
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Genet. , 26 February 2025
Sec. Genomic Assay Technology
Volume 16 - 2025 | https://doi.org/10.3389/fgene.2025.1548006
This article is part of the Research Topic Integrative Studies on Genomic Variation in Transitional Medicine: Exploring Antithrombotic Gene Expression and Pharmacogenomic Implications View all articles
Leeches are well known for blood-feeding habits and are widely used for medicinal purposes as they secrete various antithrombotic substances. However, some leeches exhibit non-hematophagous habits and their significance for medicinal use is controversial. Here we provide the chromosome-level genomes of two non-hematophagous leeches, Whitmania acranulata and Whitmania laevis, and, in combination with previous results from Whitmania pigra, we compared these genomes with an emphasis on antithrombotic biomolecules. All three species had the same chromosome number of 11. The genome size, repeat site percentage, and number of protein-coding genes of W. laevis (173.87 Mb, 28.28%, 23,818) were similar to those of W. pigra (169.37 Mb, 27.02%, 24,156), whereas these values of W. acranulata (181.72 Mb, 29.55%, 27,069) were higher than those of the other two leeches. W. laevis was a monophyletic clade of W. pigra, whereas W. acranulata had a paraphyletic relationship with W. pigra. The number of antithrombotic genes in W. laevis (N = 76) was similar to that of W. pigra (N = 79), whereas W. acranulata (N = 102) had apparently more such genes. Of the 21 gene families, 9 and 11 were differentially expressed in W. acranulata and W. laevis compared to W. pigra, respectively. The expression profiles of the antithrombotic gene families were more similar between W. acranulata and W. laevis. Although there were several cases of gene loss or pseudogenization, most antithrombotic genes of the three Whitmania species were intact and transcribable. These results provide valuable insights into the evolution of non-hematophagous leeches and development of antithrombotic drugs.
Leeches, which belong to the class Hirudinea, are renowned for their blood-sucking habits and are found all over the world except Antarctica (Sawyer, 1986). Many leeches target the blood of vertebrates and release an array of antithrombotic biomolecules, including anticoagulants, antiplatelet agents, fibrinolysis enhancers, and tissue penetration enhancers, which are essential to overcome the host’s natural hemostatic mechanisms (Sig et al., 2017). Dozens of such biomolecules involving more than 20 protein families (e.g., hirudin, antistasin, guamerin, eglin, bdellin, saratin, destabilase, and hyaluronidase) have been identified in various leech species (Babenko et al., 2020; Kvist et al., 2020; Liu et al., 2023; Liu et al., 2024; Zhao et al., 2024). Some biomolecules have played an important role in medical and pharmaceutical developments. For example, the derivatives/analogues (lepirudin, desirudin, and bivalirudin) of hirudin, which is the most potent natural thrombin inhibitor found to date, have been available for clinical use for decades (Warkentin, 2004). In addition, antistasin, the first Factor Xa inhibitor, has inspired the development of several well-known anticoagulants such as rivaroxaban and otamixaban (Perzborn et al., 2011).
Although sanguivory is the most prominent aspect of leech behavior in certain species, other feeding modes are common in the remaining species (Borda and Siddall, 2004). Of note is the genus Whitmania, which is the sister clade of the genus Hirudo (Phillips and Siddall, 2009; Zhao et al., 2021). At least three species of Whitmania have been found: Whitmania pigra (Whitman, 1884), Whitmania laevis (Baird, 1869), and Whitmania acranulata (Whitman, 1886). W. pigra is a large species of leech, typically 60–130 mm long and 13–20 mm wide. The maximum body length and body width could reach to 250 and 40 mm, respectively (Yang, 1996), with body weight over 50 g. W. pigra are specific predators of snails, although they have retained tooth plates in their jaws, the blood-feeding ability have been lost. W. laevis have a median body size, typically 32–81 mm long and 5–12 mm wide. They have a broader diet, including snails and insect larvae, and interestingly, no tooth plates are detected in W. laevis, suggesting that the blood-sucking trait is more thoroughly degraded in W. laevis than in W. pigra. (Qiao et al., 2013). W. acranulata have a small body size, only 28–67 mm long and 3.5–8 mm wide. Although W. acranulata retain tooth plates in their jaws, they are swallowers and feed on aquatic earthworms and insect larvae (Yang, 1996).
According to the Pharmacopoeia of the People’s Republic of China (PPRC) (Commission, 2020), which is an integral part of China’s drug laws and regulations, three species of leeches (H. nipponia, W. pigra, and W. acranulata) have been considered as legal materials for “Shuizhi” products. For the hematophagous H. nipponia, scientists are convinced of its medicinal value, while for W. pigra and W. acranulata, there has been a long-standing debate as to whether they can serve as the basic source of “Shuizhi” due to their non-blood-sucking habits (He et al., 2021). Some researchers argue that the two Whitmania species are no longer valid “medicinal leeches” because they have completely lost their blood-sucking behavior. As a result, their antithrombotic capabilities were likely lost during their habit change and should therefore be excluded from the PPRC (Song and Zhou, 1997). Conversely, other researchers argue that these two species should be included in the PPRC due to their close relationship with the genus Hirudo (Phillips and Siddall, 2009; Zhao et al., 2021). They believe that most of the antithrombotic proteins remain active. The latter view seems to be more accurate, since the anticoagulant properties (Ding et al., 2016; Zhang et al., 2020) and antiplatelet aggregation capabilities (Li et al., 1997; Wang et al., 2019) have been repeatedly confirmed in the two Whitmania species. Recent studies support that at least one type of hirudin from W. pigra has anticoagulant activity (Müller et al., 2022). It should be noted that, in the Medicinal Fauna of China (Li et al., 2013), another authoritative publication on Chinese traditional medicine, W. laevis is also considered as a legal material for “Shuizhi” products. A previous study showed that this species also possesses some anticoagulant and antiplatelet activates (Li et al., 1997).
With the rapid development of high-throughput sequencing technology, genomes of several medicinal leeches have been described: Hirudo medicinalis (Babenko et al., 2020; Kvist et al., 2020), Hirudinaria manillensis (Guan et al., 2019; Zheng et al., 2022), W. pigra (Tong et al., 2022; Zheng et al., 2022; Liu et al., 2024), and Hirudo nipponia and Hirudo tianjinensis (Zhao et al., 2024). Interestingly, a total of 79 antithrombotic genes were identified from the hematophagous W. pigra, even more than the typical blood-feeding H. manillensis, which had 72 antithrombotic genes (Liu et al., 2023). Combined with RNA-Seq-based gene expression analyses, our recent study showed that the number and expression level of antithrombotic genes of a non-hematophagous leech are not always lower than those of a hematophagous leech (Liu et al., 2024). The unique life history of non-hematophagous leech species provides an excellent opportunity to understand the evolution of antithrombotic-related genes in leeches. Here, we provide a chromosome-scale genome of W. acranulata and W. laevis from which we identified potential antithrombotic genes of the two species. Furthermore, in combination with RNA-Seq data, we calculated the expression levels of the antithrombotic genes. Combined with our previous results from W. pigra (Liu et al., 2024) we aim to systematically compare the compositions and expression of antithrombotic genes among the three non-hematophagous Whitmania leeches.
