- 1Biomedical Informatics Research Lab, School of Basic Medicine and Clinical Pharmacy, China Pharmaceutical University, Nanjing, China
- 2Cancer Genomics Research Center, School of Basic Medicine and Clinical Pharmacy, China Pharmaceutical University, Nanjing, China
The 2019 novel coronavirus (SARS-CoV-2) pandemic has caused a global health emergency. The outbreak of this virus has raised a number of questions: What is SARS-CoV-2? How transmissible is SARS-CoV-2? How severely affected are patients infected with SARS-CoV-2? What are the risk factors for viral infection? What are the differences between this novel coronavirus and other coronaviruses? To answer these questions, we performed a comparative study of four pathogenic viruses that primarily attack the respiratory system and may cause death, namely, SARS-CoV-2, severe acute respiratory syndrome (SARS-CoV), Middle East respiratory syndrome (MERS-CoV), and influenza A viruses (H1N1 and H3N2 strains). This comparative study provides a critical evaluation of the origin, genomic features, transmission, and pathogenicity of these viruses. Because the coronavirus disease 2019 (COVID-19) pandemic caused by SARS-CoV-2 is ongoing, this evaluation may inform public health administrators and medical experts to aid in curbing the pandemic's progression.
Introduction
The 2019 novel coronavirus (SARS-CoV-2), severe acute respiratory syndrome coronavirus (SARS-CoV), Middle East respiratory syndrome coronavirus (MERS-CoV), and influenza A viruses are major pathogens that primarily target the human respiratory system. Diseases associated with their infections vary from mild respiratory illness to acute pneumonia and even respiratory failure. Since 1918, the influenza A viruses have caused four pandemics. The first and most severe pandemic in recent history, known as “Spanish influenza,” occurred in 1918 and was caused by an H1N1 influenza A virus (IAV) strain (1). Approximately 500 million people were infected, and 50 million people died during this pandemic. The second pandemic, known as “Asian influenza,” occurred in 1957, was caused by an H2N2 IAV strain, and resulted in ~1.1 million deaths worldwide (2). The third pandemic, known as “Hong Kong flu,” occurred in 1968 and was caused by an H3N2 IAV strain, resulting in ~1 million deaths worldwide (3). The fourth pandemic was caused by the influenza A (H1N1) pdm09 virus, also known as the “novel influenza A virus,” and resulted in 151,700–575,400 deaths worldwide from 2009 to 2010 (4, 5). Since that time, the novel influenza A virus has continued to spread as a seasonal flu virus. From September 2019 to February 2020, this virus caused at least 34 million flu illnesses and 20,000 deaths. In November 2002, before the fourth influenza A pandemic, an epidemic caused by a betacoronavirus (SARS-CoV) and known as severe acute respiratory syndrome (SARS) began in South China and spread to 29 countries. The SARS outbreak caused ~8,000 infections and 774 deaths before it was contained in July 2003, with a case fatality rate (CFR) of 9.6% (the CFR was ~50% among patients 65 or older) (6). However, since 2004, there have not been any SARS cases reported anywhere in the world. In September 2012, Saudi Arabia reported the first case of Middle East respiratory syndrome (MERS), which was caused by another type of betacoronavirus (MERS-CoV). MERS-CoV spread to 27 countries and caused 2,519 infections and 866 deaths by January 2020, with a CFR of 34.4% (7).
In December 2019, cases of the new coronavirus disease 2019 (COVID-19), caused by a new betacoronavirus (SARS-CoV-2), were first reported in Wuhan, China (8). These cases were characterized by acute pneumonia-associated symptoms, such as fever, dry cough, chills, shortness of breath, and muscle pain (9). The SARS-CoV-2 outbreak rapidly spread worldwide. It has infected more than 14 million individuals and resulted in more than 500,000 deaths as of 20 July 2020. In comparison with the other two coronaviruses, SARS-CoV-2 appears to be much more contagious and infectious; it has rapidly resulted in a pandemic constituting a global health emergency (Figures 1A–C).
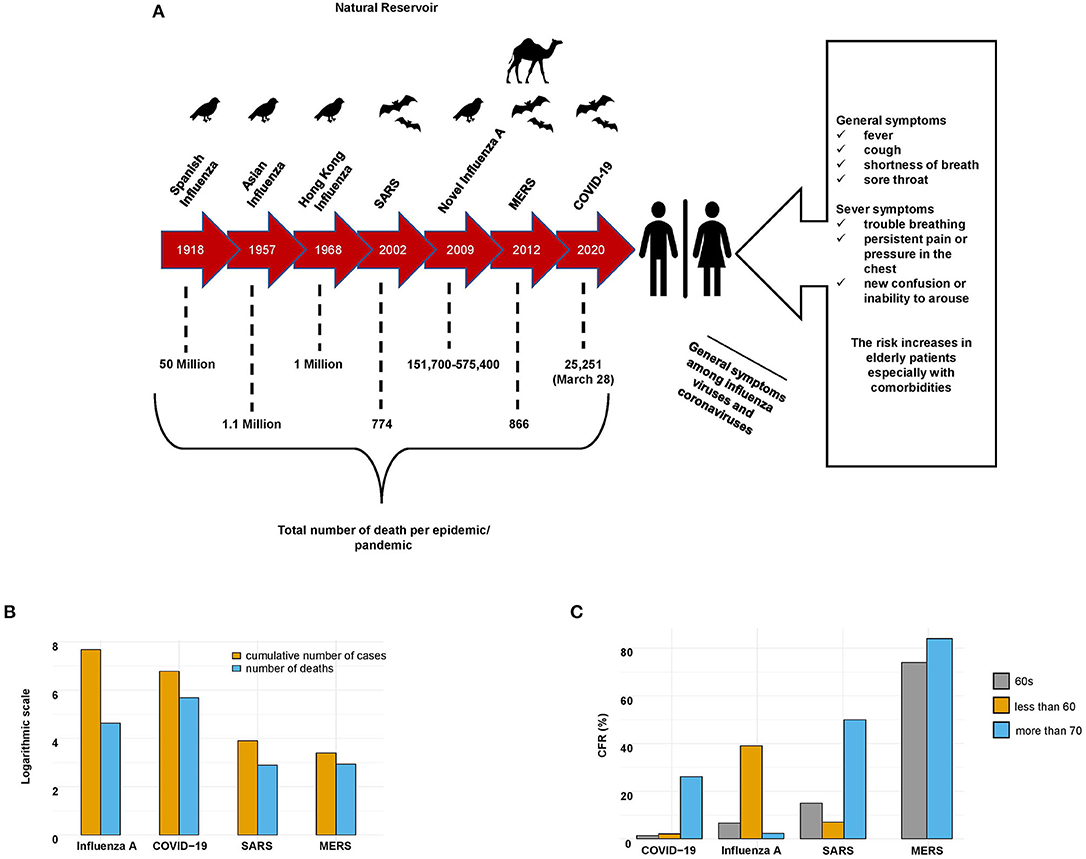
Figure 1. General characteristics of SARS-CoV-2, SARS-CoV, MERS-CoV, and influenza A viruses. (A) Epidemics of SARS-CoV-2, SARS-CoV, MERS-CoV, and influenza A viruses. The timeline, natural reservoirs, total number of deaths, and symptoms of the patients infected with these viruses. (B) Cumulative numbers of cases and deaths caused by SARS-CoV-2, SARS-CoV, MERS-CoV, and influenza A (during the last seasonal flu 2019–2020) viruses. Influenza A virus infected the most people, while SARS-CoV-2 caused the most deaths. (C) Case-fatality rate (CFR) of patients infected with SARS-CoV-2, SARS-CoV, MERS-CoV, and influenza A (the last seasonal flu 2019–2020) viruses stratified by age.
To better understand the current COVID-19 pandemic caused by SARS-CoV-2, we have performed a comparative study between SARS-CoV-2 and past epidemic/pandemic viral infections that primarily affect the respiratory system: the influenza A viruses (H3N2 and H1N1 strains) and the two coronaviruses SARS-CoV and MERS-CoV. We have explored the genomic characteristics, transmission, reservoirs, and pathogenesis of these four pathogens. We have also considered the preventive and control measures conducted by the World Health Organization (WHO) against the spread of these pathogens. Additionally, we have elucidated how these viruses attack the immune system and the associated host immune system response. This comparative study will aid in informing public health administrators and medical experts on how to adequately distinguish between these viruses and identify the preventive and control measures recommended by the WHO against the spread of SARS-CoV-2.
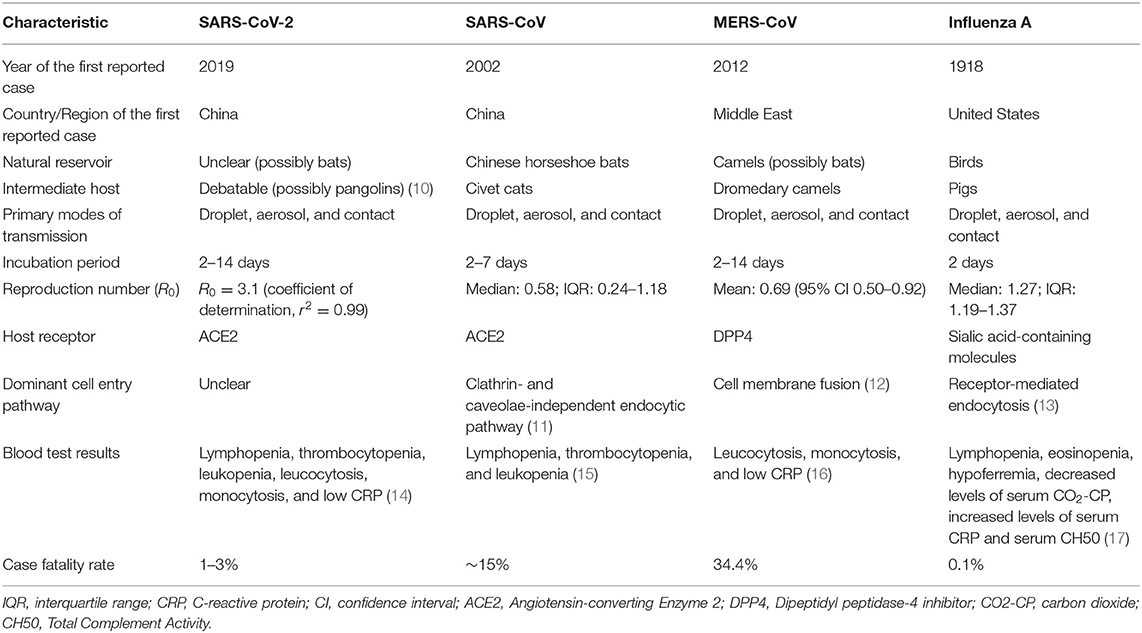
A brief comparison between the four pathogenic viruses, including their characteristics, pathogenesis, and transmission, is summarized in Table 1.
