
94% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Microbiol., 19 November 2020
Sec. Microbiological Chemistry and Geomicrobiology
Volume 11 - 2020 | https://doi.org/10.3389/fmicb.2020.536535
 Sean W. Mullin1
Sean W. Mullin1 Greg Wanger2†
Greg Wanger2† Brittany R. Kruger3
Brittany R. Kruger3 Joshua D. Sackett3
Joshua D. Sackett3 Scott D. Hamilton-Brehm4
Scott D. Hamilton-Brehm4 Rohit Bhartia2†
Rohit Bhartia2† Jan P. Amend5
Jan P. Amend5 Duane P. Moser3
Duane P. Moser3 Victoria J. Orphan1*
Victoria J. Orphan1*The microbial ecology of the deep biosphere is difficult to characterize, owing in part to sampling challenges and poorly understood response mechanisms to environmental change. Pre-drilled wells, including oil wells or boreholes, offer convenient access, but sampling is frequently limited to the water alone, which may provide only a partial view of the native diversity. Mineral heterogeneity demonstrably affects colonization by deep biosphere microorganisms, but the connections between the mineral-associated and planktonic communities remain unclear. To understand the substrate effects on microbial colonization and the community response to changes in organic carbon, we conducted an 18-month series of in situ experiments in a warm (57°C), anoxic, fractured carbonate aquifer at 752 m depth using replicate open, screened cartridges containing different solid substrates, with a proteinaceous organic matter perturbation halfway through this series. Samples from these cartridges were analyzed microscopically and by Illumina (iTag) 16S rRNA gene libraries to characterize changes in mineralogy and the diversity of the colonizing microbial community. The substrate-attached and planktonic communities were significantly different in our data, with some taxa (e.g., Candidate Division KB-1) rare or undetectable in the first fraction and abundant in the other. The substrate-attached community composition also varied significantly with mineralogy, such as with two Rhodocyclaceae OTUs, one of which was abundant on carbonate minerals and the other on silicic substrates. Secondary sulfide mineral formation, including iron sulfide framboids, was observed on two sets of incubated carbonates. Notably, microorganisms were attached to the framboids, which were correlated with abundant Sulfurovum and Desulfotomaculum sp. sequences in our analysis. Upon organic matter perturbation, mineral-associated microbial diversity differences were temporarily masked by the dominance of putative heterotrophic taxa in all samples, including OTUs identified as Caulobacter, Methyloversatilis, and Pseudomonas. Subsequent experimental deployments included a methanogen-dominated stage (Methanobacteriales and Methanomicrobiales) 6 months after the perturbation and a return to an assemblage similar to the pre-perturbation community after 9 months. Substrate-associated community differences were again significant within these subsequent phases, however, demonstrating the value of in situ time course experiments to capture a fraction of the microbial assemblage that is frequently difficult to observe in pre-drilled wells.
The continental deep biosphere represents approximately 5–15% of the biomass on Earth, or about 27–64 Gt of carbon, yet the ecological conditions that govern community structure and activity remain poorly constrained (McMahon and Parnell, 2014; Bar-On et al., 2018; Magnabosco et al., 2018). One factor that complicates our understanding of this vast habitat is that microbial community composition is often predicated on highly heterogeneous local mineralogy and geochemistry (Orcutt et al., 2011; Sylvan et al., 2013; Toner et al., 2013; Jones and Bennett, 2014). Most cells in the deep subsurface are attached to surfaces, not planktonic, with attached cells estimated to outnumber free-living cells by a factor of 10–103 (Wanger et al., 2007; McMahon and Parnell, 2014). Evidence suggests that not only do the mineralogical composition and crystallographic axes affect which microbes attach via electrostatic interactions (Yee et al., 2000; Edwards and Rutenberg, 2001), but also that microbes actively weather minerals to access scarce nutrients or by way of their central metabolism (Lovley et al., 1989; Rogers et al., 1998; Caccavo and Das, 2002; Roberts, 2004). These effects can often be challenging to detect in situ due to the difficulty of accessing underground fracture networks, and because mineralogy typically varies concurrently with changes in geochemistry, temperature, porosity, and pressure along a depth axis. In situ experiments in terrestrial boreholes drilled from the surface are an attractive option because they provide direct access to rock surfaces in the native subsurface biogeochemical context. Subseafloor biosphere studies, such as (Orcutt et al., 2010), which deployed flow-through columns containing mineral colonization substrates of interest, can provide a good model for terrestrial experiments. In situ incubation of a variety of mineral substrates enables the study of microorganisms adhering to differing mineral surfaces and further reveals the differences in composition between mineral-adhered and free-living microbial populations. Although drilling in terrestrial environments is comparatively more accessible than in marine systems as described in other studies (Orcutt et al., 2010), such experiments are not without their own challenges (Wilkins et al., 2014). Drilling always carries a risk of contamination, but drilling contamination can be mitigated in pumped or free-flowing boreholes by flushing with local groundwater to remove contaminants and re-establish the indigenous community (Moser et al., 2003; Davidson et al., 2011).
Describing the ecology of the subsurface can also prove difficult because of the extremely slow growth and metabolism of many resident microorganisms. In situ doubling times for virtually all deep biosphere organisms are unknown and most studies rely on biomass turnover rates instead, calculated from cell-specific activity measurements and amino acid racemization studies (Kieft and Phelps, 1997; Morono et al., 2011; Onstott et al., 2014; Braun et al., 2017; Trembath-Reichert et al., 2017). These studies suggest biomass turnover rates in the deep biosphere range from several months to thousands of years. In fact, in the most oligotrophic environments such as the ocean gyre subseafloor, reactive transport models indicate that resident cells, if they are growing, either have energy requirements below the reported power usage minimum required for cellular maintenance (LaRowe and Amend, 2015) or are not growing at all, but slowly dying (Bradley et al., 2018). In less austere environments, such as the coastal marine subsurface or the terrestrial deep biosphere, microbial community analyses do not indicate that these environments only host poorly-adapted surface organisms. Instead, these environments are home to their own endemic, uncultivated, and previously unknown organisms such as the aptly named Deep Sea Archaeal Group, novel branches of Chloroflexi, or Candidate Phyla (Takai et al., 2001; Inagaki et al., 2006; Jørgensen and Boetius, 2007; Osburn et al., 2014). The terrestrial deep biosphere, accessed via boreholes or mines, has proven especially fruitful for the discovery of novel microbial diversity, including from the site in this study (Figure 1; Chivian et al., 2008; Sahl et al., 2008; Teske and Sørensen, 2008; Breuker et al., 2011; Osburn et al., 2014; Momper et al., 2017, 2018; Sackett, 2018; Sackett et al., 2019). Little is known, however, about the changes in deep subsurface systems of yearly or longer timescales. It may be possible that deep biosphere organisms persist with little to no growth until a rare nutrient or energy delivery, perhaps mediated by tectonic disturbance (Riedinger et al., 2015). Controlled substrate addition experiments performed in situ offer a novel experimental approach to understand the influence of transient energy influx on deep biosphere community structure and help constrain the energy requirements and maximum metabolic rates of these organisms.
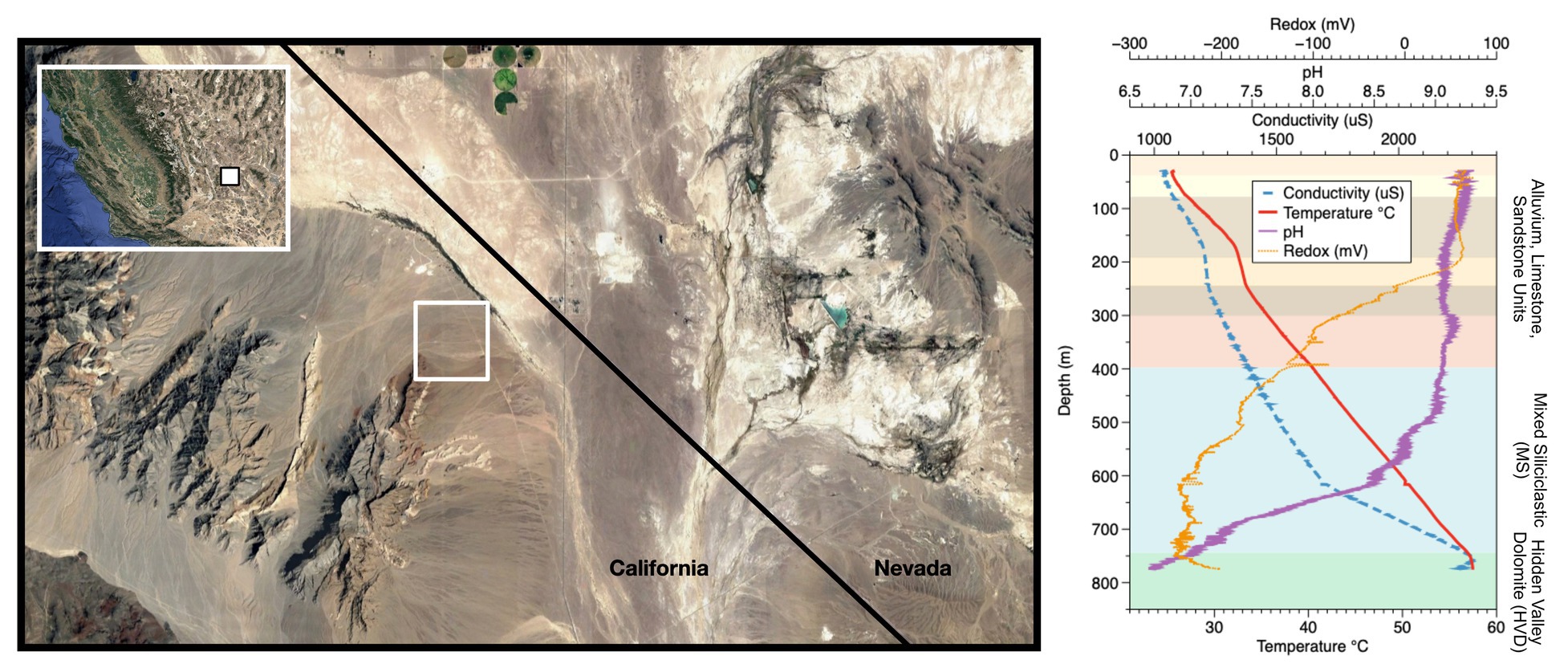
Figure 1. Site location and well profile. (Left) Inyo-BLM 1 is located northeast of Bat Mtn. in the Funeral Mountain Range. Map data: Google, Maxar Technologies. (Right) Geochemical profile of the well overlaid with lithological units as described in Fridrich et al. (2012). The well itself is cased in steel down to 750 m depth, at which point it is open to the Hidden Valley Dolomite (HVD) unit. Geochemical data from the groundwater within the well, which begins at around 28 m depth from surface, are displayed. Although the borehole was originally drilled to 884 m depth, infilling of soft sediments now limits access to approximately 755 m. Map data copyright: attribution of the map data follows Google Earth guidelines.
Here, we describe an 18-month series of five in situ microbial colonization experiments to characterize and constrain the microbial community and activity of a warm, continental biosphere from a ~775 m-deep borehole (Inyo-BLM 1) hosted in a fractured, Paleozoic carbonate beneath the Mojave Desert, CA, United States (Figure 1). We demonstrate that not only is the mineral-hosted microbial community significantly different from the planktonic community, but that mineral-hosted communities vary based on the mineralogy. We also demonstrate that a controlled organic substrate addition strongly affected the in situ microbial community composition and revealed subsurface taxa that were poised to take advantage of introduced complex organic matter.
The Inyo-BLM 1 well was completed in 2007 to a total depth of 884 m below land surface (mbls) by The Hydrodynamics Group, LLC (Edmonds, WA) on behalf of Inyo County, California (Belcher et al., 2009; Bredehoeft and King, 2010). The well pad is on the eastern flank of the Funeral Mountains in the Amargosa Valley, proximal to the California-Nevada border near Death Valley National Park (36°24'04.19N/116°28'06.58W). Inyo-BLM 1 was drilled to test hypotheses concerning deep groundwater flow across the Funeral Mountains and intersects a variety of lithologies (e.g., sandstone and tuff), ultimately encountering Hidden Valley Dolomite (HVD; Silurian-Devonian, 423-393 Ma, in age) at 744 mbls (Fridrich et al., 2012), which is associated with the Lower Carbonate Aquifer (LCA), a warm, fractured rock aquifer in the discharge zone of the Death Valley Regional Flow System (Belcher et al., 2001; Bredehoeft et al., 2008; Figure 1). Inyo-BLM 1 was pumped for development and hydrologic testing in both 2007 and 2011 by pumping out approximately 1.6 million liters of water (~100 times total volume of groundwater in Inyo-BLM 1) and then allowed to re-equilibrate with the surrounding anoxic aquifer. Other than these two pumping events in 2007 and 2011, the hole sat undisturbed apart from continuous monitoring of the static water level by the National Park Service to measure water table elevation until our sampling began in 2014. Although originally drilled to a total depth of 884 m, at the time of sampling, Inyo-BLM 1 was only accessible to 755 mbls, presumably due to collapse of unstable rock below the casing. Inyo-BLM 1 is continuously cased in unscreened low-carbon steel and cemented in a step-down manner (Bredehoeft et al., 2008); caliper measurements were completed in 2015 showing that the inner diameter decreases from 14.5–15 cm diameter (0–610 mbls) to 8.4–8.6 cm diameter (610–750 mbls), with the final 5 m open to the host rock and the warm (57°C) reducing fluids of the LCA. The borehole passes through several lithologies, including sandstone, siltstone, and mixed siliciclastic and tuff lake sediments; however, the open hole section is entirely hosted within the HVD within the Lost Burro Formation, roughly Silurian-Devonian (423–393 Ma) in age (Fridrich et al., 2012). Although the Inyo-BLM 1 drill pad is located hundreds of meters above the valley floor (694 m above sea level), hydrostatic pressure maintains the groundwater at approximately 28 m below the surface (USGS, 2018). A study by Thomas et al. (2013) indicated that although this hole likely shares the same hydrological recharge area as other wells in the area, including Ash Meadows springs, it is likely primarily sourced from a deeper and older portion of aquifer. A thermal flowmeter was deployed on August 21, 2015 to test for flow within the well, but none was detected (i.e., <0.08 L per min; not shown).