W. acranulata and W. laevis samples were collected in Yutai County, Shandong, China (E 116°28′55″, N 35°4′17″). After removal of the digestive tracts, total genomic DNA or RNA was isolated from fresh anterior body tissues. Similar as our recent study on H. nipponia and H. tianjinensis (Zhao et al., 2024), one Nanopore library was constructed respectively for W. acranulata and W. laevis and was sequenced using an ONT sequencer (Oxford Nanopore Technologies, Oxford, UK). At the meantime, one Hi-C, one Survey, and three RNA-Seq libraries for W. acranulata and W. laevis were constructed and were sequenced using the Illumina HiSeq 2000 sequencing platform (Illumina Inc., San Diego, CA, United States). Four individuals were used for each species: one for the Nanopore, Hi-C and Survey libraries, and the other three for each of the three RNA-Seq libraries.
The genome was assembled with ONT reads using NextDenovo v2.5.0 (parameters: read_type = ont, genome_size = 200 m, sort_options = -m 50 g -t 30, and also other default settings) (Hu et al., 2024), refined using NextPolish v1.4.0 (Hu et al., 2020) with Survey reads, and were then integrated with Hi-C reads using YaHS v1.1a (Zhou et al., 2023). The mitochondrial genome was assembled with Survey reads using GetOrganelle v1.7.7.0 with “-F animal_mt” and other default settings (Jin et al., 2020). BUSCO v4.1.4 (with the eukaryota_odb10 database) (Seppey et al., 2019) and Merqury v1.3 (Rhie et al., 2020) were used to assess the thoroughness and quality of the genomes, respectively. RepeatMasker v4.1.2-pl (Flynn et al., 2020) were used to search for repetitive sequences, and the repeat-masked scaffolds were used for gene prediction using a BRAKER-plus strategy (Liu et al., 2023).
Combined with previously reported genome of W. pigra (Liu et al., 2024), H. medicinalis (Kvist et al., 2020), H. manillensis (Liu et al., 2023), H. nipponia and H. tianjinensis (Zhao et al., 2024), and Dinobdella ferox (Gao et al., 2023), we tested the phylogenetic relationship among the three Whitmania species. The orthologs of the coding sequences (CDS) of all species were detected using OrthoFinder v2.3.11 with “-S diamond” and other default settings (Emms and Kelly, 2019), aligned using MACSE (Ranwez et al., 2011), and were concatenated using Seqkit v0.10.2 (Shen et al., 2016). IQ-TREE (Nguyen et al., 2015) was used to reconstruct trees using 1,000 bootstrap replicates (Dinobdella ferox was set as the outgroup).
We also used NGenomeSyn v1.41 (He et al., 2023) to visualize microcolinearity between chromosomes of the three Whitmania genomes. First, pairwise comparisons were performed using the GetTwoGenomeSyn.pl scripts with parameters set as “-MinLenA 5,000,000, -MinLenB 5,000,000, -MinAlnLen 5000, -MappingBin minimap2”. The results of the pairwise comparisons were then integrated using the main program NGenomeSyn.
The relative expression levels of antithrombotic genes of the three Whitmania species were estimated using the RNA-Seq reads. The total predicted CDS of each species were used as mapping references. The TPM values of antithrombotic genes were calculated using salmon v1.0.0 with a kmer size of 31 (Patro et al., 2017). Finally, the total TPM of all members within each antithrombotic gene family was calculated for interspecies comparison.
The TPM of each gene family of W. acranulata vs. W. pigra and W. laevis vs. W. pigra were compared using the non-parametric Mann-Whitney U test in SPSS v25.0 (IBM Corp., Armonk, NY, United States). In addition, to show the overall similarity of antithrombotic gene expression patterns among the three Whitmania species, the gene expression spectra of all samples were clustered using hierarchical cluster analysis (the TPM values of each sample were prior rescaled to the interval [0, 1]) in the R package pheatmap v1.0.12. The use of the above software or programs, if not specified, means that they have been run with default settings and parameters.
After assembly and polishing, 45 and 88 contigs with N50 of 9.40 and 11.76 Mb were obtained for W. acranulata and W. laevis, respectively. Coincidentally, 17 scaffolds with a total length of 181.72 and 173.87 Mb were obtained for both the W. acranulata and W. laevis genome assemblies after consolidation by Hi-C reads. For both species, the length of the first 11 longest scaffolds and the remaining 6 scaffolds showed a highly discontinuous distribution (Figures 1A, B). The well-resolved Hi-C maps further indicated that the 11 longest scaffolds are predicted chromosomes (Figures 1C, D), resulting in a chromosomal anchoring rate over 99.5%. In addition, a circular complete mitochondrial genome was obtained for both species (Table 1).

Figure 1. Assembly information of Whitmania acranulata and Whitmania laevis genomes. Scaffold 187 length distribution of (A) W. acranulata and (B) W. laevis (the red dots indicate the 11 chromosomes; the green dots indicate the remaining short scaffolds). Hi-C links among the chromosomes of (C) W. acranulata and (D) W. laevis (darker red color indicates higher contact probability; Wacr and Wlae are abbreviations of W. acranulata and W. laevis, respectively; the lengths of the chromosomes Wacr01 ∼ Wacr11 and Wlae01 ∼ Wlae11 decrease sequentially).
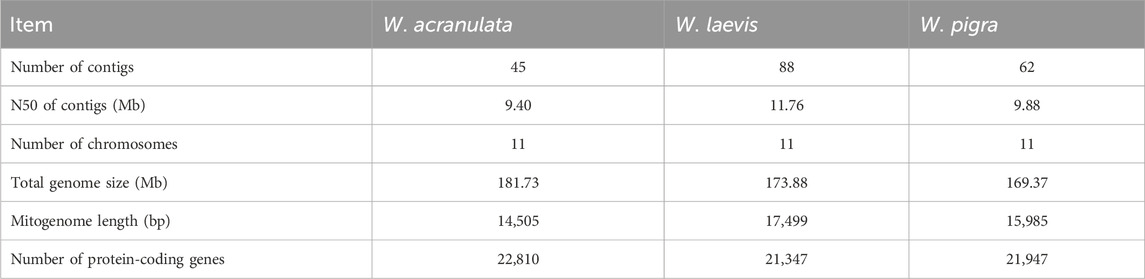
Table 1. Basic information of genomes and protein-coding genes of Whitmania acranulata, Whitmania laevis, Whitmania pigra (Liu et al., 2024).
