
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Immunol. , 13 March 2025
Sec. Cancer Immunity and Immunotherapy
Volume 16 - 2025 | https://doi.org/10.3389/fimmu.2025.1542499
This article is part of the Research Topic Advances in skin immunology View all 6 articles
 Xiaopeng Sun1
Xiaopeng Sun1 Margaret L. Axelrod2
Margaret L. Axelrod2 Paula I. Gonzalez-Ericsson3
Paula I. Gonzalez-Ericsson3 Violeta Sanchez1,3
Violeta Sanchez1,3 Yu Wang4
Yu Wang4 Jonathan L. Curry5
Jonathan L. Curry5 Elizabeth J. Phillips1
Elizabeth J. Phillips1 Yaomin Xu4
Yaomin Xu4 Douglas B. Johnson1
Douglas B. Johnson1 Justin M. Balko1*
Justin M. Balko1*Purpose: Immune checkpoint inhibitors (ICIs) are increasingly used to treat advanced malignancy but can induce immune-related adverse events (irAE). The mechanisms behind these sporadic and sometimes life-threatening irAEs remain largely unexplored. Here, we present a case report and in-depth molecular analysis of an erythema nodosum (EN) like irAE occurring in a melanoma patient with isolated brain metastasis, aiming to explore the potential mechanism of this irAE.
Methods: We performed RNA and T cell receptor (TCR) sequencing on the patient’s resected brain metastasis and biopsy of EN-like irAE. Single cell RNA/TCR sequencing was conducted on the patient’s peripheral blood mononuclear cells (PBMC) at baseline, 3 weeks after ipilimumab and nivolumab combination therapy, during EN toxicity and after resolution.
Results: The site of EN-like irAE showed a distinct accumulation of pro-inflammatory immune cells, accompanied by the upregulation of inflammatory and interferon response signatures. In addition, clonal expansion and activation of irAE-associated CD8 T cells and upregulation of monocyte-specific interferon signatures occurred concurrently with irAE onset.
Conclusion: The unique immune landscape at the EN-like irAE could indicate that this irAE is distinct from anti-tumor immune and analogous non-ICI autoimmune milieus. Our data also suggests that systemic immune activation induced by ICI treatment, as reflected in PBMC, may help monitor the patient’s treatment response and access irAE risk.
Immune checkpoint engagement is a key component in limiting autoimmune inflammation, maintaining fetal tolerance during pregnancy, and preventing the rejection of transplanted organs; however, it is also a common immune suppression mechanism that tumor cells can hijack to avoid immune surveillance. Immune checkpoint inhibitors (ICI), such as anti-PD-1/PD-L1 or anti-CTLA-4, can bind co-inhibitory immune checkpoint receptors and reactivate anti-tumor immunity. Currently, ICI has profoundly changed the treatment in 20 different cancer types, increasing response rate from 10-50% across various solid tumors (1, 2).
ICI can also cause immune-related adverse events (irAEs). Common ICI-induced irAEs include dermatitis, endocrinopathies, colitis, hepatitis, and pneumonitis, which are all thought to arise from aberrant activation of autoreactive T cells (3, 4). The rate of irAEs and severity vary by treatment regimen. From previous clinical experience in melanoma, CTLA-4 inhibition results in a high incidence of dose-dependent toxicities (high-grade toxicities in 38.6% and 57.9% of patients with metastatic melanoma receiving ipilimumab 3 mg/kg or 10 mg/kg, respectively) (5), while only 10-15% of patients receiving PD-1/PD-L1 experienced high-grade toxicity (6). Concurrent ICI use, such as combined anti-CTLA-4+anti-PD-1, also augments the risk of autoimmune toxicities, resulting in almost two-fold increased incidences of high-grade irAEs (7).
The molecular and cellular mechanisms of irAEs remain largely unclear, with a few limited studies on lichenoid and bullous pemphigoid irAEs (8, 9). To gain insights into these mechanisms, we describe a case of erythema nodosum (EN)-like irAE from a patient treated with combination ipilimumab and nivolumab and apply deep molecular analysis of the tumor, the EN-like toxicity, and longitudinal analysis of peripheral blood.
A man in his 40s without a history of known cutaneous melanoma presented with headaches and seizures and was found to have a hemorrhagic right parietal lobe mass. He underwent surgical resection which showed metastatic melanoma, and post-operative radiation therapy. Imaging showed no other intra- or extra-cranial disease. The patient received a combination of ipilimumab (anti-CTLA-4) and nivolumab (anti-PD-1) complicated by hypothyroidism and elevated liver enzymes after his second dose, which was treated by levothyroxine and prednisone, followed by mycophenolate mofetil, respectively. Following the resolution of liver enzymes, he received maintenance therapy with a single agent, nivolumab, without worsening liver function and with no evidence of a new metastatic disease (Figure 1A).
