
94% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
MINI REVIEW article
Front. Endocrinol., 23 May 2018
Sec. Translational and Clinical Endocrinology
Volume 9 - 2018 | https://doi.org/10.3389/fendo.2018.00250
This article is part of the Research TopicClassic and Pleiotropic Actions of Vitamin DView all 18 articles
The vitamin D3 metabolite 1α,25-dihydroxyvitamin D3 [1,25(OH)2D3] is the exclusive high-affinity ligand of the vitamin D receptor (VDR), a transcription factor with direct effects on gene expression. Transcriptome- and epigenome-wide data obtained in THP-1 human monocytes are the basis of the chromatin model of vitamin D signaling. The model describes, how VDR’s spatio-temporal binding profile provides key insight into the pleiotropic action of vitamin D. The transcription of some 300 primary target genes is significantly modulated through the action of genomic VDR binding sites in concert with the pioneer transcription factor PU.1 and the chromatin organizer CTCF. In parallel, the short-term vitamin D intervention study VitDbol (NCT02063334) was designed, in order to extrapolate insight into vitamin D signaling from in vitro to in vivo. Before and 24 h after a vitamin D3 bolus chromatin and RNA were prepared from peripheral blood mononuclear cells for epigenome- and transcriptome-wide analysis. The study subjects showed a personalized response to vitamin D and could be distinguished into high, mid, and low responders. Comparable principles of vitamin D signaling were identified in vivo and in vitro concerning target gene responses as well as changes in chromatin accessibility. In conclusion, short-term vitamin D supplementation studies represent a new type of safe in vivo investigations demonstrating that vitamin D and its metabolites have direct effects on the human epigenome and modulate the response of the transcriptome in a personalized fashion.
The energy of sunlight-derived UV-B (290–315 nm) is used in human skin to convert the ubiquitous cholesterol precursor 7-dehydrocholesterol into pre-vitamin D3 that isomerizes in a non-enzymatic reaction to the secosteroid vitamin D3 (1). The hydroxylation of vitamin D3 to 25-hydroxyvitamin D3 [25(OH)D3] and then to 1,25(OH)2D3 is necessary to generate the biologically most active metabolite (2). Lifestyle choices, such as preferential indoor activities and coverage by textile outdoors, in combination with climatic and seasonal changes are the main reasons for insufficient UV-B exposure of the majority of today’s human populations (3). In order to avoid deficiency due to this low endogenous production, vitamin D3 needs to be taken up by diet or supplementation with pills.
Well-known physiological roles of vitamin D are (i) control of intestinal absorption of calcium and phosphorus from diet, (ii) renal reabsorption of calcium, and (iii) remodeling of bones (4). However, vitamin D and its receptor, the transcription factor vitamin D receptor (VDR), are involved in far more functions than maintaining calcium homeostasis and bone integrity (5). Highest VDR gene expression is found in metabolic tissues, such as intestine, kidneys, and bone, but low to moderate VDR levels can be observed in more than half of the some 400 tissues and cell types forming the human body (www.proteinatlas.org/ENSG00000111424-VDR/tissue). For example, vitamin D modulates the responsive of both the innate and the adaptive immune system, i.e., it supports the human body in its fight against infections and in parallel prevents autoimmune disorders (6). Accordingly, vitamin D deficiency results not only in problems with bones, which are rickets in children or increased fracture risk for adults, but also weakens vitamin D’s protective role of against diseases like tuberculosis, multiple sclerosis, and type 1 diabetes (7).
The lipophilic structure of vitamin D3 and its metabolites allows the molecules passing through biological membranes. Thus, gene regulation by vitamin D is more direct and less complex than that of peptide hormones, growth factors, cytokines, and other hydrophilic signaling molecules. Since VDR is the only protein binding 1,25(OH)2D3 with high-affinity (8), the physiological effects of vitamin D are largely identical to those of its receptor. Thus, comprehensive insight into vitamin D signaling requires the understanding of VDR’s molecular actions.
Vitamin D receptor belongs the nuclear receptor superfamily, most members of which are activated by small lipophilic molecules (9). Within VDR’s ligand-binding domain some 40 amino acids, which are mostly non-polar, form a ligand-binding pocket that fixes 1,25(OH)2D3 with high specificity and affinity (10). The binding of ligand induces a change in the conformation of the ligand-binding domain, so that VDR’s protein–protein interaction profile alters from that of a repressor to that of an activator (11, 12). Thus, VDR functions as a vitamin D-sensitive switch that attracts a set of nuclear proteins, like co-factors and chromatin modifying enzymes, to its thousands genomic binding sites (Figure 1). This leads to local changes in chromatin accessibility at many genomic loci, i.e., the epigenome responds to vitamin D.