The genomes of W. acranulata and W. laevis are available in Supplementary Files S1 and S2, respectively. As shown in Table 1, the three Whitmania species had the same number of chromosomes (N = 11). The genome size of W. acranulata were a bit larger than those of W. laevis and W. pigra. In contrast, the mitogenome length of W. acranulata was smaller than those of W. laevis and W. pigra.
The BUSCO completeness analysis showed that, for W. acranulata, 250 of the 255 BUSCOs were captured, while 5 (2.0%) BUSCOs were missed. For W. laevis, 248 BUSCOs were captured, while 7 (2.8%) BUSCOs were missed. We also used Merqury (Rhie et al., 2020) to assess the quality of our genome assembly and obtained quality value scores of 36.54 and 39.92 for the W. acranulata and W. laevis genomes, respectively.
We searched the genomes for repeat sequences using RepeatMasker (Flynn et al., 2020). A total of 29.55% sites were identified as repeats in the W. acranulata genome: Retroelements had the highest percentage, followed by Unclassified repeats and DNA transposons. As to W. laevis, a total of 28.28% of the sites were identified as repeats: Unclassified repeats had the highest percentage, followed by Unclassified repeats and DNA transposons. The total percentage of repeat sites in W. acranulata was higher than those in W. laevis and W. pigra (Liu et al., 2024), mainly due to the apparently higher percentage of Retroelements in W. acranulata (Table 2).
A total of 4,493 orthologs were identified among the eight leech species. Phylogenetic analysis based on the concatenated sequences (5,335,974 bp in length) produced a highly confident consensus tree, i.e., all of the nodes had a bootstrap percentage of 100%. Unexpectedly, the three Whitmania species did not form into a monophyletic group. Instead, W. pigra and W. laevis formed into a subclade that was more closely related to a Hirudo subclade (H. nipponia and H. tianjinensis) than to W. acranulata (Figure 2A).
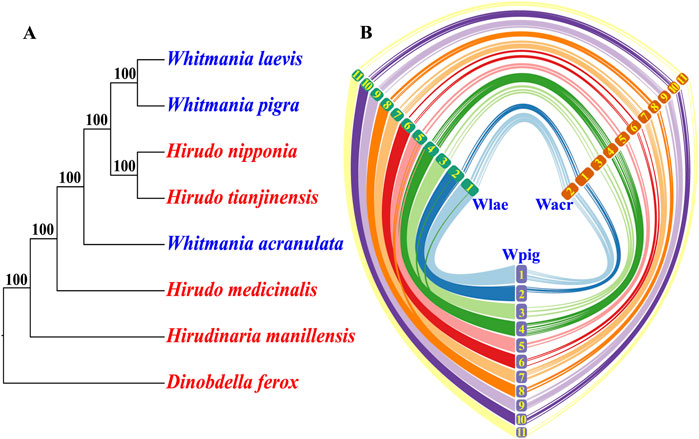
Figure 2. Genomic relationships among the three Whitmania leeches. (A) Phylogenomics of the three Whitmania species as well as other related leech taxa (Dinobdella ferox has been set as the 12 outgroup); (B) Chromosome syntenic relationships among the three Whitmania species (more curves indicate higher collinearity levels; the numbers indicate the chromosomes of each species; larger numbers represent smaller chromosomes).
Chromosome syntenic analysis showed that the chromosomes of W. pigra and W. laevis were perfectly matched. The chromosome length order is identical and there were massive syntenic segments between the two species. In contrast, there was an inconsistency in chromosome length order between W. acranulata and the other two Whitmania species, i.e., the longest chromosome (Wacr01) of W. acranulata corresponded to the second longest chromosome of W. pigra (Wpig02) and W. laevis (Wlae02). Furthermore, the syntenic segments between the W. acranulata and the other two Whitmania species were obviously fewer than those between W. pigra and W. laevis (Figure 2B).
Based on the BRAKER-plus strategy, 22,810 and 21,347 protein-coding genes were predicted for W. acranulata and W. laevis, respectively. The number of protein-coding genes in W. laevis and W. pigra were similar, but were smaller than those in W. acranulata (Table 1). The GFF files and all predicted CDS (including isoforms) of the two species (Supplementary Files S3–S6) were available as Supplementary Material. A total of 102 and 76 antithrombotic genes were identified from the genomes of W. acranulata and W. laevis, respectively. According to our previous studies (Liu et al., 2023; Liu et al., 2024), these genes could be classified into different gene families (Table 3; for more information on the functions of these protein families, see also the two previous papers and the references therein).
In contrast to W. pigra, which had two thrombin inhibitor families (hirudin and progranulin), only the progranulin was identified in W. acranulata and W. laevis, while the hirudin family was completely lost in the latter two species. Three factor Xa inhibitor families (antistasin, lefaxin, and therostasin) were recovered in both W. acranulata and W. laevis. As in W. pigra, most of the remaining coagulation inhibitors (hirustasin, hirustasin-like, guamerin, piguamerin, bdellastasin, eglin, bdellin, LDTI, and HMEI) were recovered in both W. acranulata and W. laevis. Of the three antiplatelet families (saratin, apyrase, and lumbrokinase), two and three families were recovered in W. acranulata and W. laevis, respectively. Finally, three families of fibrinolysis enhancers (lumbrokinase, destabilase, GGT, and LCI) and one family of tissue penetration enhancers (hyaluronidase) were also recovered in both W. acranulata and W. laevis (Table 3).
There was massive genetic variation among the members in each of the antithrombotic gene/protein families (Supplementary Figures S1–S17), including several pseudogenization cases (Table 2). Although hirudin is the first identified and most representative antithrombotic bioactive protein in leeches (Krezel et al., 1994), their coding genes were completely lost from the genomes of W. acranulata and W. laevis (Supplementary Figure S1), making it impossible to show their interspecific genetic variation. One or more members of the gene families therostasin (therostasin_Wlae, Supplementary Figure S5, Figure 3), piguamerin (piguamerin_Wpig, Supplementary Figure S6), HMEI (HMEI_Wacr25, HMEI_Wacr28, HMEI_Wpig07; Supplementary Figure S10), saratin (saratin_Wlae4 and saratin_Wpig03, Supplementary Figure S11), and GGT (GGT_Wpig2, Supplementary Figure S15) were pseudogenized.