To characterize the rock-associated microbial community, several sterilized solid substrates of defined or native materials were incubated in situ within polytetrafluoroethylene (PTFE) open-ended cartridges (2 cm inner diameter, approximately 18 cm long) screened with 1 mm nylon mesh. Cartridges were filled with pre-characterized substrates, which consisted of either mineral standards or cuttings of the appropriate lithologic units obtained during the drilling process and archived at the USGS Core Library and Data Center (Mercury, NV). Incubated standard substrates included glass wool (Sigma Aldrich #18421), dolomite (Ward’s Science #470025-558), pyrite (Ward’s Science #470025-736), optically clear calcite (Ward’s Science 470025-522), pyrrhotite (Ward’s 470025-750), and 3 mm diameter low-carbon steel ball bearings (McMaster-Carr 96455K49) as a proxy for the well casing. Natural lithologies tested included HVD, a fine-grained dark-gray dolomite with nodular chert, and mixed siliciclastic lake sediments and tuff (MS). Crushed minerals ranged in size from 2 to 5 mm. In situ incubations containing dolomite, HVD, calcite, or MS were always conducted in duplicate (n = 2 cartridges) for each time point. Cartridges were assembled into bundles of three, secured with 200-test Dyneema™ high-strength line (Supplementary Figure S2). Duplicate cartridges were always assembled into separate bundles. Three 5 mm PTFE rods acted as a cage for the bundles. The entire assembly was autoclaved and maintained in a sterile bag until deployment. During the deployment of the cartridges, an A-frame was constructed over the well cap, and the cartridge string was lowered into the well affixed to both 200-test and 500-test Dyneema™ fishing line as a failsafe. At regular intervals (~75–100 m) a series of polyurethane (PUR) foam blocks (Identi-plug™, Fisher Scientific Inc.) were attached to the fishing line with nylon cable ties (McMaster-Carr 70215K96) following Imachi et al. (2011). The incubation package was suspended in the open hole portion of the borehole at ~752 m depth, within the HVD unit, for periods of 3–6 months before retrieval with five total incubations over 18 months (Table 1). Upon retrieval, cartridges were stored at ambient temperature in oxygen impermeable mylar bags with sterile oxygen scrubbing packets (Oxy-Sorb; Harrisburg, NC) until they could be disassembled and subsampled under sterile conditions in a laboratory anaerobic chamber the following day. Subsamples were frozen at −80°C for DNA analysis or fixed individually in electron microscopy-grade 2% paraformaldehyde or 2.5% glutaraldehyde (Electron Microscopy Sciences; Hatfield, PA) for microscopy. The dates and mineral substrates included for each deployment are provided in Table 1.
In the period following retrieval of Incubation 2 and prior to deployment of Incubation 3, the borehole was chemically and physically characterized using calipers to measure the interior diameter, a thermal flow meter, and a QL40-Ocean Idronaut wellbore logging multiprobe (Idronaut, S.R.L.; Brugherio, Italy) that concurrently measures temperature, pH, relative oxygen reduction potential (ORP), and conductivity over the entire depth profile. Several liters of water from 579 to 752 m were also collected at this time by successive collection at ambient pressure with gas-tight discrete samplers (“bailers,” Comprobe, Inc.; Fort Worth, TX) for extensive aqueous geochemical and dissolved gas characterization, with the shallower depth sampled first to prevent mixing the water column. Dissolved metals and ions were quantified by ACZ Laboratories (Steamboat, CO), and dissolved organic carbon (DOC), particulate organic carbon (POC), and total organic carbon (TOC) were quantified by Anatek Labs (Spokane, WA), both of which are accredited environmental testing facilities. The methodologies and standard error values for these analyses are included in Supplementary Table S2, with the error values from ACZ Laboratories generated from duplicate analyses of a standard and the error values from Anatek Labs from duplicate sample analyses. Samples from two depths were analyzed (579 and 752 mbls). Sulfide was measured using the methylene blue method with a field detection kit (detection limit ~0.02 mM; ChemMet, LLC.; Buffalo, New York), following the manufacturer’s protocol. We also conducted a qualitative in situ test for dissolved sulfide using silver nitrate-coated film, following (Fike et al., 2017). The environmental detection limit of this method has not been well-quantified, but data suggests that 20 min of exposure to a 0.9 mM sulfide solution is more than sufficient for visualization. The film was attached to the discrete water sampler, lowered to 752 m depth in the uncased portion of the borehole, left stationary for 2 min, and then recovered to the surface, for a total time of ~45 min.
To measure dissolved gases in the well waters (Supplementary Table S2), sterilized 160 ml serum bottles were prepared with 50 μl saturated (0.07 g/ml) HgCl2 solution and capped with butyl rubber stoppers. The HgCl2 solution was then evaporated away to dryness under constant vacuum, leaving only the 3.5 mg of HgCl2 behind. Water samples were collected by draining the first ~50 ml fluid fractions from three separate 750 m depth water samples through platinum-cured silicone tubing (Masterflex LS-24) and a 25 G needle into the pre-evacuated serum bottles. The filled vials were stored upside down until their analysis within 8 weeks. Headspace concentrations of H2, O2, N2, CO2, CO, ethane, and propane were measured using a Shimadzu GC-2014ATF headspace gas chromatograph (GC) equipped with a Haysep 80/100 (5 m) and MS-5A 60/80 (2.5 m) molecular sieve columns and TCD and FID detectors. Samples were measured twice (relative percent differences listed in Supplementary Table S2) and were compared against a 1% standard mixture (Restek; Bellefonte, PA). See (Rowe et al., 2017) for details.
Microbial cell counts of the groundwater were performed in two ways (live and fixed) on fluids from 579 to 752 mbls collected August 20–21, 2015. Samples collected from three separate casts for each depth were counted in each way. Live samples were stored at 4°C until they were processed (August 21). Samples were also fixed in the field with 1% paraformaldehyde and stored at 4°C until they were processed on August 23, 2015. Live samples were counted with a Petroff-Hausser counting chamber, and fixed samples were stained with 4',6-diamidino-2-phenylindole (DAPI) and counted on fluorescence microscope (Olympus BX51, Olympus Scientific Solutions; Waltham, MA) at 1,000× magnification (Supplementary Table S3).
As part of a separate experiment, five natural sea sponges (Constantia; Hawthorne, CA) were attached to the line directly above the mineral cartridges (~752 mbls) during Incubation 3 as well as higher up on the suspension line at 579 mbls. All sponges from the 752 mbls set, closest to the experimental cartridges, were consumed by microbial activity by the time of recovery; one sponge at 579 mbls remained at the end of the experiment, however, this sponge was noticeably degraded with a gelatinous consistency. This degraded material was salvaged from the line and frozen for microbial community analysis.
To approximate the amount of organic carbon and nitrogen introduced into the deep aquifer and potentially consumed by in situ microorganisms, the composition of the sponges was estimated. The purchased sea sponges deployed in Incubation 3 belong to the family Dictyoceratida, which have minimal or no calcareous spicules in their skeleton and are primarily composed of spongin, a collagenous halogenated glycoprotein with high thermal stability up to ~360°C (Ehrlich et al., 2003, 2010; Green et al., 2003; Norman et al., 2016). Prior to deployment, the sponges were washed, bleached with hydrogen peroxide and rinsed, and sterilized by autoclaving for 30 min. Spongin’s chemical formula is unknown, but the carbon and nitrogen content has been quantified (carbon = 47.44% and nitrogen = 16.15%; Gamgee, 1880). Each sponge was on average 0.37 g, and based on the observation that all sponges from the region proximal to the Incubations appeared to be entirely degraded by the time Incubation 3 was recovered, we estimate the total carbon and nitrogen added to the system in the proximity of the mineral cartridges was ~0.88 and ~0.30 g, respectively.
We developed a rough diffusion-based model to approximate the time for degradation and dispersal of the sponge-derived organic matter. As no vertical flow was detected within the well, we assumed dispersal of the solubilized, degraded sponge substrate occurred through molecular diffusion. This model was simplified to a one-dimensional diffusion in two directions, described by the thin-film (Gaussian) solution to Fick’s Second Law:
where C(x,t) is the concentration of the molecule of interest at position x and time t, N is the moles of the molecule per unit area at x0 and t0 (12.58 mol C/m2, given well diameter of ~8.6 cm), and D is the diffusion coefficient (Mehrer, 2007). We bounded the time range for the diffusion of the sponge nutrients by setting t0 as either the moment the sponges are in place or halfway through Incubation 3, which assumes that the sponge organics do not diffuse for the first 37 days and then become completely diffusible after that time (37 ≤ t0 ≤ 74). Diffusion coefficients, D, are normally only determined empirically, which was not possible here. As an approximation, we used the D for lactose at 55°C: 1.06 × 10−9 m2/s (Ribeiro et al., 2006). Based on the cartridge bundle dimensions, which extended from approximately 0.25–2.0 m from the sponges, and accounting for the well casing with an average diameter of 0.086 m, we estimate 10.6 L of water surrounded the cartridge bundle. Using all of these parameters, we determined a rough estimate of the total moles of organic carbon released into the water surrounding the cartridges by integrating C(x) over the distance at either t = 37 days or t = 74 days (depending on which t0 is used) and multiplying by the volume, which translates to values in the range of 160 micro µ mol carbon to 3.01 mmol carbon. We thus assume that retrieval of the experiment would mix any remaining dissolved carbon species upward into the remaining 733 m of the water column; diffusion downward from these distances over the remaining time scale of our experiments is trivial.
Hydrogen production from the steel alloy used in the well casing was verified (see Supplementary Figure S5) by adding one of several Fe-bearing substrates to a defined medium (Supplementary Table S1) designed to approximate well fluids at 579 mbls as determined by the geochemistry suite from 8/21/15 described above (Supplementary Table S2). The substrates tested for hydrogen production were low-carbon steel analogous to the well casing material (McMaster-Carr 96455K49), 304 stainless steel (McMaster-Carr 9291K43), 316 stainless steel (McMaster-Carr 96415K71), and hematite (Ward’s Science 470025). Three replicate vials were measured for each substrate. For steel incubations, three steel ball bearings were added to the sterilized defined medium, and for hematite, 2–5 mm crushed rock was weighed to be volumetrically equivalent to the steel. The headspace of each tube was measured with the same headspace gas chromatograph program as was used to measure the natural samples, as described above.
DNA was extracted from environmental samples with the Qiagen PowerSoil DNA Extraction Kit (Qiagen; Hilden, Germany; 12888) according to the kit protocol except for the following changes during the lysis step, performed with the intention of extracting DNA from as much of the community as possible. Following the addition of 0.25–0.5 g of sample to the lysis tubes, the tubes were flash frozen in liquid nitrogen. The samples were thawed at room temperature and then vortexed for 1 min. The lysis buffer (30 μl C1 buffer) was then added and the samples were incubated in a 65°C water bath for 30 min, with each sample vortexed for 1 min every 10 min. The samples were then placed in a bead beater (FastPrep FP120, Thermo Fisher Scientific; Waltham, MA) at 5.5 m/s speed for 45 s. The samples were then centrifuged at 10,000 × g for 30 s and processed according to the rest of the manufacturer’s instructions. To keep extraction conditions consistent, filtered and unfiltered water samples were extracted with the same kit. Filtered water samples consisted of 250 ml of raw water passed through a 0.22 μm PES filter (UX-06730-43, Thermo Scientific; Waltham, MA). Approximately 0.25 g of this filter was then processed as above. To prevent detection bias against cells smaller than 0.22 μm, unfiltered water was also extracted by adding 250 μl of Inyo-BLM 1 water to the PowerSoil lysis tubes and then processed with the same alterations to the extraction protocol.
Microbial diversity was assessed by Illumina iTag sequencing of the V4 region of the 16S rRNA gene, using a protocol recommended by the Earth Microbiome Project (Caporaso et al., 2012) and sequenced with Illumina Tag sequencing by Laragen, Inc. (Los Angeles, CA) as previously described (Caporaso et al., 2012; Case et al., 2015). The V4 region of the 16S rRNA gene was amplified using degenerate 515F and 806R primer pairs (5'-GTGCCAGCMGCCGCGGTAA-3' and 5'-GGACTACHVGGGTWTCTAAT-3'). Each sample was amplified in duplicate and the resulting amplicons were pooled. Each reaction was 15 μl total: 7.5 μl Q5 Hot Start High-Fidelity 2x Master Mix (New England BioLabs, Inc.), 5 μl DNAse-free water, 0.75 μl of each primer at 10 μM, and 1 μl of gDNA extract (~0.5 ng/μl) from the sample. Reactions were held at 98°C for 2 min to denature the DNA, with amplification proceeding for 30 cycles at 98°C for 10 s, 54°C for 20 s, 72°C for 20 s; a final extension of 72°C for 2 min was added for complete amplification. These PCR products are checked for purity via gel electrophoresis and the DNA concentration quantified by Qubit HS (Bio-Rad, Inc.) before they are barcoded for sequencing. Barcoding primers include the original primer sequence as well as linkers, adapters, and on the reverse primer, a unique barcode as described previously (Mason et al., 2015). The reaction used 5 μl of the 16S rRNA DNA product, 12.5 μl of Q5 Host Start Master Mix, 1.25 μl of 10 μM forward primer, 5 μl of 2.5 μM reverse primer, and 1.25 μl DNAse-free water. Reaction temperatures were the same as above.
Extraction controls and no-template controls were included in every sequencing run; controls were amplified with 45 reaction cycles in the initial PCR reaction to produce a quantifiable PCR product. Autoclaved control minerals that were not incubated in situ were also extracted and sequenced after 45 cycles of amplification (control minerals amplified 30 times did not produce a detectable gel electrophoresis band) to control for DNA that may have survived sterilization. Supplementary Figure S1 is a comparison of mineral controls vs. incubated minerals. Sequencing sample metadata is listed in Supplementary Table S5, and the resulting OTU table with the associated sequence and taxonomic identification is listed in Supplementary Table S6. Raw sequenced data is available at the NCBI SRA database with accession number PRJNA605006.
In-house amplicon processing was completed with QIIME 1.8.0 and included joining paired-ends, quality trimming, 99% OTU clustering, singleton removal, and 0.01% relative abundance threshold removal (Caporaso et al., 2010). Remaining OTUs were then manually checked against sequenced extraction and no-template controls and non-incubated mineral controls to remove contaminating sequences. OTUs with relative abundance ≥0.05% in the controls were removed from the dataset. A total of 118 OTUs were removed during this step, leaving 520 OTUs remaining after all filtering steps. The average number of reads per sample remaining after quality-score filtering and removal of singletons, threshold filtering, and manual contaminant filtering was 23,742, with a range of 5,287–47,225. The Shannon diversity index and Chao1 richness estimate were computed for each sample and are listed in Supplementary Table S5. Although some studies examining high-throughput sequencing data rarefy each sample to the lowest sequence sample, we chose not to rarefy due to several recent studies that indicate that this transformation is not appropriate and can impede detection of differential abundance (McMurdie and Holmes, 2014; Weiss et al., 2017). These studies indicate that transforming sequence counts into relative abundance is more effective at detecting true differential abundance and does not require discarding valid data. Taxonomic assignments for each OTU were generated by comparing against the Silva 138 database (90% cut-off). Further statistical analyses, including non-metric multidimensional scaling (NMDS), analysis of similarity (ANOSIM), similarity percentage (SIMPER), and one-way ANOVA were conducted in R (R Core Team, 2014) with the vegan ecological statistics package. NMDS analyses were conducted after performing a fourth root transformation to the relative abundance data, following studies indicating that this is a simple but effective way to evaluate differential abundance in microbial communities (McMurdie and Holmes, 2014; Weiss et al., 2017). These analyses construct de novo ordinations based on ranked dissimilarities, and the fit of the ordination is measured by the “stress.” In general, stress values below 0.2 are acceptable, and all NMDS ordinations presented here have stress <0.2. In our ANOSIM tests, p < 0.05 were considered significant. ANOSIM R values indicate the strength of dissimilarity between groups from 0 to a maximum of 1. SIMPER tests were applied in R to identify specific OTUs with different relative abundances between sample groups, such as between mineral types or between individual incubations. OTUs identified by SIMPER that contributed most to sample differentiation were included for presentation, with the significance of the differential abundance of individual OTUs assessed by ANOVA. Venn diagrams of community overlap (Supplementary Figure S6) were made with Venny (Oliveros, 2007–2015).
