- 1Wuxi Mental Health Center, Nanjing Medical University, Wuxi, China
- 2Shanghai Mental Health Center, Shanghai Jiao Tong University School of Medicine, Shanghai, China
It is well known that Alzheimer’s disease (AD) is one of the most common progressive neurodegenerative diseases; it begins gradually, and therefore no effective medicine is administered in the beginning. Thus, early diagnosis and prevention of AD are crucial. The present study focused on comparing the plasma protein changes between patients with AD and their healthy counterparts, aiming to explore a specific protein panel as a potential biomarker for AD patients in Han Chinese. Hence, we recruited and collected plasma samples from 98 AD patients and 101 elderly healthy controls from Wuxi and Shanghai Mental Health Centers. Using a Luminex assay, we investigated the expression levels of fifty plasma proteins in these samples. Thirty-two out of 50 proteins were found to be significantly different between AD patients and healthy controls (P < 0.05). Furthermore, an eight-protein panel that included brain-derived neurotrophic factor (BDNF), angiotensinogen (AGT), insulin-like growth factor binding protein 2 (IGFBP-2), osteopontin (OPN), cathepsin D, serum amyloid P component (SAP), complement C4, and prealbumin (transthyretin, TTR) showed the highest determinative score for AD and healthy controls (all P = 0.00). In conclusion, these findings suggest that a combination of eight plasma proteins can serve as a promising diagnostic biomarker for AD with high sensitivity and specificity in Han Chinese populations; the eight plasma proteins were proven important for AD diagnosis by further cross-validation studies within the AD cohort.
Introduction
Recently, the Centers for Disease Control and Prevention in the United States reported a 54.5% increase from the 1999 rate of 16.5 deaths per 100,000 patients with Alzheimer’s disease (AD) (Taylor et al., 2017), which is one of the most common progressive neurodegenerative diseases and is characterized by the interaction of both genetic and environmental factors, resulting in memory dysfunction and behavioral changes (Hooli and Tanzi, 2009). In 2013, the number of older people in China was almost 200 million, and the proportion of the population that was aged 65 years and older increased to 14.3% (Sun et al., 2015). The World Alzheimer Report 2015 updated the estimates of the global prevalence, China is the region with the most people living with dementia (9.5 million), and the prevalence for the population aged 60 and older is 6.19% (Prince et al., 2015; Wu et al., 2018). Currently, disease-modifying therapy and an ideal diagnostic tool for AD are largely lacking, severely influencing patients’ quality of life and leading to a heavy financial burden for patients’ families.
Preclinical Alzheimer’s, the newly defined disease stage, demonstrates that brain changes are progressively initiated 10 to 20 years before the onset of dementia symptoms (Bateman et al., 2012). Thus, identifying biomarkers for preclinical Alzheimer’s contributed to the early recognition and prediction of the progression of AD. Notably, the diagnostic tool should be inexpensive, easy to perform, and non-invasive (The Ronald and Nancy Reagan Research Institute of the Alzheimer’s Association and the National Institute on Aging Working Group, 1998). Recent scientific evidence suggests some potential diagnostic biomarkers such as β-amyloid (Aβ) (Blennow et al., 2015) and tau (Meredith et al., 2013) accumulation in cerebrospinal fluid (CSF), Pittsburgh compound B positron emission tomography (PiB-PET) (Leuzy et al., 2015), and peripheral blood protein (Zhao et al., 2015) and microRNA (miRNA) expression (Galimberti et al., 2014), among which blood-derived biomarkers have been extensively studied due to their less invasive source (Blennow, 2017; Kitamura et al., 2017; O’Bryant et al., 2017). Although previous studies have demonstrated plasma protein profiles may be a valuable diagnostic biomarker for the early stage of AD, the findings have not been widely replicated in different races (Kiddle et al., 2014; Shi et al., 2018).
Therefore, based on these findings, the present study focuses on plasma protein differences in healthy individuals and patients with AD, exploring a specific protein panel as a potential diagnostic biomarker for AD patients in Han Chinese.
Materials and Methods
Study Population
We recruited 1,105 older people aged 56–95 years from 2015 to 2017. Eventually, 98 AD patients diagnosed with the criteria of the National Institute of Neurological and Communicative Disorders and Stroke and the Alzheimer’s disease and Related Disorders Association (NINCDS-ADRDA) were recruited from Wuxi and Shanghai Mental Health Centers. The cognitive function of these participants was assessed by two skilled professional psychiatrists using the Mini-Mental State Examination (MMSE). The absence of depression was documented on the basis of a score of 10 or less on the Hamilton Rating Scale for Depression (HAMD). A brain computed tomography (CT) or magnetic resonance imaging (MRI) scan excluded other structural brain diseases, and neurologic examination showed no significant abnormalities. A total of 101 elderly healthy subjects were recruited through advertisements, and their demographic characteristics were carefully recorded.
Exclusion Criteria
Patients with any dementia other than AD, such as vascular dementia, dementia with Lewy bodies, frontotemporal dementia or Parkinson’s disease, were excluded. In addition, a history of stroke or cerebrovascular disease, bone marrow transplantation, major psychiatric disorder, and a history of alcohol or drug abuse were causes for exclusion from the study.
Study Design
Plasma samples from AD and elderly healthy controls from Shanghai and Wuxi in China were obtained. In total, we examined plasma samples from 199 subjects: 98 with AD and 101 elderly controls with no dementia. Multiple protein differences were explored using Luminex xMAP technology. Statistical analysis was performed to assess the relative importance of these protein biomarkers for the diagnosis of AD.
This study was approved by the Ethics Committees of the Wuxi Health Mental Center. Either patients or their guardians signed informed consent. If participants failed to fill out the consent form more than twice, their guardians were asked to fill out the consent form on the patients’ behalf.
Luminex Assays
Luminex xMAP technology (Austin, TX, United States) uses a solid phase approach to analyze multiple proteins. In brief, the xMAP technology is a flow cytometric-based platform that uses microspheres inserted with a ratio of two different fluorescent dyes. In theory, up to 100 differently colored beads can be generated with a theoretical multiplex capacity of up to 100 assays per well of a 96-well plate. The capture antibody is covalently coupled to the bead, and immunoassays are run under standard sandwich immunoassay formats (Hye et al., 2014).