Taxonomy, Structure, and Genomic Properties of the Viruses
Influenza A
Influenza A viruses that infect humans mainly consist of two strains (H1N1 and H3N2). Both strains are characterized as enveloped, negative-sense, single-stranded RNA viruses with a total genome size of ~13.5 kb (18, 19). The influenza A virus genome consists of eight different segments, with each segment containing a region that encodes one or two proteins with specific functions, including hemagglutinin (HA), polymerase basic protein 2 (PB2), nucleoprotein (NP), polymerase basic protein 1 (PB1), neuraminidase (NA), matrix (M), nonstructural protein (NS1), and polymerase acidic protein (PA) (20, 21).
The HA protein of influenza A viruses binds to the glycoprotein terminal sialic acid and glycolipid receptors, which contain α-2,6 and α-2,3 sialic acid groups attached to galactose. Although HA is considered to be a more crucial antigenic determinant than NA, both proteins are potentially restrictive factors for viral evolution (20, 22). In addition, there are three viral polymerase proteins, PB1, PB2, and PA, encoded on segments 1, 2, and 3, respectively; these polymerase proteins form an enzyme complex that plays a role in transcription and replication. Finally, the NP protein encoded on segment 5 is used as a model to generate additional copies (23, 24).
Influenza A viruses exhibit antigenic drift/shift properties, allowing them to avoid the host immune response. The Centers for Disease Control and Prevention (CDC) defines antigenic drift as genetic variation that occurs in antigen structures owing to point mutations in the HA and NA genes over time, whereas antigenic shift is the result of a sudden genetic reassortment between two or more closely related influenza viral strains (23, 24). A well-known example of the antigenic shift phenomenon is the triple reassortment that occurred in the influenza A pdm09 virus and caused the 2009 pandemic as a result of the replacement of the hemagglutinin H2 and polymerase PB1 genes of the avian H2N2 virus with two new avian H3 and PB1 genes (25, 26) (Figure 2A). These antigenic drift/shift properties can potentially reduce the effectiveness of vaccines and become a considerable challenge in antiviral therapy (27, 28).
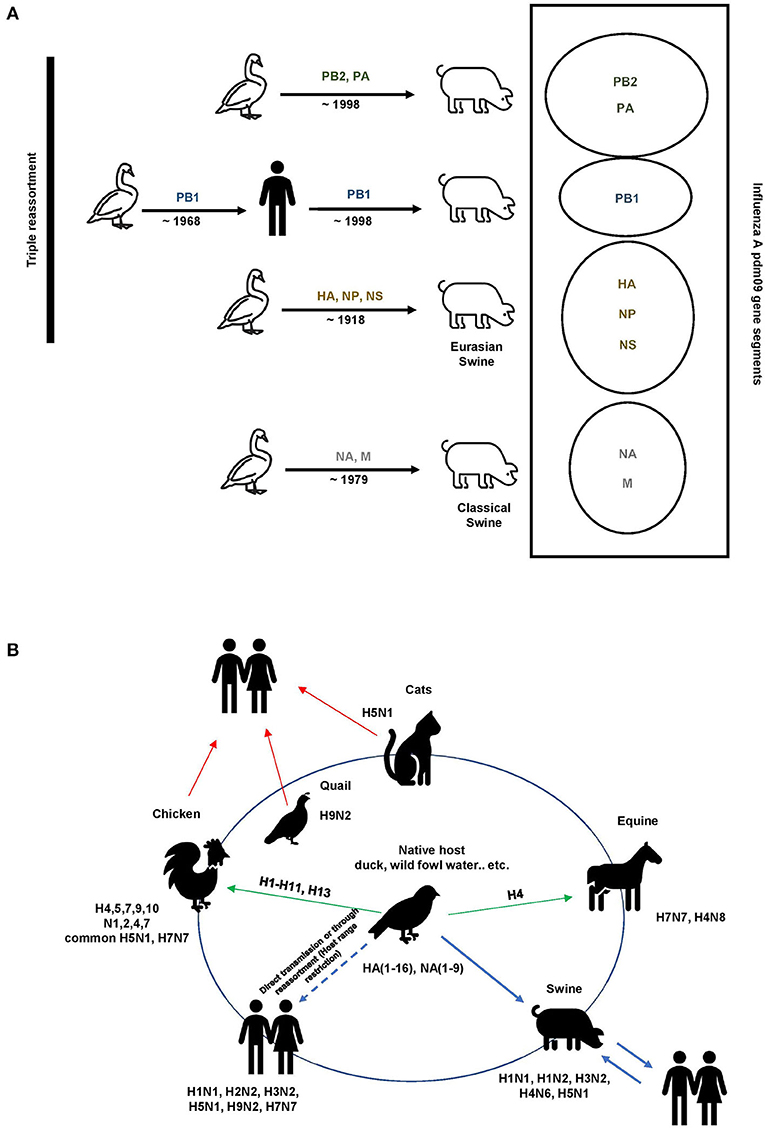
Figure 2. Influenza A evolution. (A) Triple reassortment influenza A viruses of the H1N1 subtype containing avian, swine, and human gene segments. The colored solid genes represent the gene segments as follows: yellow, classical swine A (H1N1) virus; green, North American avian virus; blue, human A (H3N2) virus; gray, Eurasian avian-like swine A(H1N1). (B) Reservoirs and interspecies transmission events of the pathogenic influenza A viruses. Wild birds, domestic birds, pigs, horses, and humans maintain their influenza A viruses. Spillover events occasionally occur, most frequently from wild birds (arrows in green).
SARS-CoV
The coronavirus family is so named because of the large spike protein molecules that are present on the virus surface and gives the virions a crown-like shape; coronavirus genomes are the largest among RNA viruses (29). This family has been classified into at least three primary genera (alpha, beta, and gamma). Within this family, seven viruses are currently known to infect humans, namely, NL63 and 229E from the alpha genus and OC43, HKU1, SARS-CoV, MERS-CoV, and SARS-CoV-2 from the beta genus. SARS-CoV is a positive-stranded RNA virus belonging to the family Coronaviridae (30), order Nidovirales, genus Betacoronavirus, lineage B (from the International Committee on Taxonomy of Viruses). It was characterized as a giant, enveloped, positive-stranded RNA virus with a genome comprising 29,727 nucleotides (~30 kb), 41% of which are guanine or cytosine. The genomic body of this virus has the original gene order of 5'-replicase (rep), which makes up approximately two-thirds of the genome and consists of the large genes ORF1a and ORF1b. ORF1a and ORF1b of the rep gene encode two large polyproteins known as pp1a (486 kDa) and pp1ab (790 kDa). In addition, the 3' structural spike (S), envelope (E), membrane (M), and nucleocapsid (N) proteins are encoded by four open reading frames (ORFs) downstream of the rep gene (31). The rep gene products are translated from genomic RNA, whereas the remaining viral proteins are translated from subgenomic mRNAs. In addition to the original genes, the SARS-CoV genome encodes another eight putative accessory proteins, known as ORFs 3a, 3b, 6, 7a, 7b, 8a, 8b, and 9b, which vary in length from 39 to 274 amino acids. Although the SARS-CoV rep gene and structural proteins have some sequence homology with other coronaviruses, the accessory proteins do not show substantial homology to the viral proteins of other coronaviruses at the amino acid level (31).
MERS-CoV
Although MERS-CoV belongs to the same family, order, and genus as SARS-CoV, it was the first betacoronavirus lineage C member identified as a “novel coronavirus” with a genome size of 30,119 nucleotides. The genome of MERS-CoV encodes 10 proteins. These 10 proteins comprise two replicase polyproteins (ORF1ab and ORF1a), four structural proteins (E, N, S, and M), and four nonstructural proteins (ORFs 3, 4a, 4b, and 5) (32). In addition to the rep and structural genes, there are accessory protein genes interspersed between the structural protein genes that may interfere with the host innate immune response in infected animals (7).
SARS-CoV-2
Although SARS-CoV-2 belongs to the same family and genus as SARS-CoV and MERS-CoV, genomic analysis revealed greater similarity between SARS-CoV-2 and SARS-CoV. Thus, researchers classified it as a member of lineage B (from the International Committee on Taxonomy of Viruses). Initially, the Coronaviridae Study Group of the International Committee on Taxonomy of Viruses identified this virus as a sister clade to the prototype human and bat severe acute respiratory syndrome coronaviruses (SARS-CoVs) of the species Severe acute respiratory syndrome-related coronavirus. Later, it was labeled as SARS-CoV-2 (33). The RNA genome size of SARS-CoV-2 is 30,000 bases in length. Among other betacoronaviruses, this virus is characterized by a unique combination of polybasic cleavage sites, a distinctive feature known to increase pathogenicity and transmissibility in other viruses (34).
Genomic analysis of SARS-CoV-2 revealed that the genome consists of six major ORFs and shares less than an 80% nucleotide sequence identity with SARS-CoV. However, the seven conserved replicase domains in the ORF1ab amino acid sequence share a 94.4% identity with those in SARS-CoV (35). Genomic analysis also revealed that the SARS-CoV-2 genome is highly similar to that of the bat coronavirus (Bat CoV RaTG13), with a sequence identity of 96.2%. Furthermore, the receptor-binding spike protein shares a 93.1% similarity to Bat CoV RaTG13 (35). Meanwhile, relative to SARS-CoV, significant differences were observed in the sequence of the S gene of SARS-CoV-2, including three short insertions in the N-terminal domain, changes in four out of five of the crucial residues in the receptor-binding motif, and the presence of an unexpected furin cleavage site at the S1/S2 boundary of the SARS-CoV-2 spike glycoprotein. This insertion is a novel feature that differentiates SARS-CoV-2 from SARS-CoV and several SARS-related coronaviruses (SARSr-CoVs) (36).