Figure 3. Alignment of therostasins and their coding sequences (red frame, early terminal codon of W. laevis therostasin).
There were many proteins whose reactive residues were predicted by functional verification tests. The arginine from the CREHC segment was predicted to be the catalytic residue in the archetypal antistasin (Dunwiddie et al., 1989), but of the eight proteins from the three Whitmania species, only one (antistasin_Wpig2) was conserved at this site (Supplementary Figure S3). The predicted catalytic residue arginine (CRIYC) in the archetypal therostasin (Chopin et al., 2000) was replaced by phenylalanine in all therostasins from the three Whitmania species (Supplementary Figure S5). The catalytic arginine (CRIRC) of the archetypal hirustasin (Mittl et al., 1997) was conserved in the hirustasins of W. pigra, but not in those of the other species (Supplementary Figure S6). The catalytic arginine (CRKYC) of the archetypal piguamerin (Kim et al., 2001) was conserved in all piguamerins except piguamerin_Wlae (Supplementary Figure S6). The catalytic methionine (CMIFC) of the archetypal guamerin (Jung et al., 1995) and the catalytic arginine (CKVKC) of the archetypal bdellastasin (Moser et al., 1998) were conserved in all guamerins and bdellastasins from the three Whitmania species (Supplementary Figure S6).
The reactive residues leucine and asparagine (McPhalen et al., 1985) in the archetypal eglin (TLDLR) were conserved in 7 of 15 eglins from the three Whitmania species (Supplementary Figure S7). The reactive residue lysine (CTKEL) in the archetypal bdellin (Kim et al., 2001) was conserved in three of five bdellins from the Whitmania species (Supplementary Figure S8). The reactive residues lysine and isoleucine (PKILK) of the archetypal LDTI (Sommerhoff et al., 1994) were conserved in the LDTIs of W. laevis and W. pigra, but not those of W. acranulata (Supplementary Figure S9). The catalytic histidine (KIHNM) of the archetypal destabilase (Marin et al., 2023) was conserved in all destabilases from the three Whitmania species, except for destabilase_Wlae5 (Supplementary Figure S14). Finally, the catalytic threonine (HGTAH) of the archetypal GGT (Castellano and Merlino, 2012) was conserved in all GGTs from the three Whitmania species (Supplementary Figure S15).
Based on the RNA-Seq data sequenced in this study we calculated the total TPM (transcripts per million) of each antithrombotic gene family of W. acranulata and W. laevis. Combined with previously reported data of W. pigra, we performed pairwise comparisons on the TPM of each gene family among the three species using the non-parametric Mann-Whitney U test. Of the 21 gene families, 9 were significantly differentiated between W. acranulata and W. pigra, and 11 were significantly differentiated between W. laevis and W. pigra (Table 4). The gene family antistasin of both W. acranulata and W. laevis had a lower express level than W. pigra. In contrast, the gene families guamerin, piguamerin, eglin, and LDTI of W. acranulata and W. laevis had higher expression levels than W. pigra. Hierarchical cluster analysis showed that the samples were grouped into three clusters, each corresponding to one of the three Whitmania species. The W. acranulata and W. laevis samples had closer expression patterns, while the expression patterns of the W. pigra samples were less similar to the other two species (Figure 4).
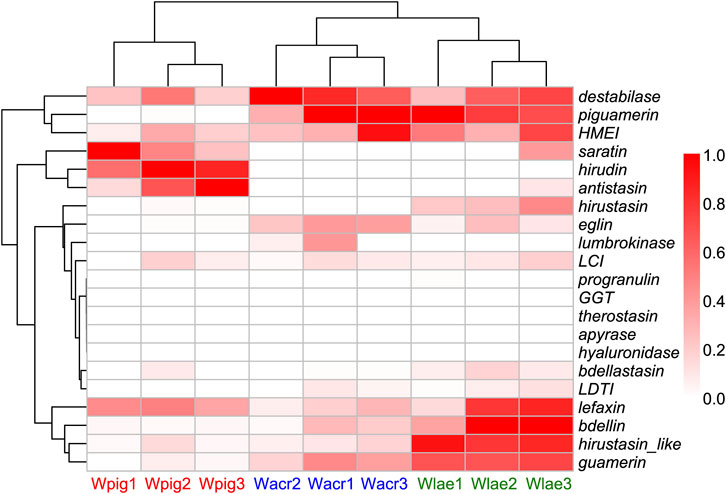
Figure 4. Heatmap of the expression level of antithrombotic genes of the three Whitmania species The TPM (transcripts per million) values of each sample was rescaled to the interval [0, 1]; darker red color indicates higher expression level. W. acranulata: Wacr1 ∼ 3, SRA No. SRR27841062 ∼ SRR27841060; W. laevis: SRA No. Wlae1 ∼ 3, SRR26541749 ∼ SRR26541747; W. pigra: Wpig1 ∼ 3, SRA No. SRR26541745 ∼ SRR26541743).
A high-quality genome is essential for molecular evolutionary analyses of organisms. Recent high-throughput sequencing and assembly technologies make it easier to obtain a chromosome-scale genome. Here, we used the third- and the next-generation sequencing methods to obtain chromosome-scale genomes of W. acranulata and W. laevis. For both species the predicted chromosomes constitute over 99.5% of the total scaffold length, indicating that the genome assemblies have high sequence continuity. The BUSCO analyses showed that over 97% of the BUSCOs were captured in the two genomes, indicating their high completeness. The Merqury analyses on the two genomes yielded quality scores above 36, also higher than the previous study on other leech species (Zheng et al., 2022). In addition, we assembled a complete circular mitochondrial genome for each of the two leeches. As a result, based on the above parameters, we have for the first time provided high quality and nearly complete genomes of the two non-hematophagous leeches.
By comparing the genome of W. pigra obtained in our previous studies (Liu et al., 2024), we showed the similarities and differences among the three non-hematophagous Whitmania genomes. Although the three species had the same chromosome numbers (N = 11), there were several substantial differences on the genomic characteristics between W. acranulata and the other two species. First, the genome size, repeat site percentage, and protein-coding gene number of W. acranulata were larger than those of W. laevis and W. pigra. Second, the chromosome syntenic analysis showed a well genome collinearity between W. laevis and W. pigra, however, the collinearity between W. acranulata and the other two Whitmania species was apparently weakened. Third, the phylogenetic analysis showed that the three Whitmania species formed into a paraphyletic group, i.e., H. nipponia and H. tianjinensis, rather than W. acranulata, became a sister clade of W. pigra and W. laevis. A previous study has found that the members of family Haemopidae (Haemopis spp. and W. laevis) were found not to be monophyletic (Phillips and Siddall, 2009). Interestingly, the present study further demonstrated the complexity of relationships among members of the Haemopidae family: even members of with the single Whitmania genus could be non-monophyletic.
