susceptible mice. Gut Microbes 11, 32–50. doi: 10.1080/19490976.2019.1599794
Ellermann, M., Huh, E. Y., Liu, B., Carroll, I. M., Tamayo, R., and Sartor, R. B. (2015). Adherent-invasive Escherichia coli production of cellulose influences iron-induced bacterial aggregation, phagocytosis, and induction of colitis. Infect. Immun. 83, 4068–4080. doi: 10.1128/IAI.00904-15
Goethel, A., Croitoru, K., and Philpott, D. J. (2018). The interplay between microbes and the immune response in inflammatory bowel disease. J. Physiol. 596, 3869–3882. doi: 10.1113/JP275396
Gogokhia, L., Buhrke, K., Bell, R., Hoffman, B., Brown, D. G., Hanke-Gogokhia, C., et al. (2019). Expansion of bacteriophages is linked to aggravated intestinal inflammation and colitis. Cell Host Microbe 25, 285–299. doi: 10.1016/j.chom.2019.01.008
Hood, M. I., and Skaar, E. P. (2012). Nutritional immunity: transition metals at the pathogen-host interface. Nat. Rev. Microbiol. 10, 525–537. doi: 10.1038/nrmicro2836
Hughes, E. R., Winter, M. G., Duerkop, B. A., Spiga, L., Furtado de Carvalho, T., Zhu, W., et al. (2017). Microbial respiration and formate oxidation as metabolic signatures of inflammation-associated dysbiosis. Cell Host Microbe 21, 208–219. doi: 10.1016/j.chom.2017.01.005.Microbial
Kaplan, G. G. (2015). The global burden of IBD: from 2015 to 2025. Nat. Rev. Gastroenterol. Hepatol. 12, 720–727. doi: 10.1038/nrgastro.2015.150
Khor, B., Gardet, A., and Xavier, R. J. (2011). Genetics and pathogenesis of inflammatory bowel disease. Nature 474, 307–317. doi: 10.1038/nature10209
Kim, S. C., Tonkonogy, S. L., Albright, C. A., Tsang, J., Balish, E. J., Braun, J., et al. (2005). Variable phenotypes of enterocolitis in interleukin 10-deficient mice monoassociated with two different commensal bacteria. Gastroenterology 128, 891–906. doi: 10.1053/j.gastro.2005.02.009
Kittana, H., Gomes-Neto, J. C., Heck, K., Sughroue, J., Xian, Y., Mantz, S., et al. (2019). Establishing the phenotypic basis of adherent-invasive Escherichia coli (AIEC) pathogenicity in intestinal inflammation. bioRxiv [Preprint]. doi: 10.1101/772012
Li, H. (2016). Minimap and miniasm: fast mapping and de novo assembly for noisy long sequences. Bioinformatics 32, 2103–2110. doi: 10.1093/bioinformatics/btw152
Li, Z., Bouckaert, J., Deboeck, F., de Greve, H., and Hernalsteens, J. P. (2012). Nicotinamide dependence of uropathogenic Escherichia coli UTI89 and application of nadB as a neutral insertion site. Microbiology 158, 736–745. doi: 10.1099/mic.0.052043-0
Liu, G., Foster, J., Manlapaz-Ramos, P., and Olivera, B. M. (1982). Nucleoside salvage pathway for NAD biosynthesis in Salmonella typhimurium. J. Bacteriol. 152, 1111–1116.