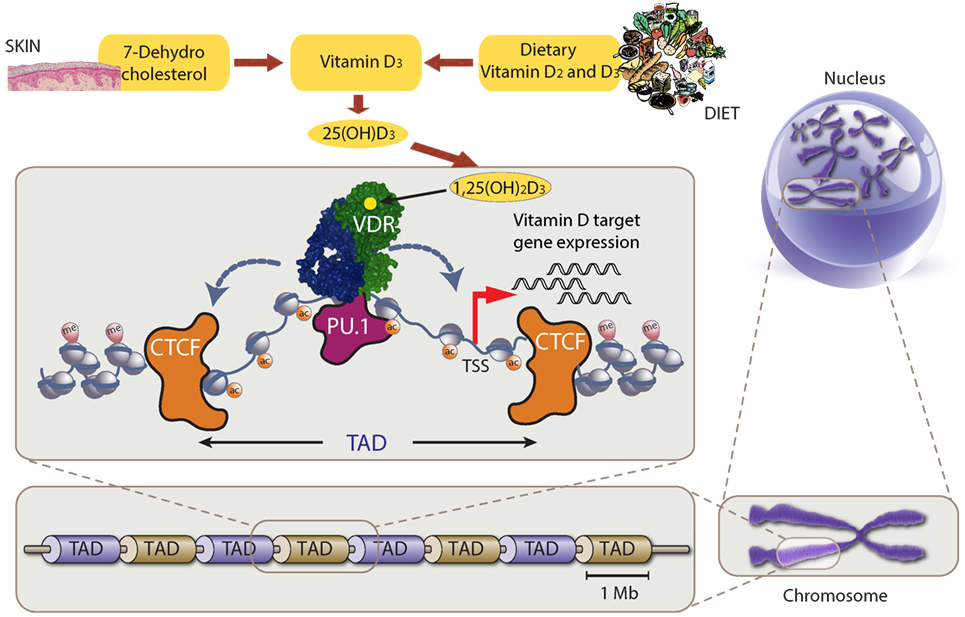
Figure 1. Chromatin model of vitamin D signaling. Top: production of vitamin D3 and its metabolites 25(OH)D3 and 1,25(OH)2D3. Center: vitamin D receptor (VDR) (green) binds accessible genomic DNA in complex with a partner protein (RXR or others, blue). VDR’s DNA binding is supported by the pioneer factor PU.1 (purple). The genomic region that can be influenced by 1,25(OH)2D3 (via binding to VDR) is restricted by CTCF proteins (orange) defining left and right topologically associated domain (TAD) borders, i.e., only vitamin D target genes within the TAD will be stimulated to produce more mRNA copies. Bottom and right: schematic illustration of TAD size on relation to chromosomes and the nucleus.
The expression of a primary vitamin D target gene is modulated, i.e., in most cases increased, when it co-locates with a prominent VDR binding site within the same higher order chromatin structure, referred to as topologically associated domain (TAD) (13). An additional condition is that the transcription start site (TSS) of the vitamin D target gene and a VDR-binding enhancer region are within accessible chromatin (12). Thus, changes in the epigenome are the first events after stimulation of a cell with vitamin D before the transcriptome gets modulated.
This review describes a transition in the understanding of vitamin D signaling. The latter was on in vitro cell culture models and now gets new insights from in vivo investigations in the context of short-term vitamin D intervention trials.
During the past years, the method chromatin immunoprecipitation sequencing (ChIP-seq) was widely used for the description of the genome-wide VDR binding pattern, the so-called “VDR cistrome” (14). VDR ChIP-seq data have been obtained in a number of human in vitro cell culture models, such as GM10855 and GM10861 human B lymphocytes (15), LS180 colorectal cancer cells (16), LX2 hepatic stellate cells (17), and lipopolysaccharide-polarized THP-1 macrophage-like cells (18). In parallel, in mouse cells VDR ChIP-seq had been performed with 3T3-L1 pre-adipocytes (19), IDG-SW3 osteocytic cells (20), pre-osteoblastic and differentiated MC3T3-E1 osteoblastic cells (21), as well as with bone marrow-derived mesenchymal stem cells differentiating into bone and fat cells (22). However, the presently most comprehensive analysis of the spatio-temporal VDR binding pattern has been performed in undifferentiated THP-1 human monocyte-like cells (13, 23).