The composition of antithrombotic genes was also different among the three Whitmania species. W. acranulata had 102 antithrombotic genes, much more than W. laevis (N = 76) and W. pigra (N = 79). Our previous studies found that the four hematophagous leeches H. manillensis H. nipponia, H. tianjinensis and H. medicinalis had 72, 86, 83 and 74 antithrombotic proteins, respectively (Liu et al., 2023; Zhao et al., 2024). These results showed that there is no significant difference between non-hematophagous and hematophagous leeches, at least in the number of antithrombotic genes. Although the molecular biology of non-hematophagous leeches has received much less attention than that of hematophagous leeches, at least three proteins from W. pigra, i.e., lumbrokinase (inferred from sequence comparison) (Jiang et al., 2020), hirudin (Müller et al., 2022) and destabilase (Cheng et al., 2025), have been shown to have antithrombotic activity, suggesting that non-haemophagous leeches also have great potential for medicinal use. There were 35 HMEIs in W. acranulata, also much more than in W. laevis (N = 21) and W. pigra (N = 20). In the eglin family, 11 were found in W. acranulata, while only five and two were found in W. laevis and W. pigra, respectively. Gene loss events seem irregularly distributed in the three Whitmania species, e.g., seven hirudins were found in W. pigra, but this gene family disappeared completely in W. acranulata and W. laevis. Similarly, 11 and six saratins were identified in W. pigra and W. laevis, respectively, whereas this gene family was completely undetectable in the W. acranulata genome. Furthermore, there were a lot of genetic variations among the members of each gene/protein family, For example, of the 11 eglins found in W. acranulata, no two proteins have the same sequence. These results indicate that the antithrombotic biomolecules in Whitmania leeches are extremely variable, while what causes these species to have such high genetic instability deserves further investigation in the future.
The hierarchical cluster analysis based on the TPM of the antithrombotic families showed that the samples from each species perfectly formed their respective clusters, indicating that the expression profiles of these gene families were apparently different among these Whitmania species. In contrast to the results of the phylogeny and chromosome synteny analyses, the antithrombotic gene expression profile showed that W. laevis was more closely related to W. acranulata than to W. pigra. Looking specifically at each gene family, there were many differences in TPM levels among Whitmania species. For example, of the 21 gene families, 9 and 11 were differentially expressed in W. acranulata and W. laevis compared to W. pigra, respectively. These results suggest that different species have evolved different expression regulation of these genes for different survival strategies. It should be noted that (as kindly suggested by the anonymous reviewers) differences in antithrombotic gene expression between species may not fully reflect differences in their medicinal activity. Firstly, the starting materials for the present study were collected in the wild and many environmental and biotic factors such as season, age or nutritional status of the leeches may influence gene expression. Secondly, it is not the genes, but the respective encoded protein, that determine the final medicinal values. Next, we will use proteomics and recombinant protein synthesis technology to assess the protein abundances and activities of these differentially expressed genes to further measure the contribution of these genes to the medicinal value of the three non-haemophagous leeches.
Leeches have been used as a medical and pharmaceutical resource for many centuries (Michalsen et al., 2007). Although Whitmania leeches cannot be used for leech therapy due to their non-hematophagous nature, these species still have pharmaceutical value in the treatment of thrombosis diseases (Commission, 2020; Yu et al., 2022). W. pigra, which has the largest body size, the most abundant resources, and is relatively easy to cultivate, has become the primary material for traditional Chinese medicine “Shuizhi” (Liu and Yang, 2014). In contrast, W. acranulata and W. laevis were less concerned due to the scarcity of resources. It has been repeatedly confirmed that, all the three Whitmania species mentioned in this study have anticoagulant and/or antiplatelet activities (Ou et al., 1996; Li et al., 1997; Guan et al., 2012; Zhang et al., 2012; Li et al., 2014; Ding et al., 2016; Cheng, 2018; Wang et al., 2019; Zhang et al., 2020; Zhong et al., 2020). However, probably due to the complexity of the composition and expression of the different antithrombotic genes, there were inconsistencies in different studies on the antithrombotic activities of the Whitmania species. For example, a previous study showed that W. pigra and W. acranulata had similar antithrombin activities (Wang et al., 2013), while some other studies indicated that the antithrombin activities of W. pigra were higher than W. acranulata (Cheng, 2018), or conversely (Zhang et al., 2012; Zhang et al., 2020). The large number of antithrombotic genes identified in this study, and the availability of complete CDSs for most of these genes (except for pseudogenes), will provide opportunities to test their function by producing recombinant proteins (Gupta et al., 2016) and performing in vitro (Liu et al., 2023) and in vivo (Tang et al., 2018) experiments.
Hirudin is the most potent thrombin-specific inhibitor identified to date and is a representative pharmacologically active substance in leeches (Markwardt, 2002; Chen et al., 2021). In the PPRC, antithrombin activity is the only standard for determining the quality of “Shuizhi” (Commission, 2020). It was not surprising that the extracts of W. pigra samples had antithrombin activities, as there were seven hirudins, of which at least one was shown to be functionally active (Müller et al., 2022). Although no hirudin was found in their genomes, W. acranulata (Li et al., 1997; Wang et al., 2013) and W. laevis (Li et al., 1997) were reported to have antithrombin activities, probably due to the antithrombin activity of other proteins such as the granulins (Hong and Kang, 1999). The current version of the PPRC listed W. pigra and W. acranulata, but not W. laevis, as a legal drug material. However, since there were still dozens of antithrombotic genes in W. laevis, and most importantly, there were 11 genes with higher expression levels than in W. pigra, we suggest that this species may also have application values for antithrombotic drug development, although further studies are still needed.
In summary, we provide two nearly complete high-quality genomes of W. acranulata and W. laevis. The genome size, repeat site percentage, and number of protein-coding genes of W. laevis were similar to those of W. pigra, whereas these indices of W. acranulata were higher than those of the other two leeches. Both the compositions and the expression profiles of antithrombotic genes were apparently different among the three leeches.
