
94% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Genet. , 07 May 2019
Sec. Applied Genetic Epidemiology
Volume 10 - 2019 | https://doi.org/10.3389/fgene.2019.00427
Microdeletions in Y-chromosomal azoospermia factor (AZF) regions have been regarded as the risk factor of spermatogenic failure (SF). However, AZF-linked duplications or complex copy number variants (CNVs) (deletion + duplication) were rarely studied. In this study, we performed multiplex ligation-dependent probe amplification (MLPA) analysis on 402 fertile healthy male controls and 423 idiopathic infertile SF patients (197 azoospermia and 226 oligozoospermia) in Han Chinese population. In total, twenty-four types of AZF-linked CNVs were identified in our study, including eleven novel CNVs (one deletion, seven duplications, and three complex CNVs). Our study revealed that AZFc-linked duplications and the instability of Y chromosome might be associated with spermatogenesis. Besides, the complex CNVs (b2/b3 deletion + DAZ1/2 duplication) were confirmed to increase genetic risks for SF in Han Chinese population. This study illustrated a spectrum of AZF-linked CNVs and presented valuable information for understanding the clinical significance of AZF-linked CNVs in male infertility.
Infertility affects an estimated 15% of couples globally at childbearing age, and male infertility is directly or indirectly responsible for about 50% of cases involving reproductive-age couples with fertility-related issues (Agarwal et al., 2015). The majority of infertile males were diagnosed with spermatogenic failure (SF) (Lo Giacco et al., 2014).
Y-chromosome microdeletions (YCM) are the second most common genetic etiology for SF after the Klinefelter syndrome (Krausz et al., 2014). In YCM, azoospermia factor (AZF) [AZFa, AZFb, AZFc(b2/b4)] deletions have resulted in SF and testing these deletions had significant clinical values both in diagnosis and prognostication of testicular sperm retrieval in azoospermic men (Pryor et al., 1997; Rozen et al., 2012), while the clinical significance of partial AZFc deletions (gr/gr, b2/b3, and b1/b3) varies in different studies (Lu et al., 2009; Bansal et al., 2016; Krausz and Casamonti, 2017). However, all previous studies on AZF-linked copy number variants (CNVs) have mainly focused on deletions rather than duplications or complex CNVs (deletions + duplications).
Owing to frequent non-allelic homologous recombination (NAHR) of AZF region, the majority of CNVs are found on Y chromosome among all human chromosomes (Freeman et al., 2006). Identifying AZF-linked CNVs have been showed to be valuable for finding the genetic etiology of SF, as well as evaluating the rate of sperm recovery after testicular sperm extraction (TESE) (Krausz et al., 2014). Traditional methods to identify AZF-linked CNVs were based on polymerase chain reaction (PCR) of sequence-tagged site (STS) markers (STS-PCR), however, STS-PCR could only determine AZF-linked deletions. To date, some AZF-linked duplications or complex CNVs have been reported (Giachini et al., 2008; Lu et al., 2011; Saito et al., 2015). However, the precise frequency and clinical significance of these CNVs have remained unclear in patients with SF.
With the development of molecular techniques, such as multiplex ligation-dependent probe amplification (MLPA) and array comparative genomic hybridization (aCGH), researchers were able to simultaneously identify multiple CNVs. Compared with aCGH, the MLPA method was relatively simple and inexpensive, and it has been suggested as a valuable technique for diagnosing genetic diseases (Quarello et al., 2012; Massalska et al., 2013; Mutlu et al., 2018).
To identify novel AZF-linked CNVs (especially duplications and complex CNVs) and their significance in spermatogenesis, we performed MLPA analysis on 402 fertile healthy male controls and 423 idiopathic infertile SF patients (197 azoospermia and 226 oligozoospermia) in Han Chinese population.
Subjects were recruited from the Nanjing Maternity and Child Health Care Hospital (Nanjing, Jiangsu province, China) between July 2016 and September 2017. This study was approved by the Ethics Committee of Nanjing Maternity and Child Health Care Hospital. All procedures involved in this study were conducted in accordance with the Declaration of Helsinki.
Semen analyses were performed based on World Health Organization criteria (2010) with the reference values parameters for semen volume of ≥ 1.5 mL, pH ≥ 7.2, total sperm count of 39 × 106 per ejaculate, sperm concentration of 15 × 106/mL, sperm total motility of 40%, and sperm with a normal morphology of 4% (Cooper et al., 2010), and each subject was examined twice to ensure the reliability of the results. In addition to semen analyses, some comprehensive andrological examinations including a series of physical examinations, scrotal ultrasound, hormone analysis and karyotype analysis were also performed. Those with varicocele, cryptorchidism, orchitis or abnormal karyotype (such as 47, XXY) were excluded from this study.
The male infertile cases, who sought treatment in the infertility clinic, were recruited into this study by a retrospective design. These cases have no child and diagnosised with idiopathic non-obstructive azoospermia (NOA) (no sperm in the ejaculate) or oligozoospermia (total sperm count < 39 × 106 per ejaculate). To distinguish between NOA and OA (obstructive azoospermia), only those azoospermic cases with soft and small testis (total testicular volume <30 ml), elevated follicular stimulating hormone in plasma were included. The controls, recruited from the same hospital during the same period, were fertile males who had normal total sperm count per ejaculate, sperm concentration, motility, morphology and had fathered at least one child without assisted reproductive. Twenty-nine infertile subjects (out of 452) were excluded, including 4 with OA, 5 with cryptorchidism, 8 with varicocele, 2 with orchitis and 10 with abnormal karyotype (7 of them with 47, XXY). Finally, 402 fertile male controls and 423 infertile patients were recruited in this study.