Cell culture models have the advantage of rather homogenous cell populations that mostly display an unlimited growth potential. This allows performing biological repeats without the risk of major variations. Moreover, growth media can be depleted from lipophilic molecules, such as vitamin D3 and its metabolites, so that a stimulation with pharmacologic doses of 1,25(OH)2D3 (10–100 nM) results in maximal induction in reference to untreated cells. Accordingly, VDR ChIP-seq datasets obtained from in vitro cell models unanimously demonstrate that stimulation with 1,25(OH)2D3 significantly increases the number of genomic VDR binding events 2- to 10-fold (18).
The cistrome of ligand-stimulated VDR comprises some 2,000–10,000 sites per cell type. The VDR binding pattern is rather cell-specific and only the small subset of some 50 sites is found in all investigated cell types (18). Therefore, most VDR expressing tissues and cell types have a rather different set of vitamin D target genes (14, 24).
In agreement with findings of the ENCODE project (25) VDR binds equally likely both up- and downstream of genes, i.e., VDR binding sites are distributed in a Gaussian fashion in relation to the TSSs of primary vitamin D target genes. Accordingly, the more distant VDR binding sites are from a TSS, the less likely they are functional for the respective gene. In addition, the VDR binding site within an enhancer region and the TSS of a primary vitamin D target gene under the control of the receptor have to be located within the same TAD. Interestingly, out of 11,600 VDR binding sites identified in THP-1 cells, the small subgroup of only 339 highly conserved persistent VDR loci is well suited for describing most vitamin D gene regulatory scenarios (13). In THP-1 cells almost all primary vitamin D target genes are located within 1,25(OH)2D3-modulated TADs (more details below). Conserved persistent VDR sites control 168 of the 311 primary vitamin D target genes, whereas 120 genes are close to transiently occupied VDR sites. The equal distribution of persistent VDR binding sites over the human genome suggests that they may be strategically positioned, in order to provide the whole genome with sensitivity to vitamin D (13). The similarly equal genomic distribution of primary vitamin D target genes (26) supports this concept. Thus, the time-dependent binding profile of a few 100 VDR loci is sufficient for regulating most primary vitamin D target genes.
Genomic DNA is not “naked” but always wrapped around nucleosomes forming a protein–DNA complex that is referred to as chromatin (Figure 1). Nucleosomes are composed of eight histone proteins that are rich in the basic amino acids lysine and arginine. In particular at the histone’s protruding amino-termini these amino acids are often post-translationally modified by methyl or acetyl groups. Such histone modifications alter the structure of chromatin by affecting the non-covalent interactions within and between nucleosomes.
The epigenome is the genome-wide representation of (i) some 100 different histone marks, (ii) the level of DNA methylation at CpG islands, and (iii) higher order chromatin organization (27). The epigenome dynamically responds to extra- and intracellular signals, such as ligand activation of the VDR (28). However, most chromatin regions are intrinsically repressed, so that the binding of transcription factors and other nuclear proteins to genomic DNA is prevented, i.e., the epigenome controls the access to the genome (29, 30). In consequence, in a differentiated cell the so-called “epigenetic landscape” is restricted to some 100–200,000 chromatin loci (Figure 2 shows an example genomic region) that are accessible to transcription factors and RNA polymerases (25). This represents less than 10% of the whole chromatin and primarily refers to regions carrying TSSs and enhancers.