The data presented in the study are deposited in the figshare repository, accession: https://doi.org/10.6084/m9.figshare.25757583. The raw data from our genome project was deposited in the SRA (Sequence Read Archive) database of National Center for Biotechnology Information with BioProject ID PRJNA1072149 and PRJNA1032729.
The manuscript presents research on animals that do not require ethical approval for their study.
FZ: Writing–original draft, Writing–review and editing. ZH: Funding acquisition, Writing–review and editing. LT: Methodology, Writing–review and editing. WZ: Investigation, Writing–review and editing. ZL: Resources, Writing–review and editing. GL: Conceptualization, Investigation, Supervision, Writing–original draft, Writing–review and editing.
The author(s) declare that financial support was received for the research, authorship, and/or publication of this article. This study was supported by the National Natural Science Foundation of China (No. 82260742), the Double Thousand Plan of Jiangxi Province (No. jxsq2020101050 and jxsq2023201063), the Science and Technology Foundation of Jiangxi Provincial Department of Education (No. GJJ190538) and the Foundation of Key Laboratory of Jiangxi Province for Functional Biology and Pollution Control in Red Soil Regions (2023SSY02051).
Author GL was employed by Shanghai Jizhi Biotechnology Corporation.
The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fgene.2025.1548006/full#supplementary-material
Babenko, V. V., Podgorny, O. V., Manuvera, V. A., Kasianov, A. S., Manolov, A. I., Grafskaia, E. N., et al. (2020). Draft genome sequences of Hirudo medicinalis and salivary transcriptome of three closely related medicinal leeches. BMC Genomics 21, 331. doi:10.1186/s12864-020-6748-0
Borda, E., and Siddall, M. E. (2004). Arhynchobdellida (Annelida: Oligochaeta: Hirudinida): phylogenetic relationships and evolution. Mol. Phylogenetics Evol. 30, 213–225. doi:10.1016/j.ympev.2003.09.002
Castellano, I., and Merlino, A. (2012). γ-Glutamyltranspeptidases: sequence, structure, biochemical properties, and biotechnological applications. Cell. Mol. Life Sci. 69, 3381–3394. doi:10.1007/s00018-012-0988-3
Chen, J., Xie, X., Zhang, H., Li, G., Yin, Y., Cao, X., et al. (2021). Pharmacological activities and mechanisms of hirudin and its derivatives - a review. Front. Pharmacol. 12, 660757. doi:10.3389/fphar.2021.660757
Cheng, B., Liu, F., Gao, T., Zhang, K., and Lu, Y. (2025). A Whitmania pigra destabilase fusion protein and its application. China Patent No CN119080950A. Beijing: China National Intellectual Property Administration.
Cheng, S. (2018). “Studies on polypeptide compositions and anticoagulant mechanism of leeches,” in Master degree. Wuhan, China: Hubei University of Chinese Medicine.
Chopin, V., Salzet, M., Baert, J., Vandenbulcke, F., Sautiére, P. E., Kerckaert, J. P., et al. (2000). Therostasin, a novel clotting factor Xa inhibitor from the rhynchobdellid leech, Theromyzon tessulatum. J. Biol. Chem. 275, 32701–32707. doi:10.1074/jbc.M909217199
Commission, C. P. (2020). Pharmacopoeia of the people’s Republic of China. Beijing: Medicine Science and Technology Press.
Ding, Y., Duan, T., Shan, Y., Wang, X., Yang, H., and Yuan, R. (2016). Comparative studies on anti-thrombin activity and anticoagulant mechanism between Whitmania pigra Whitman and Hirudinaria manillensis. China Pharm. 19, 1621–1624.
Dunwiddie, C., Thornberry, N. A., Bull, H. G., Sardana, M., Friedman, P. A., Jacobs, J. W., et al. (1989). Antistasin, a leech-derived inhibitor of factor Xa. J. Biol. Chem. 264, 16694–16699. doi:10.1016/s0021-9258(19)84761-0
Emms, D. M., and Kelly, S. (2019). OrthoFinder: phylogenetic orthology inference for comparative genomics. Genome Biol. 20, 238. doi:10.1186/s13059-019-1832-y
Flynn, J. M., Hubley, R., Goubert, C., Rosen, J., Clark, A. G., Feschotte, C., et al. (2020). RepeatModeler2 for automated genomic discovery of transposable element families. Proc. Natl. Acad. Sci. U. S. A. 117, 9451–9457. doi:10.1073/pnas.1921046117
Gao, J. W., Sun, J. W., Tong, X. R., Wang, H., Hu, Q. M., Cao, Y. R., et al. (2023). Chromosome-level Dinobdella ferox genome provided a molecular model for its specific parasitism. Parasites and Vectors 16, 322. doi:10.1186/s13071-023-05837-7
Guan, D. L., Yang, J., Liu, Y. K., Li, Y., Mi, D., Ma, L. B., et al. (2019). Draft genome of the Asian buffalo leech Hirudinaria manillensis. Front. Genet. 10, 1321. doi:10.3389/fgene.2019.01321
Guan, S., Yuan, Z., Zhou, Y., Zhang, Y., Ye, X., and Hu, B. (2012). Comparative studies on anti-thrombus and anti-coagulation effects of Hirudo of different species. Chin. J. Hosp. Pharm. 32, 1093–1096. doi:10.13286/j.cnki.chinhosppharmacyj.2012.14.005
Gupta, V., Sengupta, M., Prakash, J., and Tripathy, B. C. (2016). Production of recombinant pharmaceutical proteins. Basic Appl. Aspects Biotechnol. 1, 77–101. doi:10.1007/978-981-10-0875-7_4
He, C., Chen, X., Zhang, X., Wang, L., and Hu, S. (2021). A textual research on Whitmania pigra Whitman as the origin of Hirudo. China Med. Her. 18, 112–115. doi:10.20047/j.issn1673-7210.2021.24.028
He, W., Yang, J., Jing, Y., Xu, L., Yu, K., and Fang, X. (2023). NGenomeSyn: an easy-to-use and flexible tool for publication-ready visualization of syntenic relationships across multiple genomes. Bioinformatics 39, btad121. doi:10.1093/bioinformatics/btad121
Hong, S. J., and Kang, K. W. (1999). Purification of granulin-like polypeptide from the blood-sucking leech, Hirudo nipponia. Protein Expr. Purif. 16, 340–346. doi:10.1006/prep.1999.1077
