
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Mar. Sci. , 12 March 2025
Sec. Marine Fisheries, Aquaculture and Living Resources
Volume 12 - 2025 | https://doi.org/10.3389/fmars.2025.1532370
This article is part of the Research Topic Towards Sustainable Marine Aquaculture: Innovations and Eco-Friendly Practices View all 8 articles
 Mohamed Ashour1*
Mohamed Ashour1* Ahmed Said Al-Souti2*
Ahmed Said Al-Souti2* Ahmed Mamoon3
Ahmed Mamoon3 Fawzia S. Ali1
Fawzia S. Ali1 Mostafa E. Elshobary4
Mostafa E. Elshobary4 Mohamed M. Mabrouk3
Mohamed M. Mabrouk3 Ahmed I. A. Mansour1
Ahmed I. A. Mansour1 Abdallah Tageldein Mansour5*
Abdallah Tageldein Mansour5* Ehab El-Haroun6
Ehab El-Haroun6 Ahmed F. Abdelhamid3
Ahmed F. Abdelhamid3This work investigated the molecular identification and potential application of the cyanobacterial strain Desertifilum tharense NIOF17/006 as a novel aquafeed additive for whiteleg shrimp (Litopenaeus vannamei) postlarvae (PLs). Morphological and molecular characterization confirmed the isolate as D. tharense, with the 16S rRNA sequence analysis showing high similarity (98.01%–98.53%) to the known strains of D. tharense. Biochemical analysis revealed that the isolate contains 37.74% protein, 5.52% lipid, and 21.25% carbohydrate, on a dry weight basis. An 8-week feeding trial for L. vannamei PLs evaluated the effects of dietary supplementation with D. tharense NIOF17/006 at doses of 0, 1, 2.5, and 5 g/kg diet. Compared with shrimp in the control group, shrimp fed D. tharense-supplemented diets had significantly higher feed utilization, growth performance, survival rate, and whole body composition. The nonspecific immunity parameters (i.e., lysozyme, superoxide dismutase, and catalase), as well as the digestive enzyme activity of amylase and lipase, were significantly enhanced in shrimp fed diets supplemented with cyanobacteria, while the malondialdehyde (MDA) levels decreased. The gene expression analysis revealed the upregulation of growth-related genes (growth hormone, insulin-like growth factor I, and insulin-like growth factor II) and the immune-related genes prophenoloxidase (proPO), superoxide dismutase (SOD), and lysozyme (Lys) in shrimp muscles with increasing cyanobacteria supplementation, particularly at doses of 2.5–5 g/kg diet. Moreover, the polynomial regression machine learning model predicts that the ideal supplementation level of the probiotic cyanobacteria D. tharense NIOF17/006 ranges from 3.4 to 4.2 g/kg diet. This study demonstrates the potential of D. tharense NIOF17/006 as a promising aquafeed additive for improvement of the growth, immunity, and overall health of L. vannamei PLs, opening a new avenue for sustainable aquaculture practices.
Microalgae are spread over an extensive variety of aquatic environments and have applications in the environment, industry, and biotechnology fields (Ende et al., 2024). Due to their valuable biochemical composition, they are recognized as a respectable source of human food supplements (Rellán et al., 2009), pharmaceuticals (El-Sapagh et al., 2023; Abbas et al., 2023), cosmetics (Mourelle et al., 2017), aquafeed additives (Ashour et al., 2023), biofertilizers (Osman et al., 2010), bioethanol (de Farias Silva and Bertucco, 2016), and biodiesel (Karatay and Dönmez, 2011). In addition, cyanobacteria can contribute to phytobioremediation processes, which aids in the purification of contaminated environments (Ashour et al., 2022; Mansour et al., 2022a).
Over the last decade, the shrimp farming industry has conducted research and projects to find alternative natural components of antibiotics to boost shrimp growth and immune system response (Mansour et al., 2022b). Natural feed additives have shown promising results in disease control, survival, immune enhancement, and growth promotion (Goh et al., 2022). Immunostimulants, such as polysaccharides, nutrients, herbs, and microorganisms, have been identified as effective biofriendly agents for the control of pathogens and promotion of growth (Wang et al., 2017). Cyanobacteria and microalgae, due to their rich nutritional profile and bioactive compound content, present a promising avenue for exploration in this context. Various cyanobacteria species, including Arthrospira platensis, have been found to mitigate the effects of stress in farmed aquatic fish and shrimp (Abdel-Latif et al., 2022). Supplementation with A. platensis in the diet of shrimp has demonstrated immunomodulatory properties, enhancing intestinal defenses and improving the survival rates against viral challenges (Pilotto et al., 2019). The genus Desertifilum, which belongs to the order Oscillatoriales, has gained attention due to its adaptability to extreme environments and its potential biotechnological application (Dadheech et al., 2012b), as well as ease of harvesting as filamentous algae compared with unicellular algae (Wang et al., 2023). However, the potential of Desertifilum tharense as an aquafeed additive remains largely unexplored.
The whiteleg shrimp (Litopenaeus vannamei) is recognized as an important aquaculture species worldwide, known for its adaptability to various cultural conditions (Naser et al., 2022). Improving the performance, feed utilization, and immune status of shrimp through dietary interventions is crucial for a sustainable and profitable shrimp aquaculture. The use of cyanobacteria as a feed additive could potentially address these challenges while providing a sustainable alternative to conventional feed ingredients. For instance, natural pigments have demonstrated significant benefits in the aquaculture industry (Mansour et al., 2022d). However, it has been observed that not all cyanobacteria have positive effects on shrimp production. Cyanobacteria have also been associated with a decreased shrimp production. These contrasting findings underscore the critical importance of the careful selection and evaluation of specific cyanobacterial strains for their potential as aquafeed additives, as their effects on shrimp health and production can vary significantly (Magouz et al., 2021). Therefore, comprehensive studies on novel cyanobacterial strains, such as D. tharense, are essential in order to identify beneficial additives that can contribute to improved shrimp aquaculture practices (Maa-Iad et al., 2023).
The present study was conducted to examine the phylogenetic identification and potential application of the cyanobacterial strain D. tharense NIOF17/006 as a novel aquafeed additive for L. vannamei postlarvae (PLs). This study seeks to provide comprehensive insights into the prospective advantages of using a cyanobacterial strain in shrimp aquaculture. The results of this study could afford the development of innovative, sustainable feed additives for the aquaculture industry, potentially reducing the dependence on fishmeal and other conventional protein sources while improving shrimp health and production efficiency. Moreover, this research aligns with the growing interest in biobased solutions for sustainable aquaculture practices and the broader applications of microalgae and cyanobacteria in biotechnology and environmental management.
Water samples were obtained from the El-Mahmoudia Canal, the surface layer of water (upper 20 cm), Alexandria, Egypt (31°12′30.07″ N, 92°85′66.44″ E). The water samples were collected using sterilized bottles and promptly taken to the laboratory. A portable pH/temperature meter (Milwaukee MW102) was used to measure the temperature and pH in the field. The water turbidity and depth were also assessed using a Secchi disc with a diameter of 35 cm. Salinity was determined using a portable conductivity meter (Oakton, Eutech Instruments, Vernon Hills, IL, USA). Under control temperature conditions (25 ± 1°C) and continuous illumination (120 μmol photons m−2 s−1), selected water samples were inoculated and purified with the agar medium method using BG11 culture medium, as previously described by Robert (2005). After 2 weeks of maintenance, healthy colonies were transferred and inoculated into sterilized test tubes using the serial dilution method (Robert, 2005), with a total culture volume of 10 ml. Thereafter, the culture volume was scaled to 100 ml in 250-ml conical flasks for further subculture. Subsequently, the culture was upscaled in a volume of 20 L. Morphological examination of the isolate was first performed using a light microscope (Olympus BX51 Light Microscope, Tokyo, Japan) following standard identification references (Zaki et al., 2021). Finally, the results obtained from the morphological examination of the isolates were validated using molecular techniques.
The standard CTAB protocol was used to extract the entire genomic DNA from a dried cyanobacterial pellet according to Grube et al. (1995), with minor modifications as described by Elshobary et al. (2015). The universal cyanobacterial primers CYA106F (5′-CGGACGGGTGAGTAACGCGTGA-3′) and CYA781R (5′-GACTACAGGGGTATCTAATCCCWTT-3′) were used to amplify the 16S rRNA gene with the following PCR conditions: initial denaturation at 94°C for 5 min, 35 cycles of denaturation at 94°C for 30 s, annealing at 55°C for 30 s, and extension at 72°C for 1 min, followed by a final extension at 72°C for 7 min. The PCR product was then purified and sequenced using Sanger sequencing. Raw sequence data were edited and assembled using BioEdit software v7.2.5. The edited sequence was analyzed using BLAST against the NCBI nucleotide database for species identification. Multiple sequence alignment with retrieved reference sequences was performed using MEGA 11 software with the ClustalW algorithm. The sequence was then deposited in GenBank under the accession number OQ147028. For phylogenetic analysis, the obtained 16S rRNA sequences were aligned with 18 reference cyanobacterial strains obtained from the NCBI Ribosomal DNA database using Aulosira sp. as an outgroup to root the tree and provide evolutionary context. The neighbor-joining method was used to construct the phylogenetic tree.
Culture volumes of 10 ml were subjected to centrifugation for 10 min at 4,000 × g, with the resulting pellet kept at −20°C for biochemical analysis after discarding the supernatant. Biochemical analysis of the microalgal biomass measurement was according to AOAC (2003).
The PLs (1.68 ± 0.05 g) of L. vannamei were purchased from a private hatchery and transferred to NIOF laboratories, where they were adapted to laboratory conditions during a period of 15 days. During this acclimatization period, PLs were fed a commercial shrimp basal diet four times a day.
A total of 600 PLs were used, which were divided into four treatment groups (with three replicates per group). After a 15-day acclimation period, each treatment group received 50 PLs. The PLs were placed in net hapas measuring 0.7 m × 0.7 m × 1 m. These hapas were secured in concrete ponds measuring 4 m × 2 m × 1 m. Throughout the experimental period, the PLs were maintained under the recommended conditions for whiteleg shrimp PLs as reported by APHA (2005). The hapas are cleaned regularly, and the water daily exchange was 10%.