Viral Origin and Evolution
Influenza A
Influenza A H1N1 and H3N2 subtype viruses are two of the three combinations known to have circulated widely in humans and to currently cause seasonal influenza; these strains originated from birds and swine. Before 1979, the only lineage detected in swine herds from Europe was the classical swine influenza virus A H1N1 lineage 1A (25). This strain shares a mutual ancestor with the virus that caused the 1918 human influenza A pandemic. However, in the early 1980s, the classical swine H1N1 strain was displaced by a new European enzootic swine influenza A viral strain: the Eurasian, avian-like H1N1 (H1avN1) lineage 1C (26). After its rapid transmission from birds to mammals, the H1avN1 virus underwent rapid and sustained adaptation in mammals. Furthermore, this virus has also undergone rapid reassortment, resulting in the appearance of multiple genotypes. The two primary enzootic subtypes are H1N2 (H1huN2) lineage IB and H3N2, which occurred through the acquisition of HA or NA gene segments originating from seasonal human influenza viruses (Figure 2B) (37).
As previously mentioned, influenza A exhibits antigenic drift/shift phenomena resulting from the HA protein's ability to undergo rapid evolution because of the plasticity of the viral RNA-dependent RNA polymerase. It is believed that mutations occurring in the HA protein, including reassortments and mutations among animals and humans, were the drivers of previous pandemics (38).
Adaptive mutations can lead to a number of phenotypic changes, including variations in antigenicity, increased diversity in viral protein sequences, the ability to avoid antibody pressure, receptor preference, virulence, altered fusion functionality, and evasion of the immune response. Rapid modifications can give rise to new strains with features that are different from any viruses that have previously been confronted, potentially causing another epidemic/pandemic (38).
SARS-CoV
In the early stages of the SARS outbreak, most of the new patient cases had animal exposure before developing the disease. Wide-ranging investigations revealed that SARS-CoV strains were transmitted to palm civets from other animals (39–41). Later, two studies reported the discovery of coronaviruses related to human SARS-CoV, which were named SARS-like coronaviruses or SARSr-CoVs, in horseshoe bats (genus Rhinolophus) (42, 43). Another study revealed that the viral strains of the SARS-like coronaviruses contain all of the genetic elements that are needed to form SARS-CoV. In particular, the bat strain WIV16, the closest relative to SARS-CoV, likely occurred through recombination of two other prevalent bat SARSr-CoV strains. These results suggest that bats may be the natural reservoirs for the virus and that palm civets are only intermediate hosts (Supplementary Figure 1) (44, 45).
Thus, the hypothesis formed was that the direct ancestor of SARS-CoV was produced by recombination within bats and then transmitted to palm civets or other mammals via fecal–oral transmission. When virus-infected civets were transported to Guangdong market, the virus spread among the civets in the market and underwent further mutations before transmission to humans (46).
MERS-CoV
Unlike the SARS cases, most of the MERS cases had previous contact with dromedary camels. The MERS-CoV strains isolated from camels were almost identical to those isolated from humans (47, 48), and the MERS-CoV isolates were found to be highly prevalent in camels from the Middle East, Africa, and Asia (49, 50). Genomic sequence analysis indicated that the Tylonycteris bat coronaviruses HKU4 and HKU5 are phylogenetically related to MERS-CoV (they are all representatives of betacoronavirus lineage C) (51). Generally, all of the related MERS-CoVs isolated from bats support the hypothesis that MERS-CoV originated from bats (Supplementary Figure 1) (46).
SARS-CoV-2
Before the epidemic outbreak of COVID-19 in late January 2020, several patients had been exposed to different animals (from wild animals to poultry) at the Huanan seafood wholesale market. When the CDC declared the situation to be an epidemic, several studies identified potential reservoirs, but at present, the origin and evolution of SARS-CoV-2 remain debatable. The earliest genomic sequence analysis of SARS-CoV-2 indicated that it is a member of the genus Betacoronavirus and falls within the subgenus Sarbecovirus, which also includes SARS-CoV (9, 35, 52–54). As mentioned above, preliminary comparisons revealed that SARS-CoV-2 has an almost 79% similarity with SARS-CoV at the nucleotide sequence level and a 96% similarity with horseshoe bat RaTG13 (55–57). Correspondingly, a comparative study between the RmYN02 virus from Rhinolophus bats in Yunan Province, China, and SARS-CoV-2 indicated that RmYN02 was the closest relative to the long replicase gene of SARS-CoV-2 (~97% nucleotide sequence similarity) (35, 36).
Even though bats are likely to be the reservoir host for this virus, their general biological differences from humans make it feasible that other mammalian species acted as intermediate hosts, in which SARS-CoV-2 obtained some or all of the mutations needed for effective human transmission. One of the suspected intermediate hosts, the Malayan pangolin, harbors coronaviruses showing high similarity to SARS-CoV-2 in the receptor-binding domain, which contains mutations believed to promote binding to the angiotensin-converting enzyme 2 (ACE2) receptor and demonstrates a 97% amino acid sequence similarity. By contrast, the genomic similarity was more divergent from SARS-CoV-2 (~91%) at the whole genome level (Supplementary Figure 1) (58, 59).
Coronaviruses have lower mutation rates than other RNA viruses, especially influenza A viruses, and high rates of viral replication within hosts because of the 3′-to-5′ exoribonuclease activity associated with the nonstructural protein nsp.14 (36, 60). This protein has an RNA proofreading function and is responsible for coronaviruses' resistance to RNA mutagens (60, 61).
Receptor Binding of Viruses
The high unpredictability among influenza A viral strains and their HAs relates to the significant discrepancy among host cells in showing different vulnerabilities to viral infection. HA plays a role in mediating the binding of influenza A viruses to sialic acid host cell receptors (62). The receptor-binding site lies at the top of the R domain of HA and contains exceptionally variable antigenic binding loops (63). Once the virus is bound to the host receptor, endocytosis of the virus element occurs. Additionally, a pH-dependent membrane fusion process is significant in controlling the viral genome's release into the host cell. Influenza A viral strains and their HAs are very variable, which contributes to the significantly different vulnerabilities of host cells to viral infection (64).
Influenza A viruses have demonstrated dominant genomic mutations, such as those within the HA 220 loop (Q223) and the D222G and D222N mutations, in which aspartic acid (D) is replaced by glycine (G) or asparagine (N), respectively. The D222G mutation is responsible for a change in receptor-binding affinity that enables the virus to bind to α-2,6 and α-2,3 sialic acid receptors on the epithelial cells of the upper respiratory tract and ciliated epithelial cells in the lower respiratory tract, respectively (65, 66).
Although HA plays a crucial role in receptor binding and concurrent mutation capabilities, NA also has a key role in removing sialic acids from cellular receptors and from the new HA and NA on budding virions, which are sialylated as part of the glycosylation processes within the host cell (67). A balance between HA and NA is essential for viral fitness. Any mutations in HA or environmental changes, such as low pH conditions, can affect NA's activity against sialoglycans (68, 69).
The SARS-CoV trimeric spike protein facilitates coronavirus entry into host cells by binding to the host receptor and subsequently fusing the viral and host membranes. The spike protein consists of three segments, one of which is the ectodomain (70). The ectodomain is composed of two subunits: S1 and S2. The S1 subunit contains two individual domains, an N-terminal domain (NTD) and a C-domain, and each NTD or C-domain (sometimes both) binds to the host receptor to function as the receptor-binding domain (RBD). ACE2 is the host cell receptor of SARS-CoV and the primary target of deactivating antibodies. Several studies have shown that the binding affinity between the RBD of each SARS-CoV strain and ACE2 positively correlates with the contagion of different SARS-CoV strains in host cells (Supplementary Figure 2) (71, 72).
The MERS-CoV spike protein subunit S1 C-domain has also been identified as the RBD (73). However, unlike SARS-CoV, MERS-CoV uses a dipeptidyl peptidase 4 (DPP4) β-propeller as its receptor. Likewise, the RBD of MERS-CoV contains an accessory subdomain that functions as the receptor-binding motif (RBM). Although the RBD core structures are remarkably analogous between MERS-CoV and SARS-CoV, their RBMs are distinct and may result in the recognition of different receptors (Supplementary Figure 2) (73).
Since the outbreak of SARS-CoV-2, several studies have analyzed its genome and compared it with other coronaviruses, such as MERS-CoV and SARS-CoV (74, 75). The results of these studies have shown that SARS-CoV-2 has a similar RBD structure to that of SARS-CoV, despite amino acid variations at some key residues (9). Genomic comparison of SARS-CoV-2 with SARS-CoV and bat SARS-like coronaviruses revealed that the S1 subunits of the spike proteins have a sequence identity of ~75%, and recent experimental studies confirmed that ACE2 is the human receptor of SARS-CoV-2 (34). Therefore, it is essential to characterize the human receptor-binding capacity of SARS-CoV-2 to evaluate its human–human transmissibility. A recent study used the protein–protein docking method to measure the interaction between the SARS-CoV-2 spike RBD and ACE2; it was revealed that the SARS-CoV-2 human receptor-binding affinity was 73% of that of SARS-CoV, which suggests that SARS-CoV-2 binds to ACE2 with intermediate affinity (76) (Supplementary Figure 2).
Host Factors, Disease Severity, and Pathogenesis
Influenza, SARS, and MERS have caused major global health threats, and now the COVID-19 pandemic is rapidly spreading worldwide and is having a widespread and profound impact. Both viral and host factors determine the severity and clinical outcomes of the diseases caused by these viruses. Host factors include host immunity, age, sex, morbidities, and genetic variations.
Influenza infections can cause high morbidity and mortality rates in the elderly (65 or older) and young populations with comorbidities (Figure 1C). Pathogenesis following influenza A infection occurs in two stages. The first stage is defined by the peak viral titer, along with the peak amount of inflammation associated with the infection, and lasts ~1 to 3 days. In the second stage, the infection progresses in some patients, and in severe cases, it may be associated with acute respiratory distress syndrome and sometimes death (77). Once a patient is infected with an influenza A virus, the humoral immune response will release neutralizing antibodies to target the influenza HA protein by blocking the binding of HA to sialic acids, thereby preventing viral fusion, inhibiting the release of offspring virions, and delaying proteolytic cleavage of HA by host receptors (78).
Once a patient is infected with SARS-CoV, MERS-CoV, or SARS-CoV-2, the host innate immune system will identify the virus by using pattern recognition receptors, such as a toll-like receptor, NOD-like receptor, or RIG-I-like receptor, to recognize pathogen-associated molecular patterns. The adaptive immune response also plays a significant antiviral role by stabilizing the host defense mechanism against pathogens and minimizing the risk of developing an autoimmune reflex response or inflammation (9, 79). In general, human coronaviruses can be classified into two types: lowly pathogenic and highly pathogenic. Viruses with low pathogenicity, including HCoV-229E, HCoV-OC43, HCoV-NL63, and HCoV-HKU, can cause mild upper respiratory tract infections. In contrast, highly pathogenic viruses, including SARS-CoV, MERS-CoV, and SARS-CoV-2, can cause lower respiratory tract infections, severe pneumonia, and sometimes fatal acute lung injury or acute respiratory distress syndrome, especially in older individuals (≥65 years old) (Figure 1C) (80).