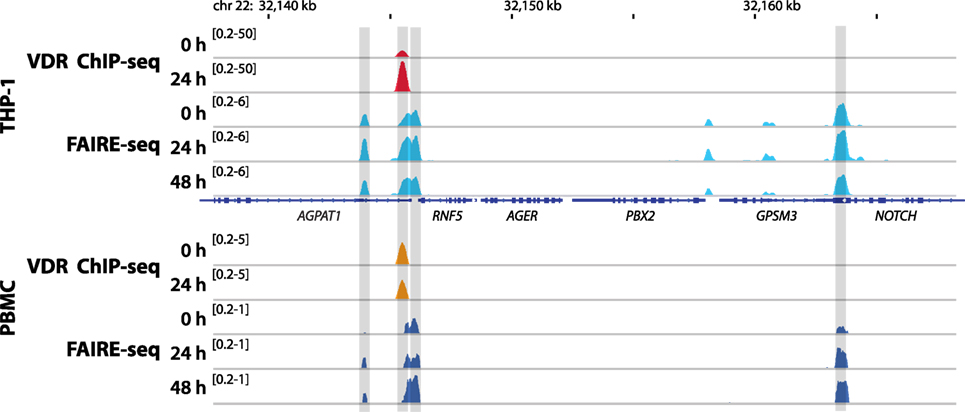
Figure 2. Vitamin D receptor (VDR) binding and chromatin opening of the 1-acylglycerol-3-phosphate O-acyltransferase 1 (AGPAT1) locus in vitro and in vivo. Top: THP-1 cells were stimulated for 0, 24, and 48 h with 1,25(OH)2D3 and VDR chromatin immunoprecipitation sequencing (ChIP-seq) and formaldehyde-assisted isolation of regulatory elements sequencing (FAIRE-seq) were performed. Bottom: in an analogous in vivo experiment (phase II context of the VitDbol study) one individual was challenged with a vitamin D3 bolus (2,000 µg). The average raise in 25(OH)D3 serum concentrations at days 1 and 2 after the vitamin D3 bolus was 11.9 and 19.4 nM, respectively. Peripheral blood mononuclear cells (PBMCs) were isolated before (day 0) and at days 1 (24 h) and 2 (48 h) and VDR ChIP-seq and FAIRE-seq were performed. The integrative genomics viewer browser was used to visualize the AGPAT1 gene locus. The peak tracks represent mergers of each three biological repeats. Gene structures are shown in blue.
Chromatin modifying and remodeling enzymes read, write, or erase chromatin marks and reposition nucleosomes, respectively (31). These enzymes are modulated in their activity by signal transduction cascades originating from intra- and extracellular signaling molecules and/or form complexes with transcription factors specifically binding to respective genomic regions (32). VDR communicates with chromatin modifying enzymes via direct and indirect interaction, such as up- and down-regulating their genes (33) or being part of the same large protein complex in the nucleus (34).
On the genome-wide level, vitamin D-induced alterations in the chromatin accessibility profile can be measured via the method formaldehyde-assisted isolation of regulatory elements sequencing (FAIRE-seq, Figure 2). In THP-1 cells, FAIRE-seq identified 62,000 accessible chromatin loci, nearly 9,000 of which are significantly modulated by 1,25(OH)2D3 (28). A 2 h stimulation with 1,25(OH)2D3 resulted at more than 3,300 genomic loci in significant changes in chromatin accessibility, after 24 h even more than 4,500 sites responded, while after 48 h only some 2,400 regions were targets of vitamin D. This suggests that maximal epigenome-wide effects occur after 24 h. In parallel, this indicates that the process of chromatin opening by vitamin D includes multiple steps. Although the exact molecular mechanisms of these vitamin D-triggered epigenome changes are not fully understood, it is obvious that they are secondary consequences of genome-wide VDR binding.
The human genome is subdivided into at least 2,000 chromatin loops (35), which segregate each chromosome into TADs. The latter are functionally independent chromatin subdomains in the size of hundreds of kilobases to a few megabases. Insulator regions separate TADs from each other (36) and contain binding sites for the transcription factor CTCF. This makes CTCF a key protein in organizing chromatin into active and inactive regions. However, from the 20,000 genome-wide CTCF loci, only some 15% are involved in forming TAD anchor regions. Interestingly, in THP-1 cells the binding of CTCF to some 1,300 sites is affected significantly by a stimulation with 1,25(OH)2D3 (37). More than half of the vitamin D-modulated CTCF sites mark one or both anchors of some 600 TADs, each of which comprises at least one VDR binding site and one vitamin D target gene. Interestingly, in the same cellular system, 587 genes are regulated significantly by 1,25(OH)2D3 (13).
In addition to the chromatin organizer CTCF, VDR also functionally associates with pioneer factors, such as PU.1 (38) or GABPA (39). A pioneer factor is a transcription factor that (i) displays many genomic binding sites, (ii) shows some promiscuity in DNA binding, and (iii) has a high diversity in protein–protein interactions (40). Accordingly, after a cellular perturbation pioneer factors are the first protein binding enhancers interacting with chromatin modifying enzymes. This makes chromatin more accessible for regular transcription factors like VDR. Interestingly, in THP-1 cells a 24 h stimulation with 1,25(OH)2D3 significantly modulated PU.1 binding at more than 5,600 sites (38).