All mineral samples were imaged both pre- and post-incubation to record any incubation-associated mineralogical changes. Glutaraldehyde-fixed mineral samples for scanning electron microscopy (SEM) were dehydrated with an ethanol dehydration series (25, 50, 75, 90, and 100%; each step 30 min), then critical-point dried with liquid CO2 to preserve cell morphology. Briefly, this involves washing out the ethanol with liquid CO2 in a pressure chamber kept cold with an ice water bath, then raising the temperature to 31°C and adjusting the internal pressure to 74 bar (the critical point of CO2), and then slowly bleeding away the remaining CO2. The samples were mounted on aluminum stubs with carbon tape and sputter-coated with 10 nm of 80:20 Platinum:Palladium to minimize charging effects. SEM was carried out on a Hitachi SU3500 microscope (Hitachi High-Technologies; Tokyo, Japan) equipped with an Oxford X-Max 150 energy dispersive x-ray spectroscopy (EDX) unit (Oxford Instruments; Abingdon, United Kingdom).
The calcite and dolomite standards, along with the HVD and MS natural samples, were first powdered manually with an aluminum oxide mortar and pestle. The resulting powder was then analyzed by X-Ray Fluorescence (XRF) with an INAM Expert 3L tabletop XRF under helium purge with a Ti target using INAM’s built-in fundamental parameters software. Data were acquired over 60 s with 15 kV excitation energy for light elements, followed by an additional 60 s with 45 kV excitation voltage for heavier elements. The resulting comparative elemental analyses (as the corresponding elemental oxides) are listed in Supplementary Table S4.
Well logging and geochemical analyses of Inyo-BLM 1 in August 2015 were consistent with well logging data collected in 2011. Over the 18-month period of our experiments, the water level in the well was 28.7 m ± 0.03 mbls (USGS, 2018). The temperature increased by 4.8°C per 100 m from 25.7°C at the surface to 57°C at the bottom of the well (Figure 1). The pH of the fluids down to 500 mbls was slightly alkaline (9.0–9.2), transitioning to circumneutral conditions (6.9) in the uncased portion of the well (depth > 750 mbls). Water samples were collected at 752 mbls (in the uncased portion) and at 579 mbls (Supplementary Table S2). Both samples showed brackish conditions [9.7 and 9.4 mM Na+ and 1.5 and 1.4 mM Cl−, respectively (222, 215, 52.4, 49.2 ppm); Supplementary Table S2]. Ferrous iron concentrations increased by 60% (0.90–1.4 μM; 0.05–0.08 ppm) and sulfate levels were three times higher (reaching 1.6 μM) in the uncased portion of the hole at 752 mbls relative to 579 mbls (Supplementary Table S2). Calcium, magnesium, and silica concentrations also increased below the casing. Sulfide was not detected by an onsite Cline assay, and a qualitative in situ test for dissolved sulfide using the silver nitrate-coated film method also indicated that sulfide concentrations were below detection (Fike et al., 2017). Continuous ORP data obtained with the Idronaut well-logging tool showed a dramatic drop in potential at about 200 mbls (or 172 m below water level), indicative of anoxia (Figure 1). Within the open hole portion at the bottom, the ORP drifts toward positive values, but these types of probes are sensitive to flocculent material in the water. We cannot determine whether the drift is due to an actual increase in the ORP or the presence of suspended particles at depth. Upon recovery of Incubation 2 on 8/21/15, average planktonic cell counts at 752 mbls were 4.1 × 106 cells per ml ± 2.8 × 106, the large standard deviation owing to the tendency of cells to form small clumps (Supplementary Table S3).
Analyses of dissolved gases in water samples obtained from 579 to 752 mbls in Inyo-BLM 1 immediately preceding the carbon amendment (start of Incubation 3) revealed up to 30-fold more dissolved hydrogen in the cased section of the well relative to the open section (200 vs. 7 μM). The increase in hydrogen co-occurred with a 10-fold increase in dissolved methane (100 vs. 10 μM; Supplementary Table S2). By contrast, the fluids at 752 mbls contained elevated dissolved CO2, consistent with a carbonate-hosted aquifer.
To test whether the steel casing could have produced hydrogen (via the Schikorr reaction), we incubated three alloys of steel as well as hematite (Fe2O3) in sterile media mimicking the water chemistry in the borehole (Schikorr, 1929; Linnenbom, 1958; Ma et al., 2013). These experiments verified that low-carbon steel similar to that used in the well casing (and used in our experimental cartridges) produced significantly more H2 than stainless steel alloys or hematite (Supplementary Figure S4). Within 13 days, the low-carbon steel produced 14.3 μmol H2, approximately 70-fold more hydrogen than the 304 stainless steel, and 700-fold more hydrogen than the 316 stainless steel alloy or hematite, which produced barely detectable hydrogen peaks during chromatography analysis.
Unincubated calcite, dolomite, HVD, and MS substrates were analyzed by XRF spectroscopy to compare their compositions (Supplementary Table S4). The elemental composition of the substrates was consistent with their mineralogical identification and in the case of the natural HVD and MS samples, with previous geological descriptions (Fridrich et al., 2012). The HVD and dolomite standards differed in their trace element composition, with the dolomite standard higher in iron and manganese and the HVD higher in silicon. The MS sample was consistent with a mixture of potassium and calcium aluminosilicates with a wide array of trace elements detected, including significant amounts of iron, manganese, and titanium.
All mineral substrates were imaged by SEM pre- and post-incubation in Inyo-BLM 1. Apart from microbial attachment, the majority of incubated minerals displayed no obvious mineralogical alteration over the course of their incubation. Notable exceptions included the two dolomitic rocks (dolomite standard and native HVD) recovered from the 3-month long Incubation 1 (2/5/15), which showed widespread precipitation of 1–5 μm diameter secondary minerals with framboidal structure (~10 per 100 μm square viewing area); EDX characterized these secondary minerals as iron sulfides (Figure 2). While large anhedral pyrite crystals were observed in both the standard dolomite and HVD samples prior to deployment, nothing resembling the framboidal precipitates were observed on any samples prior to deployment, indicating that they formed in situ. These secondary minerals were typically associated with 1–3 microbial cells. Sulfide framboid generation was only detected in the first set of incubations, prior to the organic matter amendment experiment. Sulfide mineral formation was also observed on minerals from the 6-month long Incubation 2 (recovered 8/21/15), in this case producing scattered zinc sulfide spheroids, rather than iron sulfide framboids, but spheroid-attached cells were not observed. In both cases, secondary mineral formation was only detected on the dolomitic minerals (HVD and Ward’s dolomite, n = 8) across two separate incubations and not on any glass wool, MS, low-carbon steel, or calcite minerals incubated in parallel (n = 14). Incubations 1 and 2 occurred prior to the organic matter addition in Incubation 3 on 8/23/15, and no further secondary mineral formation was observed via SEM following the organic matter addition.
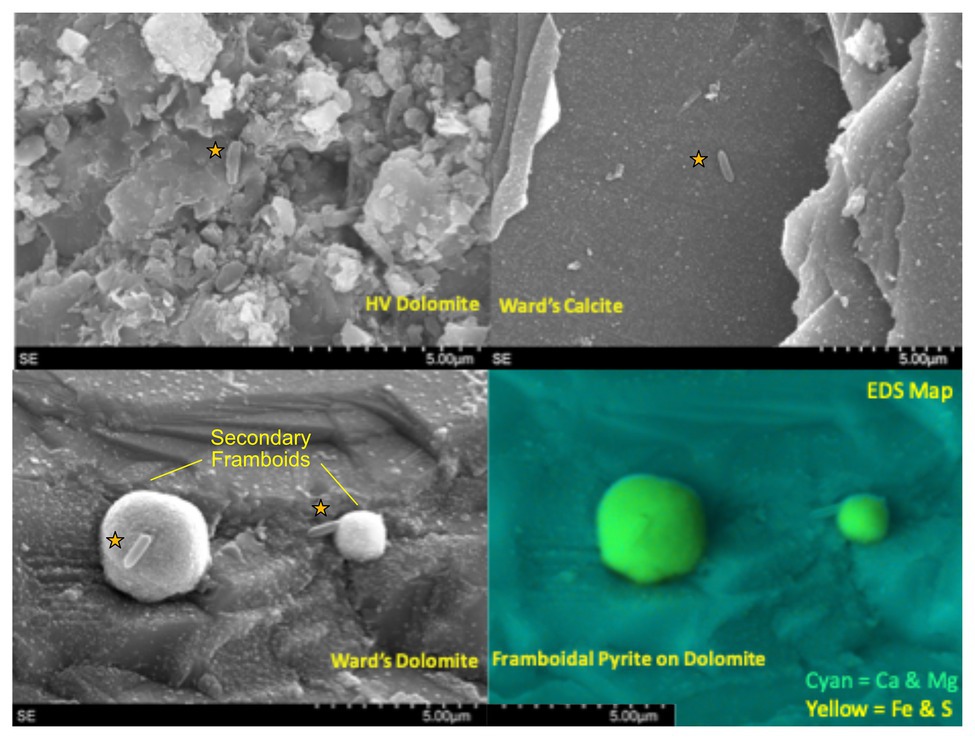

Figure 2. Scanning electron micrograph (SEM) and energy dispersive x-ray spectroscopy (EDX) overlays of carbonates from Inc. 1, with microorganisms marked with orange stars. (Top Left) Natural dolomite sample (HVD) with putative cell in center. The texture of the rock presented difficulties in distinguishing cells from surroundings. (Top Right) Although calcite samples were the easiest to image, few cells and no iron sulfide framboids were visible. (Bottom Left) Cells attached to secondary framboidal structures on the dolomite standard. (Bottom right) EDX map of bottom left panel which confirms that secondary framboids are iron sulfide.
After removing potential contaminants found in the sequencing controls, analysis of the Illumina 16S rRNA gene amplicons (iTag) of all samples taken from the site, including incubated mineral substrates from cartridges suspended at 752 mbls, PUR foam blocks from the suspension line, and water from 579 to 752 mbls, revealed a system dominated by Firmicutes, Nitrospira, Alphaproteobacteria and Betaproteobacteria, and Candidate Division KB-1 (also known as Candidate Division Acetothermia), with a moderate relative abundance of Chloroflexi, Epsilonproteobacteria, and Euryarchaeota, including two putative methanogens of the Methanobacteriales and an OTU related to the South African Gold Mine Group of Thermoplasmata (see Supplementary Table S6). The samples demonstrated low diversity overall; the maximum observed Shannon Index was 4.03, and the maximum Chao1 richness was 360. Reflecting the low diversity, the dominant OTU in 60 of the 71 samples represented >15% of the total community for that sample. Across all samples and incubations, two broad groups appeared when visualizing the microbial community by NMDS: one group consisting of water (including unfiltered and filtered water) and the other consisting of solid substrates, including minerals and glass (Supplementary Figure S4). PUR samples occupied ordination space between water samples and mineral samples but contained no unique OTUs. Instead, the PUR-colonized community represented a mixture of planktonic and surface-attached microorganisms (Supplementary Figure S5).
Analysis of similarity (ANOSIM) analyses of the samples were conducted to assess which parameters correlated most strongly with community variance; ANOSIM R values range from 0 to 1 with increasing correlation. When examined across the entire dataset, the most important parameter in the variation between samples was the Incubation number (R = 0.4535, p = 0.001), with type of substrate showing secondary importance for community differences (R = 0.2667, p = 0.001). Interestingly, carbonate samples, which included calcite, dolomite, and HVD, grouped together in all but one incubation (Incubation 1). Across all incubations, the communities colonizing carbonate minerals were not significantly different from each other (p = 0.558). We also assessed the difference between filtered and unfiltered water samples and found no significant difference (p = 0.2). Although we had originally hypothesized that the community colonizing the PUR samples would group with the water samples, these samples were significantly different from the water samples (p = 0.001). Based on the ANOSIM analyses and the similarity between the sample types, we therefore grouped the calcite, HVD, and dolomite samples into one “carbonate” bin, and the water samples (filtered and unfiltered) into one “background” bin, which increased the contribution of substrate to the variance (R = 0.3235, p = 0.001). The apparent dominance of the incubation reference number in describing the community structure was largely due to the organic matter amendment during Incubation 3. On an individual Incubation basis, colonizing substrate (e.g., carbonate, MS, and silica glass wool) was generally a strong determinant of variability (Figure 4). The ANOSIM R statistic and significance values for mineralogical variation when comparing within individual Incubations were as follows: Incubation 1 (R = 0.9733, p = 0.005), Incubation 2 (R = 0.5774, p = 0.006), Incubation 3 (not significant, p = 0.053), Incubation 4 (R = 0.6406, p = 0.026), and Incubation 5 (R = 0.3900, p = 0.026). ANOSIM analysis comparing Incubations 1 and 2 (which occurred prior to organic amendment) revealed the strong effect of substrate type (using the carbonate bin) over Incubation (R = 0.7669, p = 0.001 vs. R = 0.2167, p = 0.006). When carbonate minerals in these two incubations were treated as independent, the effect was still significant but less impactful (R = 0.6764, p = 0.001).
To understand which OTUs were most affected by mineralogical variation, we focused on the subset of data least affected by the organic matter amendment (Incubations 1, 2, and 5). After confirming that the substrate bins were still a significant contributor to the variance in these three incubations (R = 0.3871, p = 0.001), we used pairwise SIMPER to identify OTUs contributing most toward the variance between substrate bins (Figure 3). The patterns in colonization of these OTUs indicate whether they were more commonly found in water samples or on one of the different lithological substrates. For example, the Candidate Division KB-1 OTU was the most abundant OTU recovered from water and PUR samples (17.3% relative abundance on average) and was moderately abundant in glass wool cartridges (8.9%), but consistently represented <5% of the community on mineral samples. Other OTUs that were enriched in planktonic samples included an Nitrospiraceae OTU (14.1%) and an OTU matching Desulforudis sp., a putative hydrogenotrophic, sulfate-reducing Firmicute discovered in deep strata accessed by a South African mine (7.7% abundance, 100% sequence ID; Moser et al., 2005; Chivian et al., 2008). Several OTUs which were abundant in water samples were also found on the low-carbon steel, including the OTU matching Desulforudis (5% average, 15.5% max abundance). Other OTUs appeared to show a preference for solid or mineral substrates. For example, an OTU identified as a Geobacillus sp. consistently represented 1–5% of the community on glass, steel, and mineral samples (up to 19% on a glass sample), but was only detected in 1 of 17 water or PUR samples at 0.08% relative abundance.