The Luminex kits were obtained from Millipore (Billerica, MA, United States) and the assays were performed according to the manufacturer’s instructions (Soares et al., 2009). Eleven Milliplex MAP multiplex panels covering 50 proteins (96-well plate format; EMD Millipore) were utilized: cat.# HCYTOMAG-60K (7-plex); cat.# HIGFBMAG-53K (2-plex); cat.# HMHEMAG-34K (2-plex); cat.# HMMP2MAG-55K (2-plex); cat.# HNDG1MAG-36K (7-plex); cat.# HNDG2MAG-36K (6-plex); cat.# HNDG3MAG-36K (10-plex); cat.# HND2MAG-39K (3-plex); cat.# HND3MAG-39K (7-plex); cat.# SKINMAG-50K (1-plex); and cat.# HKI6MAG-99K (3-plex). Properly diluted plasma samples were incubated with the antibody-coupled microspheres and then with biotinylated detection antibody before the addition of streptavidin-phycoerythrin. The captured bead complexes were measured with a FLEXMAP 3D system (Luminex Corporation, Austin, TX, United States) using the following instrument settings: events/bead, 50; sample size, 50 μL; discriminator gate, 8000–15,000. The raw data (mean fluorescence intensity) were collected and further processed for calculating protein concentration (Zhi et al., 2014; Al-Daghri et al., 2018).
Data Processing
Quality checks (QC) based on standard curve linearity, intraassay coefficient of variation, interassay coefficient of variation for reference sample, and percentage of missing data were performed to examine the performance of each assay followed by measuring median fluorescent intensity (MFI) using xPONENT 5.1 (Luminex Corporation). This was further exported into Milliplex Analyst 5.1 (VigeneTech, United States) to calculate protein concentrations by a five-parameter logistic fit. Afterward, all analytes were subjected to the statistical analysis.
Statistical Analysis
All analysis results were expressed as the mean ± SD. Statistical analyses were performed using the Statistical Package for the Social Sciences software version 20.0 (SPSS Inc., Chicago, IL, United States). Pearson’s chi-square test was used to compare gender between control subjects and AD patients. All 50 blood protein levels were non-normally distributed, and subsequently underwent ln or square root transformation. The results of transforming the variables are displayed in the table. An independent-sample t-test was used to compare age, education and MMSE score and overall protein differences between control subjects and AD patients. Discriminant analysis was performed to assess the relative importance of these biomarkers in classifying AD and controls. In the case of the stepwise method, Wilk’s lambda method was used to build the prediction model. The discriminant analysis used a partial F-test (F to enter 3.84; F to remove 2.71) and a stepwise method (maximum number of steps = 64) to sequentially incorporate the set of 32 significant variables into the canonical discriminant function. To check the reliability of our analysis, leave-one-out cross-validation was used. Receiver operating characteristic (ROC) analyses were conducted under the non-parametric distribution assumption for standard error of area to determine the performance of the models for discriminating AD from controls.
Results
Study Participants
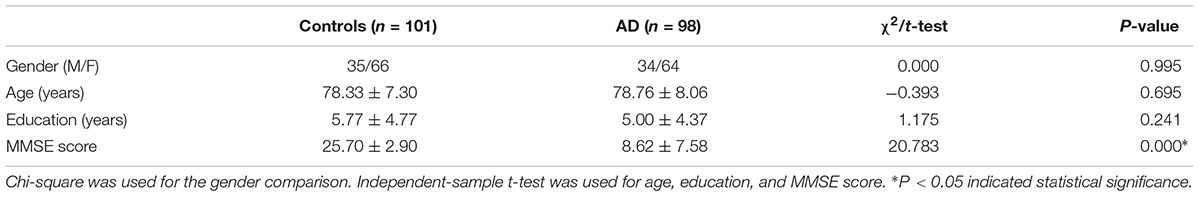
The demographic and clinical characteristics of healthy controls and AD patients are presented in Table 1. Briefly, the mean age of AD patients and their healthy counterparts were 78.76 ± 8.06 and 78.33 ± 7.30 years, respectively. No significant difference was found in age, gender or education between these two groups. As expected, patients with AD had significantly lower MMSE scores than healthy controls (P = 0.000).
Differentially Expressed Protein
In this study, we applied Luminex assay technology to determine the expression profiles of 50 proteins in the plasma from AD patients and healthy controls. Of the fifty candidates, thirty-two proteins were found to be differentially expressed, with statistical significance between AD patients and healthy controls (P < 0.05) (Table 2).
Stepwise Discriminant Function Analysis
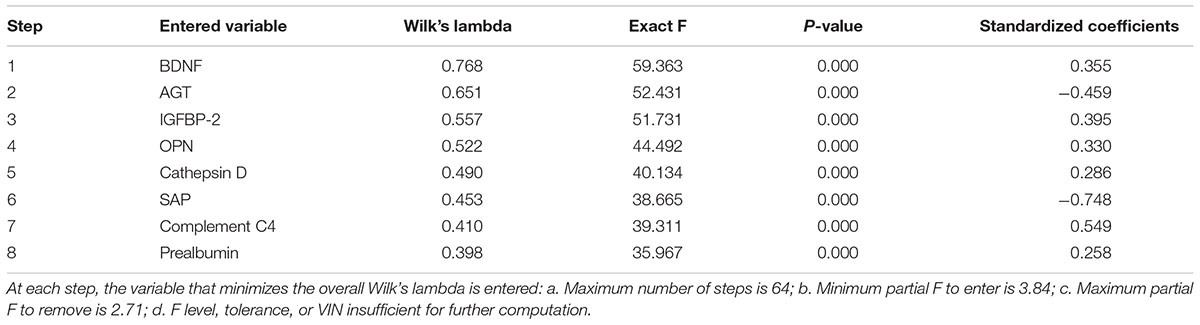
Afterward, we performed a stepwise discriminant function analysis to further determine how effectively AD patients and healthy controls can be distinguished based on the expressed protein levels and to assess the differential contribution to the diagnosis. Of the 32 significant plasma markers, a feature group of eight most discriminative proteins, including brain-derived neurotrophic factor (BDNF), angiotensinogen (AGT), insulin-like growth factor binding protein 2 (IGFBP-2), osteopontin (OPN), cathepsin D, serum amyloid P component (SAP), complement C4, and prealbumin (transthyretin, TTR), was sorted out by stepwise discriminant analysis (Table 3, all P = 0.00), indicating their potential contributions to diagnosis. To detect whether this 8-protein panel was efficient in differentiating AD from healthy controls, we carried out both original- and cross-validation, correctly classifying 86.7 and 84.7% of the cases, respectively (Table 4).