In summary, in the THP-1 model system the epigenome-wide outcomes of a 24 h stimulation with 1,25(OH)2D3 are (i) VDR binding at more than 10,000 sites, (ii) chromatin opening at some 4,500 loci, (iii) changes in CTCF-based TAD anchors affecting some 600 chromatin loops, and (iv) increased PU.1 pioneer factor binding at more than 5,000 regions. This led to the chromatin model of vitamin D signaling (Figure 1). In this model, VDR already binds, in the absence of ligand, to a limited number of loci within accessible chromatin, while 1,25(OH)2D3 stimulation increases, via support of pioneer factors like as PU.1, the number of DNA-bound VDR molecules. This VDR binding leads to changes in chromatin accessibility, which increases the binding strength of TAD anchor forming CTCF sites upstream and downstream of prominent VDR binding loci (41).
Some 300 conserved persistent VDR sites act as key nodes, at which not only primary contacts of VDR ligands with the genome are established, but also functional consequences of vitamin D induction are coordinated throughout the whole stimulation period (13). For more than half of all primary vitamin D target genes a regulatory scenario applies, where each gene is controlled by one or more conserved persistent VDR sites being located within the same TAD. In addition, a few 100 transient VDR sites mediate more tissue-specific primary functions of vitamin D, such as immune system regulation (13). In total, five TAD classes are distinguished that differ in the number of persistent and transient VDR sites and contain sets of genes that represent different physiological functions of vitamin D. Most of the remaining VDR sites are involved in mediating secondary effects of vitamin D.
In vitro cell culture models, such as THP-1 cells, use experimental setups that are designed for obtaining maximal effects of 1,25(OH)2D3 in short time periods, such as 24 h, but may not reflect the reality of the endocrinology of vitamin D in vivo (2, 42). In fact, the genetic origin of today’s populations from East Africa and respective minor changes in physiology in the limited time, since the exodus some 50,000 years ago, suggest that humans are still primarily adapted to a constant vitamin D levels rather to changes in 25(OH)D3 serum concentration between winter and summer (43). This raises the question of how far results from in vitro experiments represent vitamin D’s actions in vivo.
The VitDbol vitamin D intervention trial (NCT02063334, ClinicalTrials.gov) studied under in vivo conditions vitamin D-dependent gene regulation in humans. From healthy young adults, peripheral blood mononuclear cells (PBMCs) were isolated at days 0, 1, and 2 after supplementation with a vitamin D3 bolus. In phase I of VitDbol, changes in chromatin accessibility were measured at selected genomic regions (44) and alterations in gene expression were determined (45). The serum 25(OH)D3 concentrations of the subjects raised in average by some 20 nM, i.e., a 20–40% increase in the vitamin D status is sufficient to open chromatin and to activate genes.
VitDbol participants differed significantly both on the level of changes in chromatin accessibility as well as on vitamin D target gene expression (44, 45). Accordingly, they could be segregated into low, mid, and high responders to vitamin D. Together with comparable results from the long-term vitamin D intervention study VitDmet (46), the VitDbol results served as the basis for the concept of the personalized vitamin D index (47). Some of the differences between individuals may be based on variations in their genome, such as SNPs, affecting the vitamin D status (48). However, in analogy to common aging-related disorders, more likely differences in the epigenome of the study participants are the main molecular explanation for alterations in the underlying traits. Accordingly, throughout their entire life a significant proportion of the human population may have a vitamin D status that is significantly lower than the needs of the respective individual for optimal function of vitamin D endocrinology.
In contrast to suggestions from classical pharmacogenetics, an individual’s health or disease status cannot be deduced reliably from a single genotyping experiment (49). Therefore, persons need to be profiled on the level of their epigenome and transcriptome in time series experiments. In phase II of VitDbol, one individual received once a month a vitamin D3 bolus three times in a row (50). Figure 2 illustrates changes in chromatin accessibility of PBMCs within 2 days. FAIRE-seq was used to detect accessible chromatin at 5,205 genomic loci, the 853 most prominent of which were categorized into early, delayed, and non-responding genomic regions. Already after 1 day 70 loci showed significant chromatin opening or closing and after 2 days 361 additional genomic sites were affected. Although in PBMCs, the number of chromatin sites with significantly changed accessibility is far lower than THP-1 cells (28), some 85% of the most prominent genomic loci are found both in vitro and in vivo (50).
The main cellular components of PBMCs, lymphocytes, and monocytes, belong to the adaptive and innate immune system, respectively. This fits well with the observation that the human leukocyte antigen (HLA) region in chromosome 6 is an epigenome “hotspot” in PBMCs (50). Interestingly, the epigenome at the HLA cluster is very responsive to vitamin D. This provides a first molecular explanation that how vitamin D may modulate actions of the immune system (51).