In addition to the lungs, coronavirus infection may damage other organs or tissues, including the gastrointestinal tract (81), spleen, lymph nodes, brain, skeletal muscles, thyroid, and heart (82, 83). The destruction of lung cells prompts a local immune response, engaging macrophages and monocytes that respond to the infection, release cytokines, and enhance adaptive T and B cell immune responses. In some cases, a dysfunctional immune response occurs, which can cause severe lung and systemic pathology. The invading coronavirus may incite host immune responses, and an excessive immune response may cause immunopathological damage (known as a cytokine storm) in patients with coronavirus infections (9, 84). Cytokine storms may enhance the infiltration of non-neutralizing antiviral proteins that facilitate viral entry into host cells, leading to increased viral infectivity (82, 85). Therefore, cytokine storms play a key role in the pathogenesis and clinical outcomes of patients with coronavirus infection.
Transmissibility and Virulence
The initiation of a pandemic requires the rise of a virus in a human population in which there is little or no pre-existing immunity, and the virus must be able to persist through human-to-human transmission (86, 87). The ability of influenza A viruses to adapt to various hosts and undergo reassortment events ensures the constant generation of new strains. These strains have variable degrees of pathogenicity, pandemic transmissibility, and reproduction numbers (R0) (Table 1) (88). However, only three subtypes of influenza A (H1–H3) have acquired the properties to cause pandemics in the last two centuries. Thus, an understanding of the capability of a virus to attain a contagious phenotype is a critical factor in evaluating the pandemic potential of novel subtypes (89, 90). The use of animal models has facilitated detailed studies of influenza A virus transmission by the contact and respiratory droplet routes. The presence of a single sick individual in a small space, such as an airplane or room, has been shown to be adequate for an outbreak among healthy individuals (Supplementary Figure 3) (91). Although infection and case fatality rates vary from one pandemic to another, the rates of influenza A virus infections in the pandemics were high, especially among people with little to no pre-existing immunity. When pandemic viruses become established in humans, their effective seasonal spread among healthy individuals eventually provides an enduring and even more significant public health issue in terms of hospitalizations and, in some cases, fatalities. Particle size (92), the distance of spread (92), disposition (92, 93), temperature (94), and relative humidity (95) are all considered to be factors that influence the rate of transmissibility of influenza A viruses. In addition, sialic acid receptors (α-2,3 and α-2,6) can affect the general species-specific cellular tropism of influenza A viruses (63).
Contaminated surfaces also play an essential role in transmission. A respiratory pathogen can survive on surfaces, be transferred to hands or other equipment, and initiate infection through contact with the eyes, nose, or mouth (Supplementary Figure 3) (96). Influenza A has been shown to survive for 24–48 h on stainless steel and plastic surfaces. Inversely, the strains survived for <8–12 h on cloth, paper, and tissues. Quantifiable amounts of influenza A viruses were observed to be transmitted from stainless steel surfaces to hands after 24 h and from tissues to hands for up to 15 min. Viruses also survive on hands for up to 5 min after transfer from environmental surfaces. These results indicate a high transmission rate for influenza A viruses (97).
SARS-CoV, MERS-CoV, and SARS-CoV-2 can survive on surfaces for extended periods, sometimes up to months. Like the influenza A viruses, the factors affecting the survival of these viruses on surfaces include the strain variation, titer, surface type, mode of deposition, temperature, humidity, and method used to determine the viability of the virus (98, 99). Several studies have indicated that SARS-CoV, MERS-CoV, and SARS-CoV-2 can survive on dry surfaces for a sufficient duration to accelerate onward transmission. Viable MERS-CoV was detected on steel and plastic surfaces after 48 h at 20°C with 40% relative humidity, with a decreased viability of about 8 h at 30°C with 80% relative humidity and of about 24 h at 30°C with 30% relative humidity. The estimated half-life of MERS-CoV ranges from ~0.5 to 1 h (98). On the other hand, another study conducted on the viability of SARS-CoVs detected on plastic surfaces and on polystyrene Petri dishes revealed that the virus survived for more than 5 days and more than 20 days, respectively, at room temperature. The viral viability was constant at lower temperatures (28°C) and lower humidity (80–89%) (100), whereas survival times ranged from 5 min to 2 days on paper, disposable gowns, and cotton gowns (99).
Since the SARS-CoV-2 outbreak began, several researchers have attempted to analyze the survival time of this virus on different surfaces. One study published in the middle of March 2020 analyzed the aerosol and surface stabilities of SARS-CoV-2 and SARS-CoV. The study utilized five different environments (aerosols, plastic, stainless steel, copper, and cardboard). The results showed that the half-lives of SARS-CoV-2 and SARS-CoV were similar in aerosols and on copper. However, on cardboard surfaces, the half-life of SARS-CoV-2 was longer than that of SARS-CoV, and the highest levels of viability for both viruses were observed on stainless steel and plastic (~5.6 h on stainless steel and 6.8 h on plastic). The researchers concluded that the differences in the epidemiological characteristics of these viruses could result from other factors and that aerosol and fomite transmission of SARS-CoV-2 is probable because the virus can remain viable and infectious in aerosols and on surfaces for hours and hours to days, respectively (101).
The effective management and control of such infections are increasingly performed with extensive contributions from mathematical modeling, which not only provides information on the nature of the infection itself but also makes predictions about the likely outcome of alternative courses of action (102). One useful mathematical model is the reproductive number R0, which is defined as the average number of secondary cases generated per typical infectious case (103). A value of R0 > 1 indicates that the infection may persist or grow in the population, whereas a value of R0 < 1 indicates that this infection will decrease in the population, although exceptions occur (103). The majority of seasonal influenza R0 values have been calculated for different populations and different continents, such as Europe and North America, with a median point estimate of R0 = 1.27 (IQR: 1.19–1.37) (104). The initial estimations of the reproduction numbers of SARS-CoV and MERS-CoV were calculated for China and the Middle East with R0 median = 0.58 (IQR: 0.24–1.18) (105) and R0 mean = 0.69 (95% CI: 0.50–0.92) (106), respectively. However, among the four viruses, SARS-CoV-2 has been calculated to be the most contagious, such as the R0 value associated with the Italian outbreak with a median point estimate of R0 = 3.1 (coefficient of determination, r2 = 0.99) (107).
Prevention, Control, and Treatment of Virus Infection
Strategies for preventing and controlling pandemic/epidemic viruses can be improved by being well-prepared. Preparedness strategies, which primarily include the quarantine of infected persons, self-protection (wearing facemasks, using disinfectants, washing hands, and disinfecting surfaces with bleach or alcohols), and social distancing are all considered to be important for a comprehensive plan that can be tested and promoted by conducting exercises to engage the whole of society.
An influenza pandemic can be catastrophic, and in a typical year of seasonal outbreaks, influenza A viruses cause as many as 5 million cases of severe illness in humans and over 500,000 deaths. After the first confirmed cases of H1N1 influenza appeared in Mexico in February 2009, cases began to spread to the United States, and by the end of April 2009, cases had been reported in several United States cities and other countries on various continents, such as Canada, the United Kingdom, and New Zealand (108). During the last pandemic, the first activation of the International Health Regulations (IHR) provisions was prompted. The discussions that led to the IHR implementation were based on the SARS outbreak experience in 2003. These regulations describe the responsibilities of individual countries and the leadership role of the WHO in declaring and managing a public health emergency of international concern, establishing systematic approaches to surveillance, promoting technical cooperation, and sharing logistic support (108). However, because of the significant diversity of influenza viruses in animal hosts, extensive experimental testing and the development of pandemic preparedness measures against all viruses is unachievable (109).
In this regard, the WHO periodically updates the influenza risk management and preparedness plan, and the latest guidance document, Pandemic Influenza Risk Management (PIRM), was released in May 2017 (110). This updated document supports national and global pandemic preparedness and risk management and utilizes lessons learned at the country, regional, and global levels (110). Furthermore, several WHO preparedness documents have been released since PIRM, such as Essential steps for developing or updating a national pandemic influenza preparedness plan (released in March 2018) and A practical guide for developing and conducting simulation exercises to test and validate pandemic influenza preparedness plans (published in September 2018) (111).
During the SARS epidemic, more than 8,000 people were infected, and 774 deaths occurred between November 2002 and December 2003. SARS is highly contagious and is transmitted primarily by respiratory droplets; the highest transmission rates of SARS occurred in healthcare facilities (112). At the end of the SARS outbreak, the cases of over 1,700 healthcare workers who had been affected were reported to the WHO, from China (19% of total cases), Canada (43%), France (29%), and Hong Kong (22%). During this epidemic, insufficient or inappropriate infection control measures, such as inconsistent use of personal protective equipment, reuse of N95 masks, and lack of adequate infection control, were related to the high risk of infection among healthcare workers (113). Thus, in 2004, after the epidemic was contained, the WHO released a framework that was prepared according to the six phases of an epidemic, moving from preparedness, planning, and routine surveillance for cases, through to the prevention of the consequent international spread, to the disruption of global transmission (114).
Since 2012, 27 countries have reported cases of MERS; Saudi Arabia has reported ~80% of human cases, and more than 50% of the cases in healthcare workers were nurses (115). The WHO, in collaboration with the Food and Agriculture Organization of the United Nations (FAO), the World Organization for Animal Health (OIE), and national governments, have been working with healthcare workers and scientists in affected countries to gather and share scientific evidence based on the previous coronavirus epidemic. This information gathering process has been beneficial for better understanding of the virus and the disease it causes and for the regulation of outbreak response priorities, treatment approaches, and clinical management tactics (113).
Although accumulated knowledge and risk preparedness from the influenza pandemics and SARS/MERS epidemics allowed researchers to examine the effectiveness of strategic plans in dealing with the ongoing pandemic of COVID-19, several challenges have been raised in preventing the spread of COVID-19, such as the lack of medical supplies and laboratory facilities for the assessment of the disease and the presentation of a high number of asymptomatic cases. In response to the announcement of the emergency, governments were bound by the IHR to disclose vital information regarding the identification and detection of COVID-19, regardless of the causative agent. Within the context of the Global Humanitarian Response Plan, a Health Cluster platform has been created to assess the response to the COVID-19 pandemic worldwide. This framework has adopted the following strategies: contain the spread of the COVID-19 pandemic and decrease morbidity and mortality; decrease the deterioration of human assets and rights, social cohesion, and livelihoods; and protect, assist, and advocate for refugees, internally displaced people, migrants, and host communities who are particularly vulnerable to the pandemic (source: WHO). The primary goal of the Health Cluster is to coordinate and support partners to fulfill essential health services to achieve the framework strategies. This goal is achieved by different roles and tasks, such as by raising awareness, alertness, and response planning at the country level and by conducting training and simulation exercises. The WHO Health Cluster framework is a gateway to useful resources to support COVID-19 preparedness and response (116).