The Classification Performance of the 8-Protein Panel
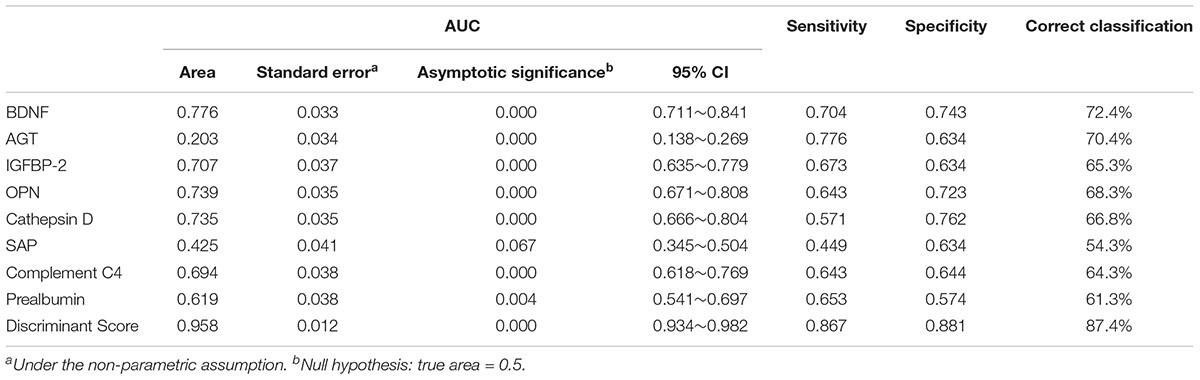
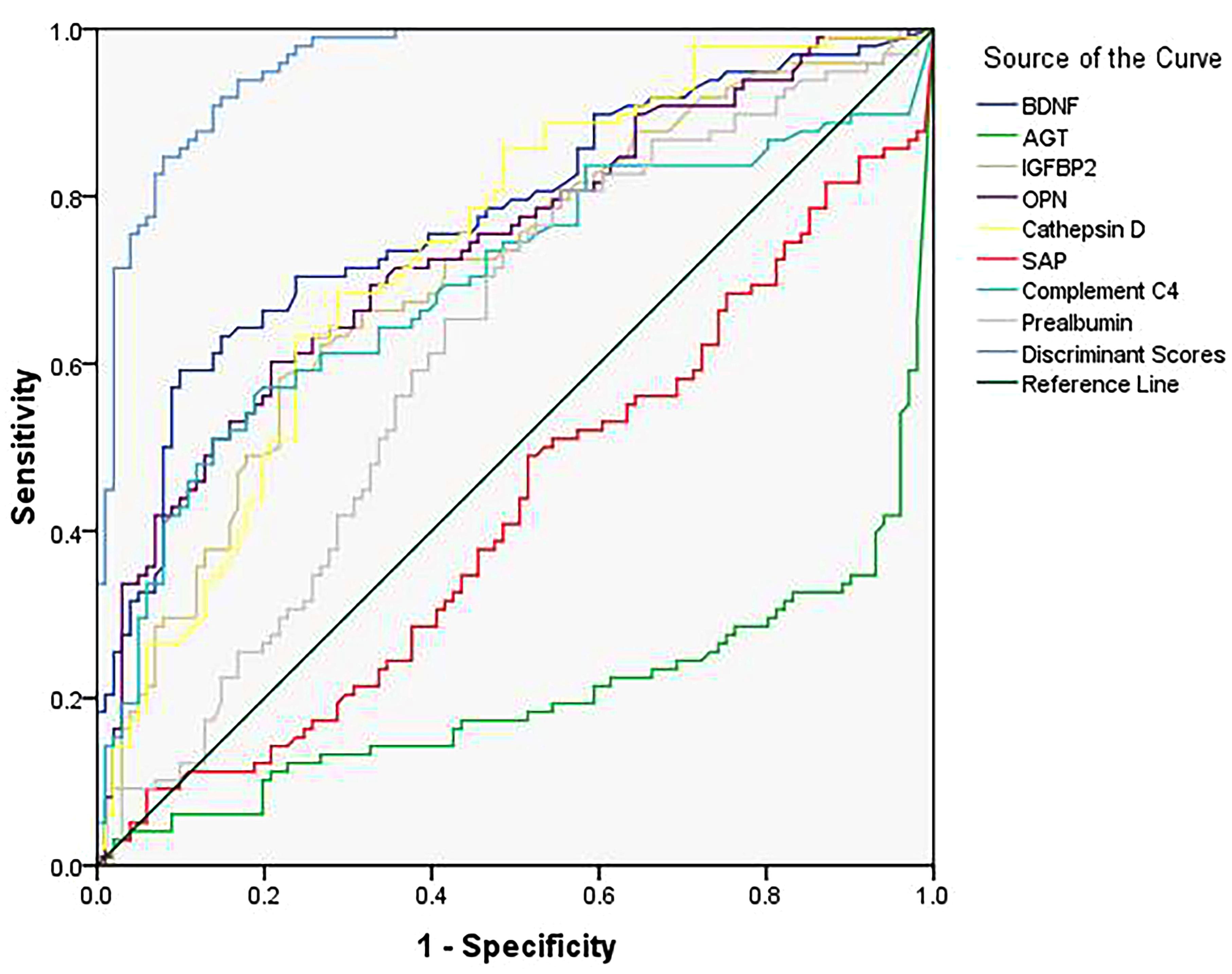
Furthermore, we decided the classification performance of the eight-protein panel and each biomarker by calculating the discriminant score (Table 5) and receiver operating characteristic (ROC) curve (Figure 1), resulting in an 87.4% correct classification for AD and control subjects with high sensitivity (86.7%) and specificity (88.1%), which suggested that the combination of these eight differentially expressed plasma proteins produced the most accurate results in a threshold classification.
Discussion
To date, disease-modifying treatments have not been successfully developed for AD. Continuous clinical trials for novel drugs have failed, suggesting early diagnosis and prevention of AD is crucial for postponing the progression of disease effectively. Approximately 500 mL of cerebrospinal fluid (CSF), which is in direct contact with the extracellular space of the brain, is absorbed into the blood daily (Davidsson et al., 2002; Davidsson and Sjogren, 2006; Hye et al., 2006). Plasma contains multiple biological components, including proteins, peptides, lipids, and metabolites, which also effectively reflect physiological activity and pathology in the central nervous system (CNS). To our knowledge, this is the first study to investigate potential plasma protein biomarkers in the Han Chinese population using a high-throughput multiplexed xMAP Luminex assay. Our present findings suggest that a set of eight plasma proteins (BDNF, AGT, IGFBP-2, OPN, cathepsin D, SAP, complement C4, and TTR) serves as a putative predictor panel for AD diagnosis with high sensitivity and specificity. Importantly, these proteins have been considered to be interesting and potentially significant in AD disease pathology in previous studies (Phillips et al., 1991; Iadecola and Davisson, 2008; Carecchio and Comi, 2011; Daborg et al., 2012; Mold et al., 2012; Tian et al., 2014; Willette et al., 2015; Buxbaum and Johansson, 2017).