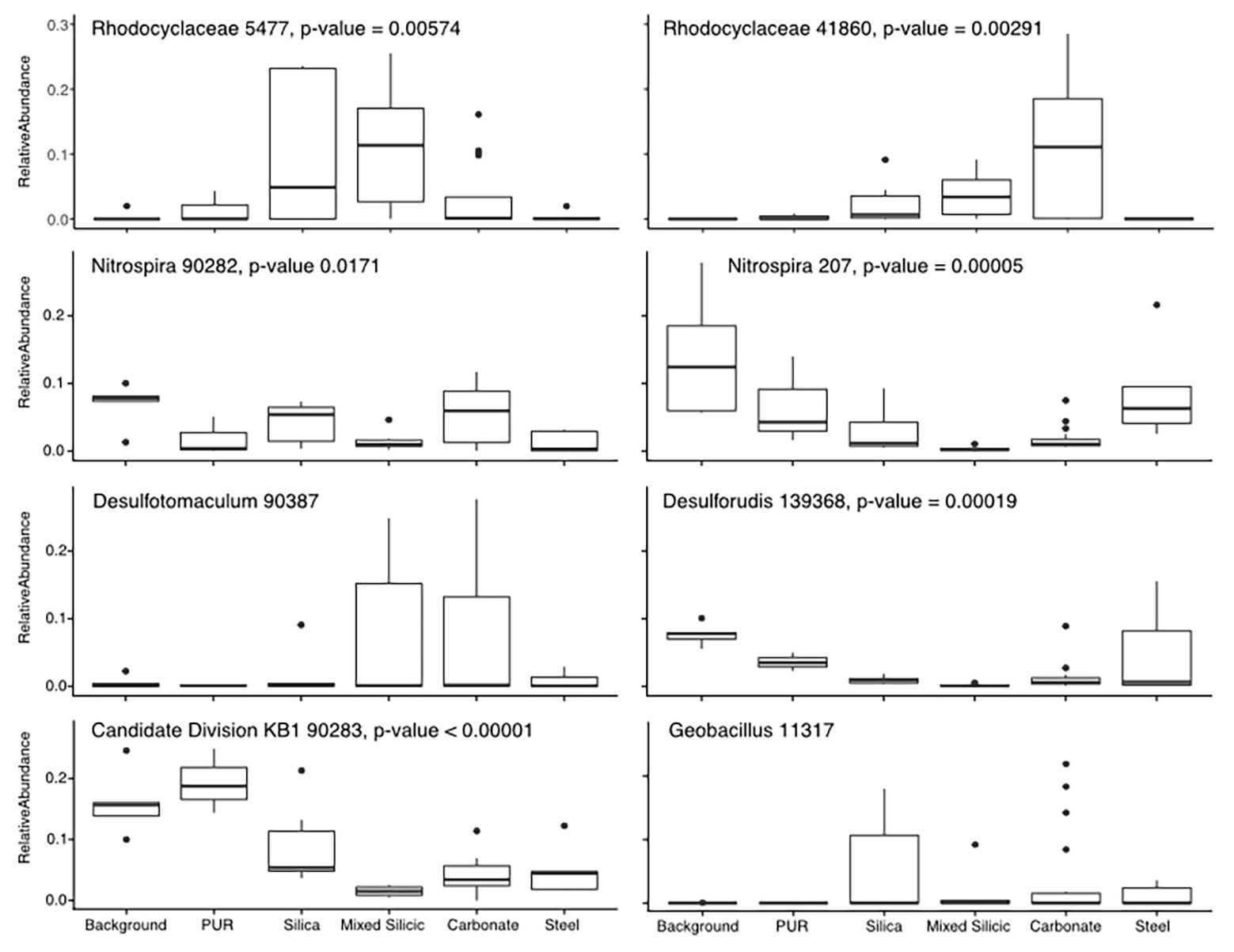
Figure 3. Tukey-style box and whiskers plot showing the average relative abundance of similarity percentage (SIMPER)-identified important taxa by substrate group. For each grouping of samples, the box represents the upper and lower quartiles surrounding the median (line in the box). Whiskers represent the remaining spread of the data, apart from outlying data points (defined as points further from the median than 1.5× Interquartile Range). We isolated 16S data from Incubations 1, 2, and 5 (since they are least impacted by the organic substrate addition) and identified the eight most common OTUs to appear in SIMPER pairwise analysis of substrates (ANOSIM by substrate: R = 0.3863, p = 0.001). The relative abundances for these OTUs were averaged across the three Incubations in order to analyze mineral preference across the microbial community. “Background” samples include bailed filtered and unfiltered water (n = 5); “PUR” indicates polyurethane foam block samples (n = 3); “Carbonates” include calcite, dolomite, and HVD samples (n = 18); “Silica” indicates glass wool samples (n = 6); “Mixed” indicates MS lithology (tuff and lake sediments; n = 6); “Steel” indicates low carbon steel ball bearings (n = 5). Plots are additionally labeled with the one-way ANOVA value of p if significant (p < 0.05). The lack of significance for the Desulfotomaculum and Geobacillus OTUs was indicative of the fact that the Desulfotomaculum population crashed after Incubation 1 and that the Geobacillus was infrequently found on minerals, but that, when present, it was often in high abundance, supporting the idea that this organism is infrequently detected but typically detected at high abundance.
In general, analysis of relative abundance values can be difficult; values can change significantly depending on total community composition. It is important, therefore, to apply a transformation to these values to minimize the effect of small number variance (e.g., fourth-root transform, as we have done) and to contextualize relative abundance shifts by examining OTU overlap between different sample bins. Overall, across all incubations and samples, 75% of the OTUs identified on lithological substrates (carbonate, MS, silica glass wool, and steel) were detected at least once on each substrate (Supplementary Figure S6). Carbonate and MS shared the most OTUs unique to two categories (3.2%) and carbonate, MS, and silica glass wool shared the most unique to three categories (8.4%). Most of the variance, therefore, between substrate types was caused by differential abundance rather than strict OTU presence or absence.
We also used these techniques to attempt to identify which, if any, OTUs were correlated with the presence of the framboidal iron sulfides found on dolomite and HVD samples in Incubation 1. By subsetting our data to Incubations 1 and 2, which contained the same substrates and differed only in length of incubation, we attempted to identify microbial community shifts concurrent with iron sulfur framboid precipitation. Variance analysis (ANOSIM) confirmed that the community colonizing dolomite samples (HVD and dolomite standard) significantly differed between Incubations 1 and 2 (R = 0.8125, p = 0.04). A SIMPER analysis identified the top two taxa responsible for driving the difference between dolomite samples in Incubation 1 and 2 as Sulfurovum sp., an Epsilonproteobacteria, and Desulfotomaculum sp., a Firmicute, which were detected in Incubation 1 at high abundances but in Incubation 2 were either completely absent (Sulfurovum) or at much lower relative abundance (Desulfotomaculum). Cultured representatives of the two genera are sulfide-oxidizers and sulfate-reducers, respectively.
Upon retrieval of the Incubation 3 deployment, organic sponges from the 752 mbls region were missing, although the zip ties used to attach them were still on the line. We assume that the 752 mbls sponges were degraded during the incubation as a significantly degraded, gelatinous sponge was still attached to the line from the 579 mbls set. The close relationship between Incubations 1 and 2 and the dramatic community shift in Incubation 3 demonstrated the profound impact that the organic matter amendment had on the microbial community. Biological replicates of substrates in 4 of the 5 incubations (1, 2, 4, and 5) were closely clustered in the NMDS, but replicates from Incubation 3, which were incubated concurrent with the organic matter amendment, displayed the greatest separation from each other as well as from samples from the other incubations (Figure 4). Changes in relative abundance of the most abundant OTUs across all incubations are summarized in Figure 5.
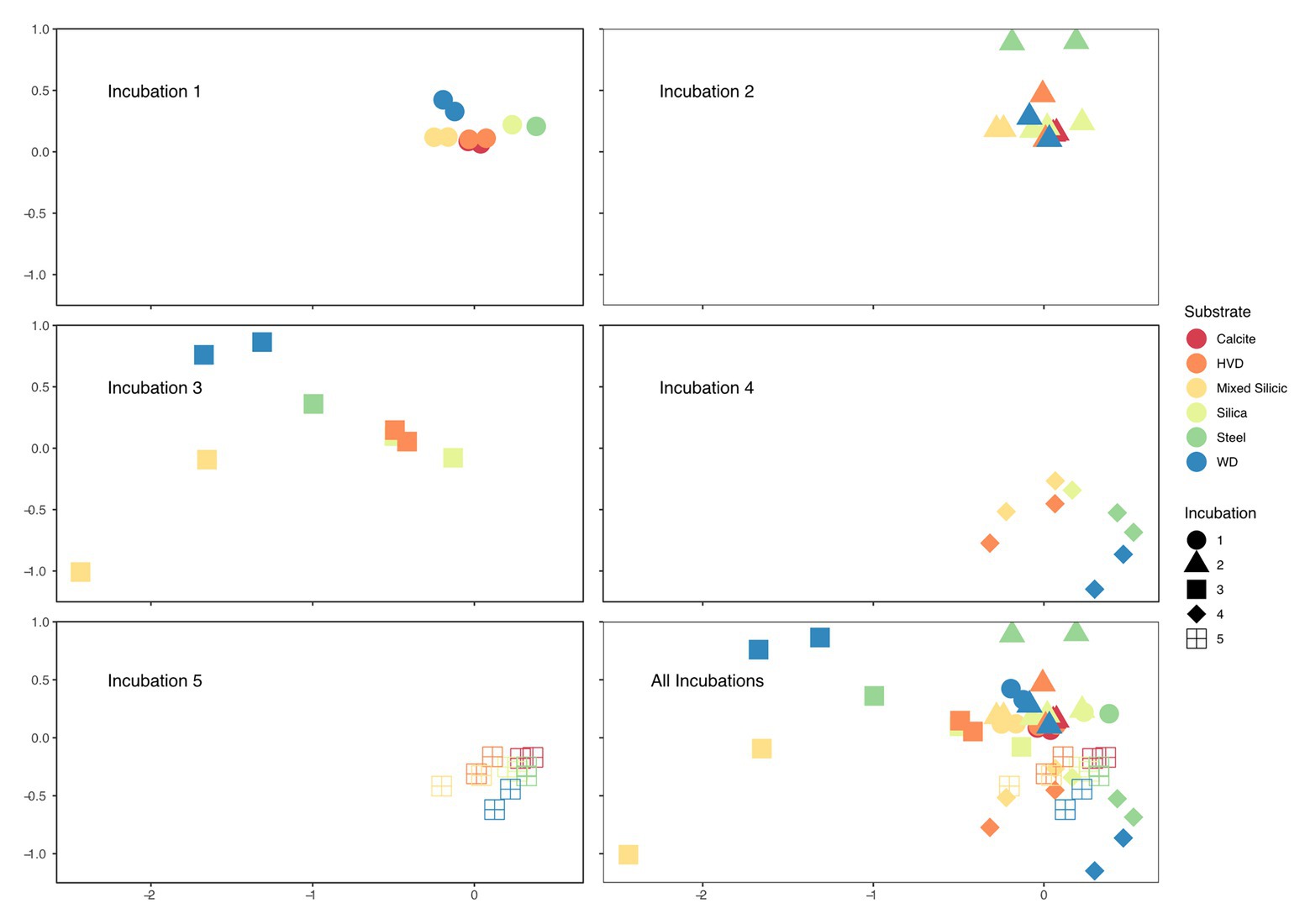
Figure 4. Faceted non-metric multidimensional scaling (NMDS) of microbial relative abundance on solid substrates used in each Incubation with combined NMDS bottom right. Each point on the plot represents an individual cartridge from an incubation; two points of the same color and shape are biological replicates of substrates from the same incubation but incubated in separated cartridges in situ. Incubations 1 and 2 are the most closely related and both occurred before the addition of the organic carbon substrate. Incubation 3 represents the first samples obtained after the carbon amendment and displays the widest variation amongst biological replicates and amongst mineral substrates. Incubation 4 is dominated by methanogenic archaea which pulls the samples into a cluster toward the bottom, and by Incubation 5, the samples are again tightly clustered and close to their starting position on the NMDS axes.
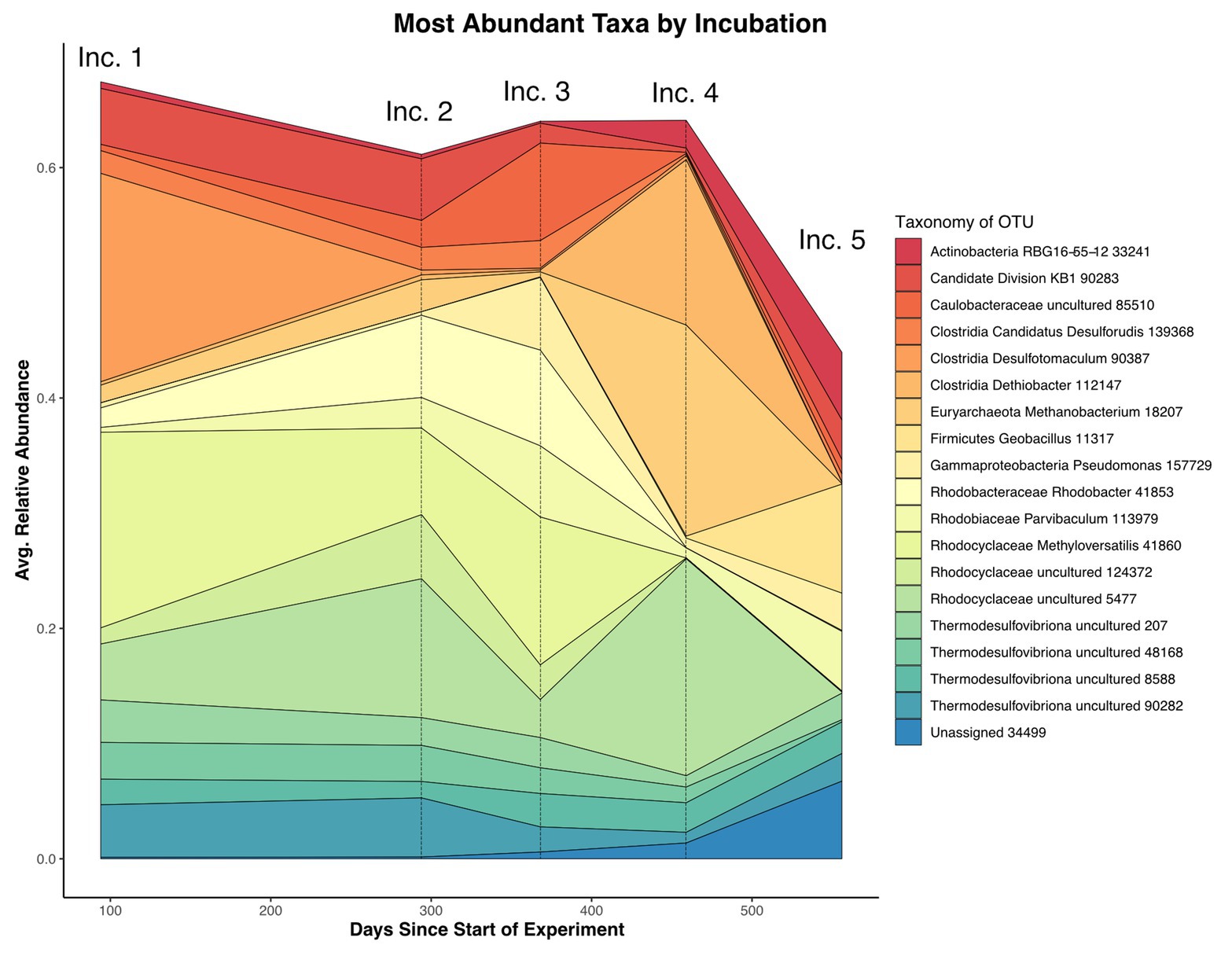
Figure 5. Average relative abundance of most abundant taxa by days since the start of the experiment, with individual incubation collection points noted above. The relative abundance as measured by 16S rRNA iTag data from mineral substrates common to all incubations was averaged at each time point. The data demonstrate the large relative abundance decrease in the Desulfotomaculum OTU co-occurent with the loss of framboidal iron sulfides in Incubation 2 as well as the large bloom in a Methanobacter OTU in Incubation 4 (6 months after organic substrate amendment). Incubation 5 displays a general increase in the proportion of rare taxa relative to the whole.