Generally, each pandemic/epidemic has presented a public health emergency of uncertain scope and effect; thus, essential elements of current approaches to pandemic preparedness and extenuation, such as the development of vaccines and stockpiling of antiviral drugs, necessitate detailed virological and immunological data on viruses with apparent pandemic potential. However, the development of vaccines against new strains is challenging. Therefore, physicians and health workers have found themselves facing the massive challenge of preventing infections or stabilizing patients' conditions. Thus, several promising attempts have been made to utilize different antiviral treatments that have already been approved by the U.S. Food and Drug Administration (FDA) for the treatment of viral pneumonia infections. A list of antiviral drugs and vaccine approaches for influenza viruses, SARS-CoV, MERS-CoV, and SARS-CoV-2 that have been used in clinics or are undergoing clinical trials are summarized in Table 2.
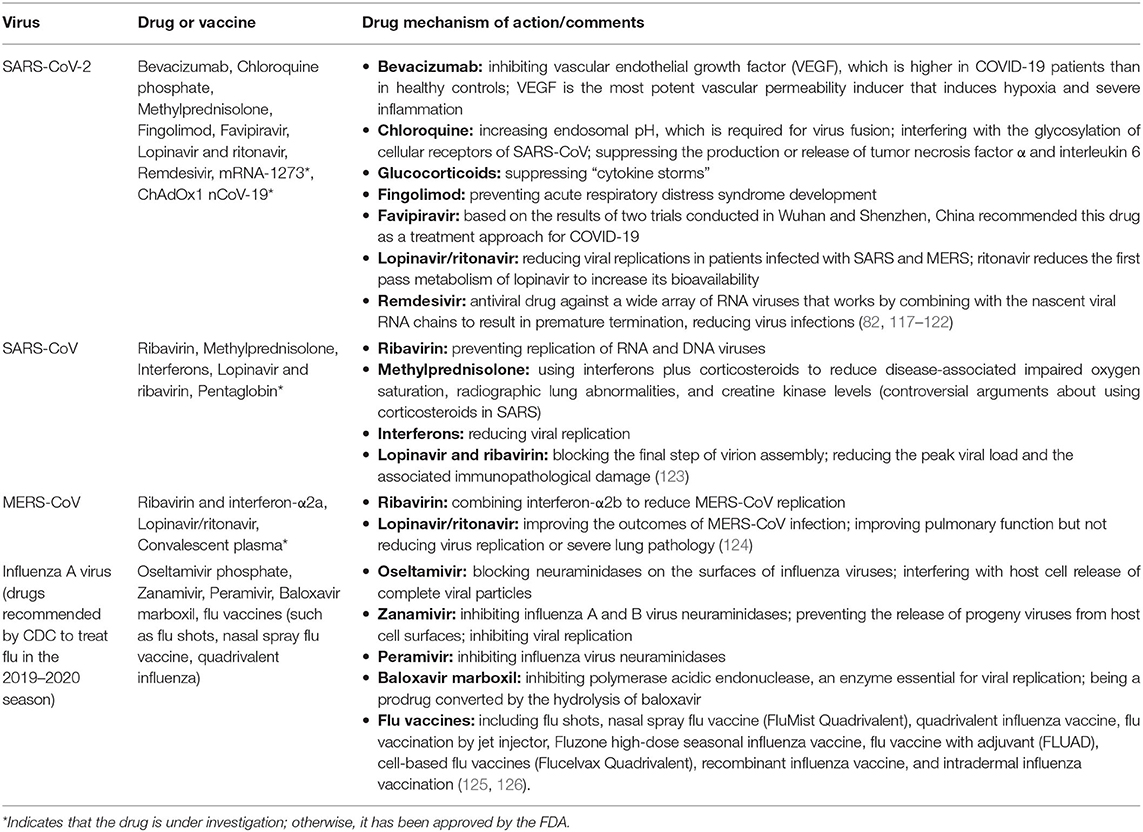
Table 2. List of antiviral drugs and vaccine approaches for SARS-CoV-2, SARS-CoV, MERS-CoV, and influenza viruses.
Discussion and Conclusion
Although the mode of transmission for SARS-CoV-2 is still somewhat unclear, all four viruses are thought to be transmitted by the same mechanism. Infection via respiratory droplets or secretions of infected individuals is the primary mode of transmission between humans. The spread of infection is occurring more rapidly for the current outbreak than in the SARS and MERS epidemics, although rates of human-to-human transmission were generally lower for MERS.
The CFRs across the four viruses range from 0.1 to 35% (Table 1), with the highest rate for MERS cases and the lowest for seasonal influenza; however, it is essential to note that the CFR for COVID-19 should be interpreted carefully because the outbreak is still ongoing.
With the exception of the influenza A viruses, the other viruses (SARS-CoV, MERS-CoV, and SARS-CoV-2) are similar in zoonotic transmission. The MERS-CoV reservoir hosts are dromedary camels, and the SARS-CoV reservoir hosts are likely bats. It is still unclear whether SARS-CoV-2 was zoonotically transmitted from an infected palm civet, snake, or other animal at the Chinese seafood market.
Regarding the origin of the virus, SARS-CoV and SARS-CoV-2 originate from China and share a high degree of similarity, including exposure to wild animals, whereas MERS-CoV and SARS-CoV-2 have shared similarities in that cases can remain asymptomatic while still spreading the disease. Furthermore, influenza A viruses and SARS-CoV-2 also have a similar characteristic when it comes to transmissibility (127).
In the setting of extensive SARS-CoV-2 transmissions, the possibility of SARS-CoV-2 should be considered in all persons with a fever or lower respiratory infection, because it is challenging to straightforwardly distinguish between seasonal influenza and COVID-19, even if an epidemiologic link cannot be readily established. Furthermore, the timely reporting of cases, updates on clinical status and disposition of patients, the real-time analysis of data, and the appropriate dissemination of information are essential for outbreak-managing decisions.
Author Contributions
ZA: conceptualization, methodology, investigation, writing—original draft, and visualization. ML: visualization. XW: conceptualization, methodology, project administration, funding acquisition, writing—review and editing, and supervision. All authors contributed to the article and approved the submitted version.
Funding
This work was supported by the China Pharmaceutical University (grant number 3150120001 to XW).
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank China Pharmaceutical University for its support and funding.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fimmu.2020.552909/full#supplementary-material
Supplementary Figure 1. The origins and intermediate hosts of SARS-CoV-2, SARS-CoV, and MERS-CoV.
Supplementary Figure 2. Virus-host interaction. Th1, T helper 1; Th17, T helper 17; ACE2, angiotensin-converting enzyme 2; INF-1, interferon 1; INFγ, interferon gamma; DPP4, dipeptidyl peptidase-4; HA, hemagglutinin; NA, neuraminidase; M2e, Matrix 2 protein; MHC-1, major histocompatibility complex class 1.
Supplementary Figure 3. Potential transmission routes of respiratory infection between infected and susceptible individuals (128). Respiratory infections with a droplet nuclei size ≤ 5 μm can travel to a distance ≥1 m. In contrast, respiratory infections with a droplet nuclei size ≥5 μm cannot travel to a distance ≥1 m. Large droplets may fall on different surfaces and infect healthy individuals through direct or indirect contact.
Abbreviations
SARS-CoV-2, severe acute respiratory syndrome coronavirus 2; SARS-CoV, severe acute respiratory syndrome coronavirus; MERS-CoV, the Middle East respiratory syndrome coronavirus; WHO, world health organization; CDC, center of disease control and prevention; nt, nucleotide; kb, kilobase; KDa, kilodalton molecular weight unit.
References
1. Jordan D. The Deadliest Flu: The Complete Story of the Discovery and Reconstruction of the 1918 Pandemic Virus. Centers for Disease Control and Prevention, National Center for Immunization and Respiratory Diseases (NCIRD), December 17 (2019). Available online at: https://www.cdc.gov/flu/pandemic-resources/1918-pandemic-h1n1.html (accessed March 19, 2020).
2. Glezen WP. Emerging infections: pandemic influenza. Epidemiol Rev. (1996) 18:64–76. doi: 10.1093/oxfordjournals.epirev.a017917
3. Viboud C, Grais RF, Lafont BA, Miller MA, Simonsen L. Multinational impact of the 1968 Hong Kong influenza pandemic: evidence for a smoldering pandemic. J Infect Dis. (2005) 192:233–48. doi: 10.1086/431150
4. Garten RJ, Davis CT, Russell CA, Shu B, Lindstrom S, Balish A, et al. Antigenic and genetic characteristics of swine-origin 2009 A(H1N1) influenza viruses circulating in humans. Science. (2009) 325:197–201. doi: 10.1126/science.1176225
5. Shieh WJ, Blau DM, Denison AM, Deleon-Carnes M, Adem P, Bhatnagar J, et al. 2009 pandemic influenza A (H1N1): pathology and pathogenesis of 100 fatal cases in the United States. Am J Pathol. (2010) 177:166–75. doi: 10.2353/ajpath.2010.100115
6. Peiris JSM, Yuen KY, Osterhaus ADME, Stöhr K. The severe acute respiratory syndrome. N Engl J Med. (2003) 349:2431–41. doi: 10.1056/NEJMra032498
7. Zumla A, Hui DS, Perlman S. Middle East respiratory syndrome. Lancet. (2015) 386:995–1007. doi: 10.1016/S0140-6736(15)60454-8
8. The Lancet. Emerging understandings of 2019-nCoV. Lancet. (2020) 395:311. doi: 10.1016/S0140-6736(20)30186-0
9. Lu R, Zhao X, Li J, Niu P, Yang B, Wu H, et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet. (2020) 395:565–74. doi: 10.1016/S0140-6736(20)30251-8
10. Xiao K, Zhai J, Feng Y, Zhou N, Zhang X, Zou JJ, et al. Isolation of SARS-CoV-2-related coronavirus from Malayan pangolins. Nature. (2020) 583:286–9. doi: 10.1038/s41586-020-2313-x
11. Wang H, Yang P, Liu K, Guo F, Zhang Y, Zhang G, et al. SARS coronavirus entry into host cells through a novel clathrin- and caveolae-independent endocytic pathway. Cell Res. (2008) 18:290–301. doi: 10.1038/cr.2008.15
12. Millet JK, Whittaker GR. Host cell entry of Middle East respiratory syndrome coronavirus after two-step, furin-mediated activation of the spike protein. Proc Natl Acad Sci USA. (2014) 111:15214–9. doi: 10.1073/pnas.1407087111
13. Lakadamyali M, Rust MJ, Zhuang X. Endocytosis of influenza viruses. Microbes Infect. (2004) 6:929–36. doi: 10.1016/j.micinf.2004.05.002
14. Xu Z, Shi L, Wang Y, Zhang J, Huang L, Zhang C, et al. Pathological findings of COVID-19 associated with acute respiratory distress syndrome. Lancet Respir Med. (2020) 8:420–2. doi: 10.1016/S2213-2600(20)30076-X
15. Yang M, Hon KL, Li K, Fok TF, Li CK. The effect of SARS coronavirus on blood system: its clinical findings and the pathophysiologic hypothesis. Zhongguo shi Yan Xue Ye Xue Za Zhi. (2003) 11:217–21.