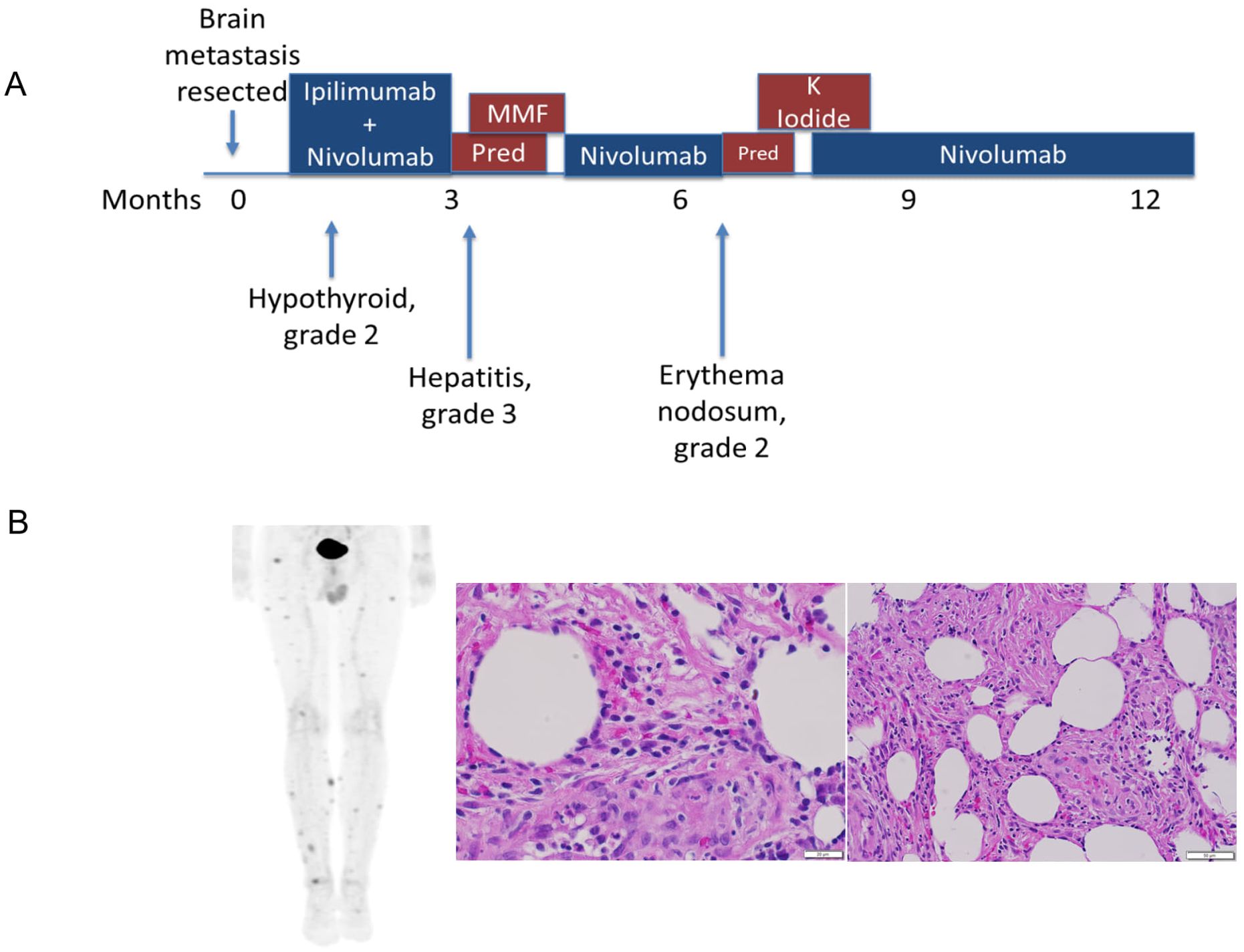
Figure 1. Clinical course of anti-PD-1-induced erythema nodosum. (A) The patient developed Erythema nodosum-like irAE approximated 6 months after metastasis resection and ICI treatment (B) PET-CT imaging demonstrating multiple FDG-avid subcutaneous nodules, which were biopsied and identified as erythema-nodosum-like panniculitis, as demonstrated by increased immune infiltration.
Approximately 4 months after resuming nivolumab, the patient developed an EN-like irAE. He developed painful subcutaneous nodules on his lower extremities and trunk. Although clinical pictures are not available for this patient, the irAE’s clinical presentation was similar to previously reported EN-like irAEs (10, 11). PET-CT showed multifocal FDG avid subcutaneous lesions concerning for metastatic disease. Biopsies of two different lesions both demonstrated adipose tissue with acute and chronic inflammation and panniculitis consistent with an EN-like reaction. IHC showed a mixture of CD4+ and CD8+ T cells, negative CD56, and rare CD20+ B cells (Figure 1B). The condition was effectively managed with potassium iodide treatment. Since EN-like irAE is not life-threatening toxicity and half of the patients experiencing irAEs do not have irAE recurrence on ICI rechallenge (12, 13), the nivolumab therapy was subsequently resumed, and the patient carefully monitored. The patient stopped therapy after approximately two years and has had no additional recurrences of tumor or irAEs approximately 5.5 years after the initial presentation. As EN is a rarely reported irAE (10, 11, 14, 15), we sought to elucidate the pathogenesis by conducting an extensive examination of samples from the patient’s blood, resected brain metastasis, and a tissue biopsy from the EN toxicity site.
We performed bulk RNA sequencing on the patient’s irAE biopsy, resected brain metastasis, and four cases of non-ICI-related skin autoimmune disease samples (three EN and one granulomatous disease [GD]) as non-ICI-induced skin condition comparators.
Compared to all other samples, the EN-like irAE demonstrated enrichment of pro-inflammatory leukocytes RNA signatures, including CD8+ T cells, memory-activated CD4+ T cells, M1 macrophages, and resting NK cells (Figure 2A). Notably, immunosuppressive M2 macrophages, which were abundant in tumor and non-ICI related EN, were nearly undetectable in the EN-like irAE (Figure 2A). Furthermore, we observed enrichment of immune activation signatures at the toxicity site, evidenced by elevated enrichment scores for Hallmark pathways “inflammatory response”, “interferon response”, and “allograft rejection” when compared to tumor and non-ICI autoimmune skin disorders (Figure 2B). Since the response to anti-PD-1 therapy has been linked to type-II interferon responses (16, 17), these findings suggest that the toxicity site is characterized by an ICI-associated immune activation pattern reminiscent of anti-tumor immune responses.
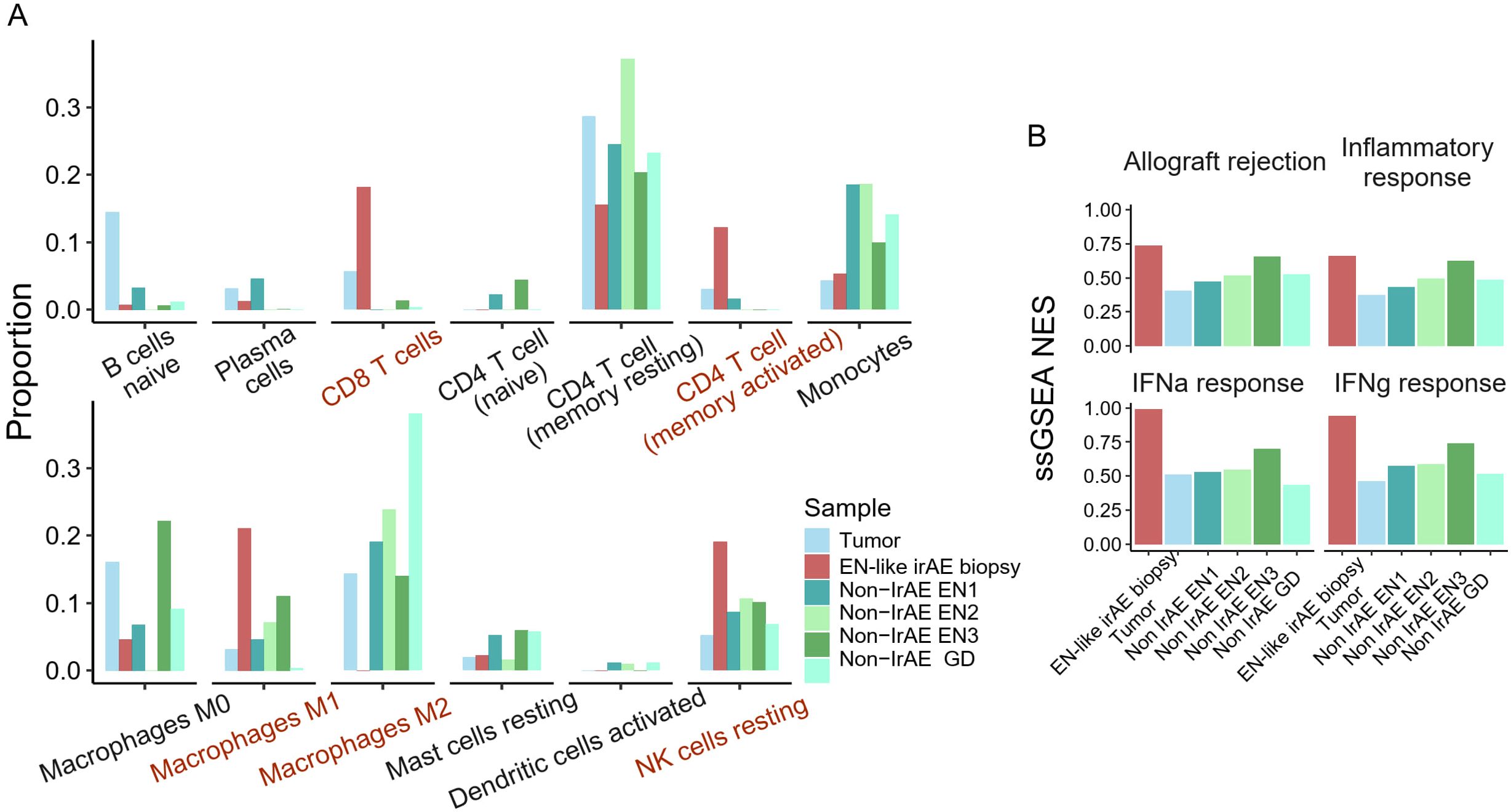
Figure 2. Site of ICI-induced erythema nodosum harbors pro-inflammatory immune cells. (A) CibersortX immune cell deconvolution using LM22 reference of the site of toxicity, tumor, and non-ICI induced skin autoimmune disease. (B) Enrichment score of Hallmark immune-related pathways.