Hu, J., Fan, J., Sun, Z., and Liu, S. (2020). NextPolish: a fast and efficient genome polishing tool for long-read assembly. Bioinformatics 36, 2253–2255. doi:10.1093/bioinformatics/btz891
Hu, J., Wang, Z., Sun, Z., Hu, B., Ayoola, A. O., Liang, F., et al. (2024). NextDenovo: an efficient error correction and accurate assembly tool for noisy long reads. Genome Biol. 25, 107. doi:10.1186/s13059-024-03252-4
Jiang, Q., Wang, L., Liu, Q., Hu, J., Li, J., and Zhang, Y. (2020). Purification and characterization of a novel fibrinolytic enzyme from Whitmania pigra Whitman. Protein Expr. Purif. 174, 105680. doi:10.1016/j.pep.2020.105680
Jin, J. J., Yu, W. B., Yang, J. B., Song, Y., Depamphilis, C. W., Yi, T. S., et al. (2020). GetOrganelle: a fast and versatile toolkit for accurate de novo assembly of organelle genomes. Genome Biol. 21, 241. doi:10.1186/s13059-020-02154-5
Jung, H. I., Kim, S. I., Ha, K. S., Joe, C. O., and Kang, K. W. (1995). Isolation and characterization of guamerin, a new human leukocyte elastase inhibitor from Hirudo nipponia. J. Biol. Chem. 270, 13879–13884. doi:10.1074/jbc.270.23.13879
Kim, Y. H., Choi, J. G., Lee, G. M., and Kang, K. W. (2001). Domain and genomic sequence analysis of bdellin-KL, a leech-derived trypsin-plasmin inhibitor. J. Biochem. 130, 431–438. doi:10.1093/oxfordjournals.jbchem.a003003
Krezel, A. M., Wagner, G., Seymour-Ulmer, J., and Lazarus, R. A. (1994). Structure of the RGD protein decorsin: conserved motif and distinct function in leech proteins that affect blood clotting. Science 264, 1944–1947. doi:10.1126/science.8009227
Kvist, S., Manzano-Marín, A., De Carle, D., Trontelj, P., and Siddall, M. E. (2020). Draft genome of the European medicinal leech Hirudo medicinalis (Annelida, Clitellata, Hirudiniformes) with emphasis on anticoagulants. Sci. Rep. 10, 9885. doi:10.1038/s41598-020-66749-5
Li, D., Huang, L., and Qu, X. (2013). Medicinal Fauna of China. Fuzhou: Fujian Science and Technology Press.
Li, J., Yu, X., Zhang, J., Zhang, T., Gong, Y., and Jin, G. (2014). Anti-thrombin activity analysis and effects of cryopreservation on anticoagulant activity of four species of leeches. Fish. Sci. 33, 591–593. doi:10.16378/j.cnki.1003-1111.2014.09.012
Li, W., Liao, F., Yin, X., Peng, J., Ou, X., Zhang, Q., et al. (1997). Experimental study on anti-platelet aggregation and anti-coagulation of seven spices of leech. Pharmacol. Clin. Chin. Materia Medica 13, 32–34.
Liu, F., and Yang, D. (2014). Study on artificial breeding model of medical leech in China. World Sci. Technol. Mod. Traditional Chin. Med. Materia Medica 10, 2170–2173. doi:10.11842/wst.2014.10.020
Liu, Z., Zhao, F., Huang, Z., He, B., Liu, K., Shi, F., et al. (2024). A chromosome-level genome assembly of the non-hematophagous leech Whitmania pigra (Whitman 1884): identification and expression analysis of antithrombotic genes. Genes 15, 164. doi:10.3390/genes15020164
Liu, Z., Zhao, F., Huang, Z., Hu, Q., Meng, R., Lin, Y., et al. (2023). Revisiting the Asian buffalo leech (Hirudinaria manillensis) genome: focus on antithrombotic genes and their corresponding proteins. Genes 14, 2068. doi:10.3390/genes14112068
Marin, E., Kornilov, D. A., Bukhdruker, S. S., Aleksenko, V. A., Manuvera, V. A., Zinovev, E. V., et al. (2023). Structural insights into thrombolytic activity of destabilase from medicinal leech. Sci. Rep. 13, 6641. doi:10.1038/s41598-023-32459-x
Markwardt, F. (2002). Hirudin as alternative anticoagulant - a historical review. Seminars Thrombosis Hemostasis 28, 405–414. doi:10.1055/s-2002-35292
Mcphalen, C. A., Schnebli, H. P., and James, M. N. (1985). Crystal and molecular structure of the inhibitor eglin from leeches in complex with subtilisin Carlsberg. FEBS Lett. 188, 55–58. doi:10.1016/0014-5793(85)80873-5
Michalsen, A., Roth, M., and Dobos, G. (2007). Medicinal leech therapy. Stuttgart: Georg Thieme Verlag.
Mittl, P. R., Di Marco, S., Fendrich, G., Pohlig, G., Heim, J., Sommerhoff, C., et al. (1997). A new structural class of serine protease inhibitors revealed by the structure of the hirustasin-kallikrein complex. Structure 5, 253–264. doi:10.1016/s0969-2126(97)00183-4
Moser, M., Auerswald, E., Mentele, R., Eckerskorn, C., Fritz, H., and Fink, E. (1998). Bdellastasin, a serine protease inhibitor of the antistasin family from the medical leech (Hirudo medicinalis) - primary structure, expression in yeast, and characterisation of native and recombinant inhibitor. Eur. J. Biochem. 253, 212–220. doi:10.1046/j.1432-1327.1998.2530212.x
Müller, C., Wang, Z., Hamann, M., Sponholz, D., and Hildebrandt, J. P. (2022). Life without blood: molecular and functional analysis of hirudins and hirudin-like factors of the Asian non-hematophagous leech Whitmania pigra. J. Thrombosis Haemostasis 20, 1808–1817. doi:10.1111/jth.15762
Nguyen, L. T., Schmidt, H. A., Von Haeseler, A., and Minh, B. Q. (2015). IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274. doi:10.1093/molbev/msu300
Ou, X., Zhang, H., Ding, J., Liu, Z., Zhang, L., and Yang, T. (1996). Studies of the anticoagulant activity of 4 kind of leeches. Nat. Prod. Res. Dev. 2, 54–56. doi:10.16333/j.1001-6880.1996.02.012
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A., and Kingsford, C. (2017). Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419. doi:10.1038/nmeth.4197
Perzborn, E., Roehrig, S., Straub, A., Kubitza, D., and Misselwitz, F. (2011). The discovery and development of rivaroxaban, an oral, direct factor Xa inhibitor. Nat. Rev. Drug Discov. 10, 61–75. doi:10.1038/nrd3185