The microbial assemblage colonizing the degraded sea sponges from the organic matter amendment experiment was distinct from that recovered on the mineral substrates during the same incubation. The top three OTUs recovered from the organic sponges are most closely related to Fervidobacterium, a Thermotogae (20.3% of recovered sequences), Anaerolinea, a Chloroflexi (19.0%), and Thermodesulfovibrio, a Nitrospiraceae (18.6%). These organisms dominated the sponge assemblage, but were only rare OTUs in samples from previous mineral incubations and water samples. We did observe these OTUs in the microbial assemblage on minerals adjacent to the organic sponge in Incubation 3 but at much lower relative abundance. For example, the next highest relative abundance of any of these sponge-dominating OTUs on any mineral substrate from this in situ deployment was an OTU related to Anaerolinea in the pyrrhotite cartridge (5.3% relative abundance). These findings indicate that a small subset of the microbial community was stimulated by spongin protein amendment but that shifts in the mineral-hosted community cannot be solely attributed to cells directly transferred from the organic matter-associated community. Mineral substrate assemblages from Incubation 3 displayed a relative increase in a few key taxa, including a putative carbon-degrading Rhodocyclales matching Methyloversatilis sp. (8.1–13.6%, 100% nucleotide identity) and an OTU identified as an uncultured Caulobacteraceae (2.3–8.5%), while the relative abundance of taxa commonly detected on minerals in previous incubations showed a marked decrease. These Methyloversatilis and Caulobacteraceae OTUs were undetected and 0.00013% of the community on the organic sponge, respectively, further suggesting that their presence on the neighboring mineral substrates in Incubation 3 was not simply due to cross-contamination from the organic sponge associated microbial assemblage.
The subsequent set of mineral cartridges in Incubation 4, recovered 6 months after the organic carbon amendment, had very few sequences of Methyloversatilis sp., with all but three cartridges (one dolomite, one HVD, and one MS) containing no detectable sequences from the Methyloversatilis OTU. Instead, a shift in community composition featuring methanogens was observed, with an OTU annotated as a Methanobacter sp. representing nearly 15% of the recovered diversity across all samples. The Methanobacter OTU, along with other archaea (Methanothermobacter, South African Gold Mine group Thermoplasmata), was detected in previous incubations but at much lower relative abundance (less than 2%; Figure 5).
The microbial assemblage recovered from Incubation 5, 9 months after the organic matter experiment, appeared to more closely resemble the original community composition in Incubations 1 and 2. Groups such as the Candidate Division KB-1, Nitrospira sp., and Sulfurovum, which decreased in their relative abundance during Incubations 3 and 4 likely due to a proportional increase in fast-growing microorganisms responding to organic matter amendment, returned to their original relative abundance in Incubation 5. Other OTUs became more prominent, and Incubation 5 samples had a comparatively higher diversity with higher Shannon indices (Supplementary Table S5). One OTU recovered from this incubation (4% average rel. abundance) matched sequences with no known phylum according to our cutoff (90% ID). This OTU had 99% similarity to a bacterial sequence recovered from hot springs (NCBI BLAST accession number KM740594.1), and was distantly related to Aminicenantia (87% ID) in the Silva database but was below our identity cutoff (National Library of Medicine, US, 1988). Other microbial lineages remained at low levels in Incubation 5, including members of the Desulfotomaculum, which was closely associated with the presence of iron-sulfide framboids in Incubation 1. The Sulfurovum OTU, which was also implicated in framboid formation, did recover to roughly the same relative abundance as in Incubation 1. Notably, the difference in Desulfotomaculum was identified in SIMPER analyses as the primary factor distinguishing communities in Incubation 1 and 5. Apart from these differences, however, the net result of these analyses indicated that much of the community composition within this borehole environment was re-established less than a year after organic matter perturbation.
Although the timescale for mineral substrate colonization in the deep biosphere is not well constrained, we infer that the substrate-attached microbial assemblages we detected in our experiments were representative of natural assemblages and likely not introduced contaminants. Prior to our experiments, the anoxic and 57°C Inyo-BLM 1 borehole sat undisturbed for 3 years, creating conditions that are generally unfavorable to the persistence of contaminating microbial organisms (Pedersen et al., 1997). A previous study in a deep South African mine borehole demonstrated that natural microbial assemblages and the in situ condition of anoxia returned within 2 months of perturbation (Moser et al., 2003). Additionally, the microorganisms observed in Inyo-BLM 1 were remarkably similar to 16S rRNA gene clone library observations originally reported in this South African gold mine (Moser et al., 2005). Conditions at this site were similar to Inyo-BLM 1, characterized by warm (54°C), anoxic, paleometeoric waters upwelling from 4 to 5 km depth into a metabasaltic unit underlying dolomite in South Africa. Clone libraries in that study detected four main taxa in the community: Desulfotomaculum, Comamonadaceae (Betaproteobacteria), Candidatus Desulforudis audaxviator, and Methanobacterium. Inyo-BLM 1 contains OTUs identified as all of these groups at high relative abundance, including Methanobacterium (100% match over length to Moser et al., 2005 sequence) as the dominant archaeon. Since their identification in 2008, the putative sulfate-reducing hydrogenotrophs Desulforudis have been reported from an array of deep biosphere environments (Chivian et al., 2008; Tiago and Verissimo, 2013; Jungbluth et al., 2017). Interestingly, many of these studies, including the original description, were performed in boreholes with steel casings, as is the case for Inyo-BLM 1. It is likely that steel casings, especially low-carbon steel, may be contributing to local H2 production that enriched these organisms. Beyond the taxa it shares with other deep biosphere studies, Inyo-BLM 1 contains a number of unusual bacterial lineages, including representatives from Candidate Divisions KB-1 and Aminicenantes (formerly OP8). Candidate Division Aminicenantes were initially described from the Obsidian Pool thermal spring in Yellowstone National Park and have since been detected in diverse marine and terrestrial subsurface environments (Hugenholtz et al., 1998). Although no cultured representatives are available, recent metagenomic analysis of Aminicenantes recovered from a Siberian deep thermal aquifer indicated a potential for complex carbohydrate and proteinaceous substrate fermentation as well as heterotrophic nitrite respiration (Kadnikov et al., 2019). Originally discovered in the Kebrit Deep brine basin in the Red Sea (Eder et al., 1999), Candidate Division KB-1 bacteria have since been identified in a number of other hypersaline environments, including other deep hypersaline brine lenses, salterns, and hypersaline lakes and sediments (Nigro et al., 2016). Today, the Inyo-BLM 1, and the LCA that feeds it, are low salinity, but the Amargosa Desert region was home to evaporitic, fluvial lakes in the Paleozoic (Khoury et al., 1982) and it is interesting to consider whether these as yet uncultured bacteria may have adapted over time to the freshening groundwater environment.
The physical and chemical environment in the terrestrial deep biosphere is often extremely heterogeneous both in terms of the mineralogy as well as the nature of the permeating fluids. While fluid sampling is common in deep rock-hosted biosphere studies through legacy boreholes and mines, the ability to study mineral-associated microbial assemblages in the deep subsurface is more challenging (Moser et al., 2003; MacLean et al., 2007; Davidson et al., 2011; Ino et al., 2016). Previous work in the shallow subsurface has demonstrated that planktonic and mineral-attached communities are distinct, with some species significantly more abundant in one fraction or the other (Flynn et al., 2013). Here, we attempted to understand the distinction between the planktonic and mineral-attached communities as well as the role that mineral heterogeneity plays on microbial colonization by incubating a set of minerals representing the in situ lithologies at Inyo-BLM 1 alongside mineral standards and comparing the attached microbial assemblages to the planktonic community recovered from the aqueous phase. Similar types of mineral colonization experiments in both terrestrial and marine environments have pointed to local mineralogy as a primary factor linked to in situ microbial diversity (Roberts, 2004; Edwards et al., 2005; Orcutt et al., 2010; Toner et al., 2013). In some cases, microbial attachment is associated with direct mineral respiration, as occurs with iron-reducing bacteria (Lovley et al., 1989; Caccavo and Das, 2002), and in other circumstances, attachment may be controlled by the surface chemistry or electrostatic interactions between cell membranes and the mineral surface (Hermansson, 1999; Yee et al., 2000; Tuson and Weibel, 2013).
Nearly all of the OTUs we recovered from Inyo-BLM 1 were recovered from the solid substrates; 27.2% of the OTUs were exclusively detected on solid substrates, whereas only 0.4% were unique to the water samples (Supplementary Figure S6). Of the OTUs that were the most common contributors to substrate differentiation, a Candidate Division KB-1 OTU (bin90283) was abundant both in the water samples and on the PUR samples, and a Geobacillus OTU (bin11317) was only abundant on solid substrates (Figure 3). Notably, the abundance of Geobacillus was not significantly different as assessed by ANOVA, because many samples, even the solid substrates, had no detectable sequences from this lineage. Its variance on solid substrates, however, was much higher, with abundance on carbonate samples ranging from 0 to 21.9%. An F-test for significantly different variance confirmed that all of the solid substrates (carbonates, MS, silica, and steel) all had significantly higher variance than water samples for the relative abundance of this OTU, implying that the Geobacillus species only rarely colonized the minerals in the cartridges, but that when this occurred, these organisms frequently occurred at relatively high abundance.
Of the 518 OTUs found on the solid substrates, 72% of the OTUs were detected on least one sample from each lithological substrate, while only 5% of the OTUs were unique to a single lithological substrate type (Supplementary Figure S6). The mineralogy seems to play a significant role in governing the relative abundance of many OTUs and recruiting specific microbial lineages, both at the class and genus level, and differentiating among predicted microbial guilds (Figure 3). Twenty-nine OTUs of Rhodocyclaceae, a family of Betaproteobacteria, were identified in our survey, with very different colonization patterns. For example, a Methyloversatilis-affiliated OTU was more than twice as abundant on average on the carbonate minerals (calcite, dolomite, and HVD) than on glass wool or the MS sediments, while another Rhodocyclaceae OTU was most abundant on the glass wool and silica-rich MS. The underlying ecological or physiological factors influencing mineral colonization preference for these uncultured groups are not known, but the persistent partitioning by mineral substrate across the different incubations is notable. In another example, two OTUs affiliated with Desulforudis sp. (bin139368) and Desulfotomaculum sp. (bin90387), both putative sulfate-reducing Firmicutes, demonstrated different patterns of colonization – Desulforudis was most abundant on the low-carbon steel and Desulfotomaculum was most abundant on carbonate minerals. Notably, the ANOVA analysis of the Desulfotomaculum species across Incubations 1, 2, and 5 did not indicate that the difference was statistically significant, because that particular OTU was detected only at very low abundance after Incubation 1. Within Incubation 1, however, that Desulfotomaculum OTU averaged 20.9% of the recovered community from the carbonates vs. 1.4% of the community on the steel. Although inferring metabolic potential from 16S rRNA data represents a tentative assignment only, sequenced genomes Desulforudis indicate that they are capable of autotrophic growth through hydrogen-coupled sulfate reduction, and we hypothesize that Desulforudis in Inyo-BLM 1 may be locally consuming H2 produced by the anaerobic corrosion of the steel (Chivian et al., 2008; Jungbluth et al., 2017; Supplementary Figure S4). Sequenced genomes and cultivated representatives of Desulfotomaculum are typically heterotrophic sulfate-reducers (Aüllo et al., 2013; Sousa et al., 2018), and although the exact reason for their preferential association with carbonates in our data is unclear, the natural tendency of those minerals to entrain organic matter as they form may be one possibility (Roberts et al., 2013). A distinct colonization pattern was observed with yet another Desulfotomaculum-identified OTU (bin86994), which was most abundant on low-carbon steel, pyrite, and pyrrhotite mineral substrates. The intra-genus colonization differences between putative sulfate-reducing Gram-positive bacteria across different mineral substrates highlights the value of examining colonization patterns at the OTU level rather than higher-order taxonomic clustering.
The three carbonate species used in our flow-through cartridges (calcite, dolomite, and HVD) demonstrated overall similar communities and were not significantly different in the iTag sequencing data when comparing across all samples. These findings differ from some studies examining authigenic calcite, aragonite, and dolomite at deep-sea methane seeps, which did observe community variations between calcite and dolomite substrates (Case et al., 2015). SEM scans did reveal, however, that both Incubation 1 dolomite mineral samples (HVD and dolomite standard) were substantially modified in situ by the precipitation of framboidal iron sulfides, whereas calcite samples from the same incubation had no observable secondary mineral precipitation (Figure 2). It is possible that the difference in colonization patterns observed in marine studies was due to increased sensitivity of marine microorganisms to carbonate mineralogy because carbonate is a more dominant lithology in marine environments than terrestrial. The colonization differences between carbonate minerals may be more subtle than the differences between carbonate phases and other mineral families. Increasing the sample size of calcite and dolomite incubations in future deployments may help resolve whether there are discernable differences between the colonization of different carbonate minerals.
The specific mechanisms underpinning mineral preferences are currently unknown, but the repeatable patterns observed at the Inyo-BLM 1 site highlight potential ecological partitioning between subsurface taxa associated with minerals. Mineralogical groupings were a significant determinant of microbial community variance in most of our samples. Aside from Incubation 3, when complex organic matter was introduced, cartridges with duplicate lithologies acted as robust biological replicates, closely associated in the NMDS analysis (Figure 4) and ANOSIM R statistics for mineral type variation for were high (>0.35). In Incubation 1, nearly all of the variation between samples was attributable to mineral type (R = 0.9733). The Incubation number, partially representing the temporal distance from the organic carbon amendment, was a more important component when considering the entire data set, but perturbations on the scale of our experiment are likely infrequent. Since the vast majority of microorganisms in the deep biosphere are presumed to be mineral-attached (McMahon and Parnell, 2014), our experiments highlight the potential for targeting certain metabolisms and organisms based on the geology of their environment.