One proposed mechanism for ICI-induced irAE is that the site of toxicity and the tumor share a common antigen(s), leading to T cells indiscriminately attacking healthy tissue upon the loss of negative modulation by immune checkpoints (18). To test whether the EN-like irAE and the patient’s original tumor harbored T cells that shared similar T cell repertoire, which could support this hypothesis, we extracted and compared T cell receptor (TCR) -beta sequences across these sites. There was minimal overlap in TCR clones between these sites (Figure 3A). Given that the brain metastasis was removed nearly six months before the onset of EN toxicity, it remains possible that the T cells had experienced clonal evolution during ICI treatment, resulting in novel TCRs.

Figure 3. Toxicity-associated TCRs were enriched in peripheral blood. (A) Venn diagram depicting overlapping TCR beta clones between the site of toxicity and the patient’s original tumor mass. (B) The percentage of peripheral blood TCRb that overlaps with clones detected on the site of toxicity(red) or tumor(blue). (C) Flow diagram of the top 30 most abundant TCRb clones detected in the longitudinal blood samples, colored by the site where the clone was detected.
Previous research demonstrated that clonal expansion of peripheral T cells is associated with the development of irAE (19). Next, we tested whether toxicity-associated TCRs could be detected in the peripheral blood and if they expanded during ICI-EN. TCR-beta sequences were extracted from longitudinal peripheral blood samples collected post-brain metastasis removal (Baseline), 3 weeks post-ICI (3W post-ICI), during EN toxicity, and after symptom improvement (Improvement). Overall, 11.6% of TCR-beta clonotypes in the baseline blood sample overlapped with the tumor. The proportion of tumor-overlapping clonotypes decreased post-surgical resection and with the initiation of ICI (Figure 3B). During EN toxicity, 42% of the clonotypes detected in the blood overlapped with those at the toxicity site. The proportion of overlapping clones slightly decreased after symptom improvement (Figure 3B). Additionally, we observed clear clonal replacement in the blood, with previously undetected, toxicity-associated clones becoming heavily enriched during the onset of toxicity (Figure 3C). Overall, the TCR data suggests that the systemic clonal dynamics in the blood may reflect the onset and resolution of irAE.
To further characterize the changes in systemic immunity during irAE onset, we performed single-cell RNA/TCR sequencing on the patient’s longitudinal PBMC samples. We observed a decrease in peripheral classical monocyte (cMono) abundance concurrent with an expansion of CD8+ T cells during EN-like irAE (Figure 4A). Differential gene expression analysis revealed a downregulation of myeloid signature genes, such as LYZ, S100A8, and S100A9, during EN-like irAE compared to pre- and post-toxicity timepoints (Figure 4B). In contrast, genes associated with CD8+ T cell cytotoxicity, such as GZMB, NKG7, and PRF1, were upregulated during EN-like irAE (Figure 4B). Additionally, our ssGSEA analysis showed an upregulation of interferon response and inflammatory response-related genes post-ICI treatment, indicating a systemic immune response induced by ICI (Figures 4C, D).