During the 8-week feeding experiment, shrimp PLs were fed the following four dietary groups: T0, a commercial shrimp diet as the control diet (containing 45% protein, 7.9% lipid, 34% carbohydrate, 3.65% fiber, and 9.1% ash; Aller-Aqua, Giza Governorate, Egypt), and three other diets (T1, T2, and T3) that were supplemented with 1, 2.5, and 5 g, respectively, of the dried powder of the microalgal isolate. The dried microalgal isolate levels were added to a specific diet according to the method previously described by Sharawy et al. (2022). In brief, four equal sections of the commercial diet, which served as the control, were crushed into a fine powder. The appropriate amount of cyanobacterial isolate was then added to each section, and the mixture was carefully mixed until homogeneous. In accordance with Sharawy et al. (2022), after dissolving the appropriate dose of cyanobacterial isolate in distilled water, it was sprayed onto the surface of the diet of the related section. An equivalent volume of distilled water without the cyanobacterial isolate was administered to the control diet (T0). To cover the mixture of diet and cyanobacterial isolate, a 5-ml/kg diet of oil (sunflower oil) was sprayed over the basal diet after drying all diets for 48 h at 40°C to maintain a moisture content of approximately 10% (Zeraatpisheh et al., 2018). Lastly, 4°C was used to maintain the diet pellets until use.
The initial weight (IW, in grams) and the final weight (FW, in grams) of PLs were measured to determine the weight gain (WG), the specific growth rate (SGR), and the feed conversion ratio (FCR). The survival rate (SR, in percent) and the protein efficiency ratio (PER) were calculated using the following equations:
From each replicate, seven PLs were collected to analyze the shrimp’s whole body chemical composition. Random shrimp samples were mixed, dried, crushed into a powder, and stored at a low temperature (−20°C) until analysis. The chemical body composition was analyzed following AOAC (2003).
From each replicate, seven PL samples were randomly chosen after a 24-h fasting period. The samples were briefly washed with sterile seawater and then cut, weighed, and frozen until analysis at −80°C. The shrimp samples were homogenized [at a pH of 7.4 in phosphate-buffered saline (PBS)] and then centrifuged (20 min at 3,000 × g). Measurement of the homogenate lysozyme activity was carried out using Lysozyme (LZM) ELISA kits following the instructions given by the manufacturer (cat no. SL0050FI; SunLong Biotech Co., Ltd., Hangzhou, China). The antioxidant enzyme activities, including malondialdehyde (MDA), catalase (CAT), and superoxide dismutase (SOD), were measured using specific kits at wavelengths of 510, 534, and 560 nm, respectively, based on the instructions given by the manufacturer (cat nos. MD2529, SD2521, and CA2517, respectively; Biodiagnostic Company, Cairo, Egypt). For digestive enzyme (amylase and lipase) activity determination, GIT-homogenized tissues were separated by careful centrifugation. The digestive enzyme activities were measured using spectrophotometric assays at wavelengths of 580 and 660 nm.
Three shrimp samples from each group were taken at the end of the experiments. Total RNA was isolated from muscle tissue using Easy-RED (Easy-RED, iNtRON, Seongnam, South Korea) according to the manufacturer’s instructions. The total RNA concentration and purity were measured using NanoDrop (Nanophotometer, NP80 touch, Implen, München, Germany). The RNA products were used for the synthesis of cDNA using a commercial kit (Thermo Scientific™ RevertAid First Strand cDNA Synthesis Kit; Thermo Fisher Scientific, Waltham, MA, USA). The cDNAs were amplified through Rotor-Gene Q thermal cycling (Hassan et al., 2023; Wang et al., 2008; Jian et al., 2013) to quantify the expression of some growth- and immunity-related genes, including growth hormone (GH), insulin-like growth factor I (IGF-I), insulin-like growth factor II (IGF-II), prophenoloxidase (proPO), superoxide dismutase (SOD), and lysozyme (Lys). The quantitative PCR (qPCR) reactions were performed in 20 μl including 10 μl of ABT 2× qPCR Mix Kit (SYBR Green/low ROX), 0.5 μl of each primer (10 μM), 4 μl (50 ng) of cDNA, and 5 μl of RNAse-free water. The reaction followed the thermal profile: initial denaturation at 95°C for 10 min, followed by 40 cycles, each of which included denaturation for 10 s at 95°C, annealing at 58–60°C for 10 s, and extension for 30 s at 72°C. Subsequently, the temperature was increased by 0.5°C from 60°C to 95°C to establish a melting curve, which was then utilized to analyze the target gene products. In these reactions, the β-actin gene was included as a housekeeping gene (Hassan et al., 2023; Wang et al., 2008; Jian et al., 2013). The primer sequences and amplicon sizes are provided in Table 1. According to the 2−ΔΔCt method, the β-actin gene was used to normalize the Ct values of the target genes (Rao et al., 2013). Statistical analysis was performed using GraphPad Prism software ver. 8.0.1 (GraphPad Prism, La Jolla, CA, USA). The results are presented as mean ± SD, with the level of significance considered at a probability value less than 0.05 (p < 0.05).
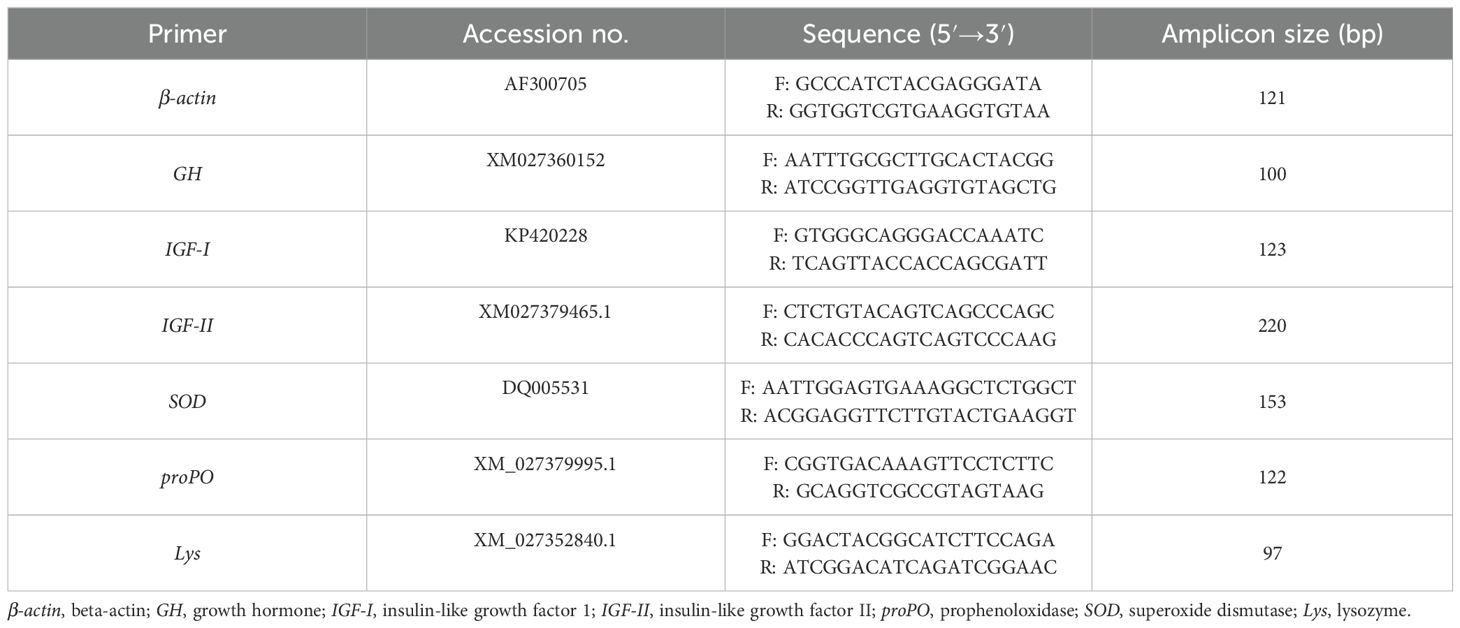
Table 1. Primers, accession numbers, sequences, and amplicon sizes (in base pairs) used for the shrimp quantitative PCR (qPCR) study.
Data were collected in three replicates (±SD). Prior to statistical analysis, Levene’s test was performed to ensure normality and homogeneity assumptions, and the results (in percent) were arcsine transformed (Zar, 1984). SPSS Statistics software was employed to perform the statistical analysis, which included one-way analysis of variance followed by the Duncan (1955) test at p ≤ 0.05. Finally, the GraphPad (Prism 8) Statistics software was employed to create the figures (Swift, 1997), while Excel software was used to conduct polynomial regression.
Figure 1 shows the morphological observations of the cyanobacterial isolate D. tharense NIOF17/006 using a light microscope (×400). The optical microscopic observations found that D. tharense NIOF17/006 is filamentous with a trichome width ranging from 2.2 to 3.6 μm.