All the participants were genetically unrelated ethnic Han-Chinese. Some influencing factors [e.g., age, body mass index (BMI), and smoking] were ruled out between patients and controls. All participants were informed about the purpose of the study, and signed the written informed consent for publication of this original research as well.
MLPA analysis was performed using SALSA MLPA probe-mix kit P360-B1 (MRC-Holland, Amsterdam, the Netherlands) containing 42 specific probes for AZF regions (Supplementary Figure S1). Briefly, 7 ul of sample DNA (20 ng/ul) were denatured for 5 min at 98°C and subsequently cooled to 25°C. Next, 5 ul denatured DNA were mixed with 3 ul hybridization master mix and heated for 1 min at 95°C, then 18 h at 60°C. After hybridization reaction, 32 ul ligase-65 master mix were added and incubated for 15 min at 54°C and 5 min for 98°C, then pause at 20°C. The ligation products were then mixed with polymerase master mix for PCR reaction. The PCR products were analyzed by ABI 3,500 using 50 cm capillaries and POP-7 polymers (Applied Biosystems, Foster City, CA, USA) and the injection mixture containing 0.9 ul PCR products, 0.1 ul LIZ 500 size standard and 9 μl HiDi formamide. The ABI 3,500 run conditions were as follows: injection voltage = 1.6 kVolt, injection time = 8 s, oven temperature = 60°C, run voltage = 19.5 kVolts, and run time = 1330 s. The relative peak area of each probe was calculated by dividing the actual peak area of the subject by the average of that of five reference samples using Coffalyser.Net Software (MRC-Holland, Amsterdam, the Netherlands). The 30% increase or decrease of the relative peak area of the probe showed duplication or deletion of the targeted region, respectively.
Results of MLPA analysis were compared with those of conventional STS-PCR based on ten STS markers in AZF region (sY84 and sY86 for AZFa, sY127 and sY134 for AZFb, sY254, sY255, sY1192, sY1191, sY1291 and sY1189 for AZFc) recommended by EAA/EMQN practice guidelines (Krausz et al., 2014).
To validate those AZF-linked CNVs not confirmed by STS-PCR, we performed real-time quantitative PCR (qPCR). Primer sequences designed for target regions were listed in Supplementary Table S2. The ACTB gene was employed as a reference. The qPCR was performed in a total volume of 20 μL consisting of 10 μL AceQ Universal SYBR qPCR Master Mix (Vazyme, China), 0.4 μL of 10 μM each primer (forward and reverse), 2 μL genomic DNA (10 ng/μL) and 7.2 μL nuclease-free water. The PCR reaction were run in ABI StepOnePlusTM (Life Technologies, USA) based on the following program: 95°C for 5 min for initial denaturation, followed by 40 cycles (95°C for 10 s and 60°C for 30 s) for amplification and fluorescence detection, and a cycle (95°C for 15 s, 60°C for 60 s, and 95°C for 15 s) for melt curve analysis. Data analysis was performed using StepOneTM Software based on the ΔΔCT method. All samples were detected in triplicates for qPCR analysis.
In this study, χ2 and Fisher's exact tests were applied to compare the statistical differences in the frequencies of CNVs between the patients and controls using SPSS 20.0 software (IBM, Armonk, NY, USA). P-value < 0.05 was statistically considered significant. We did not correct for multiple testing considering that the sample size for those novel AZF-linked CNVs were relatively small and adjustments for making multiple comparisons might miss possibly important findings.
In this study, we recruited 402 fertile healthy male controls and 423 infertile SF patients (197 azoospermia and 226 oligozoospermia). According to the achieved results, we found that there was no significant difference between the control group and the SF group in selected characteristics including age, BMI, and smoking (Table 1).
Overall, twenty-four types of AZF-linked CNVs (eight deletions, eleven duplications and five complex CNVs) were identified in our study population. Among eight deletions, four classical deletions (AZFa deletion, AZFb deletion, AZFc complete deletion (b2/b4 deletion), and AZFb + c deletion) recommended by EAA/EMQN practice guidelines (Krausz et al., 2014) were identified by MLPA analysis and verified by STS-PCR (Figures 1, 4). These four types of deletions were identified in SF patients, but not observed in controls (Table 2). Consistent with the previous reports, the classical deletions were the genetic causes of SF (Krausz et al., 2014).