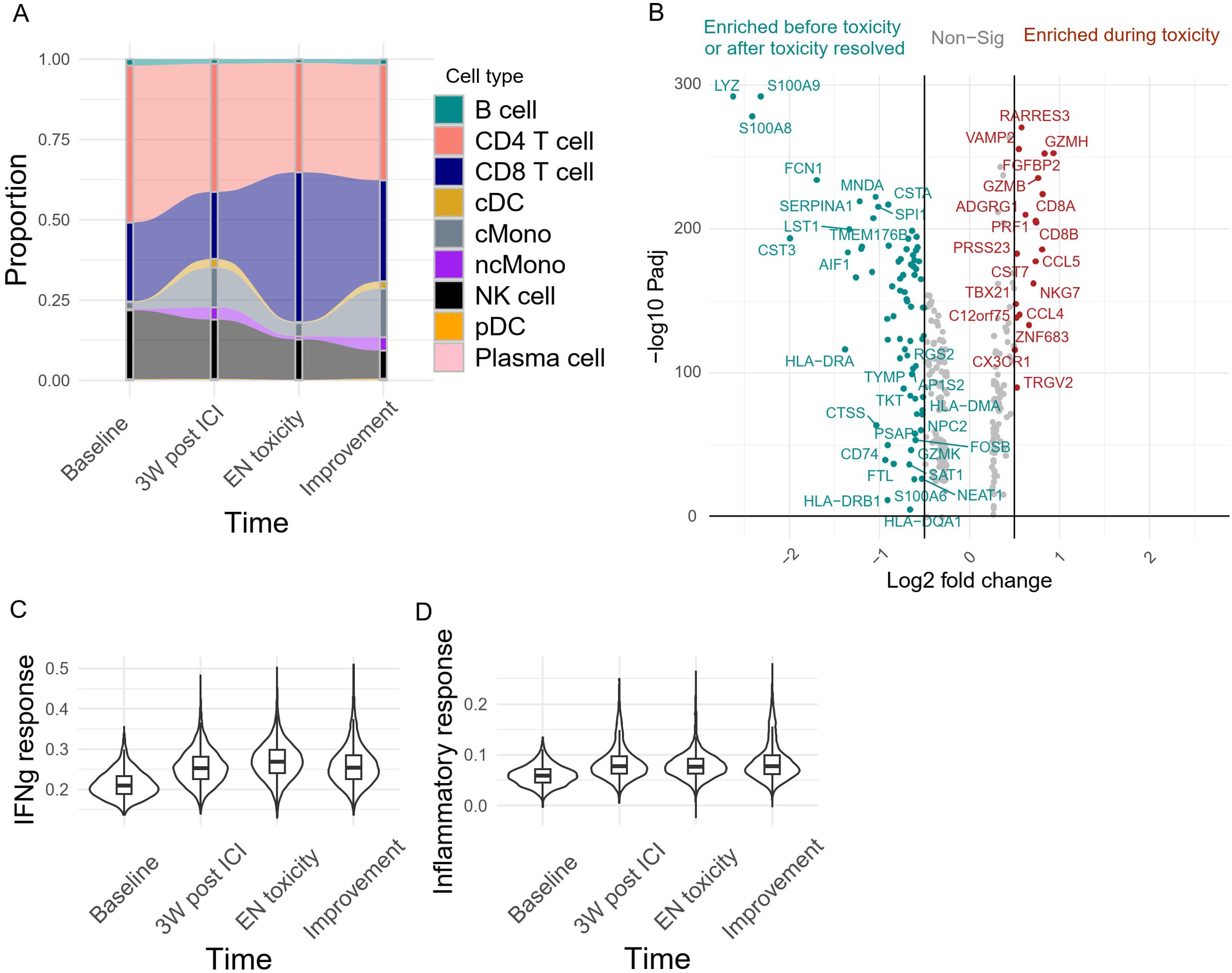
Figure 4. Systemic immune dynamics during irAE onset (A) Peripheral blood immune cell abundance monitored longitudinally. (B) Differential gene expression analysis of the peripheral blood capturing genes upregulated during EN toxicity (right, red) and pre/post-toxicity (left, green). Adjusted P value <0.01 and absolute log2 fold change >0.5. The per-cell enrichment score of (C) Hallmark interferon gamma response, and (D) Inflammatory response pathway across 4 different time points in the peripheral blood.
Sub-clustering of the CD8+ T cells resulted in eight functionally distinct clusters, including naïve/central memory, newly activated, cytotoxic, and effector memory T cells (Figure 5A). There was a dominant population of GZMB+ cytotoxic CD8 T cells, carrying TCRs that were also detected at the site of EN-like irAE (Figure 5B), alongside a smaller subset that shared nearly identical functional markers but was uniquely enriched for specific TRBV and TRAV sequences; these T cells had TCRs that were detected in both the tumor and EN-like irAE biopsy (Figure 5B). Longitudinal analysis of CD8+ T cells revealed that ICI treatment induced a significant expansion of GZMK+ early activated CD8+ T cells (Figure 5C), potentially reflecting early T cell priming. As EN toxicity developed, GZMB+ cytotoxic CD8+ T cells became the dominant phenotype (Figure 5C), indicating peripheral CD8+ T cell activation and potential clonal expansion. Furthermore, during EN toxicity, most clonal T cells in the peripheral blood carrying EN associated TCRs were the activated, GZMB+ cytotoxic subtype (Figure 5D). This suggests that the clonal expansion and activation of CD8+ T cells, particularly the GZMB+ cytotoxic subtype, are closely associated with the development of EN toxicity.
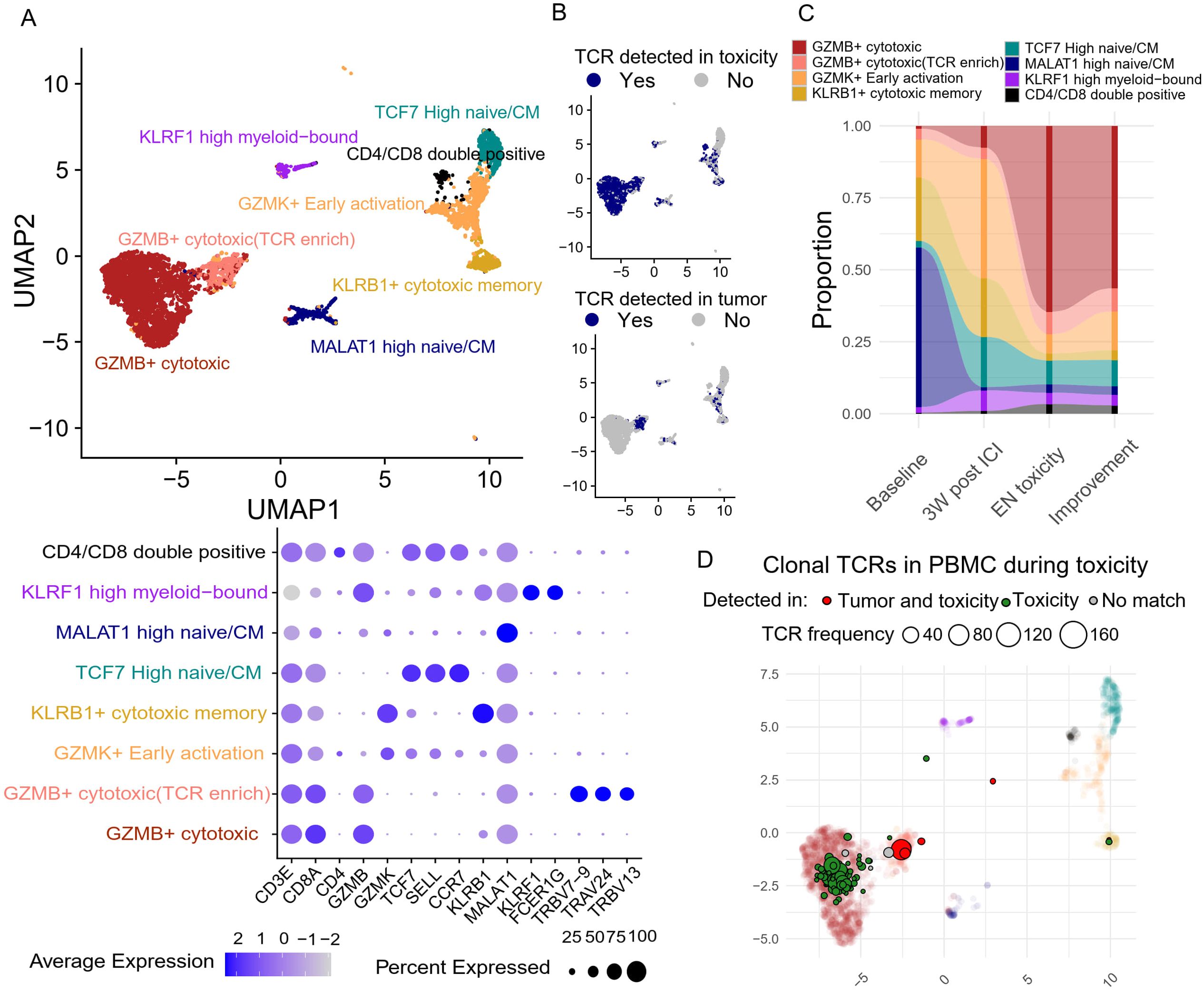
Figure 5. Peripheral blood CD8+ T cell dynamics and their association with tumor and site of toxicity based on TCR clone overlap (A) Unsupervised clustering based on transcriptomic features generates eight distinct CD8 T cell clusters, with their corresponding gene features presented in the dot plot. (B) TCR clones of each T cell are compared against tumor and site of toxicity to identify overlapping clones. (C) The abundance of each CD8 T cell cluster was tracked longitudinally. (D) The center location of each clonal (>3 T cell sharing same TCR) TCR clones on the Umap during toxicity onset. Toxicity-overlapping TCRs are colored green, and tumor+toxicity-overlapping TCRs are colored red.