Vitamin D is known as a molecule that controls calcium homeostasis and bone formation, but in humans VDR’s genome-wide actions are investigated primarily in the hematopoietic system. This emphasizes the impact of vitamin D in innate and adaptive immunity. The VitDbol study demonstrated that the human epigenome responds already within 1–2 days to vitamin D. Importantly, the design of VitDbol allows safe human in vivo experiments. Nevertheless, such in vivo investigations cannot provide the same level of reproducibility than in vitro cell culture experiments, in which conditions, such as nutrient availability and temperature, are far more constant.
This is a single author mini-review. CC has written all text and created both figures.
The author declares that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author thanks the Academy of Finland for support.
1,25(OH)2D3, 1α,25-dihydroxyvitamin D3; 25(OH)D3, 25-hydroxyvitamin D3; AGPAT1, 1-acylglycerol-3-phosphate O-acyltransferase 1; ChIP-seq, chromatin immunoprecipitation sequencing; FAIRE-seq, formaldehyde-assisted isolation of regulatory elements sequencing; HLA, human leukocyte antigen; PBMC, peripheral blood mononuclear cell; TAD, topologically associated domain; TSS, transcription start site; VDR, vitamin D receptor.
1. Holick MF. Sunlight and vitamin D for bone health and prevention of autoimmune diseases, cancers, and cardiovascular disease. Am J Clin Nutr (2004) 80:1678S–88S. doi:10.1093/ajcn/80.6.1678S
2. Norman AW. From vitamin D to hormone D: fundamentals of the vitamin D endocrine system essential for good health. Am J Clin Nutr (2008) 88:491S–9S. doi:10.1093/ajcn/88.2.491S
3. Carlberg C. Molecular approaches for optimizing vitamin D supplementation. Vitam Horm (2016) 100:255–71. doi:10.1016/bs.vh.2015.10.001
4. van de Peppel J, van Leeuwen JP. Vitamin D and gene networks in human osteoblasts. Front Physiol (2014) 5:137. doi:10.3389/fphys.2014.00137
5. Carlberg C. The physiology of vitamin D-far more than calcium and bone. Front Physiol (2014) 5:335. doi:10.3389/fphys.2014.00335
6. Chun RF, Liu PT, Modlin RL, Adams JS, Hewison M. Impact of vitamin D on immune function: lessons learned from genome-wide analysis. Front Physiol (2014) 5:151. doi:10.3389/fphys.2014.00151
8. Haussler MR, Haussler CA, Jurutka PW, Thompson PD, Hsieh JC, Remus LS, et al. The vitamin D hormone and its nuclear receptor: molecular actions and disease states. J Endocrinol (1997) 154(Suppl):S57–73.
9. Carlberg C, Molnár F. Current status of vitamin D signaling and its therapeutic applications. Curr Top Med Chem (2012) 12:528–47. doi:10.2174/156802612799436623
10. Molnár F, Peräkylä M, Carlberg C. Vitamin D receptor agonists specifically modulate the volume of the ligand-binding pocket. J Biol Chem (2006) 281:10516–26. doi:10.1074/jbc.M513609200
11. Moras D, Gronemeyer H. The nuclear receptor ligand-binding domain: structure and function. Curr Opin Cell Biol (1998) 10:384–91. doi:10.1016/S0955-0674(98)80015-X
12. Carlberg C, Campbell MJ. Vitamin D receptor signaling mechanisms: integrated actions of a well-defined transcription factor. Steroids (2013) 78:127–36. doi:10.1016/j.steroids.2012.10.019
13. Neme A, Seuter S, Carlberg C. Selective regulation of biological processes by vitamin D based on the spatio-temporal cistrome of its receptor. Biochim Biophys Acta (2017) 1860:952–61. doi:10.1016/j.bbagrm.2017.07.002