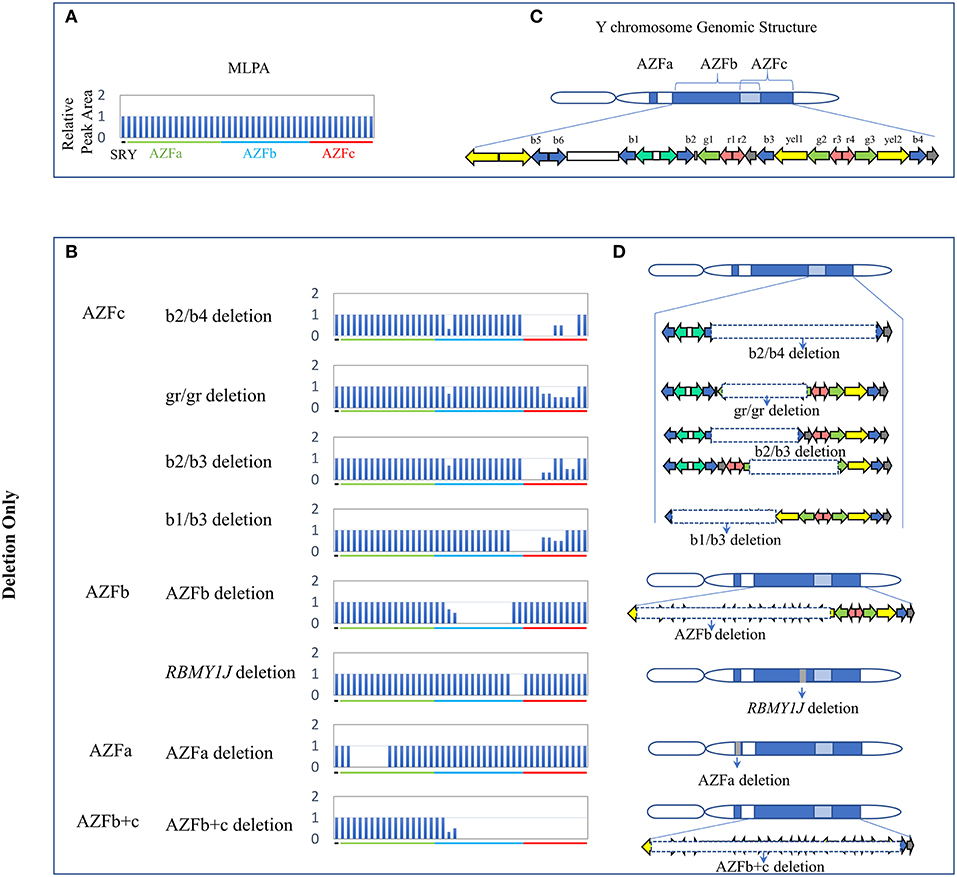
Figure 1. CNVs (Deletion Only) in AZF regions of the human Y chromosome identified by MLPA. (A,B) Results of MLPA analysis. The relative peak area of each probe was calculated by dividing the actual peak area of the subject by the average of that of five reference samples. Decreased relative peak areas mean copy-number deletion. (C,D) Corresponding structures and rearrangements of AZF regions.
The frequency of three AZFc partial deletions (gr/gr, b2/b3, and b1/b3) identified by MLPA and validated by STS-PCR showed no significantly statistical difference between the patients and the controls (Table 2), suggesting that these partial deletions might not considered to be genetic risk factors for SF in our study population.
Besides, we found a novel and de novo AZFb partial deletion (RBMY1J deletion) in a patient with oligozoospermia whose father has a normal RBMY1J copy in the same region (Table 2). The de novo deletion might be associated with oligozoospermia, and further study is required to confirm whether the RBMY1J deletion may cause the disease.
The traditional STS-PCR method could only detect AZF-linked deletions, while MLPA method could detect not only deletions, but also duplications. In this study, we identified eleven types of AZF-linked duplications in 3.98% of controls (16/402) and 7.09% of SF patients (30/423). Most of duplications (72.7%, 8/11) were located in the AZFc region likely owing to the frequent non-allele homologous recombination (NAHR) in this region (Skaletsky et al., 2003). Seven novel duplications, four in AZFc region, one in AZFb region, and two in AZFa region, were identified in our study (Figure 2, Table 3).
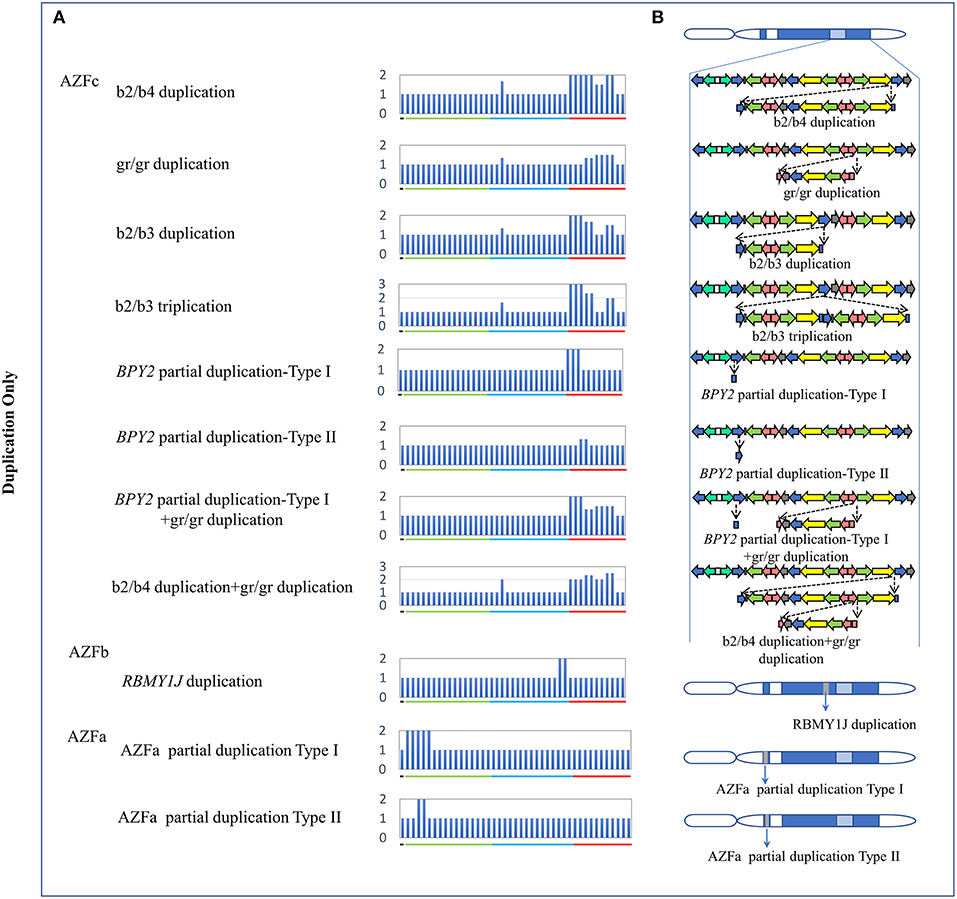
Figure 2. CNVs (Duplication Only) in AZF regions of the human Y chromosome identified by MLPA. (A) Results of MLPA analysis. The relative peak area of each probe was calculated by dividing the actual peak area of the subject by the average of that of five reference samples. Increased relative peak areas mean copy-number duplications. (B) Corresponding structures and rearrangements of AZF regions.