Recently, blood-derived biomarkers for AD have been widely considered for being relatively painless, inexpensive and having diagnostic accuracy. Using predictive analysis of microarrays, a previous study demonstrated that a panel of 18 plasma proteins (CCL18, CCL15, CCL7, CXCL8, ICAM-1, TRAIL-R4, G-CSF, GDNF, EGF, CCL5, M-CSF, IL-3, IL-1α, TNF-α, PDGF-BB, IL-11, ANG-2, and IGFBP-6) achieved a diagnostic accuracy of 90% in distinguishing AD and mild cognitive impairment (MCI) (Ray et al., 2007), which was reported to be unable to distinguish patients with AD from the populations through enough diagnostic precision (Bjorkqvist et al., 2012). In addition, it also failed to reach comparable accuracy in discriminating patients with AD and healthy controls using 16 and 8 plasma proteins derived from this 18-protein panel, respectively (Soares et al., 2009; Marksteiner et al., 2011), which might be attributed to only three of these 18 plasma proteins being differentially expressed between AD and healthy controls in an independent replication study. Afterward, Thambisetty et al. (2008) also performed a subsequent study to investigate 26 proteins that had been identified as potential AD biomarkers, including the 18-protein panel reported by Ray et al. (2007). Although only two proteins were found to be significantly different between AD and controls, they identified that a 10-protein panel (TTR, CLU, cystatin C, A1AcidG, ICAM-1, CC4, pigment epithelium-derived factor, A1AT, RANTES, and ApoC3) could predict the progression of MCI to AD with high diagnostic accuracy. However, of these proteins, ICAM-1 was a unique protein that overlapped with Ray’s study. O’Bryant and colleagues used a panel from RBM to identify a list of 30 biomarkers to detect AD (O’Bryant et al., 2010, 2011).
It is well known that the pathological mechanism of AD is diverse. The fifty proteins selected in the present study, including those proteins that were reported in previous studies, are involved in the various signaling pathways associated with AD, including the immune response (Heneka et al., 2015), inflammatory (Harrison, 2013; Heneka et al., 2015) and antioxidant (Andrieu et al., 2015) processes, and metabolism (Arnoldussen et al., 2014; Counts et al., 2017). Of this 8-protein panel, three proteins, CC4, TTR, and IGFBP-2, overlapped with previous studies (Bennett et al., 2012; Uchida et al., 2015; McLimans et al., 2017) and affect immunology, Aβ fibril formation, DNA synthesis, and cell proliferation and death in AD.
The complement system is considered to be highly involved in the inflammatory response as a powerful component of innate immunity, consisting of more than 30 fluid-phase and cell-associated proteins as well as a wide range of specific receptors that interact to trigger the inflammatory response (Ricklin et al., 2010; Wagner and Frank, 2010). Recent genome-wide association studies (GWAS) have identified the association of complement receptor 1 with AD (Harold et al., 2009; Lambert et al., 2009; Seshadri et al., 2010; Naj et al., 2011). However, whether these complement molecules have any diagnostic value has not been fully elucidated. Elevated levels of complement 3 and 4 (C3 and C4) in cerebrospinal fluid (CSF) were found in AD, compared with MCI, patients (Daborg et al., 2012). Moreover, increased plasma levels of C4a protein were found in the plasma of AD patients (Bennett et al., 2012). Consistently, complement C4 was highly expressed in the plasma of AD participants in this study. We further showed that complement C4 had 64.3% accuracy for distinguishing AD from healthy individuals with 64.3% sensitivity and 64.4% specificity, indicating that complement C4 alone was inadequate as a biomarker for AD diagnosis.
Transthyretin (TTR) is a transport protein, also known as prealbumin, which has been found in cerebrospinal fluid (CSF) as an Aβ-binding protein and suppresses the toxicity of oligomers. Thus, the sequester protein, as an efficient inhibitor of Aβ fibril formation, may delay the pathologic progression of AD via the Aβ clearance signaling pathway (Buxbaum and Johansson, 2017). A previous study showed that TTR concentration was substantially decreased in the peripheral blood of individuals with aMCI and AD, indicating that a set of sequester proteins, including TTR, can discriminate individuals with mild cognitive decline from healthy controls (Uchida et al., 2015). In contrast to this result, we found a significant increase in TTR concentration in AD patients when compared with normal controls, which might be attributed to the different ethnic populations revealing a number of influencing factors, such as heredity, environment, and lifestyle.
Among the candidate proteins, IGFBP2 was the one most frequently chosen. IGFBP-2 can influence DNA synthesis, cell proliferation and death, as well as glucose and amino acid uptake in cells by inhibiting IGF functions (Jones and Clemmons, 1995; Reyer et al., 2015). Overexpression of IGFBP-2 in mice led to decreased weights of the hippocampus, cerebellum, olfactory bulb, and prefrontal cortex (Schindler et al., 2017). In addition, IGFBP-2 was also reported to significantly increase in the serum of AD participants (Tham et al., 1993; McLimans et al., 2017), which was consistent with our findings. In addition, other insulin-like growth factors and the corresponding binding proteins were differentially expressed in both the CSF and serum from AD patients (Salehi et al., 2008), in which the expression of IGF-1 was reduced in serum (Vidal et al., 2016), while IGF-2 was highly expressed in CSF (Aberg et al., 2015). However, it has also been reported that serum, instead of CSF of IGF-1 and IGFBP3, was increased in AD (Johansson et al., 2013), both of which were more closely associated with AD in men than in women (Duron et al., 2012). Using a longitudinal case-control study, O’Bryant et al. (2010) found that serum protein-based biomarkers involving IGFBP-2 protein can be combined with clinical information to accurately classify AD (O’Bryant et al., 2010). Subsequently, using the multiplex panel, Doecke et al. (2012) identified an 18-plasma biomarker panel including IGFBP-2 that is useful for the diagnosis of AD (Doecke et al., 2012). These findings suggested that IGFBP-2 may be a key factor in a panel of protein biomarkers for the diagnosis of AD.
In addition, some proteins on the panel, including SAP, have been associated with hippocampal atrophy and the rate of change and progression to dementia (Thambisetty et al., 2010; Sattlecker et al., 2016). Importantly, we performed bioinformatics analysis and identified a close and interactive network among these proteins (data not shown), partly supporting the intimate relationship of the 8-protein panel with AD.