16. Park GE, Kang CI, Ko JH, Cho SY, Ha YE, Kim YJ, et al. Differential cell count and CRP level in blood as predictors for middle east respiratory syndrome coronavirus infection in acute febrile patients during nosocomial outbreak. J Korean Med Sci. (2017) 32:151–4. doi: 10.3346/jkms.2017.32.1.151
17. Perera RA, Wang P, Gomaa MR, El-Shesheny R, Kandeil A, Bagato O, et al. Seroepidemiology for MERS coronavirus using microneutralisation and pseudoparticle virus neutralisation assays reveal a high prevalence of antibody in dromedary camels in Egypt, June 2013. Euro Surveill. (2013) 18:20574. doi: 10.2807/1560-7917.ES2013.18.36.20574
18. Lee N, Le Sage V, Nanni AV, Snyder DJ, Cooper VS, Lakdawala SS. Genome-wide analysis of influenza viral RNA and nucleoprotein association. Nucleic Acids Res. (2017) 45:8968–77. doi: 10.1093/nar/gkx584
19. Influenza A Model Receives A Face-Lift. (2020). Available online at: https://www.contagionlive.com/news/influenza-a-model-receives-a-face-lift
20. Vincent A, Awada L, Brown I, Chen H, Claes F, Dauphin G, et al. Review of influenza A virus in swine worldwide: a call for increased surveillance and research. Zoonoses Public Health. (2014) 61:4–17. doi: 10.1111/zph.12049
21. Tian J, Zhang C, Qi W, Xu C, Huang L, Li H, et al. Genome sequence of a novel reassortant H3N2 avian influenza virus in southern China. J Virol. (2012) 86:9553–4. doi: 10.1128/JVI.01523-12
22. Bouvier NM, Palese P. The biology of influenza viruses. Vaccine. (2008) 26(Suppl. 4):D49–53. doi: 10.1016/j.vaccine.2008.07.039
23. Ahn I, Jeong BJ, Bae SE, Jung J, Son HS. Genomic analysis of influenza A viruses, including avian flu (H5N1) strains. Eur J Epidemiol. (2006) 21:511–9. doi: 10.1007/s10654-006-9031-z
24. Yoon SW, Webby RJ, Webster RG. Evolution and ecology of influenza A viruses. Curr Top Microbiol Immunol. (2014) 385:359–75. doi: 10.1007/82_2014_396
25. Reperant LA, Kuiken T, Osterhaus ADME. Influenza viruses. Hum Vaccines Immunother. (2012) 8:7–16. doi: 10.4161/hv.8.1.18672
26. Anderson TK, Macken CA, Lewis NS, Scheuermann RH, Van Reeth K, Brown IH, et al. A phylogeny-based global nomenclature system and automated annotation tool for H1 hemagglutinin genes from swine influenza A viruses. mSphere. (2016) 1:e00275-16. doi: 10.1128/mSphere.00275-16
27. Guarnaccia T, Carolan LA, Maurer-Stroh S, Lee RT, Job E, Reading PC, et al. Antigenic drift of the pandemic 2009 A(H1N1) influenza virus in A ferret model. PLoS Pathog. (2013) 9:e1003354. doi: 10.1371/journal.ppat.1003354
28. Tewawong N, Prachayangprecha S, Vichiwattana P, Korkong S, Klinfueng S, Vongpunsawad S, et al. Assessing antigenic drift of seasonal influenza A(H3N2) and A(H1N1)pdm09 viruses. PloS ONE. (2015) 10:e0139958. doi: 10.1371/journal.pone.0139958
29. Pellett PE, Mitra S, Holland TC. Chapter 2 - basics of virology. In: Tselis AC, Booss J, editors. Handbook of Clinical Neurology. 123: Michigan City, IN: Elsevier (2014). p. 45–66.
30. Torres J, Maheswari U, Parthasarathy K, Ng L, Liu DX, Gong X. Conductance and amantadine binding of a pore formed by a lysine-flanked transmembrane domain of SARS coronavirus envelope protein. Prot Sci. (2007) 16:2065–71. doi: 10.1110/ps.062730007
31. Tan Y-J, Lim SG, Hong W. Understanding the accessory viral proteins unique to the severe acute respiratory syndrome (SARS) coronavirus. Antiviral Res. (2006) 72:78–88. doi: 10.1016/j.antiviral.2006.05.010
32. Chung YS, Kim JM, Man Kim H, Park KR, Lee A, Lee NJ, et al. Genetic characterization of middle east respiratory syndrome coronavirus, South Korea, (2018) Emerg Infect Dis. (2019) 25:958–62. doi: 10.3201/eid2505.181534
33. Grifoni A, Weiskopf D, Ramirez SI, Mateus J, Dan JM, Moderbacher CR, et al. Targets of T cell responses to SARS-CoV-2 coronavirus in humans with COVID-19disease and unexposed individuals. Cell. (2020) 181:1489–501.e15. doi: 10.1016/j.cell.2020.05.015
34. Walls AC, Park YJ, Tortorici MA, Wall A, McGuire AT, Veesler D. Structure, function, and antigenicity of the SARS-CoV-2 spike glycoprotein. Cell. (2020) 181:281–92.e6. doi: 10.1016/j.cell.2020.02.058
35. Chen J, Liu D, Liu L, Liu P, Xu Q, Xia L, et al. A pilot study of hydroxychloroquine in treatment of patients with common coronavirus disease-19 (COVID-19). J Zhejiang Univ Med Sci. (2020) 49:215–19. doi: 10.3785/j.issn.1008-9292.2020.03.03
36. Andersen KG, Rambaut A, Lipkin WI, Holmes EC, Garry RF. The proximal origin of SARS-CoV-2. Nat Med. (2020) 26:450–2. doi: 10.1038/s41591-020-0820-9
37. Brown IH. History and epidemiology of Swine influenza in Europe. Curr Top Microbiol Immunol. (2013) 370:133–46. doi: 10.1007/82_2011_194
38. Castelán-Vega JA, Magaña-Hernández A, Jiménez-Alberto A, Ribas-Aparicio RM. The hemagglutinin of the influenza A(H1N1)pdm09 is mutating towards stability. Adv Appl Bioinform Chem. (2014) 7:37–44. doi: 10.2147/AABC.S68934
39. Guan Y, Zheng BJ, He YQ, Liu XL, Zhuang ZX, Cheung CL, et al. Isolation and characterization of viruses related to the SARS coronavirus from animals in southern China. Science. (2003) 302:276–8. doi: 10.1126/science.1087139
40. Tu C, Crameri G, Kong X, Chen J, Sun Y, Yu M, et al. Antibodies to SARS coronavirus in civets. Emerg Infect Dis. (2004) 10:2244–8. doi: 10.3201/eid1012.040520
41. Wang M, Xu HF, Zhang ZB, Zou XZ, Gao Y, Liu XN, et al. [Analysis on the risk factors of severe acute respiratory syndromes coronavirus infection in workers from animal markets]. Zhonghua Liu Xing Bing Xue Za Zhi. (2004) 25:503–5.