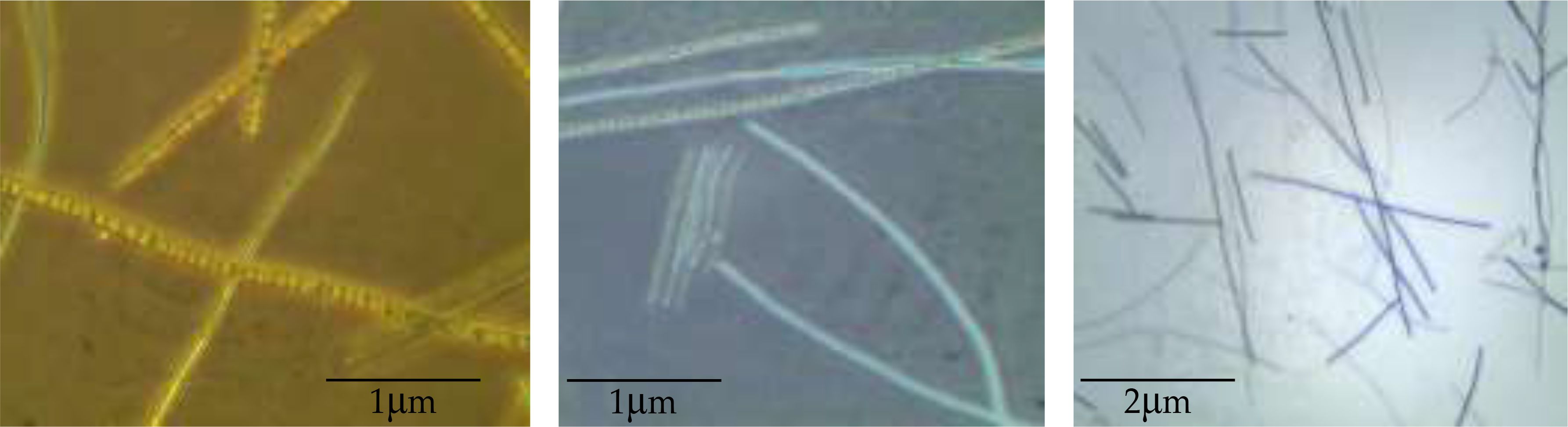

Figure 1. Optical light microscopic observations of the cyanobacterial isolate Desertifilum tharense NIOF17/006 at different magnifications.
Based on the recent taxonomic literature by Komárek (2005), the phenotypic characteristics of D. tharense NIOF17 classified it within the order Oscillatoriales. The common phenotypic features include a thin, pale to bright blue-green thallus. The filaments are either solitary or densely entangled, varying in length. This strain exhibited motility through gliding and oscillation movements, with tapered ends. The sheath surrounding the trichome is thin, colorless, and attached (agglutinated). The cylindrical cells have a uniform cell content, while the apical cells are long-conical with a rounded apex. The shape of the strains shows variability. Similar observations have been previously reported by Dadheech et al. (2012a), who isolated four strains of the cyanobacteria D. tharense: PD2001/TDC7, PD2001/TDC4, PD2001/TDC17T, and PD2001/TDC14.
A phylogenetic tree (Figure 2) of the 16S rRNA gene was constructed using PCR-based nucleotide sequences that spanned more than 1,363 bp to provide a substantial amount of genetic information, allowing for a high-resolution analysis. This approach aligns with best practices in molecular phylogenetics, where longer sequences generally yield more reliable results (Patwardhan et al., 2014).
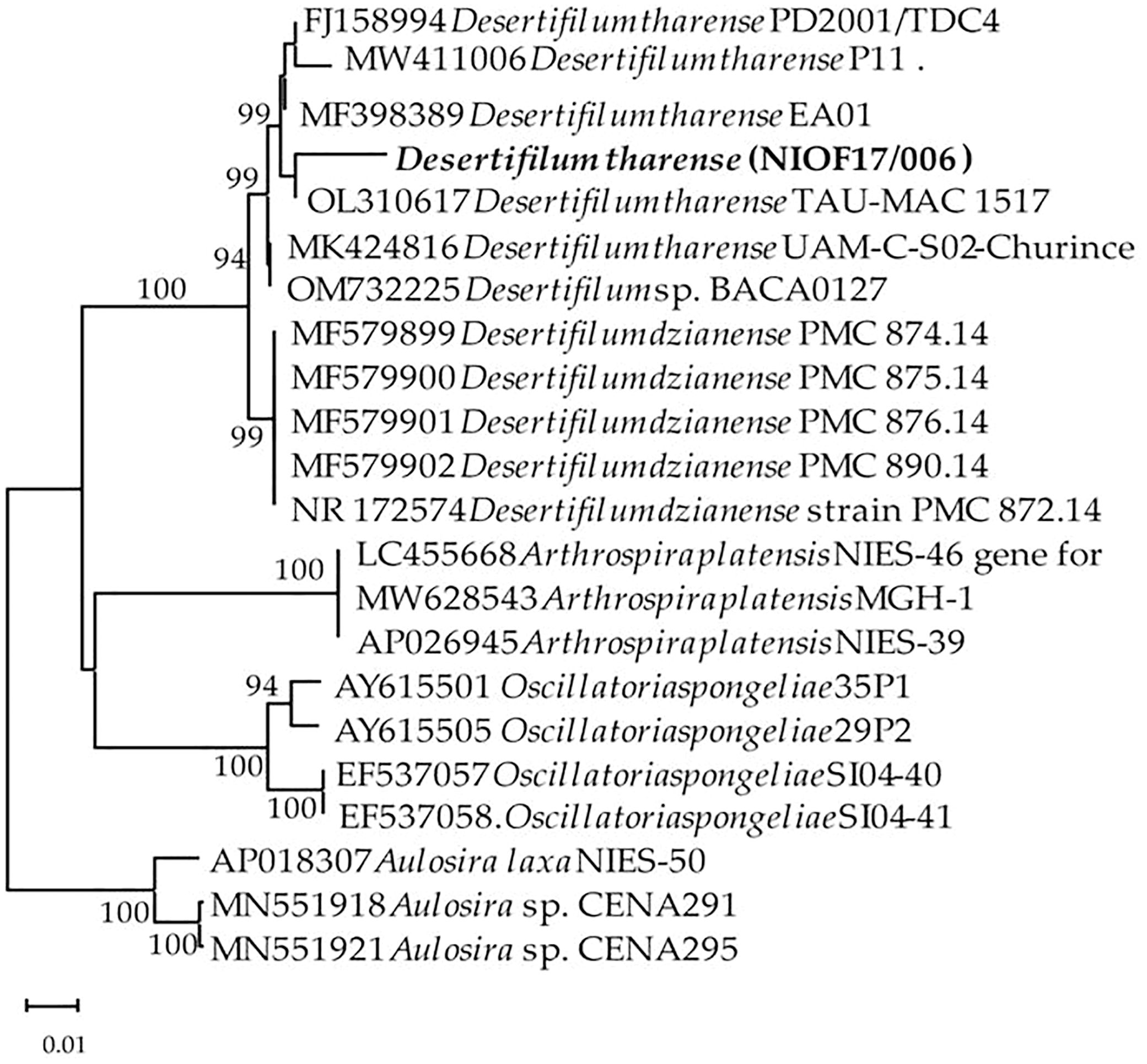
Figure 2. Neighbor-joining (NJ) dendrograms revealing the cyanobacteria-isolated strains based on 16S rRNA nucleotide sequences. Bootstrap values ≥70 are displayed on the trees.
The neighbor-joining method was used for the construction of the phylogenetic tree, which is known for its efficiency and accuracy in depicting evolutionary relationships (Trees, 1987). The obtained 16S gene sequences were aligned with the 16S sequences of 18 cyanobacterial strains from the GenBank Ribosomal DNA database to ensure a comprehensive comparison, while the use of Aulosira sp. as an outgroup helps to root the tree and provide evolutionary context (Huelsenbeck et al., 2002). This methodological approach strengthens the reliability of the phylogenetic reconstruction. The resulting phylogenetic tree (Figure 2) showed that each species formed a distinct clade, and the isolated strain was identified as D. tharense, which strongly matches D. tharense FJ158994, MK424816, and MW411006 with high similarity values of 98.53%, 98.46%, and 98.01%, respectively. It is worth noting that while 16S rDNA analysis is highly informative, complementary approaches such as whole-genome sequencing or multilocus sequence typing could provide even more comprehensive insights into the genetic makeup and evolutionary history of a strain (Huelsenbeck et al., 2002).
In the current study, the biochemical composition of the isolate D. tharense from cyanobacteria NIOF17/006 was examined, focusing on its protein, lipid, and carbohydrate contents. Figure 3 shows the biochemical composition of the cyanobacterial isolate D. tharense NIOF17/006 at the end of the late exponential growth phase. The average values for total protein, total lipid, and total carbohydrate were 37.74% ± 1.2%, 5.52% ± 0.75%, and 21.25% ± 0.87% (on a dry weight basis), respectively. Comparing these findings to the previous work by Hernández–Martínez et al. (2023), it is evident that D. tharense exhibits notable variations in its biochemical composition.

Figure 3. Biochemical composition of the cyanobacterial isolate Desertifilum tharense NIOF17/006 (percent of dry weight basis).
Table 2 illustrates the growth, survival, and nutrient utilization parameters of shrimp fed diets supplemented with the cyanobacterial strain. In this feeding trial, significant improvements (p < 0.05) were observed in the FW, WG, FCR, and SR between the control group (T0) and the groups supplemented with cyanobacteria (T1, T2, and T3). This improvement increased with increasing levels of cyanobacterial supplementation. However, no significant improvements (p < 0.05) were observed in other parameters of growth and nutrient utilization, i.e., SGR and PER.
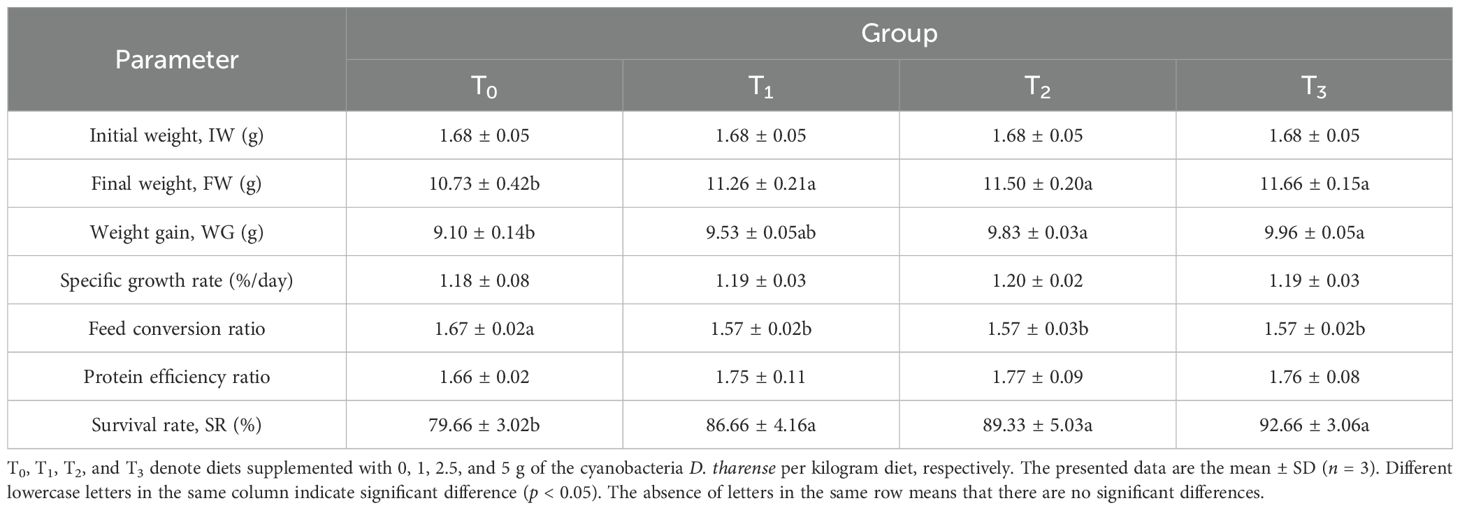
Table 2. Growth performance and feed utilization of shrimp Litopenaeus vannamei fed diets supplemented with different levels of the cyanobacteria Desertifilum tharense NIOF17/006.