Lopez, L. R., Bleich, R. M., and Arthur, J. C. (2021). Microbiota effects on carcinogenesis: Initiation, promotion and progression. Annu. Rev. Med. 72, 243–261. doi: 10.1146/annurev-med-080719-091604
Martin, H. M., Campbell, B. J., Hart, C. A., Mpofu, C., Nayar, M., Singh, R., et al. (2004). Enhanced Escherichia coli adherence and invasion in Crohn’s disease and colon cancer. Gastroenterology 127, 80–93. doi: 10.1053/j.gastro.2004.03.054
Martinez-Medina, M., Denizot, J., Dreux, N., Robin, F., Billard, E., Bonnet, R., et al. (2014). Western diet induces dysbiosis with increased E coli in CEABAC10 mice, alters host barrier function favouring AIEC colonisation. Gut 63, 116–124. doi: 10.1136/gutjnl-2012-304119
Martinez-Medina, M., and Garcia-Gil, L. J. (2014). Escherichia coli in chronic inflammatory bowel diseases: an update on adherent invasive Escherichia coli pathogenicity. World J. Gastrointest. Pathophysiol. 5, 213. doi: 10.4291/wjgp.v5.i3.213
Mirsepasi-Lauridsen, H. C., Vallance, B. A., Krogfelt, K. A., and Petersen, A. M. (2019). Escherichia coli pathobionts associated with inflammatory bowel disease. Clin. Microbiol. Rev. 32, e00060–18. doi: 10.1128/CMR.00060-18
Monk, J. M., Charusanti, P., Aziz, R. K., Lerman, J. A., Premyodhin, N., Orth, J. D., et al. (2013). Genome-scale metabolic reconstructions of multiple Escherichia coli strains highlight strain-specific adaptations to nutritional environments. Proc. Natl. Acad. Sci. U.S.A. 110, 20338–20343. doi: 10.1073/pnas.1307797110
Moran, N. A. (2002). Microbial minimalism: genome reduction in bacterial pathogens. Cell 108, 583–586. doi: 10.1016/S0092-8674(02)00665-7
Nadeem, M. S., Kumar, V., Al-Abbasi, F. A., Kamal, M. A., and Anwar, F. (2019). Risk of colorectal cancer in inflammatory bowel diseases. Semin. Cancer Biol. 64, 51–60. doi: 10.1016/j.semcancer.2019.05.001
Nash, J. H. E., Villegas, A., Kropinski, A. M., Aguilar-Valenzuela, R., Konczy, P., Mascarenhas, M., et al. (2010). Genome sequence of adherent-invasive Escherichia coli and comparative genomic analysis with other E. coli pathotypes. BMC Genomics 11:667. doi: 10.1186/1471-2164-11-667
Nayfach, S., Shi, Z. J., Seshadri, R., Pollard, K. S., and Kyrpides, N. C. (2019). New insights from uncultivated genomes of the global human gut microbiome. Nature 568, 505–510. doi: 10.1038/s41586-019-1058-x
Ormsby, M. J., Johnson, S. A., Carpena, N., Meikle, L. M., Goldstone, R. J., McIntosh, A., et al. (2020). Propionic acid promotes the virulent phenotype of Crohn’s disease-associated adherent-invasive Escherichia coli. Cell Rep. 30, 2297–2305. doi: 10.1016/j.celrep.2020.01.078
Patwa, L. G., Fan, T. J., Tchaptchet, S., Liu, Y., Lussier, Y. A., Sartor, R. B., et al. (2011). Chronic intestinal inflammation induces stress-response genes in commensal Escherichia coli. Gastroenterology 141, 1842–1851. doi: 10.1053/j.gastro.2011.06.064
Perry, R. D., and Fetherston, J. D. (2011). Yersiniabactin iron uptake: mechanisms and role in Yersinia pestis pathogenesis. Microbes Infect. 13, 808–817. doi: 10.1016/j.micinf.2011.04.008.Yersiniabactin
Prunier, A. L., Schuch, R., Fernández, R. E., Kohler, H., McCormick, B. A., and Maurelli, A. T. (2007a). nadA and nadB of Shigella flexneri 5a are antivirulence loci responsible for the synthesis of quinolinate, a small molecule inhibitor of Shigella pathogenicity. Microbiology 153, 2363–2372. doi: 10.1099/mic.0.2007/006916-0
Prunier, A. L., Schuch, R., Fernández, R. E., and Maurelli, A. T. (2007b). Genetic structure of the nadA and nadB antivirulence loci in Shigella spp. J. Bacteriol. 189, 6482–6486. doi: 10.1128/JB.00525-07
Rieder, F., Brenmoehl, J., Leeb, S., Schölmerich, J., and Rogler, G. (2007). Wound healing and fibrosis in intestinal disease. Gut 56, 130–139. doi: 10.1136/gut.2006.090456
Sambrook, J., and Russell, D. W. (2006). The Condensed Protocols from Molecular Cloning?: A Laboratory Manual. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press.