42. Lau SK, Woo PC, Li KS, Huang Y, Tsoi HW, Wong BH, et al. Severe acute respiratory syndrome coronavirus-like virus in Chinese horseshoe bats. Proc Natl Acad Sci USA. (2005) 102:14040–5. doi: 10.1073/pnas.0506735102
43. Li W, Shi Z, Yu M, Ren W, Smith C, Epstein JH, et al. Bats are natural reservoirs of SARS-like coronaviruses. Science. (2005) 310:676–9. doi: 10.1126/science.1118391
44. Hu B, Zeng LP, Yang XL, Ge XY, Zhang W, Li B, et al. Discovery of a rich gene pool of bat SARS-related coronaviruses provides new insights into the origin of SARS coronavirus. PLoS Pathog. (2017) 13:e1006698. doi: 10.1371/journal.ppat.1006698
45. Wang MN, Zhang W, Gao YT, Hu B, Ge XY, Yang XL, et al. Longitudinal surveillance of SARS-like coronaviruses in bats by quantitative real-time PCR. Virol Sin. (2016) 31:78–80. doi: 10.1007/s12250-015-3703-3
46. Cui J, Li F, Shi Z-L. Origin and evolution of pathogenic coronaviruses. Nat Rev Microbiol. (2019) 17:181–92. doi: 10.1038/s41579-018-0118-9
47. Raj VS, Farag EA, Reusken CB, Lamers MM, Pas SD, Voermans J, et al. Isolation of MERS coronavirus from a dromedary camel, Qatar, (2014) Emerg Infect Dis. (2014) 20:1339–42. doi: 10.3201/eid2008.140663
48. Chu DKW, Hui KPY, Perera R, Miguel E, Niemeyer D, Zhao J, et al. MERS coronaviruses from camels in Africa exhibit region-dependent genetic diversity. Proc Natl Acad Sci USA. (2018) 115:3144–9. doi: 10.1073/pnas.1718769115
49. Alagaili AN, Briese T, Mishra N, Kapoor V, Sameroff SC, Burbelo PD, et al. Middle East respiratory syndrome coronavirus infection in dromedary camels in Saudi Arabia. mBio. (2014) 5:e00884-14. doi: 10.1128/mBio.01002-14
50. Harcourt JL, Rudoler N, Tamin A, Leshem E, Rasis M, Giladi M, et al. The prevalence of Middle East respiratory syndrome coronavirus (MERS-CoV) antibodies in dromedary camels in Israel. Zoonoses Public Health. (2018) 65:749–54. doi: 10.1111/zph.12482
51. Lau SK, Li KS, Tsang AK, Lam CS, Ahmed S, Chen H, et al. Genetic characterization of Betacoronavirus lineage C viruses in bats reveals marked sequence divergence in the spike protein of pipistrellus bat coronavirus HKU5 in Japanese pipistrelle: implications for the origin of the novel Middle East respiratory syndrome coronavirus. J Virol. (2013) 87:8638–50. doi: 10.1128/JVI.01055-13
52. Wu F, Zhao S, Yu B, Chen YM, Wang W, Song ZG, et al. A new coronavirus associated with human respiratory disease in China. Nature. (2020) 579:265–9. doi: 10.1038/s41586-020-2008-3
53. Wu F, Zhao S, Yu B, Chen YM, Wang W, Song ZG, et al. Author correction: a new coronavirus associated with human respiratory disease in China. Nature. (2020) 580:E7. doi: 10.1038/s41586-020-2202-3
54. Zhu N, Zhang D, Wang W, Li X, Yang B, Song J, et al. A novel coronavirus from patients with pneumonia in China, (2019) N Engl J Med. (2020) 382:727–33. doi: 10.1056/NEJMoa2001017
55. Zhang YZ, Holmes EC. A genomic perspective on the origin and emergence of SARS-CoV-2. Cell. (2020) 181:223–7. doi: 10.1016/j.cell.2020.03.035
56. Coutard B, Valle C, de Lamballerie X, Canard B, Seidah NG, Decroly E. The spike glycoprotein of the new coronavirus 2019-nCoV contains a furin-like cleavage site absent in CoV of the same clade. Antiviral Res. (2020) 176:104742. doi: 10.1016/j.antiviral.2020.104742
57. Wrapp D, Wang N, Corbett KS, Goldsmith JA, Hsieh CL, Abiona O, et al. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science. (2020) 367:1260–3. doi: 10.1126/science.abb2507
58. Zhang T, Wu Q, Zhang Z. Probable pangolin origin of SARS-CoV-2 associated with the COVID-19 outbreak. Curr Biol. (2020) 30:1346–51.e2. doi: 10.1016/j.cub.2020.03.022
59. Kasibhatla SM, Kinikar M, Limaye S, Kale MM, Kulkarni-Kale U. Understanding evolution of SARS-CoV-2: a perspective from analysis of genetic diversity of RdRp gene. J Med Virol. (2020). doi: 10.1002/jmv.25909. [Epub ahead of print].
60. Minskaia E, Hertzig T, Gorbalenya AE, Campanacci V, Cambillau C, Canard B, et al. Discovery of an RNA virus 3'->5' exoribonuclease that is critically involved in coronavirus RNA synthesis. Proc Natl Acad Sci USA. (2006) 103:5108–13. doi: 10.1073/pnas.0508200103
61. Agostini ML, Andres EL, Sims AC, Graham RL, Sheahan TP, Lu X, et al. Coronavirus susceptibility to the antiviral remdesivir (GS-5734) is mediated by the viral polymerase and the proofreading exoribonuclease. mBio. (2018) 9e00221–18. doi: 10.1128/mBio.00221-18
62. Xiong X, Martin SR, Haire LF, Wharton SA, Daniels RS, Bennett MS, et al. Receptor binding by an H7N9 influenza virus from humans. Nature. (2013) 499:496–9. doi: 10.1038/nature12372
63. Skehel JJ, Wiley DC. Receptor binding and membrane fusion in virus entry: the influenza hemagglutinin. Ann Rev Biochem. (2000) 69:531–69. doi: 10.1146/annurev.biochem.69.1.531
64. Mair CM, Ludwig K, Herrmann A, Sieben C. Receptor binding and pH stability - how influenza A virus hemagglutinin affects host-specific virus infection. Biochim Biophys Acta. (2014) 1838:1153–68. doi: 10.1016/j.bbamem.2013.10.004
65. Baldo V, Bertoncello C, Cocchio S, Fonzo M, Pillon P, Buja A, et al. The new pandemic influenza A/(H1N1)pdm09 virus: is it really “new” J Prev Med Hyg. (2016) 57:E19–22.
66. Yang J, Li M, Shen X, Liu S. Influenza A virus entry inhibitors targeting the hemagglutinin. Viruses. (2013) 5:352–73. doi: 10.3390/v5010352
67. Reiter-Scherer V, Cuellar-Camacho JL, Bhatia S, Haag R, Herrmann A, Lauster D, et al. force spectroscopy shows dynamic binding of influenza hemagglutinin and neuraminidase to sialic acid. Biophys J. (2019) 116:1577. doi: 10.1016/j.bpj.2019.03.032
68. Lai JCC, Karunarathna H, Wong HH, Peiris JSM, Nicholls JM. Neuraminidase activity and specificity of influenza A virus are influenced by haemagglutinin-receptor binding. Emerg Microbes Infect. (2019) 8:327–38. doi: 10.1080/22221751.2019.1581034
69. Byrd-Leotis L, Cummings RD, Steinhauer DA. The interplay between the host receptor and influenza virus hemagglutinin and neuraminidase. Int J Mol Sci. (2017) 18:1541. doi: 10.3390/ijms18071541
70. Li F, Berardi M, Li W, Farzan M, Dormitzer PR, Harrison SC. Conformational states of the severe acute respiratory syndrome coronavirus spike protein ectodomain. J Virol. (2006) 80:6794–800. doi: 10.1128/JVI.02744-05
71. Li F, Li W, Farzan M, Harrison SC. Structure of SARS coronavirus spike receptor-binding domain complexed with receptor. Science. (2005) 309:1864–8. doi: 10.1126/science.1116480
72. Liu Y, Childs RA, Matrosovich T, Wharton S, Palma AS, Chai W, et al. Altered receptor specificity and cell tropism of D222G hemagglutinin mutants isolated from fatal cases of pandemic A(H1N1) 2009 influenza virus. J Virol. (2010) 84:12069–74. doi: 10.1128/JVI.01639-10
73. Wang N, Shi X, Jiang L, Zhang S, Wang D, Tong P, et al. Structure of MERS-CoV spike receptor-binding domain complexed with human receptor DPP4. Cell Res. (2013) 23:986–93. doi: 10.1038/cr.2013.92
74. Cao Y, Li L, Feng Z, Wan S, Huang P, Sun X, et al. Comparative genetic analysis of the novel coronavirus (2019-nCoV/SARS-CoV-2) receptor ACE2 in different populations. Cell Discov. (2020) 6:11. doi: 10.1038/s41421-020-0147-1
75. Gorbalenya AE, Baker SC, Baric RS, de Groot RJ, Drosten C, Gulyaeva AA, et al. The species severe acute respiratory syndrome-related coronavirus: classifying 2019-nCoV and naming it SARS-CoV-2. Nat Microbiol. (2020) 5:536–44. doi: 10.1038/s41564-020-0695-z
76. Huang Q, Herrmann A. Fast assessment of human receptor-binding capability of 2019 novel coronavirus (2019-nCoV). bioRxiv. (2020) 2020:2020.02.01.930537. doi: 10.1101/2020.02.01.930537
77. Li M, Li L, Zhang Y, Wang X. An Investigation of the Expression of 2019 Novel Coronavirus Cell Receptor Gene ACE2 in a Wide Variety of Human Tissues. Research Square (2020).
78. Gounder AP, Boon ACM. Influenza pathogenesis: the effect of host factors on severity of disease. J Immunol. (2019) 202:341–50. doi: 10.4049/jimmunol.1801010
79. Chen IY, Moriyama M, Chang MF, Ichinohe T. Severe acute respiratory syndrome coronavirus viroporin 3a activates the NLRP3 inflammasome. Front Microbiol. (2019) 10:50. doi: 10.3389/fmicb.2019.00050
80. Rockx B, Kuiken T, Herfst S, Bestebroer T, Lamers MM, Oude Munnink BB, et al. Comparative pathogenesis of COVID-19, MERS, and SARS in a nonhuman primate model. Science. (2020) 2020:eabb7314. doi: 10.1126/science.abb7314
81. Holshue ML, DeBolt C, Lindquist S, Lofy KH, Wiesman J, Bruce H, et al. First case of 2019 novel coronavirus in the United States. N Engl J Med. (2020) 382:929–36. doi: 10.1056/NEJMoa2001191
82. Channappanavar R, Perlman S. Pathogenic human coronavirus infections: causes and consequences of cytokine storm and immunopathology. Semin Immunopathol. (2017) 39:529–39. doi: 10.1007/s00281-017-0629-x
83. Huang C, Wang Y, Li X, Ren L, Zhao J, Hu Y, et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet. (2020) 395:497–506. doi: 10.1016/S0140-6736(20)30183-5
84. Zhang W, Zhao Y, Zhang F, Wang Q, Li T, Liu Z, et al. The use of anti-inflammatory drugs in the treatment of people with severe coronavirus disease 2019 (COVID-19): the perspectives of clinical immunologists from China. Clin Immunol. (2020) 214:108393. doi: 10.1016/j.clim.2020.108393
85. Tufan A, Avanoglu Guler A, Matucci-Cerinic M. COVID-19, immune system response, hyperinflammation and repurposing antirheumatic drugs. Turkish J Med Sci. (2020) 50:620–32. doi: 10.3906/sag-2004-168
86. Xu R, Ekiert DC, Krause JC, Hai R, Crowe JE Jr, et al. Structural basis of preexisting immunity to the 2009 H1N1 pandemic influenza virus. Science. (2010) 328:357–60. doi: 10.1126/science.1186430
87. Itoh Y, Shinya K, Kiso M, Watanabe T, Sakoda Y, Hatta M, et al. In vitro and in vivo characterization of new swine-origin H1N1 influenza viruses. Nature. (2009) 460:1021–5. doi: 10.1038/nature08260
88. Schrauwen EJ, de Graaf M, Herfst S, Rimmelzwaan GF, Osterhaus AD, Fouchier RA. Determinants of virulence of influenza A virus. Eur J Clin Microbiol Infect Dis. (2014) 33:479–90. doi: 10.1007/s10096-013-1984-8
89. Chen W, Calvo PA, Malide D, Gibbs J, Schubert U, Bacik I, et al. A novel influenza A virus mitochondrial protein that induces cell death. Nat Med. (2001) 7:1306–12. doi: 10.1038/nm1201-1306
90. Webster RG, Bean WJ, Gorman OT, Chambers TM, Kawaoka Y. Evolution and ecology of influenza A viruses. Microbiol Rev. (1992) 56:152–79. doi: 10.1128/MMBR.56.1.152-179.1992
91. Moser MR, Bender TR, Margolis HS, Noble GR, Kendal AP, Ritter DG. An outbreak of influenza aboard a commercial airliner. Am J Epidemiol. (1979) 110:1–6. doi: 10.1093/oxfordjournals.aje.a112781
92. Verreault D, Moineau S, Duchaine C. Methods for sampling of airborne viruses. Microbiol Mol Biol Rev. (2008) 72:413–44. doi: 10.1128/MMBR.00002-08
93. Nicas M, Nazaroff WW, Hubbard A. Toward understanding the risk of secondary airborne infection: emission of respirable pathogens. J Occup Environ Hyg. (2005) 2:143–54. doi: 10.1080/15459620590918466
94. Lowen A, Palese P. Transmission of influenza virus in temperate zones is predominantly by aerosol, in the tropics by contact: a hypothesis. PLoS Curr. (2009) 1:Rrn1002. doi: 10.1371/currents.RRN1002
95. Polozov IV, Bezrukov L, Gawrisch K, Zimmerberg J. Progressive ordering with decreasing temperature of the phospholipids of influenza virus. Nat Chem Biol. (2008) 4:248–55. doi: 10.1038/nchembio.77
96. Otter JA, Yezli S, French GL. The role played by contaminated surfaces in the transmission of nosocomial pathogens. Infect Control Hosp Epidemiol. (2011) 32:687–99. doi: 10.1086/660363
97. Bean B, Moore BM, Sterner B, Peterson LR, Gerding DN, Balfour HH Jr. Survival of influenza viruses on environmental surfaces. J Infect Dis. (1982) 146:47–51. doi: 10.1093/infdis/146.1.47
98. van Doremalen N, Bushmaker T, Munster VJ. Stability of Middle East respiratory syndrome coronavirus (MERS-CoV) under different environmental conditions. Euro Surveill. (2013) 18:20590. doi: 10.2807/1560-7917.ES2013.18.38.20590
99. Duan SM, Zhao XS, Wen RF, Huang JJ, Pi GH, Zhang SX, et al. Stability of SARS coronavirus in human specimens and environment and its sensitivity to heating and UV irradiation. Biomed Environ Sci. (2003) 16:246–55.