Since the proportion of cMono decreased after EN toxicity, we decided to further investigate whether the development of EN-like irAEs also coincides with a change in monocyte phenotype. Monocytes were further divided into six clusters based on their transcriptomic features, specifically by the expression of MHC-II transcripts, S100s, and interferon response genes (Figure 6A). Two clusters also expressed T cell-associated genes, which may represent monocyte-T cell doublets or physiological interacting cells. Apart from the increase in T cell-bound monocytes during toxicity, no significant changes were observed in other monocyte subclusters (Figure 6B). However, differential gene expression analysis revealed a reduction in S100 gene expression, which was previously reported to be a marker of immunosuppressive phenotypes (20, 21), and a concurrent increase in interferon response element expression (Figure 6C). Further GSEA analysis supported the observation of increased interferon and cytokine responses during EN toxicity (Figure 6D). These findings suggest a shift in the monocyte landscape towards a more pro-inflammatory state, potentially contributing to the pathogenesis of ICI-induced EN.
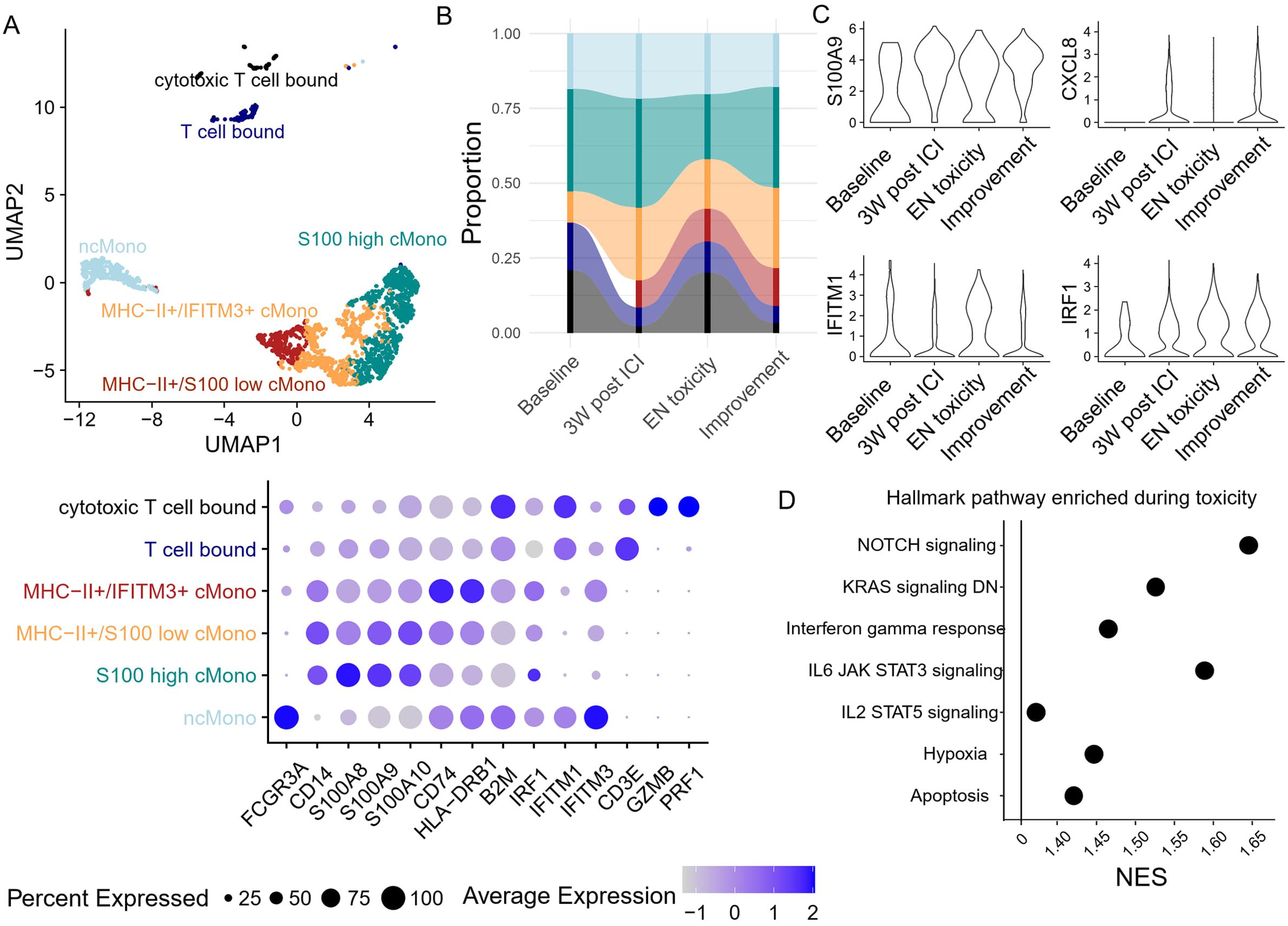
Figure 6. Peripheral blood monocyte dynamics and transcriptomic changes during toxicity onset (A) Unsupervised clustering based on transcriptomic features generates six monocyte clusters, with their corresponding gene features in the dot plot. (B) The abundance of each monocyte cluster was tracked longitudinally. (C) The expression of S100A9, CXCL8, and two interferon response-related genes in monocytes was monitored longitudinally. (D) Gene set enrichment analysis based on the differentially expressed genes between toxicity onset vs. pre/post toxicity. Only pathways with adjusted p-values <0.05 are presented.