Framboidal iron sulfides, especially pyrite, have long been considered a potential biomarker, owing to the fact that they are often found in areas of high microbial activity, such as peat bogs and biofilms (Popa et al., 2004; Picard et al., 2016). It remains unknown, however, whether framboids are formed as a direct result of microbial activity or abiotically due to geochemical conditions (Sweeney and Kaplan, 1973; Graham and Ohmoto, 1994). As MacLean noted (MacLean et al., 2008), however, laboratory abiotic syntheses have only been accomplished at high-temperatures (above 60°C), well above the temperatures at nutrient- and biomass-rich systems where framboids are frequently found (Sawlowicz, 2000; Popa et al., 2004). It has been demonstrated that organic matter may play a role in aggregating the individual crystals into the framboid formation (MacLean et al., 2007) and MacLean et al. (2008) observed individual crystals developing within an organic matrix. Particularly interesting in our study was the correlation between the presence of secondary iron sulfide framboids on two independent dolomite incubations and two OTUs corresponding to Desulfotomaculum, a presumed sulfate-reducer, and Sulfurovum, a putative sulfide-oxidizing bacterium. Mineralogical heterogeneity of the substrate did not appear to explain the observed variation. Although both dolomites contained potential mineralogical sources for the framboids (higher iron content in the dolomite standard and micron-scale, anhedral iron- and sulfur-rich regions in the HVD), the secondary framboids formation was only observed during Incubation 1, although the same minerals were used for the incubations throughout the 18-month experimental series. Precipitation of iron sulfides also could not be attributed to incubation time, as under optimal conditions, pyrite framboids have been shown to form within days (Rickard, 2019). Even if the organic matter amendment during Incubation 3 disrupted the ecological conditions, therefore, we would have expected to observe framboid formation in the Incubation 2 (197 days long). While sulfide concentrations were below detection in the water recovered from the borehole, it is possible that localized sulfur metabolism by Desulfotomaculum and/or Sulfurovum directly influenced framboid precipitation. The groundwater feeding Inyo-BLM 1 does contain trace iron and zinc (1.4 and 0.06 μM, respectively; Supplementary Table S2), which can react with sulfide and may contribute to undetectable aqueous sulfide concentrations in our water samples.
In addition to their ability to respire sulfate, it has been shown that some Desulfotomaculum are capable of growth as fermentative syntrophs coupled with hydrogenotrophic methanogens, rather than heterotrophic sulfate reducers (Imachi et al., 2000, 2006; Plugge et al., 2002; Moser et al., 2005). Based on our 16S rRNA data alone, it is difficult to assess the physiology of these taxa in Inyo-BLM 1. On one hand, the detection of hydrogenotrophic methanogens affiliated with Methanobacterium in Inyo-BLM 1, the common partner of Desulfotomaculum when growing syntrophically (Imachi et al., 2000; Moser et al., 2005), suggests a similar syntrophic interaction could occur in our system; however, the co-occurrence of Desulfotomaculum (bin90387) with iron sulfides also supports a possible role in sulfur metabolism. Our laboratory experiments with steel alloys confirmed that the steel we added in each of the in situ incubations could have resulted in elevated H2 via anaerobic corrosion (the Schikorr reaction, e.g., ; Schikorr, 1929; Linnenbom, 1958; Ma et al., 2013). Therefore, we cannot dismiss the possibility that these organisms are syntrophs, that their disruption following Incubation 1 was due to inhibitory concentrations of hydrogen, and that their correlation with pyrite formation was related to their role in producing small organic carbon molecules which aid in aggregating iron sulfide framboids rather than the production of sulfide.
In most environments, oxidation of organic matter proceeds by the sequential reduction of O2, NO3−, Fe3+, SO42−, and CO2, in order of decreasing electron potential (Ponnamperuma, 1972). Because organic matter is thought to be move extremely slowly through the deep biosphere, most of these electron acceptors are long depleted at depth (Canfield, 1994; Hedges and Keil, 1995; Arndt et al., 2013; Osburn et al., 2014; Bradley et al., 2020). This results in a typically reduced system with low energy availability to the inhabiting organisms; Inyo-BLM 1 both has low dissolved organic carbon levels (0.431 ppm DOC) and few available electron acceptors (No detectable nitrate or nitrite and <2 mM sulfate; Supplementary Table S2). Deep biosphere microorganisms are therefore thought to have low energy requirements (LaRowe and Amend, 2015) and grow extremely slowly, with estimated turnover rates varying orders of magnitude from months to thousands of years (Morono et al., 2011; Xie et al., 2013; Onstott et al., 2014; Trembath-Reichert et al., 2017). Because of the long growth times and slow metabolic rates, ecological principles such as succession, competition, or dispersal, are likely reduced in impact and difficult to constrain (Biddle et al., 2011). By taking advantage of the addition of organic sponges to our system, we were able to observe the community response and some of these ecological forces at work in an unusual system. The microorganisms colonizing the sponges themselves were significantly different from the mineral-associated community, and in fact, the microbial diversity recovered from the incubated sponge mostly did not include OTUs that were abundant on the minerals, suggesting that the changes in the mineral-hosted community were driven by downstream degradation products of the sponges, analogous to the detrital matter that the deep biosphere likely subsists upon. The results imply that the planktonic Inyo-BLM 1 assemblage is a reservoir for microorganisms that are capable of colonizing and rapidly metabolizing complex organic matter. The mineral-colonizing community, by contrast, may be taking advantage of the small organic molecules released during the breakdown of the proteinaceous sponge biomass.
This organic matter amendment experiment also provides an opportunity to estimate community-wide carbon oxidation rates. The natural sponges introduced during Incubation 3 provided ~0.9 g of proteinaceous organic carbon and 0.3 g of fixed nitrogen. While tracing the organic degradation process and associated end-product production in this open system was not possible, we used the shifts in the microbial community as a general indicator of the ecological conditions after the organic matter addition. In our rough model, we assume during Incubation 3, when the sponges are first added to the well, that the microbial community is exposed to the most labile organic carbon species. A dramatic enrichment in Methyloversatilis sp. on the mineral substrates (Figure 3), but not on the sponge itself, indicates that this group perhaps is capable of rapid colonization and metabolism of organic molecules released by the sponge degradation. Cultured members of Methyloversatilis are methylotrophs that can oxidize a wide variety of C1 compounds coupled with oxygen or nitrate respiration (Lu et al., 2012; Doronina et al., 2014) and have been recovered in high abundances from oil reservoirs and hot springs (Lenchi et al., 2013; Doronina et al., 2014). The sudden dominance of these organisms in Incubation 3 and decrease in their relative abundance in subsequent incubations (Figure 5) suggests that they may represent a subset of the community poised to take advantage of carbon spikes to the system with a boom-bust lifestyle.
To our knowledge, this dolomitic aquifer is the first time organisms from Candidate Division KB-1 have been found in abundance in a terrestrial, low-salinity system. KB-1 is now classified as a subgroup of phylum Acetothermia (formerly OP1), which has been detected in anaerobic digesters as well as hot springs and other deep biosphere studies (Takami et al., 2012; Zaitseva et al., 2017; Hao et al., 2018), although the recovered diversity of related organisms in our system fall within the more narrowly defined KB-1 clade (sometimes referred to as class Acetothermiia) and we have therefore retained that name here. The only sequenced and published genome for Candidate Division KB-1, recovered from the Red Sea, points to a central metabolism based on glycine betaine degradation, a common N-containing organic osmolyte in hypersaline environments but less likely to be found in abundance in non-marine environments (Nigro et al., 2016). Although it is difficult to predict the metabolism of the as-yet uncultured KB-1 organisms in the groundwater due to the lack of extensive genomic characterization of this clade, the dynamics of community changes during the in situ organic matter addition, where the KB-1 OTUs dropped to very low abundance values over Incubations 3 and 4, suggest they were not stimulated (Figure 5). There are a number of possibilities for the observed decrease in KB-1 relative abundance, including that the KB-1 strains in this environment do not metabolize amino acids or that they perhaps were outcompeted by other microbes following the perturbation. Acetothermia and KB-1 organisms detected in other studies have revealed that the clade is capable of thriving in a range of organic matter concentrations, but if the KB-1 organisms in Inyo-BLM 1 are indeed heterotrophic (as are all the physiologically described representatives of this clade), our results imply that they are likely better adapted to in situ organic levels than to the excess provided during our amendment (Nigro et al., 2016; Zaitseva et al., 2017; Hao et al., 2018).
The recovery of Incubation 4 occurred 6 months after the introduction of the sponges to the well, and at this time, the microbial assemblage appeared to transition to a methanogenic phase. A small number of methanogen-affiliated sequences were detected in Incubation 1, including Methanobacterium, Methanothermobacter, and Methanolobus; however, samples from Incubation 4 were dominated by the OTU affiliated with Methanobacterium. All characterized isolates belonging to the Methanobacterium genus are hydrogenotrophic or formate-utilizing methanogens. Unexpectedly, although many OTUs associated with syntrophic clades (e.g., Syntrophomonadaceae, Syntrophobacterales) were observed during the organic enrichment in Incubation 3, a few were recovered from Incubation 4, suggesting that much of the complex fermentable carbon was consumed. By the time Incubation 5 was recovered (9 months after organic matter addition), the diversity of organisms colonizing the experimental substrates was largely consistent to the community composition observed during Incubations 1 and 2 prior to organic matter amendment, suggesting that the community had reset to the state prior to the perturbation (Figure 4).
The timing of the community shifts can provide bounds for order-of-magnitude activity estimates of the whole community in the borehole. Estimates for total carbon added to the system via diffusion were calculated above: 0.160–3.01 mmol. The lower activity estimate used the lower estimate for carbon diffusion (0.160 mmol C) and assumed that the methanogens detected were primarily using formate. Thus, the total estimated time for carbon oxidation to CO2 is 177 days, extending from midway through Incubation 3 through midway through Incubation 5 (the midpoints of the incubations were used approximately to allow time for methanogen abundance to decrease). Based on a cell count of 4.1 × 106 cells/ml from Incubation 3, each cell is estimated to oxidize 0.02 fmol C per day on average. To calculate the upper bound on activity, we used the higher diffused carbon value (3.01 mmol) and instead assumed that the detected methanogen, Methanobacterium, is hydrogenotrophic, thus assuming that the maximum value for methanogen relative abundance is a proxy for the maximum in hydrogen concentration the exhaustion of reduced carbon species. Based on these assumptions, we estimate that the complete oxidation of the carbon amendment occurred halfway between the recovery of Incubation 3 and 4, which translates to a total oxidation time of approximately 120 days and an average activity rate of 0.577 fmol C per cell per day. This exercise serves as a rough order-of-magnitude estimation and predicts oxidation rates that are high, but within the margin of error as compared to carbon oxidation rates measured from other deep biosphere systems using more direct methods (0.002–0.0067 fmol C per cell per day; Morono et al., 2011; Trembath-Reichert et al., 2017; Moreno-Ulloa et al., 2019).
The terrestrial deep biosphere is characterized by its phylogenetic, physiological, and geological heterogeneity. Mineralogical changes occur both regionally, as in lithological units, and at the microscale, with secondary mineral formation such as iron sulfides, potentially a product of the local micro-environment. This heterogeneity, along with the massive volume of the subsurface biome and difficulty in obtaining samples has meant that it is difficult to generate predictions as to the microbial population size, structure, or activity (Magnabosco et al., 2018). Although efforts are ongoing to address these questions, experiments performed in the laboratory are generally unable to capture all of the complexity of the subsurface. Our experiments serve as a wide-ranging in situ exploration of the factors controlling microbial populations and microbial colonization in one deep, warm dolomitic aquifer and enabled us to distinguish between populations of microorganisms living planktonically and those that preferentially adhere to a mineral surface. These observations provide some preliminary insight into the niche partitioning that occurs within the deep biosphere and emphasize the need for more comprehensive sampling of subsurface ecosystems. In addition to examining mineral-microbe associations, we utilized a complex N-rich organic matter amendment in situ to explore the response of the deep biosphere community to stimulation at this location. By observing this environment over a period of 9 months after the carbon amendment, we observed this community oxidize the organic carbon in several phases, with putative heterotrophic organisms closest to the carbon source, putative syntrophic organisms on neighboring mineral surfaces 3 months after the organic carbon were first deployed, and the dominance of methanogenic organisms 6 months after the original amendment. After 9 months, the community returned to the assemblage that closely resembled its undisturbed state. Our experiments indicate that the microbial community is highly responsive to shifts in geochemistry and nutrient availability; tracking these community changes can even provide coarse estimates for the activity of these microorganisms. More precise activity measurements could be obtained, perhaps utilizing stable isotopic tracers, but avoidance of contamination of the slow-moving environmental system may require any such measurements be ex situ. In any case, careful consideration of the activity of deep biosphere organisms will aid in our understanding of the dynamics of this massive and understudied system.
The datasets generated for this study can be found in the NCBI SRA # PRJNA605066.
SM coordinated field experiment set-up and sampling and performed DNA sequencing, sequence analysis, microscopy, and gas chromatography. GW coordinated field sampling, including the design of the in situ suspension line and provided microscopy data. BK, JS, and SH-B performed field sampling. Additionally, BK and JS coordinated retrieval and processing of geochemistry analyses. SH-B performed phase contrast cell counts. DM supervised BK, JS, and SH-B, identified and secured the field site, co-wrote the funding proposal, and organized downhole logging. RB, JA, and VO co-wrote the proposal and supervised field activities. VO helped conceive the experimental design and supervised SM. All authors contributed to the article and approved the submitted version.
This work was supported by the NASA Astrobiology Institute “Life Underground” project (NNA13AA92A) and the Center for Dark Energy Biosphere Investigations (C-DEBI). SM received additional support from an NIH Training Grant. JS was partially supported by a NASA Space Grant Consortium Fellowship.
Two authors declared commercial interests at the time of publication but not during the period of data collection: GW is employed by Oberland Agriscience, Inc., and RB is employed by Photon Systems, Inc.
The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Samples were obtained under scientific research permit DEVA-2013-SCI-0069, issued to DM from the U.S. National Park Service (NPS), and we thank Terry Fisk, Richard Friese, Josh Hoines, Genne Nelson, and Kevin Wilson of the NPS and Alisa Lembke of the Inyo County Planning Commission for site access. We extend our thanks also to John Healey, Brad Lyles, and Chuck Russell of the Desert Research Institute, John Bredehoeft and Michael King of The Hydronamics Group, LLC for logistical assistance; and Rachel Gates, Stephanie Connon, Haley Sapers, Daan Speth, Erin Bertrand, Ted Present, and Elizabeth Trembath-Reichert for occasional field and lab assistance. Thanks also to Bill Willborn and others from the DOE UGTA program for allowing use of their downhole logging system.