Two types of BPY2 duplication (Types I and II) and “AZFa partial dup Type I” were found in both patients and controls, suggesting they might be benign CNVs. The CNVs named “BPY2 dup Type I + gr/gr dup,” “b2/b4 dup + gr/gr dup,” “RBMY1J dup,” and “AZFa partial dup Type II” were all de novo CNVs and only identified in patient group (Table 3). Additional cases should be considered to investigate the clinical relevance of these findings.
Five types of complex CNVs involved both deletions and duplications were identified in our study (Figure 3, Table 4). Among these complex CNVs, the “b2/b3 deletion + DAZ 1/2 duplication” was confirmed by analyzing the restriction enzyme digestion site of single-nucleotide variation (SNV) at sY587 to discriminate DAZ 1/2 from DAZ 3/4 (Machev et al., 2004; Supplementary Table S1). Consistent with the result reported by Lu et al. (2011), the frequency of the CNV was significantly higher in patients than that in controls (OR = 5.34, P = 3.00 × 10−2) (Table 4). The CNV named “two gr/gr deletions + b2/b4 duplications” was identified in one SF patient by Saito et al. (2015), and this CNV was considered to be benign in our patients (Table 4).
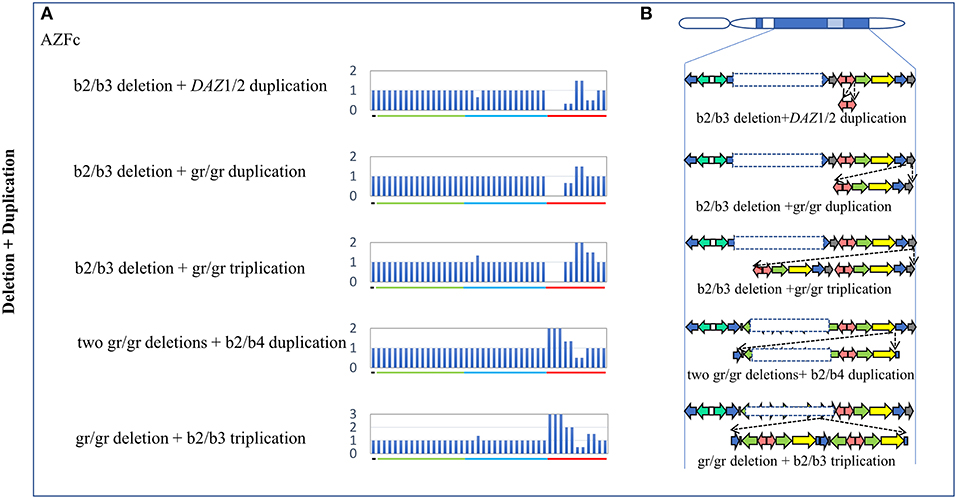
Figure 3. Complex CNVs (Deletion + Duplication) in AZF regions of the human Y chromosome identified by MLPA. (A) Results of MLPA analysis. The relative peak area of each probe was calculated by dividing the actual peak area of the subject by the average of that of five reference samples. Decreased and increased relative peak areas mean copy-number deletions and duplications, respectively. (B) Corresponding structures and rearrangements of AZF regions.
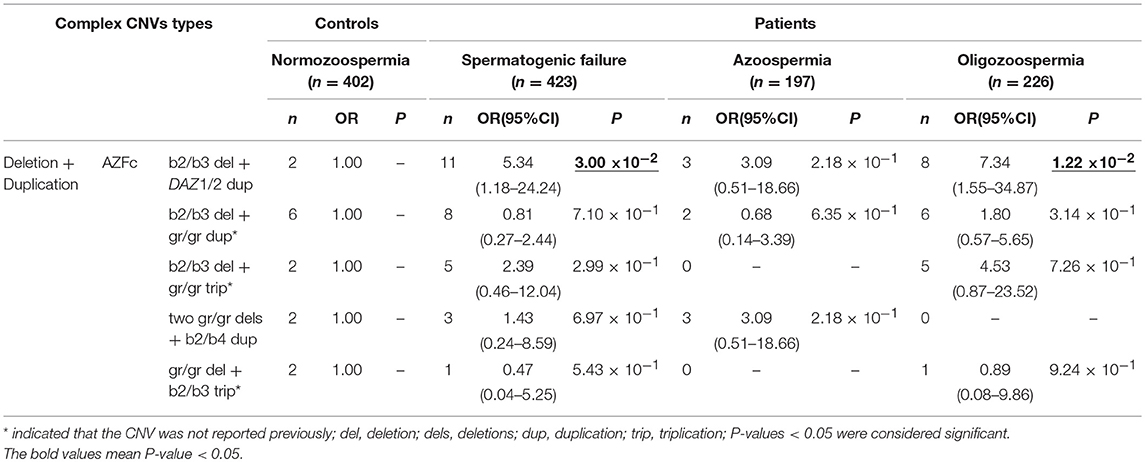
Table 4. Distribution of known and novel complex CNVs (Deletion+Duplication) in AZF regions and their effects on SF.