Polynomial regression is a basic model in statistical machine learning (ML). It enables researchers to model how the predictor parameters and the outcome variables are related. Polynomial regression is an extension of a standard linear regression model. The nonlinear relationship between a predictor and an outcome variable is modeled using polynomial regression. Nonlinear relationships can be accurately modeled using polynomial regression (Maulud and Abdulazeez, 2020). Figure 4 shows the polynomial regression model of the dietary supplementation levels (in grams) of the cyanobacterial isolate D. tharense NIOF17/006 and the WG and FCR of the whiteleg shrimp L. vannamei. Figure 4 illustrates that, with the increase of D. tharense NIOF17/006 supplementation in the diet, the WG polynomial regression increased (r2 = 0.814) while the FCR polynomial regression decreased (r2 = 0.994). In conclusion, based on Figure 4, the explanation lines of the highest and the lowest peaks of WG and FCR, respectively, presented in the polynomial regression model of ML predict that the ideal supplementation level of the probiotic cyanobacteria D. tharense NIOF17/006 ranges from 3.4 to 4.2 g/kg diet. This ideal range exists between groups T2 and T3 (2.5 and 5 g/kg, respectively).
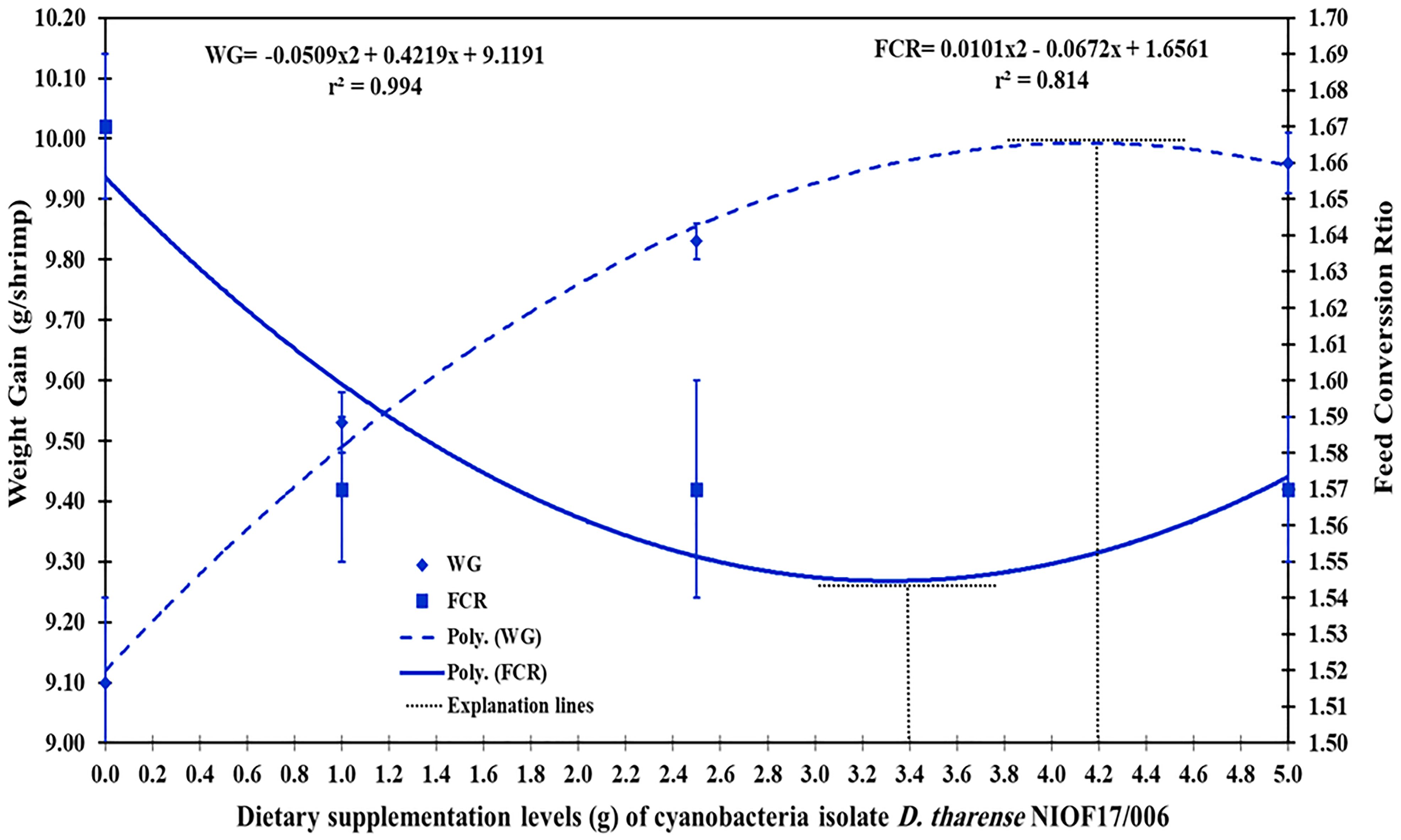
Figure 4. Polynomial regression of shrimp weight gain (WG), feed conversion ratio (FCR), and dietary supplementation levels (in grams) of the cyanobacterial isolate Desertifilum tharense NIOF17/006.
In the aquaculture domain, El-Khodary et al. (2021) conducted a study on the growth, survival, and pigmentation of the larvae of Solea aegyptiaca using different microalgal species, underscoring the importance of microalgae in fish nutrition and development. The same findings were observed by Sharawy et al. (2022), who utilized Tetraselmis suecica as an aquafeed additive for L. vannamei. To the best of our knowledge, this is the first work to report on the potential application of the cyanobacterial isolate D. tharense NIOF17/006 as an aquafeed additive for L. vannamei. In the current study, the improvements in shrimp WG, FW, FCR, and SR may be due to the nutrient composition of the probiotic cyanobacterial isolate D. tharense NIOF17/006. However, it is commonly known that several cyanobacterial strains, particularly A. platensis, have high concentrations of multiple biologically active components that support the growth performance, immunity, and antioxidant capabilities of a variety of aquatic animals, such as the shrimp L. vannamei (Sharawy et al., 2022), gilthead sea bream (Galafat et al., 2022), Nile tilapia (Mabrouk et al., 2022), hybrid red tilapia (El-Sheekh et al., 2014), and common carp (Suantika et al., 2016).
Table 3 illustrates the biochemical composition analysis of shrimp fed the diet supplemented with different levels of the cyanobacteria-isolated strain D. tharense NIOF17/006. As presented in the table, significant differences (p < 0.05) were found in the total composition (protein, fat, dry matter, and ash) of L. vannamei in the supplementation groups (T1, T2, and T3) compared with the control group (T0). The percentages of carcasses increased with increasing supplementation doses. These improvements may be due to the high contents of protein (37.74%), lipid (5.52%), and carbohydrate (21.25%) of the strain of D. tharense isolated from cyanobacteria NIOF17/006. This is the first time this strain was used as an aquafeed additive for the whiteleg shrimp L. vannamei. The data shown in Table 3 are consistent with the findings by several authors who concluded that high supplementation levels of the cyanobacterial strain A. platensis significantly increased the shrimp carcass composition even in the whiteleg shrimp L. vannamei (Ashour et al., 2024) and the freshwater prawn Macrobrachium rosenbergii (Radhakrishnan et al., 2016).
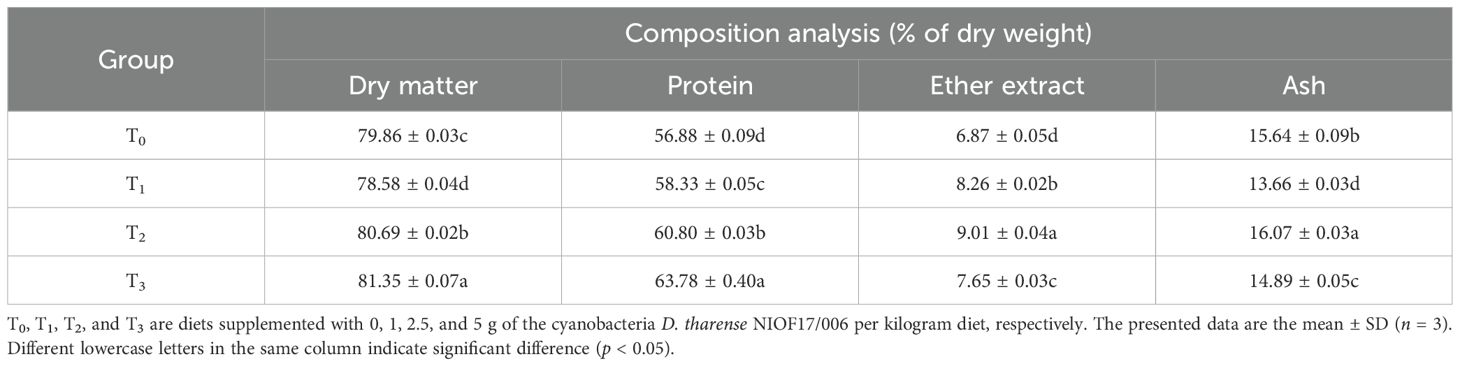
Table 3. Composition analysis (in percent) of the shrimp Litopenaeus vannamei fed diets supplemented with different levels of the cyanobacteria Desertifilum tharense NIOF17/006.