100. Chan KH, Peiris JS, Lam SY, Poon LL, Yuen KY, Seto WH. The effects of temperature and relative humidity on the viability of the SARS coronavirus. Adv Virol. (2011) 2011:734690. doi: 10.1155/2011/734690
101. van Doremalen N, Bushmaker T, Morris DH, Holbrook MG, Gamble A, Williamson BN, et al. Aerosol and surface stability of SARS-CoV-2 as compared with SARS-CoV-1. N Engl J Med. (2020) 382:1564–7. doi: 10.1056/NEJMc2004973
102. Wearing HJ, Rohani P, Keeling MJ. Appropriate models for the management of infectious diseases. PLoS Med. (2005) 2:e174. doi: 10.1371/journal.pmed.0020174
103. Heffernan JM, Smith RJ, Wahl LM. Perspectives on the basic reproductive ratio. J R Soc Interface. (2005) 2:281–93. doi: 10.1098/rsif.2005.0042
104. Biggerstaff M, Cauchemez S, Reed C, Gambhir M, Finelli L. Estimates of the reproduction number for seasonal, pandemic, and zoonotic influenza: a systematic review of the literature. BMC Infect Dis. (2014) 14:480. doi: 10.1186/1471-2334-14-480
105. Chowell G, Castillo-Chavez C, Fenimore PW, Kribs-Zaleta CM, Arriola L, Hyman JM. Model parameters and outbreak control for SARS. Emerg Infect Dis. (2004) 10:1258–63. doi: 10.3201/eid1007.030647
106. Breban R, Riou J, Fontanet A. Interhuman transmissibility of Middle East respiratory syndrome coronavirus: estimation of pandemic risk. Lancet. (2013) 382:694–9. doi: 10.1016/S0140-6736(13)61492-0
107. D'Arienzo M, Coniglio A. Assessment of the SARS-CoV-2 basic reproduction number, R0, based on the early phase of COVID-19 outbreak in Italy. Biosaf Health. (2020) 2:57–9. doi: 10.1016/j.bsheal.2020.03.004
108. Fineberg HV. Pandemic preparedness and response — lessons from the H1N1 influenza of 2009. N Engl J Med. (2014) 370:1335–42. doi: 10.1056/NEJMra1208802
109. Russell CA, Kasson PM, Donis RO, Riley S, Dunbar J, Rambaut A, et al. Science Forum: improving pandemic influenza risk assessment. eLife. (2014) 3:e03883. doi: 10.7554/eLife.03883
110. WHO. Pandemic Influenza Risk Management (PIRM). (2017). Available online at: https://www.who.int/influenza/preparedness/pandemic/influenza_risk_management/en/
111. Prem K, Liu Y, Russell TW, Kucharski AJ, Eggo RM, Davies N, et al. The effect of control strategies to reduce social mixing on outcomes of the COVID-19 epidemic in Wuhan, China: a modelling study. Lancet Public Health. (2020) 5:e261–70. doi: 10.1101/2020.03.09.20033050
112. McDonald LC, Simor AE, Su IJ, Maloney S, Ofner M, Chen KT, et al. SARS in healthcare facilities, Toronto and Taiwan. Emerg Infect Dis. (2004) 10:777–81. doi: 10.3201/eid1005.030791
113. Suwantarat N, Apisarnthanarak A. Risks to healthcare workers with emerging diseases: lessons from MERS-CoV, Ebola, SARS, and avian flu. Curr Opin Infect Dis. (2015) 28:349–61. doi: 10.1097/QCO.0000000000000183
114. WHO. Emergencies Preparedness, Response. (2004). Available online at: https://www.who.int/csr/resources/publications/WHO_CDS_CSR_ARO_2004_2/en/
115. Memish ZA, Cotten M, Meyer B, Watson SJ, Alsahafi AJ, Al Rabeeah AA, et al. Human infection with MERS coronavirus after exposure to infected camels, Saudi Arabia, 2013. Emerg Infect Dis. (2014) 20:1012. doi: 10.3201/eid2006.140402
116. Peeri NC, Shrestha N, Rahman MS, Zaki R, Tan Z, Bibi S, et al. The SARS, MERS and novel coronavirus (COVID-19) epidemics, the newest and biggest global health threats: what lessons have we learned? Int J Epidemiol. (2020) 49:717–26. doi: 10.1093/ije/dyaa033
117. Devaux CA, Rolain JM, Colson P, Raoult D. New insights on the antiviral effects of chloroquine against coronavirus: what to expect for COVID-19? Int J Antimicrob Agents. (2020) 2020:105938. doi: 10.1016/j.ijantimicag.2020.105938
118. Cortegiani A, Ingoglia G, Ippolito M, Giarratano A, Einav S. A systematic review on the efficacy and safety of chloroquine for the treatment of COVID-19. J Crit Care. (2020) 57:279–83. doi: 10.1016/j.jcrc.2020.03.005
119. Savarino A, Boelaert JR, Cassone A, Majori G, Cauda R. Effects of chloroquine on viral infections: an old drug against today's diseases? Lancet Infect Dis. (2003) 3:722–7. doi: 10.1016/S1473-3099(03)00806-5
120. Huang Z, Liu H, Zhang X, Wen G, Zhu C, Zhao Y, et al. Transcriptomic analysis of lung tissues after hUC-MSCs and FTY720 treatment of lipopolysaccharide-induced acute lung injury in mouse models. Int Immunopharmacol. (2018) 63:26–34. doi: 10.1016/j.intimp.2018.06.036
121. Zhang Z, Li W, Heng Z, Zheng J, Li P, Yuan X, et al. Combination therapy of human umbilical cord mesenchymal stem cells and FTY720 attenuates acute lung injury induced by lipopolysaccharide in a murine model. Oncotarget. (2017) 8:77407–14. doi: 10.18632/oncotarget.20491
122. Wu W, Wang JF, Liu PM, Chen WX, Yin SM, Jiang SP, et al. [Clinical features of 96 patients with severe acute respiratory syndrome from a hospital outbreak]. Zhonghua Nei Ke Za Zhi. (2003) 42:453–7.
123. Stockman LJ, Bellamy R, Garner PJ. SARS: systematic review of treatment effects. PLoS Med. (2006) 3:e343. doi: 10.1371/journal.pmed.0030343
124. Sharif-Yakan A, Kanj SS. Emergence of MERS-CoV in the Middle East: origins, transmission, treatment, and perspectives. PLoS Pathog. (2014) 10:e1004457. doi: 10.1371/journal.ppat.1004457
125. Jefferson T, Jones M, Doshi P, Spencer EA, Onakpoya I, Heneghan CJ. Oseltamivir for influenza in adults and children: systematic review of clinical study reports and summary of regulatory comments. BMJ. (2014) 348:g2545. doi: 10.1136/bmj.g2545
127. Cheng VC, Wong S-C, To KK, Ho P, Yuen K-Y. Preparedness and proactive infection control measures against the emerging novel coronavirus in China. J Hosp Infect. (2020) 104:254–5. doi: 10.1016/j.jhin.2020.01.010
Keywords: SARS-CoV-2, SARS-CoV, MERS-CoV, influenza A virus, COVID-19
Citation: Abdelrahman Z, Li M and Wang X (2020) Comparative Review of SARS-CoV-2, SARS-CoV, MERS-CoV, and Influenza A Respiratory Viruses. Front. Immunol. 11:552909. doi: 10.3389/fimmu.2020.552909
Received: 17 April 2020; Accepted: 24 August 2020;
Published: 11 September 2020.
Edited by:
Alexander Rodriguez-Palacios, Case Western Reserve University, United StatesReviewed by:
Shuvojit Banerjee, Case Western Reserve University, United StatesRachel Graham, The University of North Carolina at Chapel Hill, United States
Copyright © 2020 Abdelrahman, Li and Wang. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Xiaosheng Wang, eGlhb3NoZW5nLndhbmdAY3B1LmVkdS5jbg==
 Zeinab Abdelrahman
Zeinab Abdelrahman Mengyuan Li
Mengyuan Li Xiaosheng Wang
Xiaosheng Wang