In addition, three novel complex CNVs were identified in our study, including “b2/b3 deletion + gr/gr duplication,” “b2/b3 deletion + gr/gr triplication,” and “gr/gr deletion + b2/b3 triplication.” The frequency of each complex CNV showed that there was no significant difference between the patient group and the control group (Table 4).
Furthermore, we comprehensively analyzed all the AZF-linked CNVs to evaluate their effects on spermatogenesis. We found that “any CNV in AZF region” was at significantly higher frequency in SF cases (OR = 2.24; 95%CI, 1.62–3.08) (P = 8.37 × 10−7), azoospermia cases (OR = 1.97; 95%CI, 1.33–3.91) (P = 7.00 × 10−4), and oligozoospermia cases (OR = 2.44; 95%CI, 1.69–3.53) (P = 1.22 × 10−2), when compared with controls (Table 5).
Similarly, the “any deletions in AZF region” was found at significantly higher frequency in SF cases (OR = 2.13; 95%CI, 1.50-3.01) (P = 2.10 × 10−5), azoospermia cases (OR = 1.89; 95%CI, 1.23–2.88) (P = 3.30 × 10−3), and oligozoospermia cases (OR = 2.30; 95%CI, 1.55–3.43) (P = 4.00 × 10−5), when compared with controls (Table 5).
For “any duplications in AZFc region”, the frequency was significantly higher in SF cases (OR = 2.22; 95%CI = 1.11–4.44) (P = 2.47 × 10−2) and oligozoospermia cases (OR = 2.31; 95%CI = 1.06–5.03) (P = 3.47 × 10−2) than controls (Table 5).
As compared with MLPA, only seven known deletions (29.2%, 7/24) were identified by STS-PCR. One deletion (4.2%, 1/24) and eleven duplications (45.8%, 11/24) were not identified, and five complex CNVs (20.8%, 5/24) were detected as simple deletions (Figure 4), demonstrating that the MLPA method possesses more powerful efficiency in detecting AZF-linked CNVs.
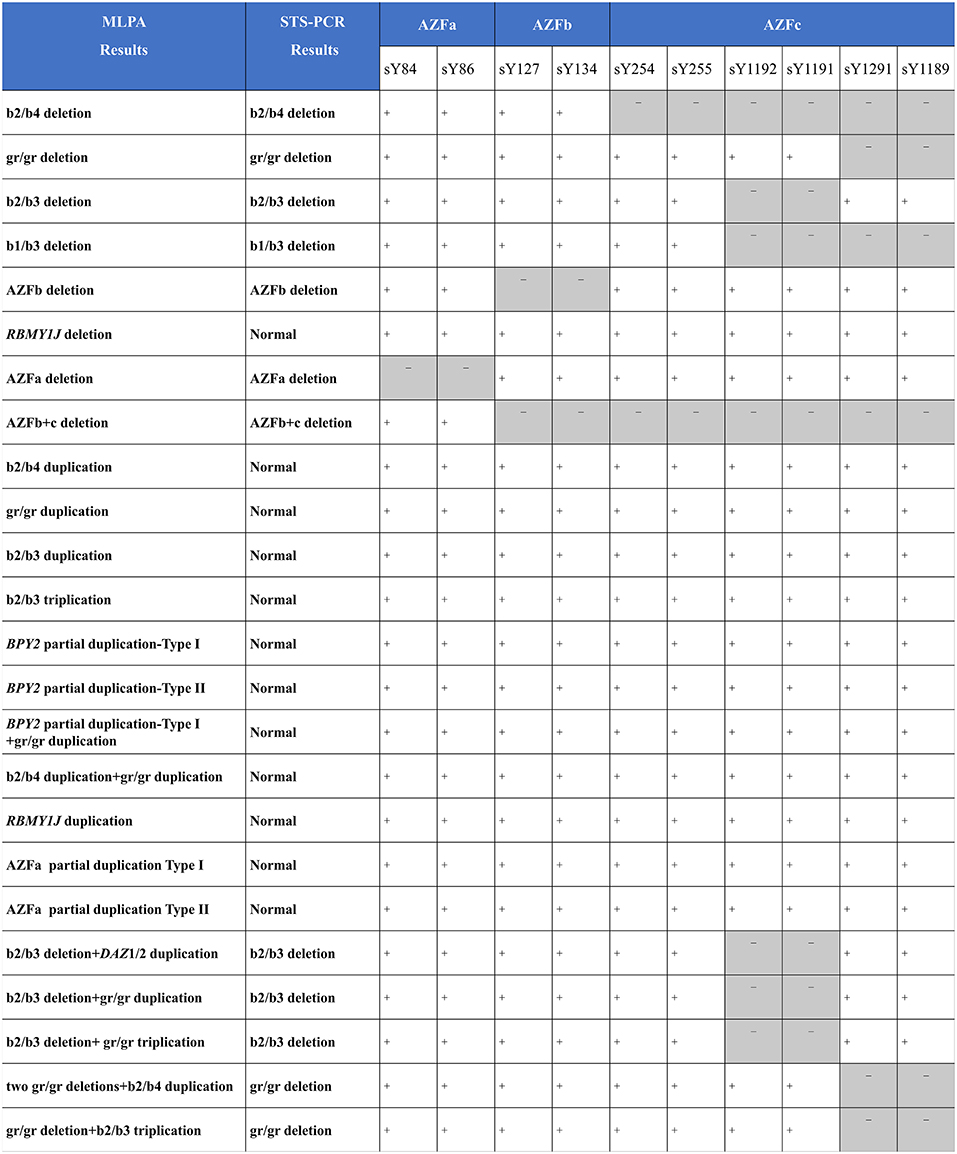
Figure 4. Comparing MLPA methods with traditional agarose electrophoresis for STS markers. +, the fragment in the STS site was not deleted; -, the fragment in the STS site was deleted.