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fmicb.2020.536535/full#supplementary-material
Arndt, S., Jørgensen, B. B., LaRowe, D. E., Middelburg, J. J., Pancost, R. D., and Regnier, P. (2013). Quantifying the degradation of organic matter in marine sediments: a review and synthesis. Earth Sci. Rev. 123, 53–86. doi: 10.1016/j.earscirev.2013.02.008
Aüllo, T., Ranchou-Peyruse, A., Ollivier, B., and Magot, M. (2013). Desulfotomaculum spp. and related Gram-positive sulfate-reducing bacteria in deep subsurface environments. Front. Microbiol. 4:362. doi: 10.3389/fmicb.2013.00362
Bar-On, Y. M., Phillips, R., and Milo, R. (2018). The biomass distribution on Earth. Proc. Natl. Acad. Sci. U. S. A. 115, 6506–6511. doi: 10.1073/pnas.1711842115
Belcher, W. R., Bedinger, M. S., Back, J. T., and Sweetkind, D. S. (2009). Interbasin flow in the Great Basin with special reference to the southern Funeral Mountains and the source of Furnace Creek springs, Death Valley, California, U.S. J. Hydrol. 369, 30–43. doi: 10.1016/j.jhydrol.2009.02.048
Belcher, W. R., Elliott, P. E., and Geldon, A. L. (2001). Hydraulic-property estimates for use with a transient ground-water flow model of the death valley regional ground-water flow system, Nevada and California. Water-Resources Investigations Report. 01.
Biddle, J. F., Sylvan, J. B., Brazelton, W. J., Tully, B. J., Edwards, K. J., Moyer, C. L., et al. (2011). Prospects for the study of evolution in the deep biosphere. Front. Microbiol. 2:285. doi: 10.3389/fmicb.2011.00285
Bradley, J. A., Amend, J. P., and LaRowe, D. E. (2018). Bioenergetic controls on microbial ecophysiology in marine sediments. Front. Microbiol. 9:180. doi: 10.3389/fmicb.2018.00180
Bradley, J. A., Arndt, S., Amend, J. P., Burwicz, E., Dale, A. W., Egger, M., et al. (2020). Widespread energy limitation to life in global subseafloor sediments. Sci. Adv. 6:eaba0697. doi: 10.1126/sciadv.aba0697
Braun, S., Mhatre, S. S., Jaussi, M., Roy, H., Kjeldsen, K. U., Pearce, C., et al. (2017). Microbial turnover times in the deep seabed studied by amino acid racemization modelling. Sci. Rep. 7:5680. doi: 10.1038/s41598-017-05972-z
Bredehoeft, J., and King, M. (2010). Potential contaminant transport in the regional carbonate aquifer beneath yucca mountain, Nevada, USA. Hydrgeol. J. 18:775. doi: 10.1007/s10040-009-0550-z
Bredehoeft, J., King, M., Jansen, J., Fridrich, C. J., and Reese, S. (2008). Death Valley Lower Carbonate Aquifer Monitoring Program wells down gradient of the proposed Yucca Mountain Nuclear Waste Repository. Inyo County Yucca Mountain Repository Assessment Office, 111. doi: 10.2172/991653
Breuker, A., Köweker, G., Blazejak, A., and Schippers, A. (2011). The deep biosphere in terrestrial sediments in the Chesapeake bay area, Virginia, USA. Front. Microbiol. 2:156. doi: 10.3389/fmicb.2011.00156
Caccavo, F., and Das, A. (2002). Adhesion of dissimilatory Fe(III)-reducing bacteria to Fe(III) minerals. Geobiol. J. 19, 161–177. doi: 10.1080/01490450252864262
Canfield, D. E. (1994). Factors influencing organic carbon preservation in marine sediments. Chem. Geol. 114, 315–329. doi: 10.1016/0009-2541(94)90061-2
Caporaso, J. G., Kucynski, J., Stombaugh, J., Bittinger, K., Bushman, F. D., Costello, E. K., et al. (2010). QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336. doi: 10.1038/nmeth.f.303
Caporaso, J. G., Lauber, C. L., Walters, W. A., Berg-Lyons, D., Huntley, J., Fierer, N., et al. (2012). Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J. 6, 1621–1624. doi: 10.1038/ismej.2012.8
Case, D. H., Pasulka, A. L., Marlow, J. J., Grupe, B. M., Levin, L. A., and Orphan, V. J. (2015). Methane seep carbonates host distinct, diverse, and dynamic microbial assemblages. MBio 6, e01348–e01315. doi: 10.1128/mBio.01348-15
Chivian, D., Brodie, E. L., Alm, E. J., Culley, D. E., Dehal, P. S., DeSantis, T. S., et al. (2008). Environmental genomics reveals a single-species ecosystem deep with Earth. Science 322, 275–278. doi: 10.1126/science.1155495
Davidson, M. M., Silver, B. J., Onstott, T. C., Moser, D. P., Gihring, T. M., Pratt, L. M., et al. (2011). Capture of planktonic microbial diversity in fractures by long-term monitoring of flowing boreholes, evander basin, South Africa. Geomicrobiol. J. 28, 275–300. doi: 10.1080/01490451.2010.499928
Doronina, N. V., Kaparullina, E. N., and Trotsenko, Y. A. (2014). Methyloversatilis thermotolerans sp. nov., a novel thermotolerant facultative methylotroph isolated from a hot spring. Int. J. Syst. Evol. Microbiol. 64, 158–164. doi: 10.1099/ijs.0.055046-0
Eder, W., Ludwig, W., and Huber, R. (1999). Novel 16S rRNA gene sequences retrieved from highly saline brine sediments of Kebrit Deep, Red Sea. Arch. Microbiol. 172, 213–218. doi: 10.1007/s002030050762
Edwards, K. J., Bach, W., and McCollom, T. M. (2005). Geomicrobiology in oceanography: microbe-mineral interactions at and below the seafloor. Trends Microbiol. 13, 449–456. doi: 10.1016/j.tim.2005.07.005
Edwards, K. J., and Rutenberg, A. J. (2001). Microbial response to surface microtopography: the role of metabolism in localized mineral dissolution. Chem. Geol. 180, 19–32. doi: 10.1016/S0009-2541(01)00303-5
Ehrlich, H., Deutzmann, R., Brunner, E., Cappellini, E., Koon, H., Solazzo, C., et al. (2010). Mineralization of the metre-long biosilica structures of glass sponges is templated on hydroxylated collagen. Nat. Chem. 2, 1084–1088. doi: 10.1038/nchem.899
Ehrlich, H., Maldonado, M., Hanke, T., Meissner, H., Born, R., Scharnweber, D., et al. (2003). Spongins: nanostructural investigations and development of biomimetic material model. VDI Ber. 1803, 287–292.
Fike, D. A., Houghton, J. L., Moore, S. E., Gilhooly, W. P., Dawson, K. S., Druschel, G. K., et al. (2017). Spatially resolved capture of hydrogen sulfide from the water column and sedimentary pore waters for abundance and stable isotopic analysis. Mar. Chem. 197, 26–37. doi: 10.1016/j.marchem.2017.10.004
Flynn, T. M., Sanford, R. A., Ryu, H., Bethke, C. M., Levine, A. D., Ashbolt, N. J., et al. (2013). Functional microbial diversity explains groundwater chemistry in a pristine aquifer. BMC Microbiol. 13:146. doi: 10.1186/1471-2180-13-146
Fridrich, C. J., Thompson, R. A., Slate, J. L., Berry, M. E., and Machette, M. N. (2012). Geologic map of the southern funeral mountains including nearby groundwater discharge sites in death valley national park, California and Nevada. Geological Survey Scientific Investigations(Map 3151), 20 p. pamphlet, 21 sheet, scale 21:50,000. doi: 10.3133/sim3151
Gamgee, A. (1880). A text-book of the physiological chemistry of the animal body, including an account of the chemical changes occurring in disease: The physiological chemistry of the elementary tissues of the animal body. New York, NY: Macmillan and Company.
Graham, U. M., and Ohmoto, H. (1994). Experimental study of formation mechanisms of hydrothermal pyrite. Geochim. Cosmochim. Acta 58, 2187–2202. doi: 10.1016/0016-7037(94)90004-3
Green, D., Howard, D., Yang, X., Kelly, M., and Oreffo, R. O. C. (2003). Natural marine sponge fiber skeleton: a biomimetic scaffold for human osteoprogenitor cell attachment, growth, and differentiation. Tissue Eng. 9, 1159–1166. doi: 10.1089/10763270360728062
Hao, L., McIlroy, S. J., Kirkegaard, R. H., Karst, S. M., Fernando, W. E. Y., Aslan, H., et al. (2018). Novel prosthecate bacteria from the candidate phylum Acetothermia. ISME J. 12, 2225–2237. doi: 10.1038/s41396-018-0187-9
Hedges, J. I., and Keil, R. G. (1995). Sedimentary organic matter preservation: an assessment and speculative hypothesis. Mar. Chem. 49, 81–115. doi: 10.1016/0304-4203(95)00008-F
Hermansson, M. (1999). The DLVO theory in microbial adhesion. Colloids Surf. B Biointerfaces 14, 105–119. doi: 10.1016/S0927-7765(99)00029-6
Hugenholtz, P., Pitulle, C., Hershberger, K. L., and Pace, N. R. (1998). Novel division level bacterial diversity in a yellowstone hot spring. J. Bacteriol. 180, 366–376. doi: 10.1128/JB.180.2.366-376.1998
Imachi, H., Aoi, K., Tasumi, E., Saito, Y., Yamanaka, Y., Saito, Y., et al. (2011). Cultivation of methanogenic community from subseafloor sediments using a continuous-flow bioreactor. ISME J. 5, 1913–1925. doi: 10.1038/ismej.2011.64
Imachi, H., Sekiguchi, Y., Kamagata, Y., Loy, A., Qiu, Y. L., Hugenholtz, P., et al. (2006). Non-sulfate-reducing, syntrophic bacteria affiliated with desulfotomaculum cluster I are widely distributed in methanogenic environments. Appl. Environ. Microbiol. 72, 2080–2091. doi: 10.1128/AEM.72.3.2080-2091.2006
Imachi, H., Sekiguchi, Y., Kamagata, Y., Ohashi, A., and Harada, H. (2000). Cultivation and in situ detection of a thermophilic bacterium capable of oxidizing propionate in syntrophic association with hydrogenotrophic methanogens in a thermophilic methanogenic granular sludge. Appl. Environ. Microbiol. 66, 3608–3615. doi: 10.1128/aem.66.8.3608-3615.2000
Inagaki, F., Kuypers, M. M. M., Tsunogai, U., Ishibashi, J., Nakamura, K., Treude, T., et al. (2006). Microbial community in a sediment-hosted CO2 lake of the southern OkinawaTrough hydrothermal system. Proc. Natl. Acad. Sci. U. S. A. 103, 14164–14169. doi: 10.1073/pnas.0606083103
Ino, K., Konno, U., Kouduka, M., Hirota, A., Togo, Y. S., Fukuda, A., et al. (2016). Deep microbial life in high-quality granitic groundwater from geochemically and geographically distinct underground boreholes. Environ. Microbiol. Rep. 8, 285–294. doi: 10.1111/1758-2229.12379
Jones, A. A., and Bennett, P. C. (2014). Mineral microniches control the diversity of subsurface microbial populations. Geomicrobiol. J. 31, 246–261. doi: 10.1080/01490451.2013.809174
Jørgensen, B. B., and Boetius, A. (2007). Feast and famine—microbial life in the deep-sea bed. Nat. Rev. Microbiol. 5, 770–781. doi: 10.1038/nrmicro1745
Jungbluth, S. P., Glavina Del Rio, T., Tringe, S. G., Stepanauskas, R., and Rappe, M. S. (2017). Genomic comparisons of a bacterial lineage that inhabits both marine and terrestrial deep subsurface systems. PeerJ 5:e3134. doi: 10.7717/peerj.3134
Kadnikov, V. V., Mardanov, A. V., Beletsky, A. V., Karnachuk, O. V., and Ravin, N. V. (2019). Genome of the candidate phylum Aminicenantes bacterium from a deep subsurface thermal aquifer revealed its fermentative saccharolytic lifestyle. Extremophiles 23, 189–200. doi: 10.1007/s00792-018-01073-5
Khoury, H. N., Eberl, D. D., and Jones, B. F. (1982). Origin of magnesium clays from the Amargos Desert, Nevada. Clays Clay Miner. 30, 327–336.
Kieft, T. L., and Phelps, T. J. (1997). “Life in the slow lane: activites of microorganisms in the subsurface” in The microbiology of the terrestrial subsurface. ed. P. S. Amy (Boca Raton: CRC Press), 137–163.