Phillips, A. J., and Siddall, M. E. (2009). Poly-paraphyly of Hirudinidae: many lineages of medicinal leeches. BMC Evol. Biol. 9, 246. doi:10.1186/1471-2148-9-246
Qiao, N., Bai, Y., Wang, G., and Xu, S. (2013). Comparative morphology study on the jaws of three leech species of the genus Whitmania Blanchard. Sichuan J. Zoology 32, 526–529. doi:10.3969/j.issn.1000-7083.2013.04.009
Ranwez, V., Harispe, S., Delsuc, F., and Douzery, E. J. (2011). MACSE: multiple Alignment of Coding SEquences accounting for frameshifts and stop codons. PLoS One 6, e22594. doi:10.1371/journal.pone.0022594
Rhie, A., Walenz, B. P., Koren, S., and Phillippy, A. M. (2020). Merqury: reference-free quality, completeness, and phasing assessment for genome assemblies. Genome Biol. 21, 245. doi:10.1186/s13059-020-02134-9
Seppey, M., Manni, M., and Zdobnov, E. M. (2019). BUSCO: assessing genome assembly and annotation completeness. Methods Mol. Biol. 1962, 227–245. doi:10.1007/978-1-4939-9173-0_14
Shen, W., Le, S., Li, Y., and Hu, F. (2016). SeqKit: a cross-platform and ultrafast toolkit for FASTA/Q file manipulation. PLoS One 11, e0163962. doi:10.1371/journal.pone.0163962
Sig, A. K., Guney, M., Uskudar Guclu, A., and Ozmen, E. (2017). Medicinal leech therapy - an overall perspective. Integr. Med. Res. 6, 337–343. doi:10.1016/j.imr.2017.08.001
Sommerhoff, C. P., Söllner, C., Mentele, R., Piechottka, G. P., Auerswald, E. A., and Fritz, H. (1994). A Kazal-type inhibitor of human mast cell tryptase: isolation from the medical leech Hirudo medicinalis, characterization, and sequence analysis. Biol. Chem. Hoppe-Seyler 375, 685–694. doi:10.1515/bchm3.1994.375.10.685
Song, J., and Zhou, Q. (1997). Discussion on the sources of shuizhi (leeches) in Chinese Pharmacopoeia. Lishizhen Med. Materia Medica Res. 8, 104.
Tang, X., Chen, M., Duan, Z., Mwangi, J., Li, P., and Lai, R. (2018). Isolation and characterization of poecistasin, an anti-thrombotic antistasin-type serine protease inhibitor from leech Poecilobdella manillensis. Toxins 10, 429. doi:10.3390/toxins10110429
Tong, L., Dai, S. X., Kong, D. J., Yang, P. P., Tong, X., Tong, X. R., et al. (2022). The genome of medicinal leech (Whitmania pigra) and comparative genomic study for exploration of bioactive ingredients. BMC Genomics 23, 76. doi:10.1186/s12864-022-08290-5
Wang, X., Gan, Q., Hu, H., Hao, J., Wu, G., Gao, Q., et al. (2019). Measurement of antiplatelet aggregation and potency of Hirudo. Acta Pharm. Sin. 54, 2178–2183. doi:10.16438/j.0513-4870.2019-0147
Wang, Y., Wang, S., Liu, Y., and Xie, Y. (2013). Effects of different processing methods on amino acid contents and anti-thrombin activity in leech. J. Traditional Chin. Med. Univ. Hunan 33, 42–45. doi:10.3969/j.issn.1674-070X.2013.11.012.042.04
Warkentin, T. E. (2004). Bivalent direct thrombin inhibitors: hirudin and bivalirudin. Best. Pract. Res. Clin. Haematol. 17, 105–125. doi:10.1016/j.beha.2004.02.002
Yu, M., Su, S., Zhou, M., Cao, M., and Feng, X. (2022). Identification of suspected species of Hirudo nipponia based on COI gene sequence. J. Southwest Univ. Nat. Sci. Ed. 44, 74–80. doi:10.13718/j.cnki.xdzk.2022.10.009
Zhang, B., Wang, B., Gong, Y., Yu, X., and Lv, J. (2012). Anticoagulant active substances extraction and anti-thrombin activity analysis of several species of leeches. Acta Sci. Nat. Univ. Sunyatseni 51, 92–96.
Zhang, X., Zhuang, Y., Li, Z., Wu, J., Zou, J., Gao, P., et al. (2020). Comparative study on enzymatic hydrolysis processes of different radicals based on anticoagulant activity. Contemp. Chem. Ind. 49, 549–554. doi:10.13840/j.cnki.cn21-1457/tq.2020.03.011
Zhao, F., Huang, Z., He, B., Liu, K., Li, J., Liu, Z., et al. (2024). Comparative genomics of two Asian medicinal leeches Hirudo nipponia and Hirudo tianjinensis: with emphasis on antithrombotic genes and their corresponding proteins. Int. J. Biol. Macromol. 270, 132278. doi:10.1016/j.ijbiomac.2024.132278
Zhao, F., Jiang, K., Su, T., He, B., Wu, Q., Lin, G., et al. (2021). Phylogenomics of common species in Hirudiniformes. J. Jinggangshan Univ. Nat. Sci. 42, 41–47. doi:10.3669/j.issn.1674-8085.2021.04.009
Zheng, J., Wang, X., Feng, T., Rehman, S. U., Yan, X., Shan, H., et al. (2022). Molecular mechanisms underlying hematophagia revealed by comparative analyses of leech genomes. Gigascience 12, giad023. doi:10.1093/gigascience/giad023
Zhong, M., Lei, Y., Li, Y., Tan, H., and Yuan, R. (2020). In vivo anticoagulant activity of different concoctions of Whitmania pigra and Hirudinaria manillensis. J. Chin. Med. Mater. 43, 1351–1353. doi:10.13863/j.issn1001-4454.2020.06.012
Keywords: Whitmania species, chromosome-level genome, antithrombotic gene, genetic variation, gene expression
Citation: Zhao F, Huang Z, Tang L, Zhang W, Liu Z and Lin G (2025) Comparative genomics of three non-hematophagous leeches (Whitmania spp.) with emphasis on antithrombotic biomolecules. Front. Genet. 16:1548006. doi: 10.3389/fgene.2025.1548006
Received: 19 December 2024; Accepted: 28 January 2025;
Published: 26 February 2025.
Edited by:
Damjan Glavač, University of Ljubljana, SloveniaReviewed by:
Christian Mueller, University of Greifswald, GermanyCopyright © 2025 Zhao, Huang, Tang, Zhang, Liu and Lin. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Zichao Liu, YWJjbHpjQGFsaXl1bi5jb20=; Gonghua Lin, bGluZ29uZ2h1YUAxNjMuY29t
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.