To validate the CNVs not verified by STS-PCR especially those AZF-linked duplications and complex CNVs, we performed real-time quantitative PCR (qPCR).
For each CNV to be tested, two reference samples (normal MLPA results) and one randomly selected positive CNV sample were examined. The qPCR results were all consistent with MLPA results (Figure 5 and Supplementary Figure S2). Those above results indicated the accuracy of the MLPA method for detecting AZF-linked CNVs ( both deletions and duplications).
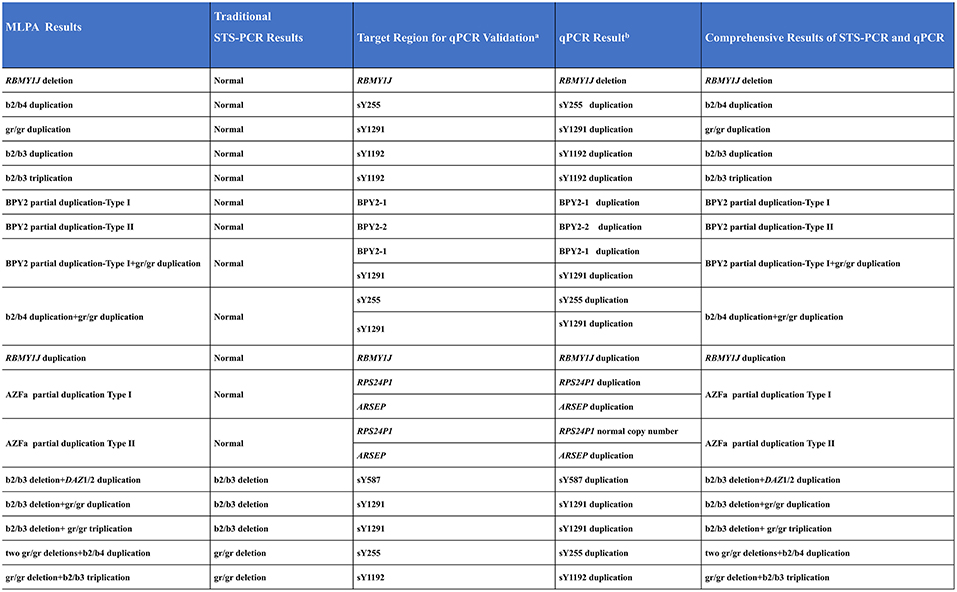
Figure 5. Comprehensive results of STS-PCR and qPCR validation. a, Special regions of target AZF-linked CNVs for qPCR validation; b, Simplified qPCR results. Detailed qPCR results see Supplementary Figure S2.
Y chromosome microdeletion was the second most frequent genetic etiology after Klinefelter syndrome (Krausz et al., 2014). In recent years, AZFa, b and c (b2/b4) deletions have been identified as the pathogenic CNVs in spermatogenesis (Krausz et al., 2014). However, AZF-linked duplications and complex CNVs were rarely studied due to limits of technical means. In this study, we revealed the relatively precise frequency and clinical significance of AZF-linked CNVs in Han Chinese population by using the MLPA method.
For AZFc partial deletions (gr/gr del, b2/b3 del, and b1/b3 del), we did not observe significantly different distributions between SF cases and control subjects. The clinical significance of these partial deletions was reported to be variable in previous studies (Lu et al., 2009; Bansal et al., 2016; Krausz and Casamonti, 2017). Hence, further study should be performed in diverse racial or ethnic groups to confirm the respective results.
Interestingly, two de novo RBMY1J CNVs, a deletion and a duplication were identified in our oligozoospermic patients. RBMY1J, a member of RBMY1 (Y-linked RNA binding motif protein family 1), is expressed in all stages of spermatogenesis and acts as a splicing factor during spermatogenesis (Elliott et al., 1997; Dreumont et al., 2010). Yan et al. revealed that RBMY1 is associated with sperm motility (Yan et al., 2017). Considering the normal sperm motility in both patients, our study suggests that RBMY1J is associated with decreased sperm numbers in clinic. Therefore, further studies are required to determine the function of RBMY1 in spermatogenesis.
Besides, Y chromosome is thought to be “fragile” owing to complex NAHR structure. In the present study, “any CNVs in AZF region” showed significantly higher frequency in SF patients than controls, and we speculated that the instability of Y chromosome might be associated with spermatogenic impairment. Additionally, the frequency of “any duplication in AZFc region” was also significantly higher in SF patients, suggesting that AZFc-linked duplications might play vital roles in spermatogenesis. Similar results were reported in the previous studies (Lin et al., 2007; Ye et al., 2013; Saito et al., 2015).
In addition, three novel complex CNVs identified in the present study expanded our knowledge on AZF-lined CNVs. Notably, all five complex CNVs were detected as simple deletions (gr/gr or b2/b3 deletion) by the traditional STS-PCR method. The technical limitation of STS-PCR would be a potential confounding factor for risk assessment in studies of gr/gr or b2/b3 deletion. It justifies why the previous research has demonstrated conflicting findings.