The mechanism behind ICI-induced irAEs remains largely case-specific. While theoretically, these toxicities can affect nearly every organ, dermatologic cases are among the most common irAEs for ICI-treated patients (22, 23). Here, we report our observations on the clinical and molecular features of ICI-induced EN in a melanoma patient. A detailed description of the patient’s peripheral immune signature during irAE highlights the role of systemic immunity during this rare cutaneous toxicity.
Several cases of EN-like irAE have been reported in ICI treated melanoma patients, with isolated cases in clear cell carcinoma, esophageal cancer, and renal cell carcinoma (10, 11, 14, 15). irAE onset ranges from 4 weeks to a year after ICI treatment. Among most cases, increased lymphocytes, histiocytes, and neutrophils were observed with no sign of infection, similar to the immune infiltration pattern observed in our case (10). In addition, prior or concomitant hypothyroidism was found in two EN toxicity cases (10), hinting at a potential association between these conditions. Although sparse HLA typing data were available for prior reported cases, upon comparing the patients’ HLA typing, our case shared HLA-B*35 and HLA-DPB1*04, two frequently carried alleles, with a previously reported EN-like irAE patient who had a prior medical history of hypothyroidism (10), suggesting the association between HLA genotype and the toxicity pathogenesis.
The basic cancer immunity cycle suggests that antigens released from tumors during cell turnover and in response to therapies drain to the lymph node, where they are acquired by professional antigen presenting cells and presented to T lymphocytes for priming. Appropriately primed T cells are ‘licensed’ to leave the lymph node and seek out sources of inflammation, whereby they must pass through the systemic peripheral circulation as a conduit (24). Similarly, irAE induced by bystander effects from activated T-cells or T cells targeting healthy organ-tumor overlapping antigens could use a similar adaptive immune response cycle (19, 25). Previous studies have shown that peripheral activated CD4 memory T cell abundance and TCR diversity are strongly associated with irAE onset (25). Using longitudinal peripheral blood, we showed that expansion of cytotoxic CD8 T cells and increased monocytic interferon response were also strongly associated with EN toxicity onset. Together, those data suggest that systemic immune activation induced by ICI treatment may also reflect the risk of irAE, providing a potential method to monitor the patient’s treatment response and irAE risk assessment. However, these results also suggest that differential mechanisms (for example dominated by CD4 or CD8 T cell responses) may exist among irAEs.
One popular mechanistic explanation of irAE is that shared antigens between the tumor and the affected organ may cause a break in self-tolerance during anti-tumor immune responses (18, 26). In this unique case, however, the TCRs detected were not shared between the site of irAE and the original brain metastasis that was resected 6 months prior. It is still possible that the TCRs we detected in the site of toxicity may target potential micrometastases that we were not able to detect and/or sample. It is important to note that the development of irAEs has been correlated with anti-tumor immunity and response to ICI (27–29). The T cell activation signatures observed in the peripheral blood support the idea that ICI could induce long-term constant immune surveillance against cancer, even when the patient appears to be disease free.
Previous analysis demonstrated that clonal T cell activation in tumors and peripheral blood is associated with better ICI treatment response in melanoma. As demonstrated by our data and other studies (19, 25, 30, 31), systemic T cell expansion is also associated with the development of irAE. Currently, one of the challenges of irAE management is early detection and biomarker development. Using peripheral biomarkers such as T cell clonality and T cell activation, it may be possible to detect patients experiencing systemic immune responses and enhance patient monitoring to mitigate potential severe irAEs. Nonetheless, further research is needed to determine whether such markers can be used to monitor patients for development of potential irAEs.
In conclusion, we present a deep molecular analysis of ICI-induced EN. In addition, the observation of systemic inflammation during toxicity onset further strengthens the importance of systemic immunity during ICI and the development of irAEs.
The patient was treated at Vanderbilt University Medical Center and consented to the clinical and biospecimen repository (IRB#100178). Peripheral blood samples were taken at baseline, early on treatment, and at time of toxicity per our protocol. FFPE samples from resected brain metastasis and skin biopsies were obtained from pathology. Samples from patients non-ICI autoimmune disorders were obtained from pathology (IRB# 150754).
Total RNA was isolated from FFPE tumor, ICI-EN and non-ICI skin autoimmune disease biopsy samples using the Promega Maxwell 16 FFPE RNA kits per the manufacturer’s protocol. mRNA enrichment and cDNA library were prepared utilizing the stranded mRNA (polyA-selected) library preparation kit. Sequencing was performed at Paired-End 150 bp on the Illumina NovaSeq 6000 targeting an average of 50M reads per sample. Demultiplexed FASTQ files were next aligned using STAR with a genome index generated from human Hg38. FeatureCount was next applied to create gene count matrix. Subsequent MultiQC was performed to ensure sample homogeneity. Raw count generated by FeatureCount was imported to R. Genes that were expressed in less than 50% of the samples were excluded from the analysis. The filtered gene count matrix was next used to generate DESeq2 objects with corresponding metadata. Raw gene counts were transformed using VST transformation followed by GSVA enrichment analysis which assigns enrichment scores of the Hallmark pathways to each sample. The transformed gene sets were deconvoluted with CIBERSORTx using the LM22 matrix to obtain immune cell composition in each biopsy.
Using the whole RNA extracted from FFPE biopsies, TCRs were sequenced using the TCR Immunoverse all chain assay per the manufacturer’s protocol (Invitae/ArcherDX). Sequencing results were evaluated using Archer Immunoverse analyser. CDR3 sequences and frequency tables were extracted from the manufacturers’ analysis platform. TCR beta sequence was extracted to identify matching clones among tumor biopsy, ICI-EN site, and peripheral blood.
Each sample (targeting 5,000–15,000 cells per sample) was processed for single-cell 5′ RNA and TCR sequencing utilizing the 10x Chromium system. Libraries were prepared following the manufacturer’s protocol. The libraries were sequenced using NovaSeq 6000 with 150 bp paired-end reads. RTA (v.2.4.11; Illumina) was used for base calling, and analysis was completed using 10x Genomics Cell Ranger software. Data were analyzed in R using the filtered h5 gene matrices in the Seurat package (32). In brief, samples were subset to include cells with >200 but <3,000 unique transcripts to exclude probable non-cellular RNA reads and doublets. Cells with >15% of reads coming from mitochondrial transcripts were also excluded as probable dying cells. General immune cell subtypes were imputed using scPred (33). CD8 T cell clusters were generated with FindClusters function after removing all CD8 T cell without TCR information. Monocyte clusters were generated in by setting FindClusters. Detailed cell subtype identify was given based on top10 differentially expressed genes in each cluster.