It is well known that recent clinical trials for the Aβ clearing strategies have failed (Galimberti and Scarpini, 2016; Mehta et al., 2017). Previous findings showed that 30% of cognitively normal elderly patients show signs of Aβ accumulation and, accordingly, a substantial number of AD patients show no signs of Aβ accumulation (Yang et al., 2012). In addition, several conflicting findings were demonstrated when using plasma Aβ peptides as markers for AD, suggesting that Aβ from peripheral blood may not reflect brain Aβ metabolism (Cummings, 2011; Hampel et al., 2012; Koyama et al., 2012). Importantly, Aβ can bind to a variety of proteins in blood, resulting in epitope masking and analytical interference (Marcello et al., 2009). In addition, methodology was also a limitation in the present study. The Milliplex MAP multiplex panels containing Aβ protein can only be used with CSF samples. To ensure the strong consistency of the results, the detection of plasma Aβ levels was not performed using other methods in this study.
Although many studies have identified plasma proteins related to AD [e.g., BDNF (Laske et al., 2007; Laske et al., 2011)], complement C4a (Bennett et al., 2012), IGFBP-2 (McLimans et al., 2017), TTR (Uchida et al., 2015), and SAP (Wilson et al., 2008), these are unlikely to be useful as a diagnostic test when used as single markers due to a lack of sensitivity and specificity. Based on the complexity of the pathogenesis of AD, as a relatively reliable biomarker for the diagnosis of AD, combinations of plasma proteins associated with various biological pathways may be necessary. Of note, accumulating evidence indicates that the replication and validation of results is urgent and important for exploratory studies. Thus, we plan to further validate our findings in a large-scale population that includes AD, MCI, and healthy controls.
The present findings should be interpreted considering some limitations. Since AD overlaps with other dementia forms, such as vascular dementia, frontotemporal dementia, and dementia with Lewy bodies, in the context of pathological traits, whether the same protein panel can accurately reflect their relationship to AD, but not to other dementia diseases, is a very important question. At present, only AD patients and healthy controls were included in this study, and no other dementia subtypes or disease stages were identified. In addition, our results may be influenced by the limited sample size. Moreover, due to the limitation of methodology, we did not detect plasma Aβ levels in the present study. Hence, more extensive studies including a large number of dementia subtypes should be conducted in the future. Furthermore, there are various factors, such as activity, diet, and medications, that can alter plasma protein levels.
Conclusion
Taken together, the present study identified a plasma 8-protein panel including BDNF, AGT, IGFBP-2, OPN, cathepsin D, SAP, complement C4 and TTR that showed the highest determinative score for AD and healthy controls. Thus, these findings suggest that a combination of eight plasma proteins is a valuable diagnostic biomarker for AD in the Chinese population, providing novel insight for the diagnosis of AD.
Author Contributions
ZC and SX conceived and designed the experiments. ZC, JY, HY, CJ, FZ, ZW, XL, YW, and TW performed the experiments and contributed to reagents, materials, and analysis tools. ZC, JY, HY, and CJ analyzed the data. ZC, JY, and SX wrote the paper.
Funding
This study was supported by a grant from Jiangsu Province Science and Technology Department (BE2015615).
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
Aberg, D., Johansson, P., Isgaard, J., Wallin, A., Johansson, J. O., Andreasson, U., et al. (2015). Increased cerebrospinal fluid level of insulin-like growth factor-II in male patients with Alzheimer’s disease. J. Alzheimers Dis. 48, 637–646. doi: 10.3233/JAD-150351
Al-Daghri, N. M., Yakout, S. M., Wani, K., Khattak, M. N. K., Garbis, S. D., Chrousos, G. P., et al. (2018). IGF and IGFBP as an index for discrimination between vitamin D supplementation responders and nonresponders in overweight Saudi subjects. Medicine 97:e0702. doi: 10.1097/MD.0000000000010702
Andrieu, S., Coley, N., Lovestone, S., Aisen, P. S., and Vellas, B. (2015). Prevention of sporadic Alzheimer’s disease: lessons learned from clinical trials and future directions. Lancet Neurol. 14, 926–944. doi: 10.1016/S1474-4422(15)00153-2
Arnoldussen, I. A., Kiliaan, A. J., and Gustafson, D. R. (2014). Obesity and dementia: adipokines interact with the brain. Eur. Neuropsychopharmacol. 24, 1982–1999. doi: 10.1016/j.euroneuro.2014.03.002
Bateman, R. J., Xiong, C., Benzinger, T. L., Fagan, A. M., Goate, A., Fox, N. C., et al. (2012). Clinical and biomarker changes in dominantly inherited Alzheimer’s disease. N. Engl. J. Med. 367, 795–804. doi: 10.1056/NEJMoa1202753
Bennett, S., Grant, M., Creese, A. J., Mangialasche, F., Cecchetti, R., Cooper, H. J., et al. (2012). Plasma levels of complement 4a protein are increased in Alzheimer’s disease. Alzheimer Dis. Assoc. Disord. 26, 329–334. doi: 10.1097/WAD.0b013e318239dcbd
Bjorkqvist, M., Ohlsson, M., Minthon, L., and Hansson, O. (2012). Evaluation of a previously suggested plasma biomarker panel to identify Alzheimer’s disease. PLoS One 7:e29868. doi: 10.1371/journal.pone.0029868
Blennow, K. (2017). A review of fluid biomarkers for Alzheimer’s disease: moving from CSF to blood. Neurol. Ther. 6, 15–24. doi: 10.1007/s40120-017-0073-9