In 2012, Bunyan et al. performed MLPA in 100 subjects (50 SF patients and 50 controls) and identified four types of simple deletions, which cannot be detected by STS-PCR (Bunyan et al., 2012). In 2014, Liu et al. performed MLPA in 199 fathers and their 228 sons, and found that assisted reproductive technology didn't increase the risk of Y-chromosome microdeletions in male offspring (Liu et al., 2014). Both studies used the MLPA analysis to detect Y-chromosome microdeletions. In 2015, Saito et al. performed MLPA in 56 SF patients and 65 control individuals from who all were Japanese, and identified both AZF-linked deletions and duplications (Saito et al., 2015). Compared with previous studies, we applied the MLPA analysis to more subjects and identified more types of AZF-linked CNVs. Besides, for those novel CNVs only detected in SF patients, we further detected if they were inherited from their fathers and found they were all de novo CNVs. In addition, we classified the patients with SF into azoospermia group and oligozoospermia group, which would be helpful to understand the clinical significance of AZF-linked CNVs in different SF types.
However, there are some limitations in the current study. Although a decent number of subjects were tested in this retrospective case-control study, it is not enough to reach conclusions on the role of those rare AZF-linked CNVs. In addition, the p-values presented were not corrected for multiple testing in the statistical analysis, which may lead to reject the null hypothesis too readily and increase the risk of false positives. In the future, we plan to establish the roles of rare AZF-linked CNVs in a larger prospective study.
In summary, this study comprehensively identified AZF-linked CNVs, especially duplications, in Han ethnicity. It provides valuable findings for understanding the roles of AZF-linked CNVs in spermatogenesis, and will be helpful in the diagnosis and treatment of SF in clinical practice.
This study was carried out in accordance with the recommendations of Nanjing Maternity and Child Health Care Hospital with written informed consent from all subjects. All subjects gave written informed consent in accordance with the Declaration of Helsinki. The protocol was approved by the Ethics Committee of Nanjing Maternity and Child Health Care Hospital.
RZ and JC performed the experiments and wrote the manuscript. DM and JT analyzed the data. YW and PH performed clinical evaluation and collected clinical data. ZX designed the study.
This study was supported by the National Natural Science Foundation of China (81701427 and 81770236).
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
We are grateful to the patients and volunteers for their help and participation in the study. Thank for the contribution made by Nanjing Maternity and Child Health Care Hospital.
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fgene.2019.00427/full#supplementary-material
Agarwal, A., Mulgund, A., Hamada, A., and Chyatte, M. R. (2015). A unique view on male infertility around the globe. Reprod. Biol. Endocrinol. 13:37. doi: 10.1186/s12958-015-0032-1
Bansal, S. K., Jaiswal, D., Gupta, N., Singh, K., Dada, R., Sankhwar, S. N., et al. (2016). Gr/gr deletions on Y-chromosome correlate with male infertility: an original study, meta-analyses, and trial sequential analyses. Sci. Rep. 6:19798. doi: 10.1038/srep19798
Bunyan, D. J., Callaway, J. L., and Laddach, N. (2012). Detection of partial deletions of Y-chromosome AZFc in infertile men using the multiplex ligation-dependent probe amplification assay. J. Reprod. Infertil. 13, 174–178.
Cooper, T. G., Noonan, E., von Eckardstein, S., Auger, J., Baker, H. W., Behre, H. M., et al. (2010). World Health Organization reference values for human semen characteristics. Hum. Reprod. Update 16, 231–245. doi: 10.1093/humupd/dmp048
Dreumont, N., Bourgeois, C. F., Lejeune, F., Liu, Y., Ehrmann, I. E., Elliott, D. J., et al. (2010). Human RBMY regulates germline-specific splicing events by modulating the function of the serine/arginine-rich proteins 9G8 and Tra2-{beta}. J. Cell Sci. 123, 40–50. doi: 10.1242/jcs.055889
Elliott, D. J., Millar, M. R., Oghene, K., Ross, A., Kiesewetter, F., Pryor, J., et al. (1997). Expression of RBM in the nuclei of human germ cells is dependent on a critical region of the Y chromosome long arm. Proc. Natl. Acad. Sci. U.S.A. 94, 3848–3853. doi: 10.1073/pnas.94.8.3848
Freeman, J. L., Perry, G. H., Feuk, L., Redon, R., McCarroll, S. A., Altshuler, D. M., et al. (2006). Copy number variation: new insights in genome diversity. Genome Res. 16, 949–961. doi: 10.1101/gr.3677206
Giachini, C., Laface, I., Guarducci, E., Balercia, G., Forti, G., and Krausz, C. (2008). Partial AZFc deletions and duplications: clinical correlates in the Italian population. Hum. Genet. 124, 399–410. doi: 10.1007/s00439-008-0561-1
Krausz, C., and Casamonti, E. (2017). Spermatogenic failure and the Y chromosome. Hum. Genet. 136, 637–655. doi: 10.1007/s00439-017-1793-8