The results showed that, compared with those in T0, shrimp in the cyanobacteria supplementation groups (T1, T2, and T3) showed the highest significant (p < 0.05) Lys, SOD, and CAT values. On the other hand, these groups exhibited the lowest significant (p < 0.05) MDA value compared with T0. The shrimp immune system primarily relies on nonspecific processes due to its lack of adaptive immunity (Farzanfar, 2006). The enzymes Lys and SOD break down the cell walls of harmful bacteria and destroy free radicals in cells (Naiel et al., 2021). MDA is commonly used as an indicator of oxidative stress as it also shows an increase in the generation of free radicals (Chen et al., 2016). These shrimp humoral substances, e.g., Lys, SOD, MDA, and CAT, play essential roles in both specific and nonspecific immunity (Ashour et al., 2024). The findings of the current work aligned with those of the study by Ashour et al. (2024), who concluded that the nonspecific immunity and antioxidant activities of L. vannamei significantly improved when fed diets containing different levels of the cyanobacteria-isolated strain A. platensis. Furthermore, several works have reported that the use of different algal strains significantly improved the nonspecific immunity and antioxidant activities of aquatic animals (Mansour et al., 2022c). According to Jerez-Cepa and Ruiz-Jarabo (2021), immune-related factors have been studied as predicted indications for the examination of shrimp health. Figure 5 illustrates the immunological responses and antioxidant activities of L. vannamei fed diets containing different levels of the cyanobacterial strain D. tharense NIOF17/006.
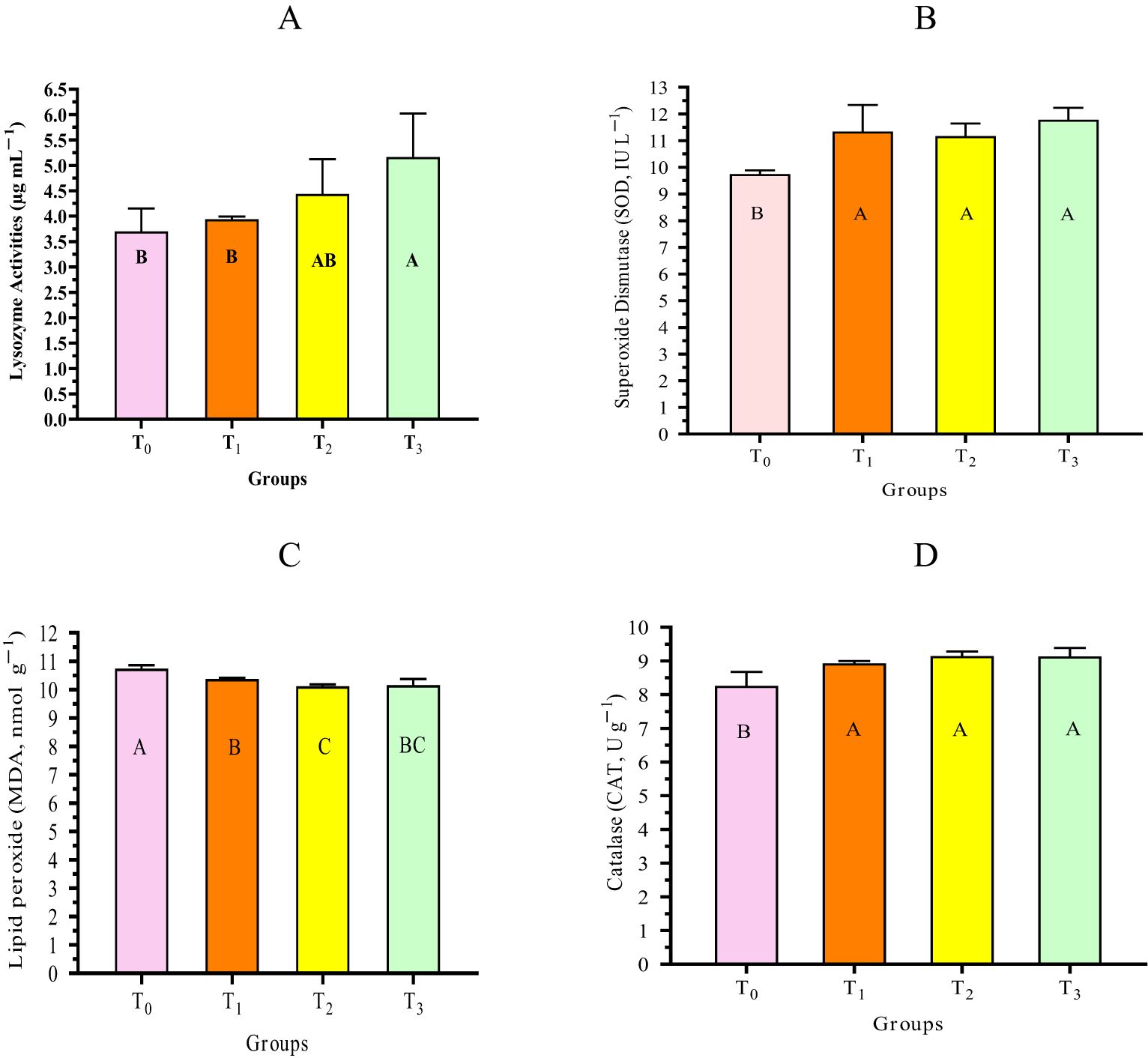
Figure 5. (A–D) Nonspecific immunity and antioxidant activity of Litopenaeus vannamei fed diets containing different levels of the cyanobacterial isolate Desertifilum tharense NIOF17/006. T0, T1, T2, and T3 denote diets supplemented with 0, 1, 2.5, and 5 g of D. tharense NIOF17/006 per kilogram diet, respectively. The presented data are the mean ± SD (n = 3). Different uppercase letters in the columns indicate significant difference (p < 0.05).
Figure 6 shows the digestive enzyme activity of L. vannamei fed diets supplemented with different doses of the cyanobacterial strain D. tharense NIOF17/006. The results revealed that shrimp in the cyanobacteria supplementation groups (T1, T2, and T3) exhibited the highest significant (p < 0.05) amylase and lipase activities compared with those fed T0. The highest significant (p < 0.05) amylase and lipase values were observed in shrimp reared in groups T3 and T2, respectively. These current findings may be attributed to the fact that the bioactive materials of D. tharense NIOF17/006 may improve the secretion of digestive enzymes and, subsequently, improve feed absorption and digestibility (Xu et al., 2018). Moreover, it could be explained by the fact that shrimp fed diets containing higher levels of D. tharense NIOF17/006 might stimulate recycling, which is believed to be the outcome of the movement of fluid in the midgut lumen and the compartmentalization caused by the peritrophic membrane (Alexandre et al., 2014). Furthermore, according to this theory, if aquafeed supplementation increases the amount of protein in the diet, the corresponding digestive enzymes will be displaced, which ultimately results in a greater recovery of these types of enzymes in the feces (Ozorio et al., 2015).
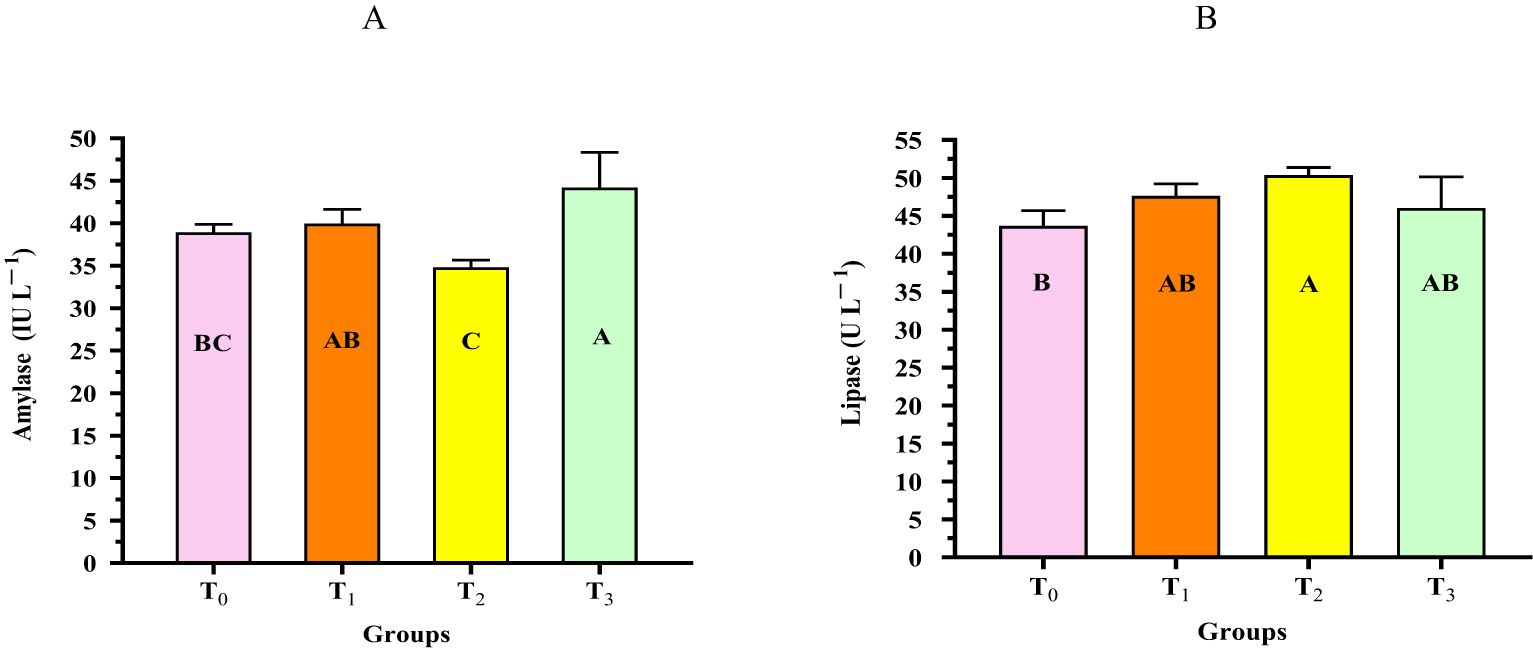
Figure 6. (A, B) Digestive enzyme (amylase and lipase) activity of the shrimp Litopenaeus vannamei fed diets supplemented with different levels of the cyanobacterial strain Desertifilum tharense NIOF17/006. T0, T1, T2, and T3 denote diets supplemented with 0, 1, 2.5, and 5 g of D. tharense NIOF17/006 per kilogram diet, respectively. The presented data are the mean ± SD (n = 3). Different letters in the columns indicate significant difference (p < 0.05).