14. Carlberg C. Genome-wide (over)view on the actions of vitamin D. Front Physiol (2014) 5:167. doi:10.3389/fphys.2014.00167
15. Ramagopalan SV, Heger A, Berlanga AJ, Maugeri NJ, Lincoln MR, Burrell A, et al. A ChIP-seq defined genome-wide map of vitamin D receptor binding: associations with disease and evolution. Genome Res (2010) 20:1352–60. doi:10.1101/gr.107920.110
16. Meyer MB, Goetsch PD, Pike JW. VDR/RXR and TCF4/beta-catenin cistromes in colonic cells of colorectal tumor origin: impact on c-FOS and c-MYC gene expression. Mol Endocrinol (2012) 26:37–51. doi:10.1210/me.2011-1109
17. Ding N, Yu RT, Subramaniam N, Sherman MH, Wilson C, Rao R, et al. A vitamin D receptor/SMAD genomic circuit gates hepatic fibrotic response. Cell (2013) 153:601–13. doi:10.1016/j.cell.2013.03.028
18. Tuoresmäki P, Väisänen S, Neme A, Heikkinen S, Carlberg C. Patterns of genome-wide VDR locations. PLoS One (2014) 9:e96105. doi:10.1371/journal.pone.0096105
19. Siersbaek R, Rabiee A, Nielsen R, Sidoli S, Traynor S, Loft A, et al. Transcription factor cooperativity in early adipogenic hotspots and super-enhancers. Cell Rep (2014) 7:1443–55. doi:10.1016/j.celrep.2014.04.042
20. St John HC, Bishop KA, Meyer MB, Benkusky NA, Leng N, Kendziorski C, et al. The osteoblast to osteocyte transition: epigenetic changes and response to the vitamin D3 hormone. Mol Endocrinol (2014) 28:1150–65. doi:10.1210/me.2014-1091
21. Meyer MB, Benkusky NA, Lee CH, Pike JW. Genomic determinants of gene regulation by 1,25-dihydroxyvitamin D3 during osteoblast-lineage cell differentiation. J Biol Chem (2014) 289:19539–54. doi:10.1074/jbc.M114.578104
22. Meyer MB, Benkusky NA, Sen B, Rubin J, Pike JW. Epigenetic plasticity drives adipogenic and osteogenic differentiation of marrow-derived mesenchymal stem cells. J Biol Chem (2016) 291:17829–47. doi:10.1074/jbc.M116.736538
23. Heikkinen S, Väisänen S, Pehkonen P, Seuter S, Benes V, Carlberg C. Nuclear hormone 1α,25-dihydroxyvitamin D3 elicits a genome-wide shift in the locations of VDR chromatin occupancy. Nucleic Acids Res (2011) 39:9181–93. doi:10.1093/nar/gkr654
24. Campbell MJ. Vitamin D and the RNA transcriptome: more than mRNA regulation. Front Physiol (2014) 5:181. doi:10.3389/fphys.2014.00181
25. ENCODE-Project-ConsortiumBernstein BE, Birney E, Dunham I, Green ED, Gunter C, et al. An integrated encyclopedia of DNA elements in the human genome. Nature (2012) 489:57–74. doi:10.1038/nature11247
26. Neme A, Nurminen V, Seuter S, Carlberg C. The vitamin D-dependent transcriptome of human monocytes. J Steroid Biochem Mol Biol (2016) 164:180–7. doi:10.1016/j.jsbmb.2015.10.018
27. Carlberg C, Molnár F. The impact of chromatin. 2nd ed. Mechanisms of Gene Regulation. Dordrecht: Springer (2016). p. 17–34.
28. Seuter S, Neme A, Carlberg C. Epigenome-wide effects of vitamin D and their impact on the transcriptome of human monocytes involve CTCF. Nucleic Acids Res (2016) 44:4090–104. doi:10.1093/nar/gkv1519
29. Beisel C, Paro R. Silencing chromatin: comparing modes and mechanisms. Nat Rev Genet (2011) 12:123–35. doi:10.1038/nrg2932
30. Smith ZD, Meissner A. DNA methylation: roles in mammalian development. Nat Rev Genet (2013) 14:204–20. doi:10.1038/nrg3354
31. Carlberg C, Molnár F. Chromatin modifiers. 2nd ed. Mechanisms of Gene Regulation. Dordrecht: Springer (2016). p. 129–45.