LaRowe, D. E., and Amend, J. P. (2015). Power limits for microbial life. Front. Microbiol. 6:718. doi: 10.3389/fmicb.2015.00718
Lenchi, N., Inceoglu, O., Kebbouche-Gana, S., Gana, M. L., Lliros, M., Servais, P., et al. (2013). Diversity of microbial communities in production and injection waters of algerian oilfields revealed by 16s rRNA gene amplicon 454 pyrosequencing. PLoS One 8:e66588. doi: 10.1371/journal.pone.0066588
Linnenbom, V. J. (1958). The reaction between iron and water in the absence of oxygen. J. Electrochem. Soc. 105, 322–324. doi: 10.1149/1.2428838
Lovley, D. R., Baedecker, M., Lonergan, D. J., Cozzarelli, I. M., Phillips, E. J. P., and Siegel, D. I. (1989). Oxidation of aromatic contaminants coupled to microbial iron reduction. Nature 339, 297–300. doi: 10.1038/339297a0
Lu, H., Kalyuzhnaya, M., and Chandran, K. (2012). Comparative proteomic analysis reveals insights into anoxic growth of Methyloversatilis universalis FAM 5 on methanol and ethanol. Environ. Microbiol. 14, 2935–2945. doi: 10.1111/j.1462-2920.2012.02857.x
Ma, M., Zhang, Y., Guo, Z., and Gu, N. (2013). Facile synthesis of ultrathin magnetic iron oxide nanoplates by Schikorr reaction. Nanoscale Res. Lett. 8:16. doi: 10.1186/1556-276X-8-16
MacLean, L. C. W., Pray, T. J., Onstott, T. C., Brodie, E. L., Hazen, T. C., and Southam, G. (2007). Mineralogical, chemical and biological characterization of an anaerobic biofilm collected from a borehole in a deep gold mine in South Africa. Geomicrobiol. J. 24, 491–504. doi: 10.1080/01490450701572416
MacLean, L. C. W., Tyliszczak, T., Gilbert, P. U. P. A., Zhou, D., Pray, T. J., Onstott, T. C., et al. (2008). A high-resolution chemical and structural study of framboidal pyrite formed within a low-temperature bacterial biofilm. Geobiology 6, 471–480. doi: 10.1111/j.1472-4669.2008.00174.x
Magnabosco, C., Lin, L. H., Dong, H., Bomberg, M., Ghiorse, W., Stan-Lotter, H., et al. (2018). The biomass and biodiversity of the continental subsurface. Nat. Geosci. 11, 707–717. doi: 10.1038/s41561-018-0221-6
Mason, O. U., Case, D. H., Naehr, T. H., Lee, R. W., Thomas, R. B., Bailey, J. V., et al. (2015). Comparison of archaeal and bacterial diversity in methane seep carbonate nodules and host sediments, eel river basin and hydrate ridge, USA. Microb. Ecol. 70, 766–784. doi: 10.1007/s00248-015-0615-6
McMahon, S., and Parnell, J. (2014). Weighing the deep continental biosphere. FEMS Microbiol. Ecol. 87, 113–120. doi: 10.1111/1574-6941.12196
McMurdie, P. J., and Holmes, S. (2014). Waste not, want not: why rarefying microbiome data is inadmissible. PLoS Comput. Biol. 10:e1003531. doi: 10.1371/journal.pcbi.1003531
Momper, L., Aronson, H. S., and Amend, J. P. (2018). Genomic description of ‘candidatus abyssubacteria,’ a novel subsurface lineage within the candidate phylum hydrogenedentes. Front. Microbiol. 9:1993. doi: 10.3389/fmicb.2018.01993
Momper, L., Reese, B. K., Zinke, L., Wanger, G., Osburn, M. R., Moser, D., et al. (2017). Major phylum-level differences between porefluid and host rock bacterial communitites in the terrestrial deep subsurface. Environ. Microbiol. Rep. 9, 501–511. doi: 10.1111/1758-2229.12563
Moreno-Ulloa, A., Diaz, V. S., Tejeada-Mora, J. A., Macias Contreras, M. I., Diaz Costillo, F., Guerrero, A., et al. (2019). Metabolic and metagenomic profiling of hydrocarbon-degrading microorganisms obtained from the deep biosphere of the Gulf of Mexico. bioRxiv [preprint]. doi: 10.1101/606806
(2011). Carbon and nitrogen assimilation in deep subseafloor microbial cells. Proc. Natl. Acad. Sci. U. S. A. 108, 18295–18300. doi: 10.1073/pnas.1107763108
Moser, D. P., Gihring, T. M., Brockman, F. J., Fredrickson, J. K., Balkwill, D. L., Dollhopf, M. E., et al. (2005). Desulfotomaculum and Methanobacterium spp. dominate a 4- to 5-kilometer-deep fault. Appl. Environ. Microbiol. 71, 8773–8783. doi: 10.1128/AEM.71.12.8773-8783.2005
Moser, D. P., Onstott, T. C., Fredrickson, J. K., Brockman, F. J., Balkwill, D. L., Drake, G. R., et al. (2003). Temporal shifts in the geochemistry and microbial community structure of an ultradeep mine borehole following isolation. Geomicrobiol. J. 20, 517–548. doi: 10.1080/01490450390249280
Nigro, L. M., Hyde, A. S., MacGregor, B. J., and Teske, A. (2016). Phylogeography, salinity adaptations and metabolic potential of the candidate division KB1 bacteria based on a partial single cell genome. Front. Microbiol. 7:1266. doi: 10.3389/fmicb.2016.01266
Norman, M., Zdarta, J., Bartczak, P., Piasecki, A., Petrenko, I., Ehrlich, H., et al. (2016). Marine spone skeleton photosensitized by copper phthalocyanine: a catalyst for Rhodamine B degradation. Open Chem. 14, 243–254. doi: 10.1515/chem-2016-0025
Oliveros, J. C. (2007–2015). Venny. An interactive tool for comparing lists with Venn’s diagrams. Available at: https://bioinfogp.cnb.csic.es/tools/venny/index.html (Accessed Novomber 17, 2019).
Onstott, T. C., Magnabosco, C., Aubrey, A. D., Burton, A. S., Dworkin, J. P., Elsila, J. E., et al. (2014). Does aspartic acid racemization constrain the depth limit of the subsurface biosphere? Geobiology 12, 1–19. doi: 10.1111/gbi.12069
Orcutt, B. N., Bach, W., Becker, K., Fisher, A. T., Hentscher, M., Toner, B. M., et al. (2011). Colonization of subsurface microbial observatories deployed in young ocean crust. ISME J. 5, 692–703. doi: 10.1038/ismej.2010.157
Orcutt, B., Wheat, C. G., and Edwards, K. J. (2010). Subseafloor ocean crust microbial observatories development of flocs (flow through osmo colonization system) and evaluation of borehole construction. Geomicrobiol. J. 27, 143–157. doi: 10.1080/01490450903456772
Osburn, M. R., LaRowe, D. E., Momper, L. M., and Amend, J. P. (2014). Chemolithotrophy in the continental deep subsurface: Sanford Underground Research Facility (SURF), USA. Front. Microbiol. 5:610. doi: 10.3389/fmicb.2014.00610
Pedersen, K., Hallbeck, L., Arlinger, J., Erlandson, A. -C., and Jahromi, N. (1997). Investigation of the potential for microbial contaminatioin of deep granitic aquifers during drilling using 16S rRNA gene sequencing and culturing methods. J. Microbiol. Methods 30, 179–192. doi: 10.1016/S0167-7012(97)00066-3
Picard, A., Gartman, A., and Girguis, P. R. (2016). What do we really know about the role of microorganisms in iron sulfide mineral formation? Front. Earth Sci. 4:68. doi: 10.3389/feart.2016.00068
Plugge, C. M., Balk, M., and Stam, A. J. M. (2002). Desulfotomaculum thermobenzoicum subsp. thermosyntrophicum subsp. nov., a thermophilic, syntrophic, propioonat-oxidizing, spore-forming bacterium. Int. J. Syst. Evol. Microbiol. 52, 391–399. doi: 10.1099/ijs.0.01948-0
Ponnamperuma, F. N. (1972). The chemistry of submerged soils. Adv. Agron. 24, 29–96. doi: 10.1016/S0065-2113(08)60633-1
Popa, R., Kinkle, B. K., and Badescu, A. (2004). Pyrite framboids as biomarkers for iron-sulfur systems. Geomicrobiol. J. 21, 193–206. doi: 10.1080/01490450490275497
R Core Tea (2014). R: A language and environment for statistical computing. Vienna, Austria: R foundation for statistical computing.
Ribeiro, A. C. F., Ortona, O., Simões, S. M. N., Santos, C. I. A. V., Prazeres, P. M. R. A., Valente, A. J. M., et al. (2006). Binary mutual diffusion coefficients of aqueous solutions of sucrose, lactose, glucose and fructose in the temperature range from (298.15-328.15) K. J. Chem. Eng. Data 51, 1836–1840. doi: 10.1021/je0602061
Rickard, D. (2019). How long does it take a pyrite framboid to form? Earth Planet. Sci. Lett. 513, 64–68. doi: 10.1016/j.epsl.2019.02.019
Riedinger, N., Strasser, M., Harris, R. N., Klockgether, G., Lyons, T. W., and Screaton, E. J. (2015). Deep subsurface carbon cycling in the Nankai Trough (Japan)-evidence of tectonically induced stimulation of a deep microbial biosphere. Geochem. Geophys. Geosyst. 16, 3257–3270. doi: 10.1002/2015gc006050
Roberts, J. A. (2004). Inhibition and enhancement of microbial surface colonization: the role of silicate composition. Chem. Geol. 212, 313–327. doi: 10.1016/j.chemgeo.2004.08.021
Roberts, J. A., Kenward, P. A., Fowle, D. A., Goldstein, R. H., Gonzalez, L. A., and Moore, D. S. (2013). Surface chemistry allows for abiotic precipitation of dolomite at low temperature. Proc. Natl. Acad. Sci. U. S. A. 110, 14540–14545. doi: 10.1073/pnas.1305403110
Rogers, J. R., Bennett, P. C., and Choi, W. J. (1998). Feldspars as a source of nutrients for microorganisms. Am. Mineral. 83, 1532–1540. doi: 10.2138/am-1998-11-1241
Rowe, A. R., Yoshimura, M., LaRowe, D. E., Bird, L. J., Amend, J. P., Hashimoto, K., et al. (2017). In situ electrochemical enrichment and isolation of a magnetite-reducing bacterium from a high pH serpentinizing spring. Environ. Microbiol. 19, 2272–2285. doi: 10.1111/1462-2920.13723
Sackett, J. D. (2018). Prokaryotic diversity and aqueous geochemistry of the subsurface environments of the death valley regional flow system. Ph.D. thesis. Las Vegas: University of Nevada.
Sackett, J. D., Kruger, B. R., Becraft, E. D., Jarett, J. K., Stepanauskas, R., Woyke, T., et al. (2019). Four draft single-cell genome sequences of novel, nearly identical kiritimatiellaeota strains isolated from the continental deep subsurface. Microbiol. Resour. Announc. 8, e01249–e01318. doi: 10.1128/MRA.01249-18
Sahl, J. W., Schmidt, R., Swanner, E. D., Mandernack, K. W., Templeton, A. S., Kieft, T. L., et al. (2008). Subsurface microbial diversity in deep-granitic-fracture water in Colorado. Appl. Environ. Microbiol. 74, 143–152. doi: 10.1128/AEM.01133-07
Schikorr, G. (1929). Uber das System Eisen-Wasser. Zeitschrift für Elektrochemie 35, 62–65. doi: 10.1002/bbpc.19290350203
Sousa, D. Z., Visser, M., van Gelder, A. H., Boeren, S., Pieterse, M. M., Pinkse, M. W. H., et al. (2018). The deep-subsurface sulfate reducer Desulfotomaculum kuznetsovii employs two methanol-degrading pathways. Nat. Commun. 9:239. doi: 10.1038/s41467-017-02518-9
Sweeney, R. E., and Kaplan, I. R. (1973). Pyrite framboid formation: laboratory synthesis and marine sediments. Econ. Geol. 68, 618–634. doi: 10.2113/gsecongeo.68.5.618
Sylvan, J. B., Sia, T. Y., Haddad, A. G., Briscoe, L. J., Toner, B. M., Girguis, P. R., et al. (2013). Low temperature geomicrobiology follows host rock composition along a geochemical gradient in lau basin. Front. Microbiol. 4:61. doi: 10.3389/fmicb.2013.00061
Takai, K., Moser, D. P., DeFlaun, M., Onstott, T. C., and Fredrickson, J. K. (2001). Archaeal diversity in waters from deep south african gold mines. Appl. Environ. Microbiol. 67, 5750–5760. doi: 10.1128/aem.67.21.5750-5760.2001
Takami, H., Noguchi, H., Takaki, Y., Uchiyama, I., Toyoda, A., Nishi, S., et al. (2012). A deeply branching thermophilic bacterium with an ancient acetyl-coa pathway dominates a subsurface ecosystem. PLoS One 7:e30559. doi: 10.1371/journal.pone.0030559
Teske, A., and Sørensen, K. B. (2008). Uncultured archaea in deep marine subsurface sediments: have we caught them all. ISME J. 2, 3–18. doi: 10.1038/ismej.2007.90
Thomas, J. M., Moser, D. P., Fisher, J. C., Reihle, J., Wheatley, A., Hershey, R. L., et al. (2013). Using water chemistry, isotopes and microbiology to evaluate groundwater sources, flow paths and geochemical reactions in the death valley flow system, USA. Proc. Earth Planet. Sci. 7, 842–845. doi: 10.1016/j.proeps.2013.03.033
Tiago, I., and Verissimo, A. (2013). Microbial and functional diversity of a subterrestrial high pH groundwater associated to serpentinization. Environ. Microbiol. 15, 1687–1706. doi: 10.1111/1462-2920.12034
Toner, B. M., Lesniewski, R. A., Marlow, J. J., Briscoe, L. J., Santelli, C. M., Bach, W., et al. (2013). Mineralogy drives bacterial biogeography of hydrothermally inactive seafloor sulfide deposits. Geomicrobiol. J. 30, 313–326. doi: 10.1080/01490451.2012.688925
Trembath-Reichert, E., Morono, Y., Ijiri, A., Hoshino, T., Dawson, K. S., Inagaki, F., et al. (2017). Methyl-compound use and slow growth characterize microbial life in 2-km-deep subseafloor coal and shale beds. Proc. Natl. Acad. Sci. U. S. A. 114, E9206–E9215. doi: 10.1073/pnas.1707525114
Tuson, H. H., and Weibel, D. B. (2013). Bacteria-surface interactions. Soft Matter 9, 4368–4380. doi: 10.1039/C3SM27705D
USGS (2018). National water information system-groundwater levels for inyo-BLM1. USGS nevada water science center. Available at: https://nwis.waterdata.usgs.gov/nwis/gwlevels/?site_no=362402116280901 (Accessed January 7, 2020).
Wanger, G., Southam, G., and Onstott, T. C. (2007). Structural and chemical characterization of a natural fracture surface from 2.8 kilometers below land surface: biofilms in the deep subsurface. Geomicrobiol. J. 23, 443–452. doi: 10.1080/01490450600875746
Weiss, S., Xu, Z. Z., Peddada, S., Amir, A., Bittinger, K., Gonzalez, A., et al. (2017). Normalization and microbial differential abundance strategies depend upon data characteristics. Microbiome 5:27. doi: 10.1186/s40168-017-0237-y
Wilkins, M. J., Daly, R. A., Mouser, P. J., Trexler, R., Sharma, S., Cole, D. R., et al. (2014). Trends and future challenges in sampling the deep terrestrial biosphere. Front. Microbiol. 5:481. doi: 10.3389/fmicb.2014.00481
Xie, S., Lipp, J. L., Wegener, G., Ferdelman, T. G., and Hinrichs, K. -U. (2013). Turnover of microbial lipids in the deep biosphere and growth of benthic archaeal populations. Proc. Natl. Acad. Sci. U. S. A. 110, 6010–6014. doi: 10.1073/pnas.1218569110
Yee, N., Fein, J. B., and Daughney, C. J. (2000). Experimental study of the pH, ionic strength, and reversibility behavior of bacteria-mineral adsorption. Geochim. Cosmochim. Acta 64, 609–617. doi: 10.1016/S0016-7037(99)00342-7
Keywords: microbial ecology, deep biosphere, fractured rock, mineral colonization, microbial succession, carbonate, pyrite, methanogenesis
Citation: Mullin SW, Wanger G, Kruger BR, Sackett JD, Hamilton-Brehm SD, Bhartia R, Amend JP, Moser DP and Orphan VJ (2020) Patterns of in situ Mineral Colonization by Microorganisms in a ~60°C Deep Continental Subsurface Aquifer. Front. Microbiol. 11:536535. doi: 10.3389/fmicb.2020.536535
Received: 20 February 2020; Accepted: 24 September 2020;
Published: 19 November 2020.
Edited by:
Trinity L. Hamilton, University of Minnesota Twin Cities, United StatesReviewed by:
Matthew Schrenk, Michigan State University, United StatesCopyright © 2020 Mullin, Wanger, Kruger, Sackett, Hamilton-Brehm, Bhartia, Amend, Moser and Orphan. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Victoria J. Orphan, dm9ycGhhbkBncHMuY2FsdGVjaC5lZHU=
†Present address: Greg Wanger, Oberland Agriscience Inc., Halifax, NS, Canada
Rohit Bhartia, Photon Systems, Inc., Covina, CA, United States
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.