The results of the gene expression analysis showed that the expression of the growth-related genes (GH, IGF-I, and IGF-II) increased almost significantly with increasing levels of supplementation with the cyanobacterial strain D. tharense NIOF17/006 in the formulated diets, except for T2, which had a lower expression compared with T1 (Figures 7A–C). The highest expression in shrimp was observed in group T3 (5 g/kg diet). Similarly, the expression of the immune-related genes (proPO, SOD, and Lys) showed the same pattern and was upregulated in the muscles of L. vannamei with increasing concentrations of the cyanobacterial strain D. tharense NIOF17/006 in the T1 and T3 formulated diets (Figures 7D–F). With regard to the growth-related genes, GH is synthesized in the shrimp’s pituitary gland and exerts its action on the target cell to stimulate shrimp growth (Chandhini et al., 2021). IGF-I is a peptide hormone that is essential for the control of growth and development in shrimp (Pang et al., 2021). IGF-II is another protein that belongs to the IGF family. It plays a role in fetal growth and development and participates in tissue repair and replacement in adult shrimp (Li et al., 2024). With regard to the immunity-related genes, ProPO is a protein and an enzyme that participates in the defense system of shrimp.
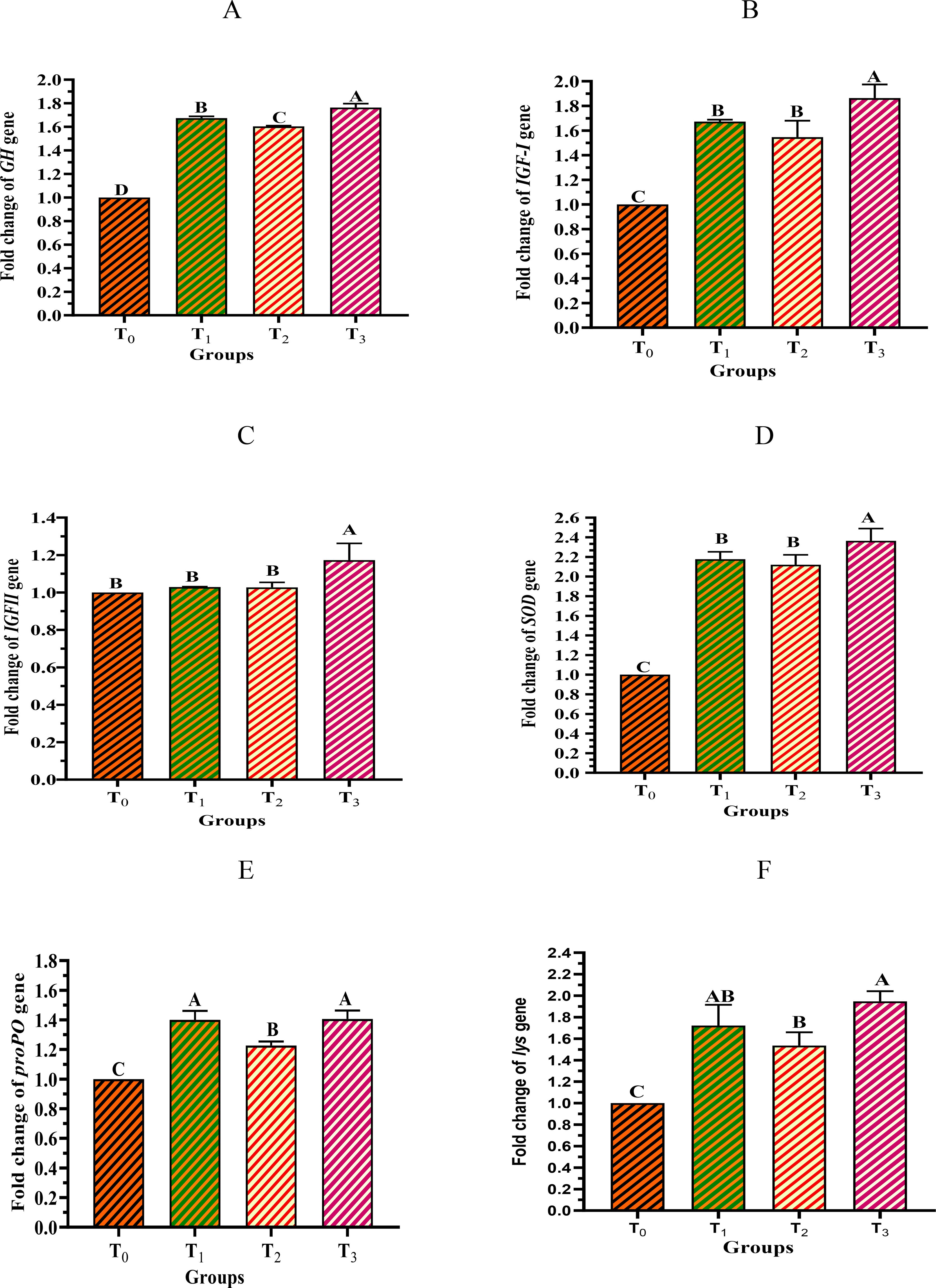
Figure 7. Expression levels of the growth-related genes—GH (A), IGF-1 (B), and IGF-II (C) and the immunity-related genes—SOD (D), ProPO (E), and Lys (F) in the muscles of Litopenaeus vannamei fed diets containing different levels of the cyanobacterial strain Desertifilum tharense NIOF17/006.
When shrimp encounter pathogens, proPO is converted into the active form and catalyzes the melanization process. Melanization assists in pulling into the body these foreign microbes and destroys them, hence acting as a barrier against diseases (Amparyup et al., 2013). SOD is one of the antioxidant enzymes, the role of which is to protect shrimp cells against damage. SOD sustains the proper work of cells and shields the shrimp body from free radicals by transforming superoxide radicals into less harmful molecules (Chirawithayaboon et al., 2020). Finally, Lys is an antimicrobial enzyme that comprise an essential part of the shrimp immune system defense due to its content in the hemolymph. It is a versatile enzyme that is capable of attacking a wide variety of bacteria, hence offering an indication of protection against bacterial infections (Ferraboschi et al., 2021). According to the literature, there are no previous studies on the potential application of the cyanobacterial strain D. tharense NIOF17/006 as a feed additive for the shrimp L. vannamei or as an aquafeed additive in general. However, the findings of the current study have been previously confirmed by several studies reporting that the dietary inclusion of micro/macroalgae positively improved the growth and immune gene expression of L. vannamei (Ashour et al., 2024), the rainbow trout Oncorhynchus mykiss (Ratti et al., 2023), and the Nile tilapia Oreochromis niloticus (Mabrouk et al., 2024).
The biotechnological applications of microalgae have attracted significant global attention. This study contributes to this growing field by identifying and evaluating the potential of the cyanobacterial strain D. tharense NIOF17/006 as a novel aquafeed additive for L. vannamei PLs. The findings demonstrate that D. tharense possesses a favorable biochemical composition suitable for shrimp nutrition. Dietary supplementation of this cyanobacteria at levels of 2.5 and 5 g/kg resulted in significant improvements in feed utilization, growth performance, survival, whole body composition, nonspecific immunity, antioxidant activity, and digestive enzyme function, as well as in the expression of the growth- and immunity-related genes. In particular, the most pronounced benefits were observed at supplementation levels of 2.5–5 g/kg. Furthermore, the polynomial regression ML model predicts that the ideal level of supplementation with the probiotic cyanobacteria D. tharense NIOF17/006 ranges from 3.4 to 4.2 g/kg diet. This study highlights the potential use of D. tharense NIOF17/006 as a valuable feed additive in shrimp aquaculture, offering a sustainable approach to enhancing shrimp health and production efficiency.
The raw data supporting the conclusions of this article will be made available by the authors, without undue reservation.
The manuscript presents research on animals that do not require ethical approval for their study. The animal study protocol was approved by the Institutional Review Board of the National Institute of Oceanography and Fisheries, Egypt.
MA: Conceptualization, Data curation, Formal analysis, Funding acquisition, Investigation, Methodology, Project administration, Resources, Software, Visualization, Writing – review & editing. ASA: Funding acquisition, Resources, Validation, Writing – review & editing, Investigation. AhM: Formal Analysis, Investigation, Methodology, Resources, Writing – review & editing. FA: Conceptualization, Data curation, Formal Analysis, Methodology, Writing – original draft. ME: Conceptualization, Investigation, Methodology, Resources, Software, Writing – original draft. MM: Conceptualization, Formal analysis, Methodology, Supervision, Validation, Writing – original draft. AIAM: Conceptualization, Data curation, Methodology, Resources, Writing – original draft. ATM: Investigation, Validation, Visualization, Writing – review & editing. EE: Resources, Software, Visualization, Writing – review & editing. AFA: Conceptualization, Investigation, Methodology, Validation, Writing – original draft.
The author(s) declare that financial support was received for the research and/or publication of this article. This work was supported by the Deanship of Scientific Research, Vice Presidency for Graduate Studies and Scientific Research, King Faisal University, Saudi Arabia (KFU250399).