32. Badeaux AI, Shi Y. Emerging roles for chromatin as a signal integration and storage platform. Nat Rev Mol Cell Biol (2013) 14:211–24. doi:10.1038/nrm3545
33. Pereira F, Barbachano A, Silva J, Bonilla F, Campbell MJ, Munoz A, et al. KDM6B/JMJD3 histone demethylase is induced by vitamin D and modulates its effects in colon cancer cells. Hum Mol Genet (2011) 20:4655–65. doi:10.1093/hmg/ddr399
34. Molnár F. Structural considerations of vitamin D signaling. Front Physiol (2014) 5:191. doi:10.3389/fphys.2014.00191
35. Dixon JR, Selvaraj S, Yue F, Kim A, Li Y, Shen Y, et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature (2012) 485:376–80. doi:10.1038/nature11082
36. Van Bortle K, Corces VG. The role of chromatin insulators in nuclear architecture and genome function. Curr Opin Genet Dev (2013) 23:212–8. doi:10.1016/j.gde.2012.11.003
37. Neme A, Seuter S, Carlberg C. Vitamin D-dependent chromatin association of CTCF in human monocytes. Biochim Biophys Acta (2016) 1859:1380–8. doi:10.1016/j.bbagrm.2016.08.008
38. Seuter S, Neme A, Carlberg C. Epigenomic PU.1-VDR crosstalk modulates vitamin D signaling. Biochim Biophys Acta (2017) 1860:405–15. doi:10.1016/j.bbagrm.2017.02.005
39. Seuter S, Neme A, Carlberg C. ETS transcription factor family member GABPA contributes to vitamin D receptor target gene regulation. J Steroid Biochem Mol Biol (2018) 177:46–52. doi:10.1016/j.jsbmb.2017.08.006
40. Zaret KS, Carroll JS. Pioneer transcription factors: establishing competence for gene expression. Gen Dev (2011) 25:2227–41. doi:10.1101/gad.176826.111
41. Carlberg C. Molecular endocrinology of vitamin D on the epigenome level. Mol Cell Endocrinol (2017) 453:14–21. doi:10.1016/j.mce.2017.03.016
42. DeLuca HF. Overview of general physiologic features and functions of vitamin D. Am J Clin Nutr (2004) 80:1689S–96S. doi:10.1093/ajcn/80.6.1689S
43. Jablonski NG, Chaplin G. The roles of vitamin D and cutaneous vitamin D production in human evolution and health. Int J Paleopathol (2018). doi:10.1016/j.ijpp.2018.01.005
44. Seuter S, Virtanen JK, Nurmi T, Pihlajamäki J, Mursu J, Voutilainen S, et al. Molecular evaluation of vitamin D responsiveness of healthy young adults. J Steroid Biochem Mol Biol (2017) 174:314–21. doi:10.1016/j.jsbmb.2016.06.003
45. Vukic M, Neme A, Seuter S, Saksa N, De Mello VD, Nurmi T, et al. Relevance of vitamin D receptor target genes for monitoring the vitamin D responsiveness of primary human cells. PLoS One (2015) 10:e0124339. doi:10.1371/journal.pone.0124339
46. Carlberg C, Seuter S, De Mello VD, Schwab U, Voutilainen S, Pulkki K, et al. Primary vitamin D target genes allow a categorization of possible benefits of vitamin D3 supplementation. PLoS One (2013) 8:e71042. doi:10.1371/journal.pone.0071042
47. Carlberg C, Haq A. The concept of the personal vitamin D response index. J Steroid Biochem Mol Biol (2018) 175:12–7. doi:10.1016/j.jsbmb.2016.12.011
48. Wang TJ, Zhang F, Richards JB, Kestenbaum B, Van Meurs JB, Berry D, et al. Common genetic determinants of vitamin D insufficiency: a genome-wide association study. Lancet (2010) 376:180–8. doi:10.1016/S0140-6736(10)60588-0
49. Carlberg C, Raunio H. From pharmacogenomics to integrated personal omics profiling: a gap in implementation into healthcare. Per Med (2014) 11:625–9. doi:10.2217/pme.14.38
50. Carlberg C, Seuter S, Nurmi T, Tuomainen TP, Virtanen JK, Neme A. In vivo response of the human epigenome to vitamin D: a proof-of-principle study. J Steroid Biochem Mol Biol (2018). doi:10.1016/j.jsbmb.2018.01.002
Keywords: vitamin D, vitamin D receptor, vitamin D target genes, vitamin D intervention trial, chromatin, epigenome, immune system
Citation: Carlberg C (2018) Vitamin D Genomics: From In Vitro to In Vivo. Front. Endocrinol. 9:250. doi: 10.3389/fendo.2018.00250
Received: 23 March 2018; Accepted: 02 May 2018;
Published: 23 May 2018
Edited by:
Pawel Pludowski, Children’s Memorial Health Institute, PolandReviewed by:
Pavel M. Marozik, National Academy of Sciences of Belarus, BelarusCopyright: © 2018 Carlberg. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Carsten Carlberg, Y2Fyc3Rlbi5jYXJsYmVyZ0B1ZWYuZmk=
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.