Blennow, K., Mattsson, N., Scholl, M., Hansson, O., and Zetterberg, H. (2015). Amyloid biomarkers in Alzheimer’s disease. Trends Pharmacol. Sci. 36, 297–309. doi: 10.1016/j.tips.2015.03.002
Buxbaum, J. N., and Johansson, J. (2017). Transthyretin and BRICHOS: the paradox of amyloidogenic proteins with anti-amyloidogenic activity for abeta in the central nervous system. Front. Neurosci. 11:119. doi: 10.3389/fnins.2017.00119
Carecchio, M., and Comi, C. (2011). The role of osteopontin in neurodegenerative diseases. J. Alzheimers Dis. 25, 179–185. doi: 10.3233/JAD-2011102151
Counts, S. E., Ikonomovic, M. D., Mercado, N., Vega, I. E., and Mufson, E. J. (2017). Biomarkers for the early detection and progression of Alzheimer’s disease. Neurotherapeutics 14, 35–53. doi: 10.1007/s13311-016-0481-z
Cummings, J. L. (2011). Biomarkers in Alzheimer’s disease drug development. Alzheimers Dement. 7, e13–e44. doi: 10.1016/j.jalz.2010.06.004
Daborg, J., Andreasson, U., Pekna, M., Lautner, R., Hanse, E., Minthon, L., et al. (2012). Cerebrospinal fluid levels of complement proteins C3. C4 and CR1 in Alzheimer’s disease. J. Neural. Transm. 119, 789–797. doi: 10.1007/s00702-012-0797-8
Davidsson, P., and Sjogren, M. (2006). Proteome studies of CSF in AD patients. Mech. Ageing Dev. 127, 133–137. doi: 10.1016/j.mad.2005.09.021
Davidsson, P., Westman-Brinkmalm, A., Nilsson, C. L., Lindbjer, M., Paulson, L., Andreasen, N., et al. (2002). Proteome analysis of cerebrospinal fluid proteins in Alzheimer patients. Neuroreport 13, 611–615. doi: 10.1097/00001756-200204160-00015
Doecke, J. D., Laws, S. M., Faux, N. G., Wilson, W., Burnham, S. C., Lam, C. P., et al. (2012). Blood-based protein biomarkers for diagnosis of Alzheimer disease. Arch. Neurol. 69, 1318–1325. doi: 10.1001/archneurol.2012.1282
Duron, E., Funalot, B., Brunel, N., Coste, J., Quinquis, L., Viollet, C., et al. (2012). Insulin-like growth factor-I and insulin-like growth factor binding protein-3 in Alzheimer’s disease. J. Clin. Endocrinol. Metab. 97, 4673–4681. doi: 10.1210/jc.20122063
Galimberti, D., and Scarpini, E. (2016). Emerging amyloid disease-modifying drugs for Alzheimer’s disease. Expert Opin. Emerg. Drugs 21, 5–7. doi: 10.1517/14728214.2016.1146678
Galimberti, D., Villa, C., Fenoglio, C., Serpente, M., Ghezzi, L., Cioffi, S. M., et al. (2014). Circulating miRNAs as potential biomarkers in Alzheimer’s disease. J. Alzheimers Dis. 42, 1261–1267. doi: 10.3233/JAD-140756
Hampel, H., Lista, S., and Khachaturian, Z. S. (2012). Development of biomarkers to chart all Alzheimer’s disease stages: the royal road to cutting the therapeutic Gordian Knot. Alzheimers Dement. 8, 312–336. doi: 10.1016/j.jalz.2012.05.2116
Harold, D., Abraham, R., Hollingworth, P., Sims, R., Gerrish, A., Hamshere, M., et al. (2009). Genome-wide association study identifies variants at CLU and PICALM associated with Alzheimer’s disease, and shows evidence for additional susceptibility genes. Nat. Genet. 41, 1088–1093. doi: 10.1038/ng.440
Harrison, N. (2013). Inflammation and mental illness. J. Neurol. Neurosurg. Psychiatry 84:e1. doi: 10.1136/jnnp-2013-306103.4
Heneka, M. T., Carson, M. J., El Khoury, J., Landreth, G. E., Brosseron, F., Feinstein, D. L., et al. (2015). Neuroinflammation in Alzheimer’s disease. Lancet Neurol. 14, 388–405. doi: 10.1016/S1474-4422(15)70016-5
Hooli, B. V., and Tanzi, R. E. (2009). A current view of Alzheimer’s disease. F1000 Biol. Rep. 1:54. doi: 10.3410/B1-54
Hye, A., Lynham, S., Thambisetty, M., Causevic, M., Campbell, J., Byers, H. L., et al. (2006). Proteome-based plasma biomarkers for Alzheimer’s disease. Brain 129(Pt 11), 3042–3050. doi: 10.1093/brain/awl279
Hye, A., Riddoch-Contreras, J., Baird, A. L., Ashton, N. J., Bazenet, C., Leung, R., et al. (2014). Plasma proteins predict conversion to dementia from prodromal disease. Alzheimers Dement. 10, 799.e2–807.e2. doi: 10.1016/j.jalz.2014.05.1749
Iadecola, C., and Davisson, R. L. (2008). Hypertension and cerebrovascular dysfunction. Cell Metab. 7, 476–484. doi: 10.1016/j.cmet.2008.03.010
Johansson, P., Aberg, D., Johansson, J. O., Mattsson, N., Hansson, O., Ahren, B., et al. (2013). Serum but not cerebrospinal fluid levels of insulin-like growth factor-I (IGF-I) and IGF-binding protein-3 (IGFBP-3) are increased in Alzheimer’s disease. Psychoneuroendocrinology 38, 1729–1737. doi: 10.1016/j.psyneuen.2013.02.006
Jones, J. I., and Clemmons, D. R. (1995). Insulin-like growth factors and their binding proteins: biological actions. Endocr. Rev. 16, 3–34. doi: 10.1210/edrv-16-1-3
Kiddle, S. J., Sattlecker, M., Proitsi, P., Simmons, A., Westman, E., Bazenet, C., et al. (2014). Candidate blood proteome markers of Alzheimer’s disease onset and progression: a systematic review and replication study. J. Alzheimers Dis. 38, 515–531. doi: 10.3233/JAD-130380
Kitamura, Y., Usami, R., Ichihara, S., Kida, H., Satoh, M., Tomimoto, H., et al. (2017). Plasma protein profiling for potential biomarkers in the early diagnosis of Alzheimer’s disease. Neurol. Res. 39, 231–238. doi: 10.1080/01616412.2017.1281195
Koyama, A., Okereke, O. I., Yang, T., Blacker, D., Selkoe, D. J., and Grodstein, F. (2012). Plasma amyloid-beta as a predictor of dementia and cognitive decline: a systematic review and meta-analysis. Arch. Neurol. 69, 824–831. doi: 10.1001/archneurol.2011.1841
Lambert, J. C., Heath, S., Even, G., Campion, D., Sleegers, K., Hiltunen, M., et al. (2009). Genome-wide association study identifies variants at CLU and CR1 associated with Alzheimer’s disease. Nat. Genet. 41, 1094–1099. doi: 10.1038/ng.439