Differential gene expression analysis was performed across all cells using FindMarkers function between different timepoints. Gene set enrichment analysis was performed to obtain pathway enrichment score by taking the log2 fold change and p-value from differential gene expression results. Besides pair-wise comparison, enrichment scores for each Hallmark pathways is generated for each cell using Escape (34).
We performed four-digit class I and II typing of HLA (with Illumina MiSeq) for the HLA antigens ABC, DR, DQ, and DP on DNA extracted from peripheral blood.
The original contributions presented in the study are included in the article/supplementary files, further inquiries can be directed to the corresponding author/s.
The studies involving humans were approved by Vanderbilt University Medical Center. The studies were conducted in accordance with the local legislation and institutional requirements. The participants provided their written informed consent to participate in this study. Written informed consent was obtained from the individual(s) for the publication of any potentially identifiable images or data included in this article.
XS: Conceptualization, Data curation, Formal Analysis, Investigation, Writing – original draft, Writing – review & editing. MA: Conceptualization, Data curation, Formal Analysis, Writing – review & editing. PG-E: Conceptualization, Data curation, Investigation, Writing – review & editing. VS: Data curation, Investigation, Writing – review & editing. YW: Data curation, Formal Analysis, Investigation, Writing – review & editing. JC: Conceptualization, Data curation, Investigation, Resources, Writing – review & editing. EP: Conceptualization, Data curation, Investigation, Resources, Writing – review & editing. YX: Data curation, Formal Analysis, Investigation, Writing – review & editing. DJ: Conceptualization, Formal Analysis, Funding acquisition, Investigation, Project administration, Resources, Supervision, Writing – original draft, Writing – review & editing. JB: Conceptualization, Funding acquisition, Investigation, Project administration, Resources, Supervision, Writing – review & editing.
The author(s) declare that financial support was received for the research and/or publication of this article. R01CA227481 (to DJ and JB), R01HL156021 (to JB and DJ)
JB receives research support from Genentech/Roche and Incyte Corporation, has received advisory board payments from AstraZeneca and Mallinckrodt and is an inventor on patents regarding immunotherapy targets and biomarkers in cancer.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
1. Yarchoan M, Hopkins A, Jaffee EM. Tumor mutational burden and response rate to PD-1 inhibition. N Engl J Med. (2017) 377:2500–1. doi: 10.1056/NEJMc1713444
2. Zhao B, Zhao H, Zhao J. Efficacy of PD-1/PD-L1 blockade monotherapy in clinical trials. Ther Adv Med Oncol. (2020) 12:1758835920937612. doi: 10.1177/1758835920937612
3. Postow MA, Sidlow R, Hellmann MD. Immune-related adverse events associated with immune checkpoint blockade. New Engl J Med. (2018) 378:158–68. doi: 10.1056/NEJMra1703481
4. Johnson DB, Chandra S, Sosman JA. Immune checkpoint inhibitor toxicity in 2018. JAMA. (2018) 320:1702–3. doi: 10.1001/jama.2018.13995
5. Tarhini AA, Lee SJ, Hodi FS, Rao UNM, Cohen GI, Hamid O, et al. Phase III study of adjuvant ipilimumab (3 or 10 mg/kg) versus high-dose interferon alfa-2b for resected high-risk melanoma: north american intergroup E1609. J Clin Oncol. (2020) 38:567–75. doi: 10.1200/JCO.19.01381
6. Weber J, Mandala M, Vecchio MD, Gogas HJ, Arance AM, Cowey CL, et al. Adjuvant nivolumab versus ipilimumab in resected stage III or IV melanoma. New Engl J Med. (2017) 377:1824–35. doi: 10.1056/NEJMoa1709030
7. Wolchok JD, Chiarion-Sileni V, Gonzalez R, Rutkowski P, Grob J-J, Cowey CL, et al. Overall survival with combined nivolumab and ipilimumab in advanced melanoma. N Engl J Med. (2017) 377:1345–56. doi: 10.1056/NEJMoa1709684
8. Curry JL, Reuben A, Szczepaniak-Sloane R, Ning J, Milton DR, Lee CH, et al. Gene expression profiling of lichenoid dermatitis immune-related adverse event from immune checkpoint inhibitors reveals increased CD14+ and CD16+ monocytes driving an innate immune response. J Cutaneous Pathol. (2019) 46:627–36. doi: 10.1111/cup.13454
9. Marques-Piubelli ML, Seervai RNH, Mudaliar KM, Ma W, Milton DR, Wang J, et al. Gene expression profiling and multiplex immunofluorescence analysis of bullous pemphigoid immune-related adverse event reveal upregulation of toll-like receptor 4/complement-induced innate immune response and increased density of T1 T-cells. J Cutaneous Pathol. (2023) 50:661–73. doi: 10.1111/cup.14442
10. Tetzlaff MT, Jazaeri AA, Torres-Cabala CA, Korivi BR, Landon GA, Nagarajan P, et al. Erythema nodosum-like panniculitis mimicking disease recurrence: A novel toxicity from immune checkpoint blockade therapy—Report of 2 patients. J Cutaneous Pathol. (2017) 44:1080–6. doi: 10.1111/cup.2017.44.issue-12
11. Pach J, Moody K, Ring N, Panse G, Zhang M, Deverapalli S, et al. Erythema nodosum-like panniculitis associated with immune checkpoint inhibitor therapy: Two cases reporting a rare cutaneous adverse event. JAAD Case Rep. (2021) 13:118. doi: 10.1016/j.jdcr.2021.05.002
12. Dolladille C, Ederhy S, Sassier M, Cautela J, Thuny F, Cohen AA, et al. Immune checkpoint inhibitor rechallenge after immune-related adverse events in patients with cancer. JAMA Oncol. (2020) 6:865–71. doi: 10.1001/jamaoncol.2020.0726
13. Pollack MH, Betof A, Dearden H, Rapazzo K, Valentine I, Brohl AS, et al. Safety of resuming anti-PD-1 in patients with immune-related adverse events (irAEs) during combined anti-CTLA-4 and anti-PD1 in metastatic melanoma. Ann Oncol. (2018) 29:250–5. doi: 10.1093/annonc/mdx642