Krausz, C., Hoefsloot, L., Simoni, M., Tuttelmann, F., and European Academy of Andrology, European Molecular Genetics Quality Network. (2014). EAA/EMQN best practice guidelines for molecular diagnosis of Y-chromosomal microdeletions: state-of-the-art 2013. Andrology 2, 5–19. doi: 10.1111/j.2047-2927.2013.00173.x
Lin, Y. W., Hsu, L. C., Kuo, P. L., Huang, W. J., Chiang, H. S., Yeh, S. D., et al. (2007). Partial duplication at AZFc on the Y chromosome is a risk factor for impaired spermatogenesis in Han Chinese in Taiwan. Hum. Mutat. 28, 486–494. doi: 10.1002/humu.20473
Liu, X. H., Yan, L. Y., Lu, C. L., Li, R., Zhu, X. H., Jin, H. Y., et al. (2014). ART do not increase the risk of Y-chromosome microdeletion in 19 candidate genes at AZF regions. Reprod. Fertil. Dev. 26, 778–786. doi: 10.1071/RD13092
Lo Giacco, D., Chianese, C., Sanchez-Curbelo, J., Bassas, L., Ruiz, P., Rajmil, O., et al. (2014). Clinical relevance of Y-linked CNV screening in male infertility: new insights based on the 8-year experience of a diagnostic genetic laboratory. Eur. J. Hum. Genet. 22, 754–761. doi: 10.1038/ejhg.2013.253
Lu, C., Zhang, F., Yang, H., Xu, M., Du, G., Wu, W., et al. (2011). Additional genomic duplications in AZFc underlie the b2/b3 deletion-associated risk of spermatogenic impairment in Han Chinese population. Hum. Mol. Genet. 20, 4411–4421. doi: 10.1093/hmg/ddr369
Lu, C., Zhang, J., Li, Y., Xia, Y., Zhang, F., Wu, B., et al. (2009). The b2/b3 subdeletion shows higher risk of spermatogenic failure and higher frequency of complete AZFc deletion than the gr/gr subdeletion in a Chinese population. Hum. Mol. Genet. 18, 1122–1130. doi: 10.1093/hmg/ddn427
Machev, N., Saut, N., Longepied, G., Terriou, P., Navarro, A., Levy, N., et al. (2004). Sequence family variant loss from the AZFc interval of the human Y chromosome, but not gene copy loss, is strongly associated with male infertility. J. Med. Genet. 41, 814–825. doi: 10.1136/jmg.2004.022111
Massalska, D., Bijok, J., Zimowski, J. G., Jozwiak, A., Jakiel, G., and Roszkowski, T. (2013). Multiplex ligation-dependent probe amplification (MLPA)–new possibilities of prenatal diagnosis. Ginekol. Pol. 84, 461–464. doi: 10.17772/gp/1605
Mutlu, E. T., Aykan, H. H., and Karagoz, T. (2018). Analysis of gene copy number variations in patients with congenital heart disease using multiplex ligation-dependent probe amplification. Anatol. J. Cardiol. 20, 9–15. doi: 10.14744/AnatolJCardiol.2018.70481
Pryor, J. L., Kent-First, M., Muallem, A., Van Bergen, A. H., Nolten, W. E., Meisner, L., et al. (1997). Microdeletions in the Y chromosome of infertile men. N. Engl. J. Med. 336, 534–539. doi: 10.1056/NEJM199702203360802
Quarello, P., Garelli, E., Brusco, A., Carando, A., Mancini, C., Pappi, P., et al. (2012). High frequency of ribosomal protein gene deletions in Italian Diamond-Blackfan anemia patients detected by multiplex ligation-dependent probe amplification assay. Haematologica 97, 1813–1817. doi: 10.3324/haematol.2012.062281
Rozen, S. G., Marszalek, J. D., Irenze, K., Skaletsky, H., Brown, L. G., Oates, R. D., et al. (2012). AZFc deletions and spermatogenic failure: a population-based survey of 20,000 Y chromosomes. Am. J. Hum. Genet. 91, 890–896. doi: 10.1016/j.ajhg.2012.09.003
Saito, K., Miyado, M., Kobori, Y., Tanaka, Y., Ishikawa, H., Yoshida, A., et al. (2015). Copy-number variations in Y-chromosomal azoospermia factor regions identified by multiplex ligation-dependent probe amplification. J. Hum. Genet. 60, 127–131. doi: 10.1038/jhg.2014.115
Skaletsky, H., Kuroda-Kawaguchi, T., Minx, P. J., Cordum, H. S., Hillier, L., Brown, L. G., et al. (2003). The male-specific region of the human Y chromosome is a mosaic of discrete sequence classes. Nature 423, 825–837. doi: 10.1038/nature01722
Yan, Y., Yang, X., Liu, Y., Shen, Y., Tu, W., Dong, Q., et al. (2017). Copy number variation of functional RBMY1 is associated with sperm motility: an azoospermia factor-linked candidate for asthenozoospermia. Hum. Reprod. 32, 1521–1531. doi: 10.1093/humrep/dex100
Keywords: male infertility, Y chromosome, azoospermia factor regions, spermatogenic failure, copy number variants
Citation: Zhou R, Cheng J, Ma D, Tan J, Wang Y, Hu P and Xu Z (2019) Identifying Novel Copy Number Variants in Azoospermia Factor Regions and Evaluating Their Effects on Spermatogenic Impairment. Front. Genet. 10:427. doi: 10.3389/fgene.2019.00427
Received: 26 November 2018; Accepted: 18 April 2019;
Published: 07 May 2019.
Edited by:
Dana C. Crawford, Case Western Reserve University, United StatesReviewed by:
Oscar Puig, Bristol Myers Squibb, United StatesCopyright © 2019 Zhou, Cheng, Ma, Tan, Wang, Hu and Xu. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Zhengfeng Xu, emhlbmdmZW5nJiN4MDAwNUY7eHUmI3gwMDA1RjtuakAxNjMuY29t
†These authors have contributed equally to this work
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.