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declared that they were an editorial board member of Frontiers, at the time of submission. This had no impact on the peer review process and the final decision.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Abbas E. M., Al-Souti A. S., Sharawy Z. Z., El-Haroun E., Ashour M. (2023). Impact of dietary administration of seaweed polysaccharide on growth, microbial abundance, and growth and immune-related genes expression of the pacific whiteleg shrimp (Litopenaeus vannamei). Life 13, 344. doi: 10.3390/life13020344
Abdel-Latif H. M., El-Ashram S., Yilmaz S., Naiel M. A., Kari Z. A., Hamid N. K. A., et al. (2022). The effectiveness of Arthrospira platensis and microalgae in relieving stressful conditions affecting finfish and shellfish species: An overview. Aquaculture Rep. 24, 101135. doi: 10.1016/j.aqrep.2022.101135
Alexandre D., Ozorio R. A., Derner R. B., Fracalossi D. M., Oliveira G. B., Samuels R. I., et al. (2014). Spatial distribution of digestive proteinases in the midgut of the Pacific white shrimp (Litopenaeus vannamei) indicates the existence of endo-ectoperitrophic circulation in Crustacea. Comp. Biochem. Physiol. Part B: Biochem. Mol. Biol. 172, 90–95. doi: 10.1016/j.cbpb.2014.04.010
Amparyup P., Charoensapsri W., Tassanakajon A. (2013). Prophenoloxidase system and its role in shrimp immune responses against major pathogens. Fish Shellfish Immunol. 34, 990–1001. doi: 10.1016/j.fsi.2012.08.019
AOAC (2003). Official Methods of Analysis of the Association of Official’s Analytical Chemists. 17th Edition. (Arlington, Virginia: Association of Official Analytical Chemists (AOAC)).
APHA (2005). APHA, Standard methods for the examination of water and wastewater (USA: Washington DC, USA: American Public Health Association).
Ashour M., Alprol A. E., Khedawy M., Abualnaja K. M., Mansour A. T. (2022). Equilibrium and kinetic modeling of crystal violet dye adsorption by a marine diatom, Skeletonema costatum. Materials 15, 6375. doi: 10.3390/ma15186375
Ashour M., Al-Souti A. S., Hassan S. M., Ammar G. A. G., Goda A., El-Shenody R., et al. (2023). Commercial seaweed liquid extract as strawberry biostimulants and bioethanol production. Life 13, 85. doi: 10.3390/life13010085
Ashour M., Mabrouk M. M., Mansour A. I. A., Abdelhamid A. F., Abdel-Kader M. F., Elokaby M. A., et al. (2024). Impact of dietary administration of Arthrospira platensis free-lipid biomass on growth performance, body composition, redox status, immune responses, and some related genes of pacific whiteleg shrimp, Litopenaeus vannamei. PLoS One 19, e0300748. doi: 10.1371/journal.pone.0300748
Chandhini S., Trumboo B., Jose S., Varghese T., Rajesh M., Kumar V. R. (2021). Insulin-like growth factor signalling and its significance as a biomarker in fish and shellfish research. Fish Physiol. Biochem. 47, 1011–1031. doi: 10.1007/s10695-021-00961-6
Chen Y.-Y., Chen J.-C., Tayag C. M., Li H.-F., Putra D. F., Kuo Y.-H., et al. (2016). Spirulina elicits the activation of innate immunity and increases resistance against Vibrio alginolyticus in shrimp. Fish shellfish Immunol. 55, 690–698. doi: 10.1016/j.fsi.2016.06.042
Chirawithayaboon P., Areechon N., Meunpol O. (2020). Hepatopancreatic antioxidant enzyme activities and disease resistance of Pacific white shrimp (Litopenaeus vannamei) fed diet supplemented with garlic (Allium sativum) extract. Agric. Natural Resour. 54, 377–386-377–386.
Dadheech P. K., Abed R. M., Mahmoud H., Mohan M. K., Krienitz L. (2012a). Polyphasic characterization of cyanobacteria isolated from desert crusts, and the description of Desertifilum tharense gen. et sp. nov.(Oscillatoriales). Phycologia 51, 260–270. doi: 10.2216/09-51.1
Dadheech P. K., Mahmoud H., Kotut K., Krienitz L. (2012b). Haloleptolyngbya alcalis gen. et sp. nov., a new filamentous cyanobacterium from the soda lake Nakuru, Kenya. Hydrobiologia 691, 269–283. doi: 10.1007/s10750-012-1080-6
de Farias Silva C. E., Bertucco A. (2016). Bioethanol from microalgae and cyanobacteria: a review and technological outlook. Process Biochem. 51, 1833–1842. doi: 10.1016/j.procbio.2016.02.016
El-Khodary G. M., El-Sayed H. S., Khairy H. M., El-Sheikh M. A., Qi X., Elshobary M. E. (2021). Comparative study on growth, survival and pigmentation of Solea aEgyptiaca larvae by using four different microalgal species with emphasize on water quality and nutritional value. Aquaculture Nutr. 27, 615–629. doi: 10.1111/anu.13211
El-Sapagh S., El-Shenody R., Pereira L., Elshobary M. (2023). Unveiling the potential of algal extracts as promising antibacterial and antibiofilm agents against multidrug-resistant pseudomonas aeruginosa: in vitro and in silico studies including molecular docking. Plants 12, 3324. doi: 10.3390/plants12183324
El-Sheekh M., El-Shourbagy I., Shalaby S., Hosny S. (2014). Effect of feeding Arthrospira platensis (Spirulina) on growth and carcass composition of hybrid red tilapia (Oreochromis niloticus x Oreochromis mossambicus). Turkish J. Fisheries Aquat. Sci. 14, 471–478. doi: 10.4194/1303-2712-v14_2_18
Elshobary M. E., Osman M. E., Abushady A. M., Piercey-Normore M. D. (2015). Comparison of lichen-forming cyanobacterial and green algal photobionts with free-living algae. Cryptogamie Algologie 36, 81–100. doi: 10.7872/crya.v36.iss1.2015.81
Ende S., Henjes J., Spiller M., Elshobary M., Hanelt D., Abomohra A. (2024). Recent advances in recirculating aquaculture systems and role of microalgae to close system loop. Bioresource Technol. 407, 131107. doi: 10.1016/j.biortech.2024.131107
Farzanfar A. (2006). The use of probiotics in shrimp aquaculture. FEMS Immunol. Med. Microbiol. 48, 149–158. doi: 10.1111/j.1574-695X.2006.00116.x
Ferraboschi P., Ciceri S., Grisenti P. (2021). Applications of lysozyme, an innate immune defense factor, as an alternative antibiotic. Antibiotics 10, 1534. doi: 10.3390/antibiotics10121534
Galafat A., Vizcaíno A. J., Sáez M. I., Martínez T. F., Arizcun M., Chaves-Pozo E., et al. (2022). Assessment of dietary inclusion of crude or hydrolysed Arthrospira platensis biomass in starter diets for gilthead seabream (Sparus aurata). Aquaculture 548, 737680. doi: 10.1016/j.aquaculture.2021.737680
Goh J. X. H., Tan L. T. H., Law J. W. F., Ser H. L., Khaw K. Y., Letchumanan V., et al. (2022). Harnessing the potentialities of probiotics, prebiotics, synbiotics, paraprobiotics, and postbiotics for shrimp farming. Rev. Aquaculture 14, 1478–1557. doi: 10.1111/raq.12659
Grube M., Depriest P., Gargas A., Hafellner J. (1995). DNA isolation from lichen ascomata. Mycological Res. 99, 1321–1324. doi: 10.1016/S0953-7562(09)81215-X
Hassan S. A., Sharawy Z. Z., Hemeda S. A., El Nahas A. F., Abbas E. M., Khalil H. S., et al. (2023). Sugarcane bagasse improved growth performance, digestive enzyme activity, microbial dynamics, and mRNA transcripts of immune-, growth, and antioxidant-related genes of Litopenaeus vannamei in a zero-water exchange system. Aquaculture Rep. 33, 101788. doi: 10.1016/j.aqrep.2023.101788
Hernández–Martínez I., González–Resendiz L., Sánchez–García L., Vigueras–Ramírez G., Arroyo–Maya I. J., Morales–Ibarría M. (2023). C-phycocyanin production with high antioxidant activity of a new thermotolerant freshwater Desertifilum tharense UAM-C/S02 strain. Bioresource Technol. 369, 128431. doi: 10.1016/j.biortech.2022.128431
Huelsenbeck J. P., Bollback J. P., Levine A. M. (2002). Inferring the root of a phylogenetic tree. Systematic Biol. 51, 32–43. doi: 10.1080/106351502753475862
Jerez-Cepa I., Ruiz-Jarabo I. (2021). Physiology: An important tool to assess the welfare of aquatic animals. Biology 10, 61. doi: 10.3390/biology10010061
Jian J.-C., Wu Z.-H., Yang S.-P. (2013). Effect of marine red yeast Rhodosporidium paludigenum on antioxidant-related gene expression in Litopenaeus vannamei. Israeli J. Aquaculture-Bamidgeh 65. doi: 10.46989/001c.20665
Karatay S. E., Dönmez G. (2011). Microbial oil production from thermophile cyanobacteria for biodiesel production. Appl. Energy 88, 3632–3635. doi: 10.1016/j.apenergy.2011.04.010
Komárek J. (2005). The modern classification of cyanoprokaryotes [Cyanobacteria. Oceanological Hydrobiological Stud. 34, 5–17.
Li F., Cui X., Mao M., Fu C., Li Z., Liu J., et al. (2024). Molecular characterization and functional identification of the insulin-like peptides receptor gene in the oriental river prawn Macrobrachium nipponense. Aquaculture Rep. 36, 102190. doi: 10.1016/j.aqrep.2024.102190
Maa-Iad K., Keawtawee T., Seel-Audom M. (2023). Effect of cyanobacteria Limnothrix strain lmTK01 on the survival rate of shrimp Litopenaeus vannamei and factors effected on lmTK01 growth. Burapha Sci. J. 28 (2), 976–990.