Laske, C., Leyhe, T., Stransky, E., Hoffmann, N., Fallgatter, A. J., and Dietzsch, J. (2011). Identification of a blood-based biomarker panel for classification of Alzheimer’s disease. Int. J. Neuropsychopharmacol. 14, 1147–1155. doi: 10.1017/S1461145711000459
Laske, C., Stransky, E., Leyhe, T., Eschweiler, G. W., Maetzler, W., Wittorf, A., et al. (2007). BDNF serum and CSF concentrations in Alzheimer’s disease, normal pressure hydrocephalus and healthy controls. J. Psychiatr. Res. 41, 387–394. doi: 10.1016/j.jpsychires.2006.01.014
Leuzy, A., Carter, S. F., Chiotis, K., Almkvist, O., Wall, A., and Nordberg, A. (2015). Concordance and diagnostic accuracy of [11C]PIB PET and cerebrospinal fluid biomarkers in a sample of patients with mild cognitive impairment and Alzheimer’s disease. J. Alzheimers Dis. 45, 1077–1088. doi: 10.3233/JAD-142952
Marcello, A., Wirths, O., Schneider-Axmann, T., Degerman-Gunnarsson, M., Lannfelt, L., and Bayer, T. A. (2009). Circulating immune complexes of Abeta and IgM in plasma of patients with Alzheimer’s disease. J. Neural. Transm. 116, 913–920. doi: 10.1007/s00702-009-0224-y
Marksteiner, J., Kemmler, G., Weiss, E. M., Knaus, G., Ullrich, C., Mechtcheriakov, S., et al. (2011). Five out of 16 plasma signaling proteins are enhanced in plasma of patients with mild cognitive impairment and Alzheimer’s disease. Neurobiol. Aging 32, 539–540. doi: 10.1016/j.neurobiolaging.2009.03.011
McLimans, K. E., Webb, J. L., Anantharam, V., Kanthasamy, A., Willette, A. A., and Alzheimer’s Disease Neuroimaging Initiative. (2017). Peripheral versus central index of metabolic dysfunction and associations with clinical and pathological outcomes in Alzheimer’s disease. J. Alzheimers Dis. 60, 1313–1324. doi: 10.3233/JAD-170263
Mehta, D., Jackson, R., Paul, G., Shi, J., and Sabbagh, M. (2017). Why do trials for Alzheimer’s disease drugs keep failing? A discontinued drug perspective for 2010-2015. Expert Opin. Investig. Drugs 26, 735–739. doi: 10.1080/13543784.2017.1323868
Meredith, J. E. Jr., Sankaranarayanan, S., Guss, V., Lanzetti, A. J., Berisha, F., Neely, R. J., et al. (2013). Characterization of novel CSF Tau and ptau biomarkers for Alzheimer’s disease. PLoS One 8:e76523. doi: 10.1371/journal.pone.0076523
Mold, M., Shrive, A. K., and Exley, C. (2012). Serum amyloid P component accelerates the formation and enhances the stability of amyloid fibrils in a physiologically significant under-saturated solution of amyloid-beta42. J. Alzheimers Dis. 29, 875–881. doi: 10.3233/JAD-2012-120076
Naj, A. C., Jun, G., Beecham, G. W., Wang, L. S., Vardarajan, B. N., Buros, J., et al. (2011). Common variants at MS4A4/MS4A6E. CD2AP, CD33 and EPHA1 are associated with late-onset Alzheimer’s disease. Nat. Genet. 43, 436–441. doi: 10.1038/ng.801
O’Bryant, S. E., Mielke, M. M., Rissman, R. A., Lista, S., Vanderstichele, H., Zetterberg, H., et al. (2017). Blood-based biomarkers in Alzheimer disease: current state of the science and a novel collaborative paradigm for advancing from discovery to clinic. Alzheimers Dement. 13, 45–58. doi: 10.1016/j.jalz.2016.09.014
O’Bryant, S. E., Xiao, G., Barber, R., Reisch, J., Doody, R., Fairchild, T., et al. (2010). A serum protein-based algorithm for the detection of Alzheimer disease. Arch. Neurol. 67, 1077–1081. doi: 10.1001/archneurol.2010.215
O’Bryant, S. E., Xiao, G., Barber, R., Reisch, J., Hall, J., Cullum, C. M., et al. (2011). A blood-based algorithm for the detection of Alzheimer’s disease. Dement. Geriatr. Cogn. Disord. 32, 55–62. doi: 10.1159/000330750
Phillips, H. S., Hains, J. M., Armanini, M., Laramee, G. R., Johnson, S. A., and Winslow, J. W. (1991). BDNF mRNA is decreased in the hippocampus of individuals with Alzheimer’s disease. Neuron 7, 695–702. doi: 10.1016/0896-6273(91)90273-3
Prince, M., Wimo, A., Guerchet, M., Ali, G., Wu, Y., and Prina, M. (2015). World Alzheimer Report 2015 The Global Impact of Dementia an analysis of Prevalence, Incidence, Cost, and Trends. London: Alzheimer’s Disease International.
Ray, S., Britschgi, M., Herbert, C., Takeda-Uchimura, Y., Boxer, A., Blennow, K., et al. (2007). Classification and prediction of clinical Alzheimer’s diagnosis based on plasma signaling proteins. Nat. Med. 13, 1359–1362. doi: 10.1038/nm1653
Reyer, A., Schindler, N., Ohde, D., Walz, C., Kunze, M., Tuchscherer, A., et al. (2015). The RGD sequence present in IGFBP-2 is required for reduced glucose clearance after oral glucose administration in female transgenic mice. Am. J. Physiol. Endocrinol. Metab. 309, E409–E417. doi: 10.1152/ajpendo.00168.2015
Ricklin, D., Hajishengallis, G., Yang, K., and Lambris, J. D. (2010). Complement: a key system for immune surveillance and homeostasis. Nat. Immunol. 11, 785–797. doi: 10.1038/ni.1923
Salehi, Z., Mashayekhi, F., and Naji, M. (2008). Insulin like growth factor-1 and insulin like growth factor binding proteins in the cerebrospinal fluid and serum from patients with Alzheimer’s disease. Biofactors 33, 99–106. doi: 10.1002/biof.5520330202
Sattlecker, M., Khondoker, M., Proitsi, P., Williams, S., Soininen, H., Kloszewska, I., et al. (2016). Longitudinal protein changes in blood plasma associated with the rate of cognitive decline in Alzheimer’s disease. J. Alzheimers Dis. 49, 1105–1114. doi: 10.3233/JAD-140669
Schindler, N., Mayer, J., Saenger, S., Gimsa, U., Walz, C., Brenmoehl, J., et al. (2017). Phenotype analysis of male transgenic mice overexpressing mutant IGFBP-2 lacking the Cardin-Weintraub sequence motif: reduced expression of synaptic markers and myelin basic protein in the brain and a lower degree of anxiety-like behaviour. Growth Horm. IGF Res. 33, 1–8. doi: 10.1016/j.ghir.2016.11.003