Sartor, R. B. (2006). Mechanisms of disease: pathogenesis of Crohn’s disease and ulcerative colitis. Nat. Clin. Pract. Gastroenterol. Hepatol. 3, 390–407. doi: 10.1038/ncpgasthep0528
Sartor, R. B. (2008). Microbial influences in inflammatory bowel diseases. Gastroenterology 134, 577–594. doi: 10.1053/j.gastro.2007.11.059
Schirmer, M., Garner, A., Vlamakis, H., and Xavier, R. J. (2019). Microbial genes and pathways in inflammatory bowel disease. Nat. Rev. Microbiol. 17, 497–511. doi: 10.1038/s41579-019-0213-6
Schmitz, J. M., Tonkonogy, S. L., Dogan, B., Leblond, A., Whitehead, K. J., Kim, S. C., et al. (2019). Murine adherent and invasive E. coli induces chronic inflammation and immune responses in the small and large intestines of monoassociated IL-10-/- mice independent of long polar fimbriae adhesin A. Inflamm. Bowel Dis. 25, 875–885. doi: 10.1093/ibd/izy386
Seif, Y., Choudhary, K. S., Hefner, Y., Anand, A., Yang, L., and Palsson, B. O. (2020). Metabolic and genetic basis for auxotrophies in Gram-negative species. Proc. Natl. Acad. Sci. U.S.A. 117, 6264–6273. doi: 10.1073/pnas.1910499117
Sharon, G., Garg, N., Debelius, J., Knight, R., Dorrestein, P. C., and Mazmanian, S. K. (2014). Specialized metabolites from the microbiome in health and disease. Cell Metab. 20, 719–730. doi: 10.1016/j.cmet.2014.10.016
Small, C. L. N., Reid-Yu, S. A., McPhee, J. B., and Coombes, B. K. (2013). Persistent infection with Crohn’s disease-associated adherent-invasive Escherichia coli leads to chronic inflammation and intestinal fibrosis. Nat. Commun. 4:1957. doi: 10.1038/ncomms2957
Tchaptchet, S., Fan, T. J., Goeser, L., Schoenborn, A., Gulati, A. S., Sartor, R. B., et al. (2013). Inflammation-induced acid tolerance genes gadAB in luminal commensal Escherichia coli attenuate experimental colitis. Infect. Immun. 81, 3662–3671. doi: 10.1128/IAI.00355-13
Tenaillon, O., Skurnik, D., Picard, B., and Denamur, E. (2010). The population genetics of commensal Escherichia coli. Nat. Rev. Microbiol. 8, 207–217. doi: 10.1038/nrmicro2298
Tronnet, S., Garcie, C., Rehm, N., Dobrindt, U., Oswald, E., and Martin, P. (2016). Iron homeostasis regulates the genotoxicity of Escherichia coli that produces colibactin. Infect. Immun. 84, 3358–3368. doi: 10.1128/IAI.00659-16
Utsumi, R., Katayama, S., Taniguchi, M., Horie, T., Ikeda, M., Igaki, S., et al. (1994). Newly identified genes involved in the signal transduction of Escherichia coli K-12. Gene 140, 73–77. doi: 10.1016/0378-1119(94)90733-1
Vaser, R., Soviæ, I., Nagarajan, N., and Šikiæ, M. (2017). Fast and accurate de novo genome assembly from long uncorrected reads. Genome Res. 27, 737–746. doi: 10.1101/gr.214270.116
Keywords: inflammatory bowel disease, adherent invasive E. coli, Escherichia coli, auxotrophy, nicotinic acid, E. coli NC101
Citation: Lopez LR, Barlogio CJ, Broberg CA, Wang J and Arthur JC (2021) A nadA Mutation Confers Nicotinic Acid Auxotrophy in Pro-carcinogenic Intestinal Escherichia coli NC101. Front. Microbiol. 12:670005. doi: 10.3389/fmicb.2021.670005
Received: 19 February 2021; Accepted: 12 April 2021;
Published: 02 June 2021.
Edited by:
Johann Heider, University of Marburg, GermanyReviewed by:
Liya Liang, University of Colorado Boulder, United StatesRongming Liu, University of Colorado Boulder, United States
Copyright © 2021 Lopez, Barlogio, Broberg, Wang and Arthur. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Janelle C. Arthur, SmFuZWxsZV9hcnRodXJAbWVkLnVuYy5lZHU=
†Present address: Cassandra J. Barlogio, Locus Biosciences, Inc., Durham, NC, United States; Christopher A. Broberg, Department of Chemistry, The University of North Carolina at Chapel Hill, Chapel Hill, NC, United States
 Lacey R. Lopez
Lacey R. Lopez Cassandra J. Barlogio
Cassandra J. Barlogio Christopher A. Broberg
Christopher A. Broberg Jeremy Wang2
Jeremy Wang2 Janelle C. Arthur
Janelle C. Arthur