Mabrouk M. M., Ashour M., Labena A., Zaki M. A. A., Abdelhamid A. F., Gewaily M. S., et al. (2022). Nanoparticles of Arthrospira platensis improves growth, antioxidative and immunological responses of Nile tilapia (Oreochromis niloticus) and its resistance to Aeromonas hydrophila. Aquaculture Res. 53, 125–135. doi: 10.1111/are.15558
Mabrouk M. M., Ashour M., Younis E. M., Abdel-Warith A.-W. A., Bauomi M. A., Toutou M. M., et al. (2024). Arthrospira platensis nanoparticles dietary supplementation improves growth performance, steroid hormone balance, and reproductive productivity of Nile tilapia (Oreochromis niloticus) broodstock. PloS One 19, e0299480. doi: 10.1371/journal.pone.0299480
Magouz F. I., Essa M. A., Matter M., Tageldein Mansour A., Alkafafy M., Ashour M. (2021). Population dynamics, fecundity and fatty acid composition of Oithona nana (Cyclopoida, Copepoda), fed on different diets. Animals 11, 1188. doi: 10.3390/ani11051188
Mansour A. T., Alprol A. E., Ashour M., Ramadan K. M., Alhajji A. H., Abualnaja K. M. (2022a). Do red seaweed nanoparticles enhance bioremediation capacity of toxic dyes from aqueous solution? Gels 8, 310. doi: 10.3390/gels8050310
Mansour A. T., Ashour M., Abbas E. M., Alsaqufi A. S., Kelany M. S., El-Sawy M. A., et al. (2022b). Growth performance, immune-related and antioxidant genes expression, and gut bacterial abundance of pacific white leg shrimp, Litopenaeus vannamei, dietary supplemented with natural astaxanthin. Front. Physiol. 13. doi: 10.3389/fphys.2022.874172
Mansour A. T., Ashour M., Alprol A. E., Alsaqufi A. S. (2022c). Aquatic plants and aquatic animals in the context of sustainability: Cultivation techniques, integration, and blue revolution. Sustainability 14, 3257. doi: 10.3390/su14063257
Mansour A. T., Ashry O. A., Ashour M., Alsaqufi A. S., Ramadan K. M., Sharawy Z. Z. (2022d). The optimization of dietary protein level and carbon sources on biofloc nutritive values, bacterial abundance, and growth performances of whiteleg shrimp (Litopenaeus vannamei) juveniles. Life 12, 888. doi: 10.3390/life12060888
Maulud D., Abdulazeez A. M. (2020). A review on linear regression comprehensive in machine learning. J. Appl. Sci. Technol. Trends 1, 140–147. doi: 10.38094/jastt1457
Mourelle M. L., Gómez C. P., Legido J. L. (2017). The potential use of marine microalgae and cyanobacteria in cosmetics and thalassotherapy. Cosmetics 4, 46. doi: 10.3390/cosmetics4040046
Naiel M. A., Farag M. R., Gewida A. G., Elnakeeb M. A., Amer M. S., Alagawany M. (2021). Using lactic acid bacteria as an immunostimulants in cultured shrimp with special reference to Lactobacillus spp. Aquaculture Int. 29, 219–231. doi: 10.1007/s10499-020-00620-2
Naser M. N., Sarker M. N., Hosain M. E. (2022). Whiteleg shrimp Litopenaeus vannamei: Current status, future prospects and opportunities for Bangladesh aquaculture. Bangladesh J. Zoology 2, 143–184. doi: 10.3329/bjz.v50i2.62051
Osman M. E. H., El-Sheekh M. M., El-Naggar A. H., Gheda S. F. (2010). Effect of two species of cyanobacteria as biofertilizers on some metabolic activities, growth, and yield of pea plant. Biol. fertility soils 46, 861–875. doi: 10.1007/s00374-010-0491-7
Ozorio R. A., Lopes R. G., Goes B. S., Da Silva C. P., Derner R. B., Fracalossi D. M. (2015). Growth and enzymatic profile of the pacific white shrimp fed with Porphyridium cruentum extract. Boletim do Instituto Pesca 41, 123–131.
Pang Y., Zhang X., Yuan J., Zhang X., Xiang J., Li F. (2021). Characterization and expression analysis of insulin growth factor binding proteins (IGFBPs) in Pacific White Shrimp Litopenaeus vannamei. Int. J. Mol. Sci. 22, 1056. doi: 10.3390/ijms22031056
Patwardhan A., Ray S., Roy A. (2014). Molecular markers in phylogenetic studies-a review. J. Phylogenet. Evolutionary Biol. 2, 131. doi: 10.4172/2329-9002.1000131
Pilotto M. R., Milanez S., Moreira R. T., Rosa R. D., Perazzolo L. M. (2019). Potential immunomodulatory and protective effects of the Arthrospira-based dietary supplement on shrimp intestinal immune defenses. Fish shellfish Immunol. 88, 47–52. doi: 10.1016/j.fsi.2019.02.062
Radhakrishnan S., Belal I. E., Seenivasan C., Muralisankar T., Bhavan P. S. (2016). Impact of fishmeal replacement with Arthrospira platensis on growth performance, body composition and digestive enzyme activities of the freshwater prawn, Macrobrachium rosenbergii. Aquaculture Rep. 3, 35–44. doi: 10.1016/j.aqrep.2015.11.005
Rao X., Huang X., Zhou Z., Lin X. (2013). An improvement of the 2ˆ (–delta delta CT) method for quantitative real-time polymerase chain reaction data analysis. Biostatistics Bioinf. biomathematics 3, 71.
Ratti S., Zarantoniello M., Chemello G., Giammarino M., Palermo F. A., Cocci P., et al. (2023). Spirulina-enriched substrate to rear black soldier fly (Hermetia illucens) prepupae as alternative aquafeed ingredient for rainbow trout (Oncorhynchus mykiss) diets: Possible effects on zootechnical performances, gut and liver health status, and fillet quality. Animals 13, 173. doi: 10.3390/ani13010173
Rellán S., Osswald J., Saker M., Gago-Martinez A., Vasconcelos V. (2009). First detection of anatoxin-a in human and animal dietary supplements containing cyanobacteria. Food Chem. Toxicol. 47, 2189–2195. doi: 10.1016/j.fct.2009.06.004
Robert A. A. (2005). Algal culturing techniques: traditional microalgae isolation techniques (USA: Elsevier Academic Press).
Sharawy Z. Z., Ashour M., Labena A., A.s. A., T. M. A., Abbas E. (2022). Effects of dietary Arthrospira platensis nanoparticles on growth performance, feed utilization, and growth-related gene expression of Pacific white shrimp, Litopenaeus vannamei. Aquaculture 551, 737905. doi: 10.1016/j.aquaculture.2022.737905
Suantika G., Muhammad H., Azizah F. F. N., Rachminiwati N., Situmorang M. L., Astuti D. I., et al. (2016). The use of Cyanobacteria Arthrospira platensis and Cladoceran Daphnia magna as complementary protein and lipid sources in transitional diet for Common Carp (Cyprinus carpio L.) Nursery. Natural Resour. 7, 423–433. doi: 10.4236/nr.2016.77037
Swift M. L. (1997). GraphPad prism, data analysis, and scientific graphing. J. Chem. Inf. Comput. Sci. 37, 411–412. doi: 10.1021/ci960402j
Wang H., Hu X., Shao C., Elshobary M., Zhu F., Cui Y., et al. (2023). Optimizing mixotrophic cultivation of oil-rich Tribonema minus using volatile fatty acids and glycerin: a promising approach for pH-controlling and enhancing lipid productivity. J. Cleaner Production 402, 136733. doi: 10.1016/j.jclepro.2023.136733
Wang W., Sun J., Liu C., Xue Z. (2017). Application of immunostimulants in aquaculture: current knowledge and future perspectives. Aquaculture Res. 48, 1–23. doi: 10.1111/are.2017.48.issue-1
Wang J., Wang W., Li R., Li Y., Tian G., Goodman L., et al. (2008). The diploid genome sequence of an Asian individual. Nature 456, 60–65. doi: 10.1038/nature07484
Xu T., Xia Q.-X., Long J., Long X. (2018). A study on multi-domain modeling and simulation of precision high-speed servo numerical control punching press. Proc. Institution Mechanical Engineers Part I: J. Syst. Control Eng. 232, 830–844. doi: 10.1177/0959651818762945
Zaki M. A., Ashour M., Heneash A. M., Mabrouk M. M., Alprol A. E., Khairy H. M., et al. (2021). Potential Applications of Native Cyanobacterium Isolate (Arthrospira platensis NIOF17/003) for Biodiesel Production and Utilization of Its Byproduct in Marine Rotifer (Brachionus plicatilis) Production. Sustainability 13, 1769. doi: 10.3390/su13041769
Zar J. H. (1984). Statistical significance of mutation frequencies, and the power of statistical testing, using the Poisson distribution. Biometrical J. 26, 83–88. doi: 10.1002/bimj.4710260116
Keywords: aquafeed additive, cyanobacteria, Desertifilum tharense NIOF17/006, molecular identification, digestive enzymes, gene expression, immunostimulants, Litopenaeus vannamei
Citation: Ashour M, Al-Souti AS, Mamoon A, Ali FS, Elshobary ME, Mabrouk MM, Mansour AIA, Mansour AT, El-Haroun E and Abdelhamid AF (2025) Cyanobacteria Desertifilum tharense NIOF17/006 as a novel aquafeed additive: effect on the growth, immunity, digestive function, and gene expression of whiteleg shrimp postlarvae. Front. Mar. Sci. 12:1532370. doi: 10.3389/fmars.2025.1532370
Received: 21 November 2024; Accepted: 05 February 2025;
Published: 12 March 2025.
Edited by:
Yafei Duan, South China Sea Fisheries Research Institute, ChinaReviewed by:
Amit Ranjan, Tamil Nadu Fisheries University, IndiaCopyright © 2025 Ashour, Al-Souti, Mamoon, Ali, Elshobary, Mabrouk, Mansour, Mansour, El-Haroun and Abdelhamid. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Ahmed Said Al-Souti, c291dGlAc3F1LmVkdS5vbQ==; Abdallah Tageldein Mansour, YW1hbnNvdXJAa2Z1LmVkdS5zYQ==; Mohamed Ashour, bWljcm9hbGdhZV9lZ3lwdEB5YWhvby5jb20=
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.