14. Choi ME, Lee KH, Won CH, Chang SE, Lee MW, Choi JH, et al. A case of erythema nodosum-like panniculitis induced by nivolumab in a patient with oesophageal cancer. Australas J Dermatol. (2019) 60:154–6. doi: 10.1111/ajd.2019.60.issue-2
15. Seban R-D, Vermersch C, Champion L, Bonsang B, Roger A, Ghidaglia J. Immune-related erythema nodosum mimicking in transit melanoma metastasis on [18F]-FDG PET/CT. Diagnostics. (2021) 11:747. doi: 10.3390/diagnostics11050747
16. Grasso CS, Tsoi J, Onyshchenko M, Abril-Rodriguez G, Ross-Macdonald P, Wind-Rotolo M, et al. Conserved interferon-γ Signaling drives clinical response to immune checkpoint blockade therapy in melanoma. Cancer Cell. (2020) 38:500–515.e3. doi: 10.1016/j.ccell.2020.08.005
17. Benci JL, Johnson LR, Choa R, Xu Y, Qiu J, Zhou Z, et al. Opposing functions of interferon coordinate adaptive and innate immune responses to cancer immune checkpoint blockade. Cell. (2019) 178:933–948.e14. doi: 10.1016/j.cell.2019.07.019
18. Johnson DB, Balko JM, Compton ML, Chalkias S, Gorham J, Xu Y, et al. Fulminant myocarditis with combination immune checkpoint blockade. N Engl J Med. (2016) 375:1749–55. doi: 10.1056/NEJMoa1609214
19. Subudhi SK, Aparicio A, Gao J, Zurita AJ, Araujo JC, Logothetis CJ, et al. Clonal expansion of CD8 T cells in the systemic circulation precedes development of ipilimumab-induced toxicities. Proc Natl Acad Sci. (2016) 113:11919–24. doi: 10.1073/pnas.1611421113
20. Zhao F, Hoechst B, Duffy A, Gamrekelashvili J, Fioravanti S, Manns MP, et al. S100A9 a new marker for monocytic human myeloid-derived suppressor cells. Immunology. (2012) 136:176–83. doi: 10.1111/j.1365-2567.2012.03566.x
21. Hao W, Zhang Y, Dou J, Cui P, Zhu J. S100P as a potential biomarker for immunosuppressive microenvironment in pancreatic cancer: a bioinformatics analysis and. Vitro study. BMC Cancer. (2023) 23:997. doi: 10.1186/s12885-023-11490-1
22. Xu C, Chen Y-P, Du X-J, Liu J-Q, Huang C-L, Chen L, et al. Comparative safety of immune checkpoint inhibitors in cancer: systematic review and network meta-analysis. BMJ. (2018) 363:k4226. doi: 10.1136/bmj.k4226
23. Arnaud-Coffin P, Maillet D, Gan HK, Stelmes J-J, You B, Dalle S, et al. A systematic review of adverse events in randomized trials assessing immune checkpoint inhibitors. Int J Cancer. (2019) 145:639–48. doi: 10.1002/ijc.v145.3
24. Hiam-Galvez KJ, Allen BM, Spitzer MH. Systemic immunity in cancer. Nat Rev Cancer. (2021) 21:345–59. doi: 10.1038/s41568-021-00347-z
25. Lozano AX, Chaudhuri AA, Nene A, Bacchiocchi A, Earland N, Vesely MD, et al. T cell characteristics associated with toxicity to immune checkpoint blockade in patients with melanoma. Nat Med. (2022) 28:353–62. doi: 10.1038/s41591-021-01623-z
26. Berner F, Bomze D, Diem S, Ali OH, Fässler M, Ring S, et al. Association of checkpoint inhibitor–induced toxic effects with shared cancer and tissue antigens in non–small cell lung cancer. JAMA Oncol. (2019) 5:1043–7. doi: 10.1001/jamaoncol.2019.0402
27. Eggermont AMM, Kicinski M, Blank CU, Mandala M, Long GV, Atkinson V, et al. Association between immune-related adverse events and recurrence-free survival among patients with stage III melanoma randomized to receive pembrolizumab or placebo: A secondary analysis of a randomized clinical trial. JAMA Oncol. (2020) 6:519–27. doi: 10.1001/jamaoncol.2019.5570
28. Maher VE, Fernandes LL, Weinstock C, Tang S, Agarwal S, Brave M, et al. Analysis of the association between adverse events and outcome in patients receiving a programmed death protein 1 or programmed death ligand 1 antibody. J Clin Oncol. (2019) 37:2730–7. doi: 10.1200/JCO.19.00318
29. Das S, Johnson DB. Immune-related adverse events and anti-tumor efficacy of immune checkpoint inhibitors. J Immunother Cancer. (2019) 7:306. doi: 10.1186/s40425-019-0805-8
30. Ostmeyer J, Park JY, Itzstein MS, von Hsiehchen D, Fattah F, Gwin M, et al. T-cell tolerant fraction as a predictor of immune-related adverse events. J Immunother Cancer. (2023) 11:e006437. doi: 10.1136/jitc-2022-006437
31. Kim KH, Hur JY, Cho J, Ku BM, Koh J, Koh JY, et al. Immune-related adverse events are clustered into distinct subtypes by T-cell profiling before and early after anti-PD-1 treatment. Oncoimmunology. (2020) 9:1722023. doi: 10.1080/2162402X.2020.1722023
32. Stuart T, Butler A, Hoffman P, Hafemeister C, Papalexi E, Mauck WM, et al. Comprehensive integration of single-cell data. Cell. (2019) 177:1888–1902.e21. doi: 10.1016/j.cell.2019.05.031
33. Alquicira-Hernandez J, Sathe A, Ji HP, Nguyen Q, Powell JE. scPred: accurate supervised method for cell-type classification from single-cell RNA-seq data. Genome Biol. (2019) 20:264. doi: 10.1186/s13059-019-1862-5
Keywords: immunotherapy, melanoma, irAE, erythema nodosum (EN), autoimmunity
Citation: Sun X, Axelrod ML, Gonzalez-Ericsson PI, Sanchez V, Wang Y, Curry JL, Phillips EJ, Xu Y, Johnson DB and Balko JM (2025) Molecular analysis of immune checkpoint inhibitor associated erythema nodosum-like toxicity. Front. Immunol. 16:1542499. doi: 10.3389/fimmu.2025.1542499
Received: 09 December 2024; Accepted: 24 February 2025;
Published: 13 March 2025.
Edited by:
Olga Simionescu, Carol Davila University of Medicine and Pharmacy, RomaniaReviewed by:
Chandra Rudi, Universitas Prima Indonesia, IndonesiaCopyright © 2025 Sun, Axelrod, Gonzalez-Ericsson, Sanchez, Wang, Curry, Phillips, Xu, Johnson and Balko. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Justin M. Balko, anVzdGluLmJhbGtvQHZ1bWMub3Jn
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.