Seshadri, S., Fitzpatrick, A. L., Ikram, M. A., DeStefano, A. L., Gudnason, V., Boada, M., et al. (2010). Genome-wide analysis of genetic loci associated with Alzheimer disease. JAMA 303, 1832–1840. doi: 10.1001/jama.2010.574
Shi, L., Baird, A. L., Westwood, S., Hye, A., Dobson, R., Thambisetty, M., et al. (2018). A Decade of Blood Biomarkers for Alzheimer’s Disease Research: an evolving field, improving study designs, and the challenge of replication. J. Alzheimes Dis. 62, 1181–1198. doi: 10.3233/JAD-170531
Soares, H. D., Chen, Y., Sabbagh, M., Roher, A., Schrijvers, E., and Breteler, M. (2009). Identifying early markers of Alzheimer’s disease using quantitative multiplex proteomic immunoassay panels. Ann. N. Y. Acad. Sci. 1180, 56–67. doi: 10.1111/j.1749-6632.2009.05066.x
Sun, R., Cao, H., Zhu, X., Liu, J.-P., and Dong, E. (2015). Current aging research in China. Protein Cell 6, 314–321. doi: 10.1007/s13238-015-0145-5
Taylor, C. A., Greenlund, S. F., McGuire, L. C., Lu, H., and Croft, J. B. (2017). Deaths from Alzheimer’s Disease - United States, 1999-2014. MMWR Morb. Mortal. Wkly. Rep. 66, 521–526. doi: 10.15585/mmwr.mm6620a1
Tham, A., Nordberg, A., Grissom, F. E., Carlsson-Skwirut, C., Viitanen, M., and Sara, V. R. (1993). Insulin-like growth factors and insulin-like growth factor binding proteins in cerebrospinal fluid and serum of patients with dementia of the Alzheimer type. J. Neural Transm. Park. Dis. Dement. Sect. 5, 165–176. doi: 10.1007/bf02257671
Thambisetty, M., Hye, A., Foy, C., Daly, E., Glover, A., Cooper, A., et al. (2008). Proteome-based identification of plasma proteins associated with hippocampal metabolism in early Alzheimer’s disease. J. Neurol. 255, 1712–1720. doi: 10.1007/s00415-008-0006-8
Thambisetty, M., Simmons, A., Velayudhan, L., Hye, A., Campbell, J., Zhang, Y., et al. (2010). Association of plasma clusterin concentration with severity, pathology, and progression in Alzheimer disease. Arch. Gen. Psychiatry 67, 739–748. doi: 10.1001/archgenpsychiatry.2010.78
The Ronald and Nancy Reagan Research Institute of the Alzheimer’s Association and the National Institute on Aging Working Group (1998). Consensus report of the working group on: “Molecular and Biochemical Markers of Alzheimer’s Disease”. Neurobiol. Aging 19, 109–116. doi: 10.1016/S0197-4580(98)00022-0
Tian, L., Zhang, K., Tian, Z. Y., Wang, T., Shang, D. S., Li, B., et al. (2014). Decreased expression of cathepsin D in monocytes is related to the defective degradation of amyloid-beta in Alzheimer’s disease. J. Alzheimers Dis. 42, 511–520. doi: 10.3233/JAD-132192
Uchida, K., Shan, L., Suzuki, H., Tabuse, Y., Nishimura, Y., Hirokawa, Y., et al. (2015). Amyloid-beta sequester proteins as blood-based biomarkers of cognitive decline. Alzheimers Dement 1, 270–280. doi: 10.1016/j.dadm.2015.04.003
Vidal, J. S., Hanon, O., Funalot, B., Brunel, N., Viollet, C., Rigaud, A. S., et al. (2016). Low serum insulin-like growth factor-i predicts cognitive decline in Alzheimer’s disease. J. Alzheimers Dis. 52, 641–649. doi: 10.3233/JAD-151162
Wagner, E., and Frank, M. M. (2010). Therapeutic potential of complement modulation. Nat. Rev. Drug Discov. 9, 43–56. doi: 10.1038/nrd3011
Willette, A. A., Bendlin, B. B., Starks, E. J., Birdsill, A. C., Johnson, S. C., Christian, B. T., et al. (2015). Association of insulin resistance with cerebral glucose uptake in late middle-aged adults at risk for Alzheimer disease. JAMA Neurol. 72, 1013–1020. doi: 10.1001/jamaneurol.2015.0613
Wilson, M. R., Yerbury, J. J., and Poon, S. (2008). Potential roles of abundant extracellular chaperones in the control of amyloid formation and toxicity. Mol. Biosyst. 4, 42–52. doi: 10.1039/b712728f
Wu, Y.-T., Ali, G.-C., Guerchet, M., Prina, A. M., Chan, K. Y., Prince, M., et al. (2018). Prevalence of dementia in mainland China, Hong Kong and Taiwan: an updated systematic review and meta-analysis. Int. J. Epidemiol. 47, 709–719. doi: 10.1093/ije/dyy007
Yang, L., Rieves, D., and Ganley, C. (2012). Brain amyloid imaging–FDA approval of florbetapir F18 injection. N. Engl. J. Med. 367, 885–887. doi: 10.1056/NEJMp1208061
Zhao, X., Lejnine, S., Spond, J., Zhang, C., Ramaraj, T. C., Holder, D. J., et al. (2015). A candidate plasma protein classifier to identify Alzheimer’s disease. J. Alzheimers Dis. 43, 549–563. doi: 10.3233/JAD-141149
Keywords: Alzheimer’s disease, biomarker, plasma protein, early diagnosis, Han Chinese
Citation: Cheng Z, Yin J, Yuan H, Jin C, Zhang F, Wang Z, Liu X, Wu Y, Wang T and Xiao S (2018) Blood-Derived Plasma Protein Biomarkers for Alzheimer’s Disease in Han Chinese. Front. Aging Neurosci. 10:414. doi: 10.3389/fnagi.2018.00414
Received: 04 April 2018; Accepted: 30 November 2018;
Published: 17 December 2018.
Edited by:
Athanasios Alexiou, Novel Global Community Educational Foundation (NGCEF), AustraliaReviewed by:
Md. Sahab Uddin, Southeast University, BangladeshGirish Kumar Gupta, Maharishi Markandeshwar University, India
Copyright © 2018 Cheng, Yin, Yuan, Jin, Zhang, Wang, Liu, Wu, Wang and Xiao. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Zaohuo Cheng, chengzaohuo@126.com Shifu Xiao, xiaoshifu@msn.com
†These authors have contributed equally to this work
 Zaohuo Cheng
Zaohuo Cheng Jiajun Yin
Jiajun Yin Hongwei Yuan1
Hongwei Yuan1 Fuquan Zhang
Fuquan Zhang Tao Wang
Tao Wang Shifu Xiao
Shifu Xiao